e2 cram
1/90
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
91 Terms
enzyme
alters a reaction rate by lowering activation energy
apoenzyme
enzyme without a cofactor
holoenzyme
enzyme with a cofactor
substrate
substance acted on by an enzyme
active site
location where substrate binds
cofactor
aids enzyme
coenzyme
organic molecule that associates with the enzyme
prosthetic group
organic molecule that covalently attaches to the enzyme
substrate level control
alters susbtrate concentrations based on Le Chatlier’s Principles
Feedback Control
accumulation of product can either increase or decrease the rate of the previous step
allosteric regulation
allosteric binding leads to conformational change of the active site
homotrophic effector
appears the same as the substrate and binds to the same site
heterotrophic effector
appears different than the substrate and binds to a site alternate to the active site
positive effector
enhances binding
negative effector
inhibits binding
6 enzyme classes
OTHLIL
oxidoreductase, transferase, hydrolase, lyase, isomerase, and ligase
oxidoreductase (1)
catalyzes oxidation/reduction reactions
A + B → C + D
transferase (2)
catalyzes the transfer of a functional group from one molecule to another
A-X + B → A + B-X
hydrolase (3)
catalyzes the hydrolysis of a chemical bond, typically involving the reaction of water with a substrate
AB + H2O → A-OH + H-B
lyase (4)
catalyzes the cleavage of chemical bonds by means other than hydrolysis or oxidation
B + C → A
isomerase (5)
catalyzes the conversion of a molecule from one isomer to another without the addition or removal of atoms
A-B → B-A
ligase (6)
catalyzes the joining of two molecules, typically using ATP
A + B + ATP → A-B + ADP + Pi
6 Reaction Mechanisms
(ACMEPT)
acid-base catalysis
covalent catalysis
metal ion catalysis
electrostatic catalysis
proximity
transition state catalysis
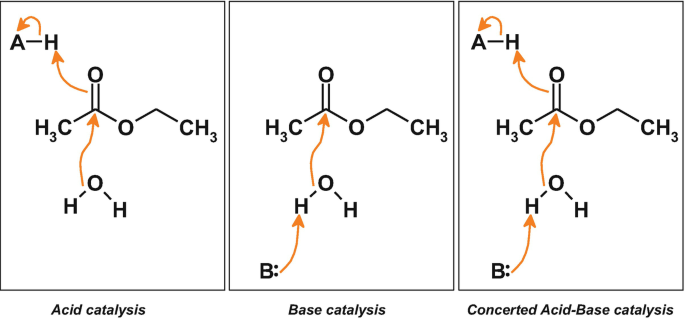
acid-base catalysis
a process where proton transfer facilitates the reaction mechanism by stabilizing charged intermediates.
a. General Acid Catalysis: An amino acid side-chain of the enzyme donates H+ to the substrate. Commonly used: Lys, Tyr, Arg, Cys.
b. General Base Catalysis: An amino acid side-chain of the enzyme abstracts H+ from the substrate. Commonly used: Glu, Asp, His.
