BIOL 271 Wolverton Genetics Exam 2
1/54
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
55 Terms
Segregation
2 Alleles at the same locus separate in gamete formation to ensure each gamete receives one allele for each gene.
Independent Assortment
Alleles at 1 locus act independently of alleles at another locus during gamete formation
Recombination
New allele combinations from independent assortment
Linked Genes
Genes located close together on same chromosome travel together during meiosis
Crossing Over
Genes switch from one homologous chromosome to another, resulting in recombination. Max rate of 50% = independent assortment. Closer loci have a lower probability.
Complete Linkage
All offspring show nonrecombinant phenotypes after testcross
Recombination Frequency
(Number of recombinant offspring) / (Total number of offspring)
Coupling
Double dominant or double recessive genes on the same chromosome. After testcross, if homozygous individuals are the majority.
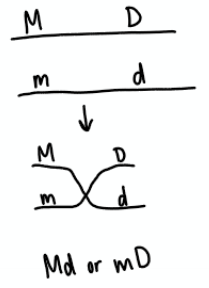
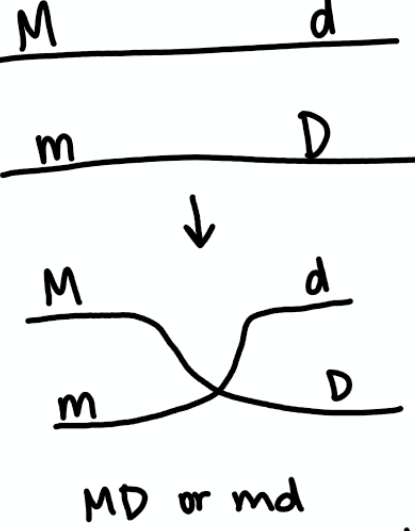
Repulsion
Doubly heterozygous chromosomes. After testcross, if heterozygous individuals are in majority.
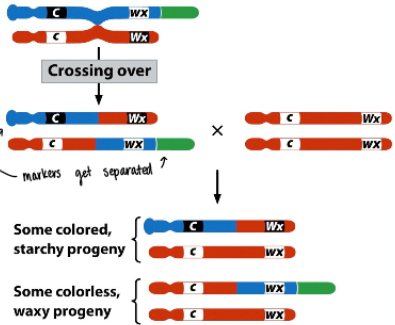
Barbara McClintock’s experiment
Demonstrated that recombination was due to physical exchange between chromosomes (crossing over) by pairing counting offspring phenotypes with staining chromosomes to track locations
Chi Square Test of Independence
χ² = Σ((O-E)² / E, but with the fancy table where E = ((row total x column total)/grand total)
Gene mapping
Genetic maps based on recombination frequencies. Double crossover always decreases the distance between genes.
1) Identify nonrecombinants (most common)
2) Identify double-crossover progeny (least common)
3) Determine the middle locus
4) Rewrite alleles in correct order
5) Determine locations of crossovers
6) Determine recombination frequencies
Supercoiling
Lowest energy state for B-DNA (10 bp/turn)
Topoisomerases rotate DNA, requiring energy input
Overrotates: Positive supercoiling
Underrotated: Negative supercoiling
Heterochromatin
Remains condensed throughout cell cycle. Less available to cell. Long-term storage
Euchromatin
Condenses and decondenses throughout cell cycle by energy input. More accessible for cell.
Histone
Proteins that DNA is wrapped around
Nucleosome
Collection of 8 wrapped histones with about 145-147 bp of DNA. 11 nm
Chromatosome
Collection of wrapped nucleosomes
30-nm fiber
Collection of chromatosomes. Main thread knitted into chromosome.
Linker DNA
DNA between histones that is cleaved and destroyed by nuclease
Does chromatin structure change during transcription? (is gene expression altered by chromatin structure?)
Experiment: Chicken DNA- Sensitivity of DNA to DNase I is correlated with gene expression, suggesting that chromatin structure changes during transcription
Control: Brain cells throughout development, which do not produce globin remained INSENSITIVE to DNase
Erythroblasts first 24 hours: before hemoglobin synthesis, no globin genes are sensitive to DNase I digestion.
Erythroblasts 5 days: After globin synthesis has begun, embryonic globin gene U is most sensitive at same time of gene expression for that region.
Erythroblasts 14 days: Adult genes are most sensitive and embryonic gene is insensitive.
Centromere
Defined location where microtubules connect. Defined by specific DNA sequences involved in topology/shape, but not informative and never encoded into mRNA. The sequence acts as a binding site for kinetochore using linking proteins. Often contains tandem repeats of G and C as they have triple H bonds.
