Ribosome & Protein Translation: Prokaryotic
1/21
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
22 Terms
Ribozyme
the ribosome is a ribozyme predominantly composed of RNA, with its catalytic activity driven by ribosomal RNA (rRNA)
ribosome is a ribozyme that catalyzes biochemical reactions
both tRNA (yellow) and mRNA are RNA molecules
consequently, translation is primarily an RNA-driven process
Intro to Translation
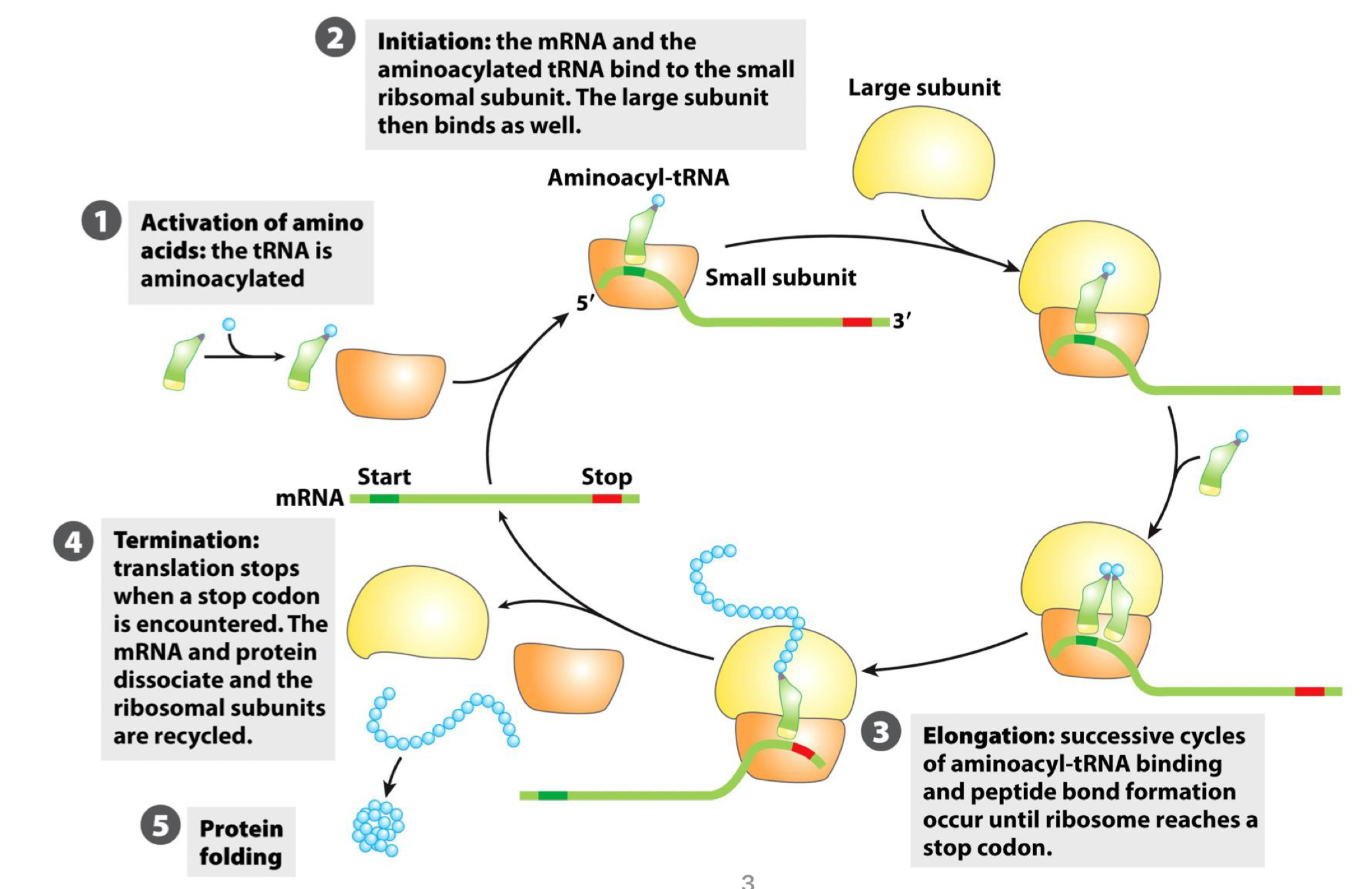
translation is the mRNA-guided process leading to protein synthesis by ribosomes, and is divided in 5 steps