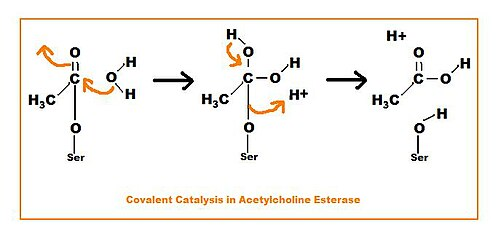
covalent catalysis
an amino acid side-chain of the enzyme forms a covalent bond with the substrate
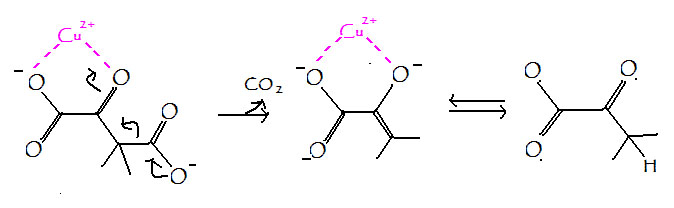
metal ion catalysis
metal ion binds substrate to stabilize negative charge
electrostatic catalysis
charged amino acid side-chain of enzyme stabilizes opposite charge on substrate
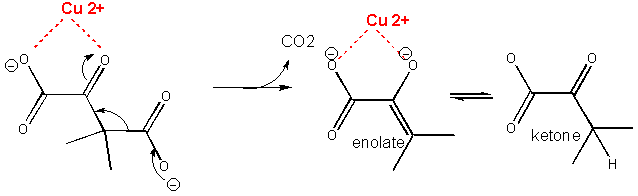
proximity and orientation effect
two substrates are brought close enough and properly oriented to each other to react
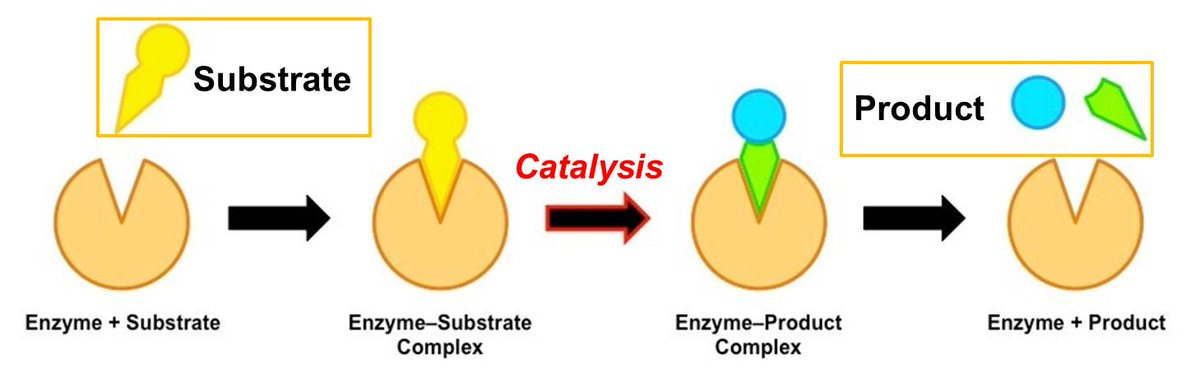
catalysis via preferential transition state binding
binding to the enzyme allows the substrate to adopt a transition state conformation, making the reaction easier
Michaelis-Menten Reaction Equation
[E]t=[E]+[ES]
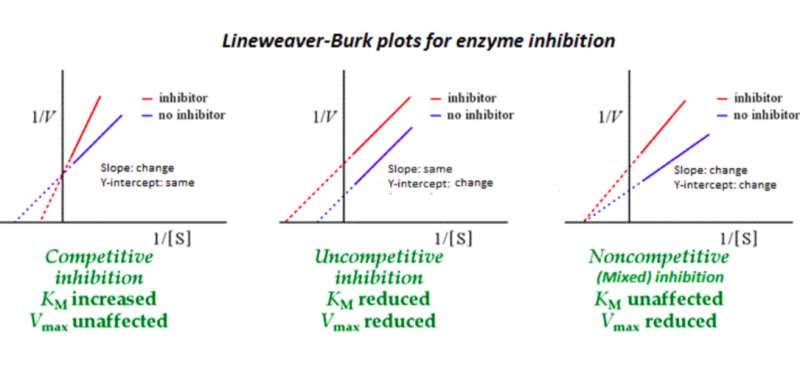
Types of Inhibition based on the Lineweaver-Burke Plot