Metacentric
Centromere in middle
Submetacentric
Centromere slightly off middle
Acrocentric
Centromere greatly off middle
Telocentric
Centromere at end
Telomeres
Caps on the ends of chromosomes. G-C rich, highly conserved region of DNA throughout eukaryotes.
Central Dogma
DNA→transcription→RNA→translation→PROTEIN
Reverse Transcription
Used by viruses RNA→DNA
Properties of Hereditary Material
1) High-density info storage
2) Faithful replication
3) Able to encode phenotype
4) Capacity to vary
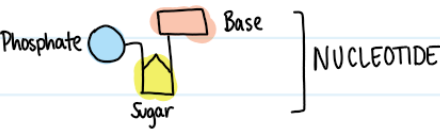
Contents of DNA
Phosphate, sugar, and nitrogenous base form nucleotide
Griffith -1928
Study of pneumonia in mice. Discovered TRANSFORMATION PRINCIPLE (genetic material is transferred/transformed)
Virulent (smooth, sneaky, alive)= dead
nonvirulent (rough, alive)= alive
heat-killed virulent (smooth, sneaky, dead) = alive
heat-killed virulent + nonvirulent (smooth, sneaky container + alive)= DEAD
Avery, MacLeod, McCarty -1944
DNA is the transforming substance! (not RNA or PROTEIN)
1) Took heat killed smooth, sneaky + dead bacteria
2) Treated with RNase, Protease, and DNase.
3) Sample with no DNA=no transformation (no smooth cells)
Hershey and Chase -1955
Changes model systems to bacteria phage viruses.
1) Infect E. coli grown in a medium containing 35S (because Sulfur is a part of proteins but not DNA)
a) 35S PROTEIN + UNLABELED BACTERIA
2) Infect E. coli grown in a medium containing 32P (because Phosphate is part of DNA and not PROTEINS)
a) 32P DNA + UNLABELED BACTERIA
3) Shear off protein coats in blender
4) Seperate protein from cells by centrifuging
5) PROTEIN BACTERIA IS UNLABELED, DNA BACTERIA IS LABELED (DNA was transmitted)
Chargaff’s Rules
( A = T ) and ( G = C )
(%A + %G)/(%T + %C) = 1
X-ray crystallography
Allows us to see DNA structure
Rosalind Franklin’s Contribution
DNA has two strands
Sugar-phosphate backbone
Atomic distance measurements
Watson and Crick -1953
DNA is double-stranded
Strands for a double helix
Strands are chemically antiparallel
DNA is complimentary
Purine
Adenine and Guanine, 2 rings
Pyrimidine
Cytosine, Thymine, and Uracil, 1 ring
DNA Primary Structure
Nucleotide
DNA Secondary Structure
Double-stranded, complimentary, antiparallel, double-helix with phospho-diester bonds on outsides and H+ bonds on insides. C-G is TRIPLE H+ bonded, A-T is DOUBLE H+ bonded. 2 nm wide. 10 bases every full revolution is about 3.4 nm. 0.34 nm between each base.
Can only add 5’→3’ on the 3’ strand
Different forms of DNA Helix
A form: more condensed
B form: occurs in high water content, less condensed
Z form: direction of helix is backwards
Chromosomal Sex Determination
Sex determined by genes encoded on sex chromosomes encoding for sexual development and can impact how autosomal influences are presented.
50:50 sex ratio maintained by segregation (XX vs XY)
X and Y chromosomes are homologous only at pseudoautosomal regions, essential for X-Y chromosome pairing in meiosis in the male (XY). Primary region at short arm end. Secondary region at long arm end.
Turner Syndrome
XO genotype (lack of Y chromosome) FEMALE, but underdeveloped traits
SRY gene
Sex-determining region on Y chromosome, acts as master regulator because it encodes for development of male sexual traits
ZZ/ZW
X/Y system, but females are the heterogametic sex.
XX/XO
Female XX, male have XO (no pair)
Hemizygous
Have 1 allele and no homologue on other chromosome (XO)
Haplodiploidy
Sex determined by number of chromosome sets (ploidy level) male: n (no fertilization) female: 2n (fertilization)
Genic Sexual Determination
No difference in chromosomes (no presence of sex chromosomes)
Sex determined by genotype at particular loci
Environmental Sexual Determination
Environmental position influences exposure to sex-determining factor (e.g. temperature, hormones) (e.g. marine mollusks that attract larvae that develop into males on female, mate, and eventually switch into females (hermaphroditism) that attract additional larvae, etc)
Genic Balance System Sexual Determination
Sex determined by ratio of autosomal and sex chromosome genes. (X:A)
X has female-producing genes
Autosomes (A) has male-producing genes
X:A ratio
Sex-linked traits
Typically encoded by gene on sex chromosome
Nondisjunction
Failure of chromosomes to separate properly during cell division, causing an abnormal number of chromosomes in daughter cells.