the ATP-dependent activation of amino acid mediated by the aminoacyl tRNA synthetase
step 2-4 that are mediated by the ribosome
last step is protein folding
Prokaryotic Translation & Polysomes
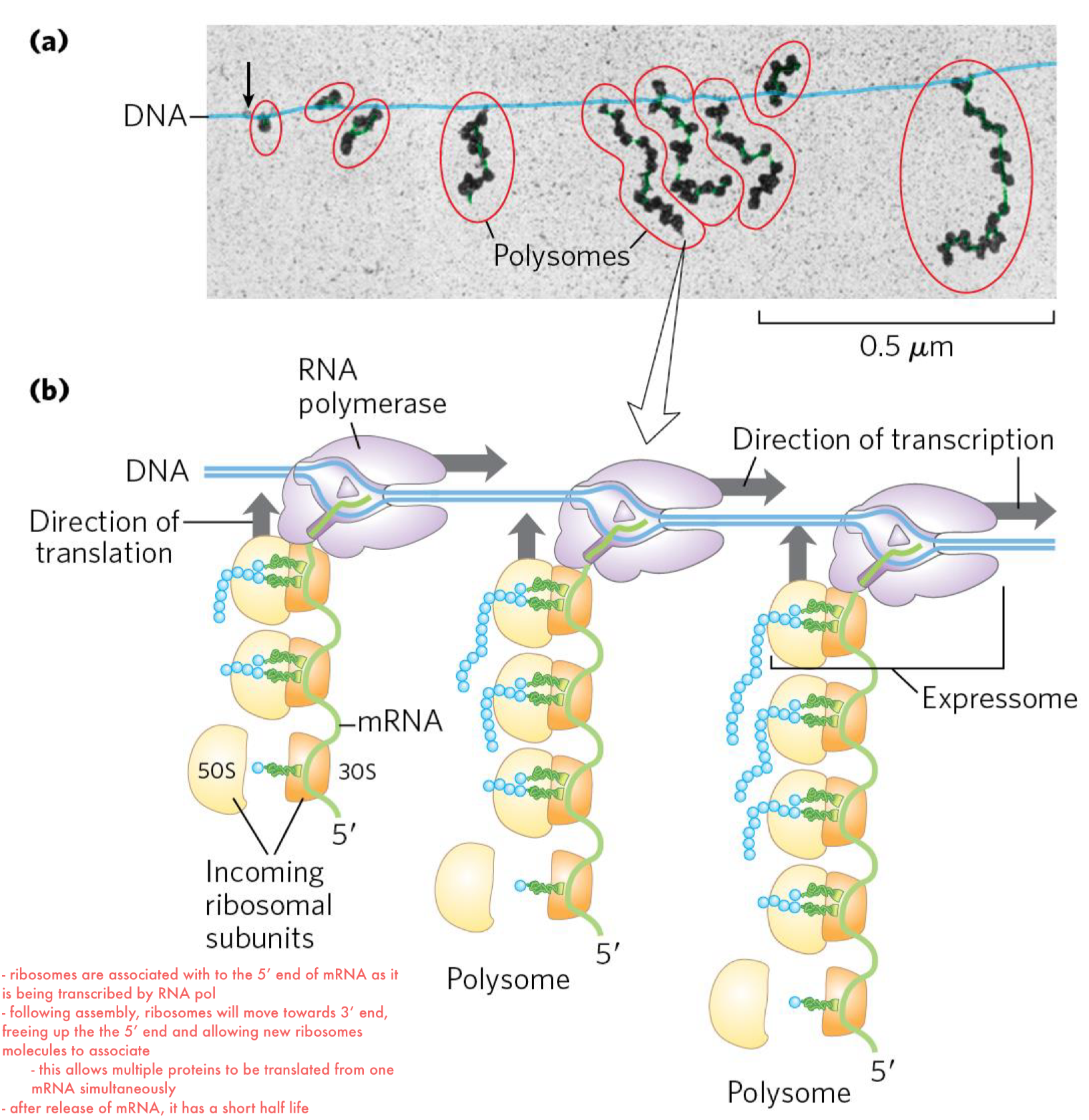
the mRNA tends to be short-lived in prokaryotes and translation is therefore tightly coupled to transcription
translation occurs on mRNA while they are still transcribed by RNA pol → forming polysomes
polysome = a cluster of ribosomes held together by a strand of messenger RNA that each ribosome is translating
multiple ribosomes are loaded on the 5’ end of mRNA forming dense structures