Competitive, Uncompetitive, and Noncompetitive
Reaction Rate
the change in concentration per unit time
Competitive Inhibition
effector binds to the free enzyme and blocks the active site so that the substrate cannot bind
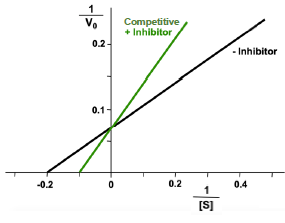
Uncompetitive Inhibition
effector binds at the allosteric site and slows down the reaction
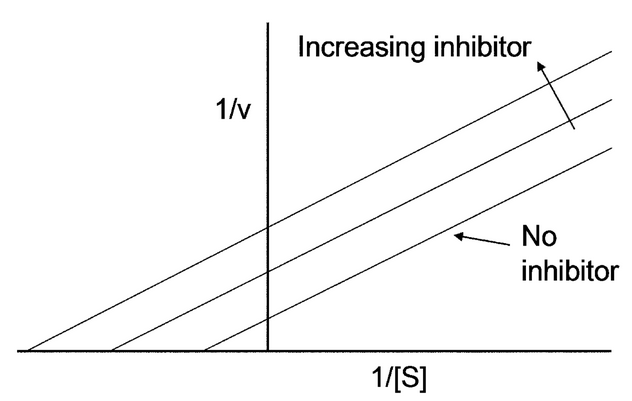
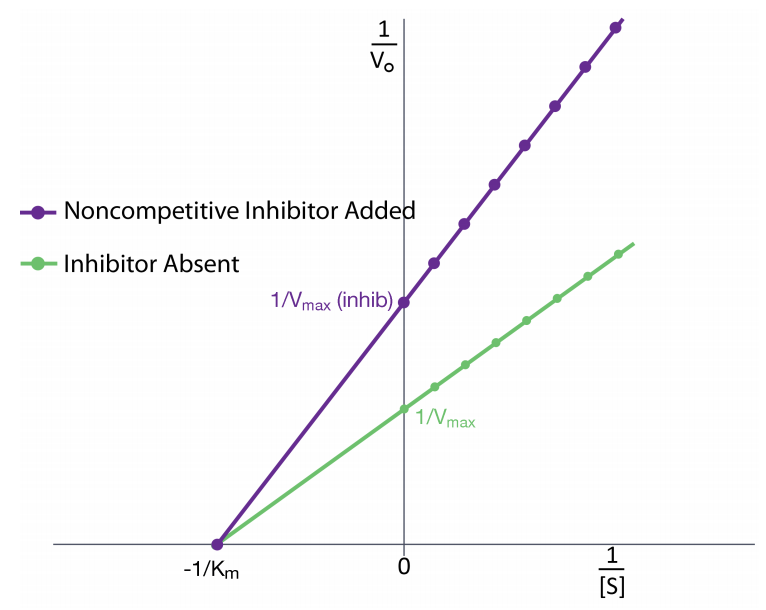
Noncompetitive Inhibition
effector binds either the E or ES at an allosteric site, slowing the reaction
Carbohydrate
monomeric or polymeric polyhydroxylated aldehyde or ketone
aldose
sugar with an aldehyde group
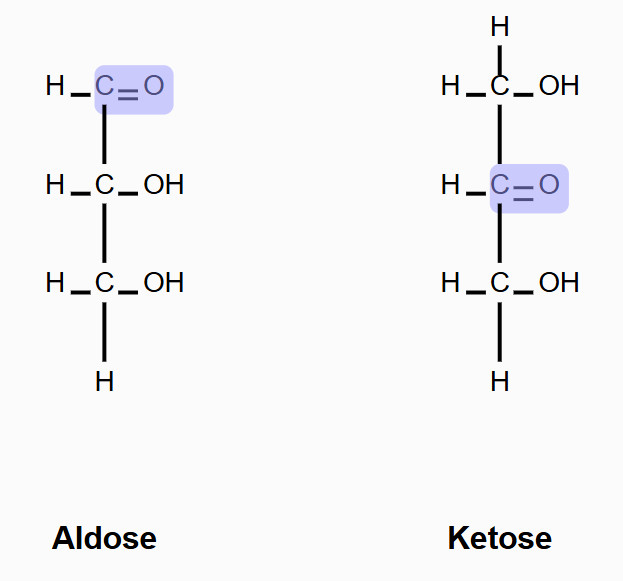
ketose
sugar with a ketone group
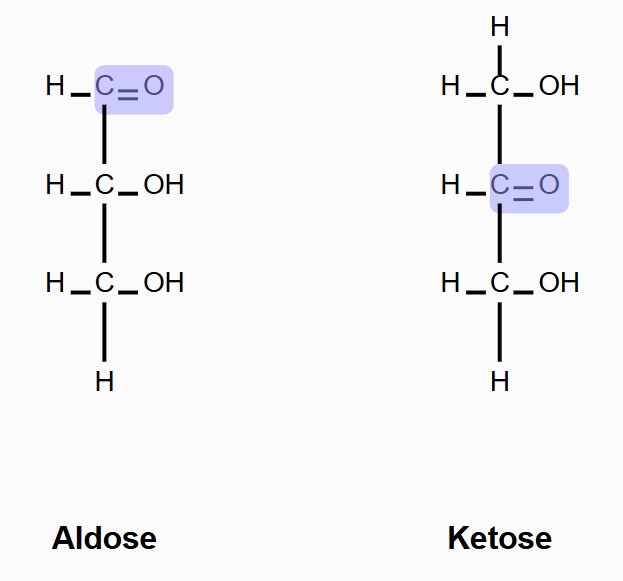
sugar acids
sugar with a carboxylic acid group (two types: aldonic and uronic acids)
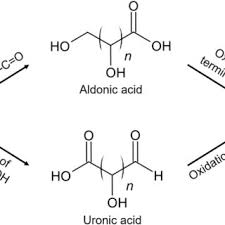
aldonic acid
Carboxylic acid at C1
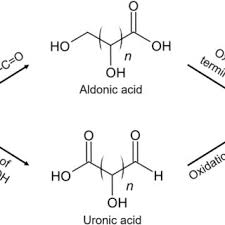
uronic acid
Carboxylic acid at last carbon
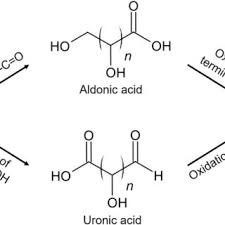
Mutarotation
change of rotation of plane-polarized light during interconversion of anomeric forms in solution
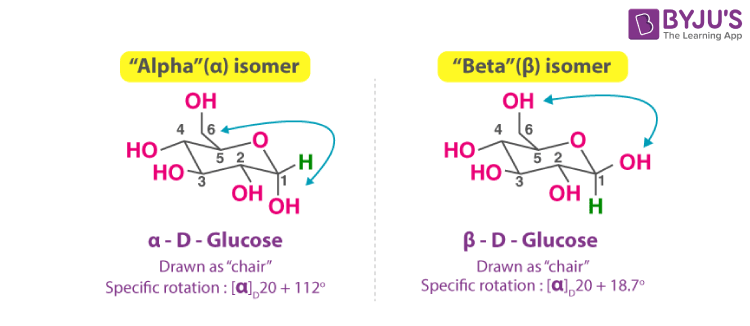
pyranose
6-membered ring form
furanose
5-membered ring form
anomers
stereoisomers that differ in orientation about the anomeric carbon , where one is typically a carbohydrate's cyclic form and the other its open-chain form. e
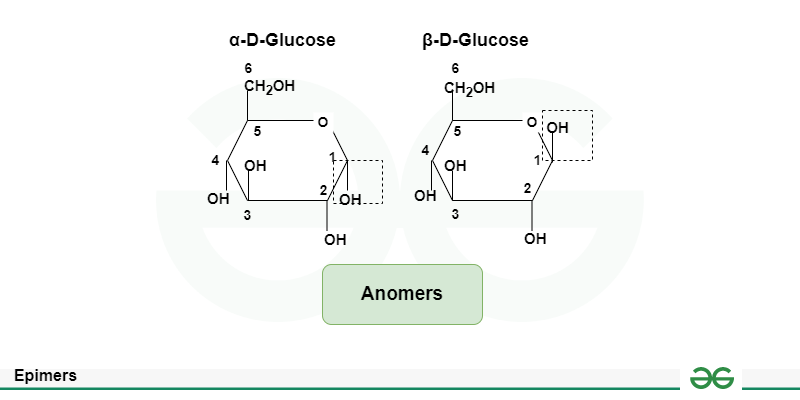
epimers
steroisomers that differ at any non-anomeric carbon
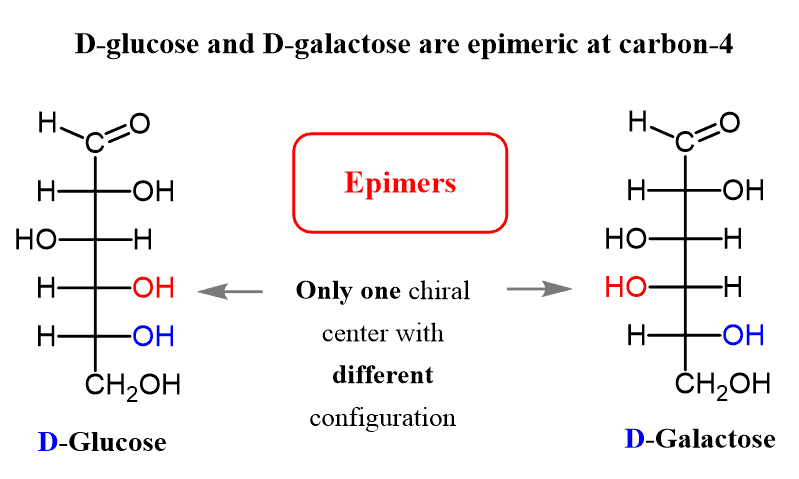
monosaccharides are…
polar and capable of hydrogen bonding
Glycosylation
reaction of sugar with alcohols yielding glycosidic linkages
5 Key sugars
(GMRGF)
Glucose
Mannose
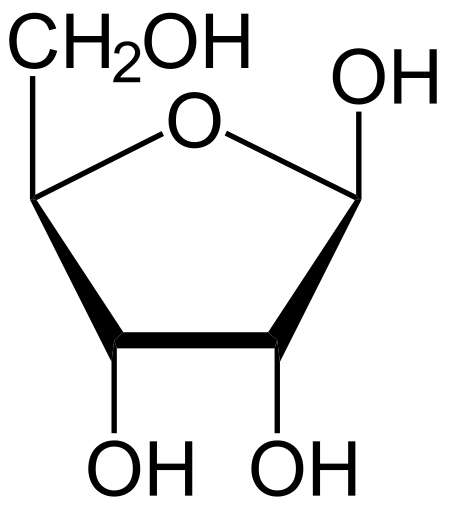
Ribose
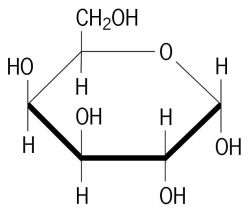
Galactose
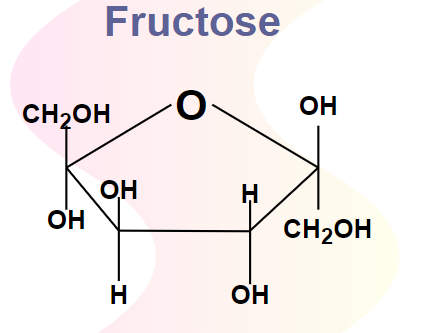
Fructose
Glucose
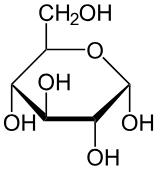
Mannose
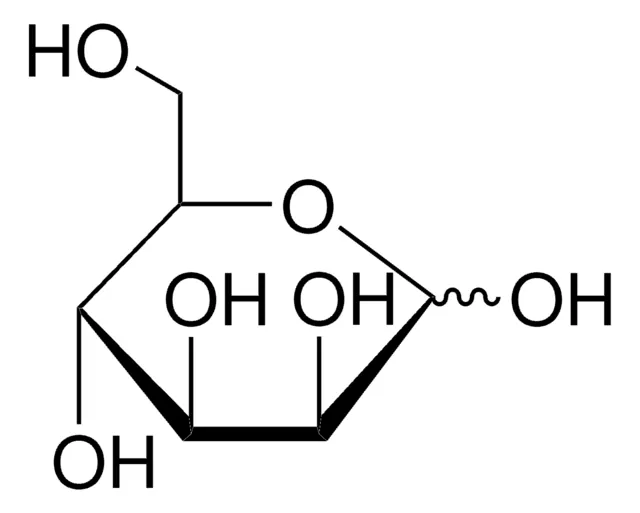
Ribose
Galactose
Fructose
2 Classifications of Polysaccharides
Structural and Storage
(STORE SG, STRIKE CC)
Storage polysaccharides
starch and glycogen
Structural polysaccharides
Cellulose, Chitin, and acidic polysaccharides
Glycolysis
conversion of hexoses into trioses with ATP production
Glycogenolysis
degradation of glycogen into glucose for glycolysis, primarily in the muscles and liver
Pentose Phosphate Pathway
degradation of glycolytic metabolites yielding NADPH and carbohydrate byproducts
Gluconeogenesis
construction of glucose from small molecules
Glycogenesis
creation of glycogen
What are the 3 Catabolic Pathways in Carbohydrate Metabolism?
(LYSIS)
Glycolysis
Glucogenolysis
Pentose Phosphate Pathway
What are the 3 Anabolic Pathways in Carbohydrate Metabolism?
(GENESIS)
Gluconeogenesis
Glycogenesis
Pentose Phosphate Pathway
Of the methods of carbohydrate metabolism, which process can serve both anabolic and catabolic functions?
The Pentose Phosphate Pathway is unique as it can serve both anabolic functions, such as nucleotide synthesis, and catabolic functions, such as generating NADPH for cellular processes.
Glycolysis has ___ phases
2
Phase 1 of Glycolysis
glucose splits into two trioses, utilizing 2 ATP
Phase 2 of Glycolysis
conversion of two trioses into pyruvate, generating 4 ATP
What are the enzymes, in order, of Glycolysis?
(HPPATGPPEP)
(“Harry Potter Pirated All The Goods!” Pleaded Eunice Politely)
(2.7, 5.3, 2.7, 4.1, 5.3, 1.2, 2.7, 5.4, 4.2, 2.7)
Hexokinase - 2.7
Phosphoglucoisomerase -b 5.3
Phosphofructokinase - 2.7
Aldolase - 4.1
Triose Phosphate Isomerase - 5.3
Glyceraldehyde-3-phosphate dehydrogenase - 1.2
Phosphoglycerate kinase - 2.7
Phosphoglycerate mutase - 5.4
Enolase - 4.2
Pyruvate kinase - 2.7
Hexokinase
Enzyme 1 - costs 1 ATP to place phosphate on glucose via SN2
Phosphoglucoisomerase
Enzyme 2 - Catalyzes isomerization via enediol/enediolate intermediates
Phosphofructokinase-1
Enzyme 3 - adds a phosphate to F6P
Aldolase
Enzyme 4 - cleaved F-1,6-bisphosphate into DHAP and G3P
Triose Phosphate Isomerase
Enzyme 5 - interconverts DHAP to G3P
glyceraldedehyde-3-phosphate dehydrogenase
enzyme 6 - catalyzes the conversion of G3P to 1,3-bisphosphoglycerate, reducing NAD+ to NADH in the process.
phosphoglycerate kinase
enzyme 7 - produces ATP from 1,3-bisphosphoglycerate
phosphoglycerate mutase
enzyme 8 - catalyzes the conversion of 2-phosphoglycerate
Enolase
enzyme 9 - catalyzes the conversion of 2-phosphoglycerate to phosphoenolpyruvate, releasing water in the process.
pyruvate kinase
enzyme 10 - forms pyruvate and ATP.
3 Regulatory Enzymes in Glycolysis
Hexokinase, Phosphofructokinase-1, and Pyruvate Kinase
How does Hexokinase regulate Glycolysis?
prevents glucose from leaving the cell and inhibits its own activity when glucose-6-phosphate levels are high.
How does Pyruvate Kinase regulate Glycolysis?
it is inhibited by ATP and alanine and activated by F1,6BP via feedforward activation
Gluconeogenesis is the reverse of…
Glycolysis
what are the key enzymes of gluconeogenesis?
(PPFG)
(Power PuFf Girls!)
Pyruvate Carboxylase
Phosphoenolpyruvate Carboxykinase
Fructose 1,6 bisphosphate
Glucose-6-Phosphatase
Pyruvate Carboxylase in Gluconeogenesis
converts pyruvate into oxaloacetate; activated by acetyl-CoA
Phosphoenolpyruvate Carboxykinase in Gluconeogenesis
converts oxaloacetate to phosphoenolpyruvate
Fructose-1,6-Bisphosphate in Gluconeogenesis
critical regulatory step inhibited by AMP and F2,6BP
Glucose-6-Phosphatase in Gluconeogenesis
final step; high Km for G6P
What enzyme ensures that gluconeogenesis only occurs when G6P levels are high?
Glucose-6-Phosphatase
What are the important enzymes in Glycogenolysis?
(GPG)
(gloopy puppies glide)
Glycogen Phosphorylase
Phosphoglucomutase
Glycogen Debranching Enzyme
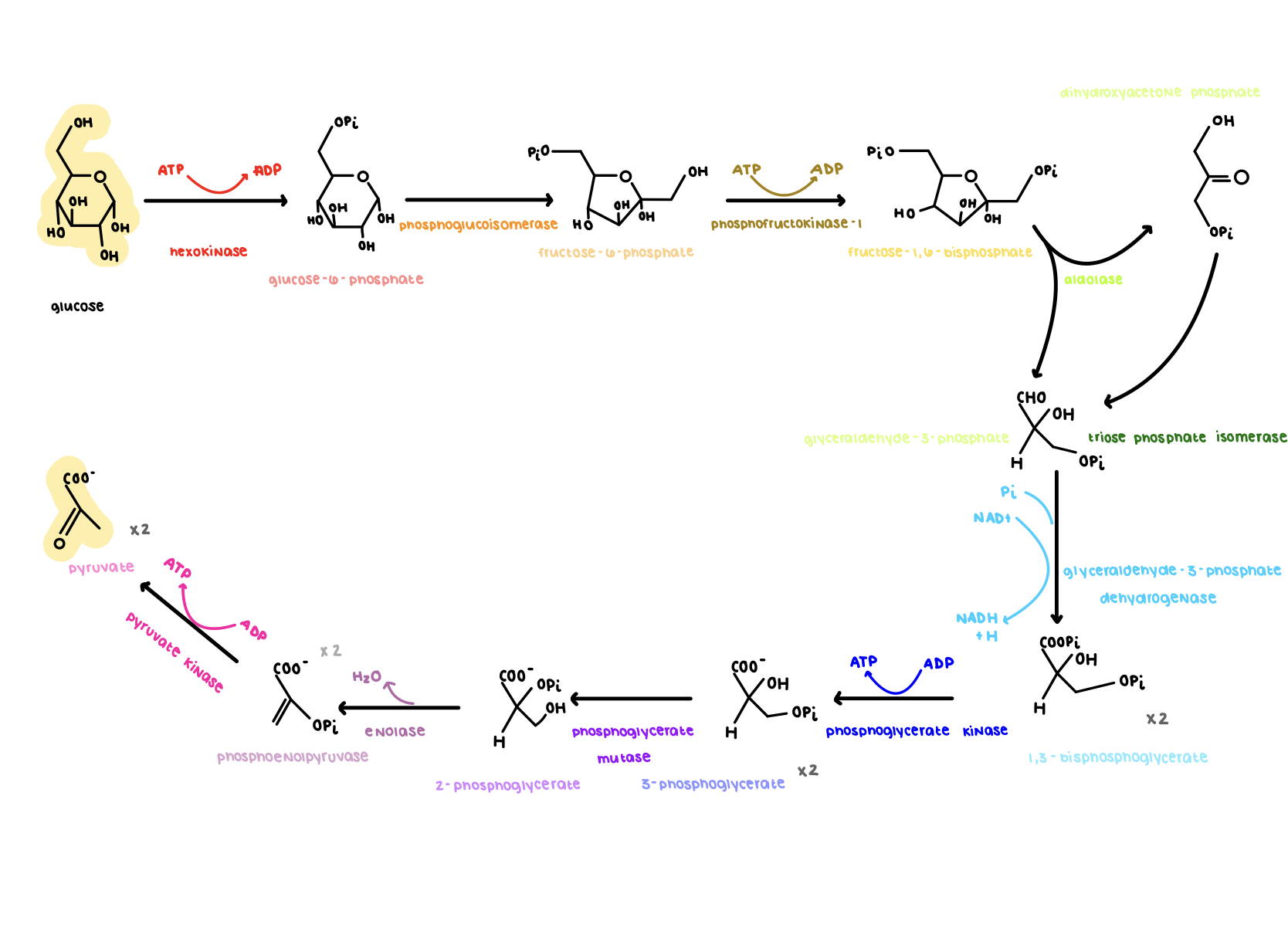
Draw out all of Glycolysis
see image