following assembly of the first ribosome on the mRNA, it moves toward the 3′ end, freeing up the 5’ end and allowing new ribosome molecules to associate
initiation is initiated near the 5’end of mRNA
Prokaryotic Translation & Polysomes FIGURE
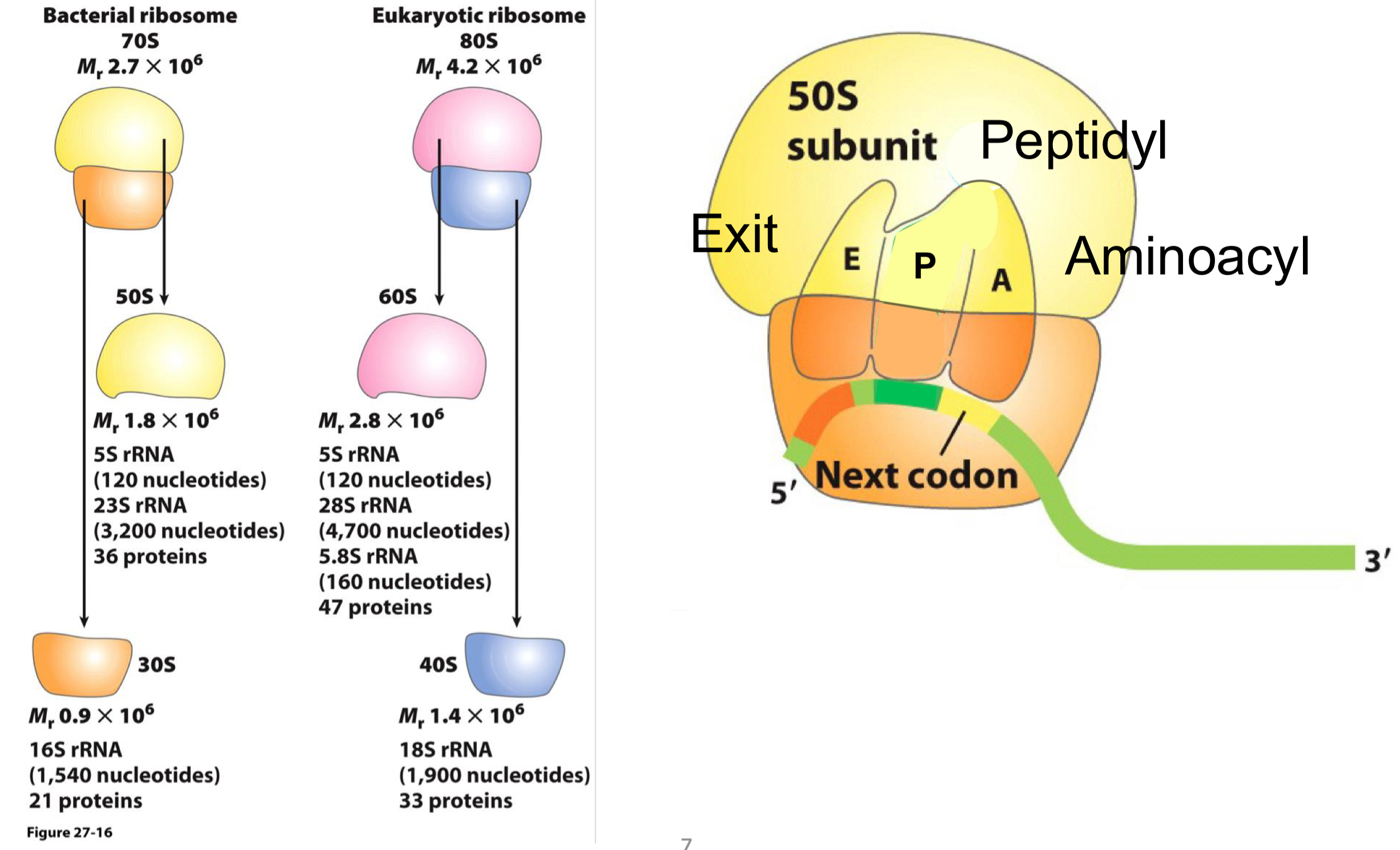
Prokaryotic Ribosomes
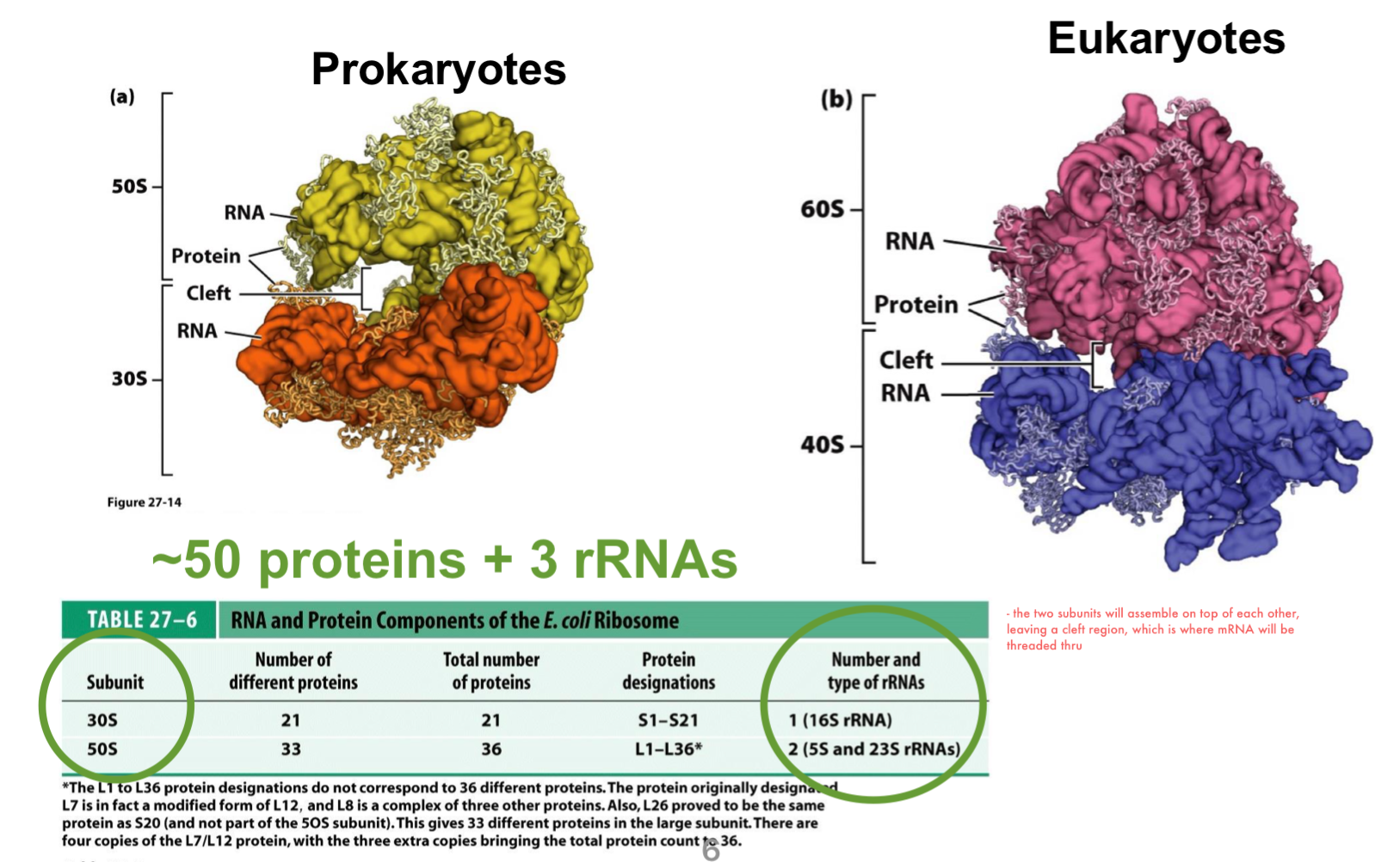
prokaryotic and eukaryotic ribosomes have similar architectures, with eukaryotes being larger
prokaryotic ribosome is formed by the 30S and 50S subunits
the two subunits will assemble on top of each other, leaving a cleft region, which is where mRNA will be threaded thru
it contains ~50 proteins and 3 major rRNA molecules: 5S and 23S in the large subunit and 16S in the small subunit
Assembled Prokaryotic Ribosome
the assembled prokaryotic ribosome forms the 70s complex
there are 3 major binding sites for tRNA:
aminoacyl (A)
peptidyl (P0
exit (E)
Initiation of Translation in Prokaryotes: Required Elements
Initiation of translation in prokaryotic cells depends on 3 elements unique to prokaryotes:
a set of initiation factors (IF 1-3)
a Shine-Dalgarno sequence on mRNA
a modified methionine fMet-tRNAfMet
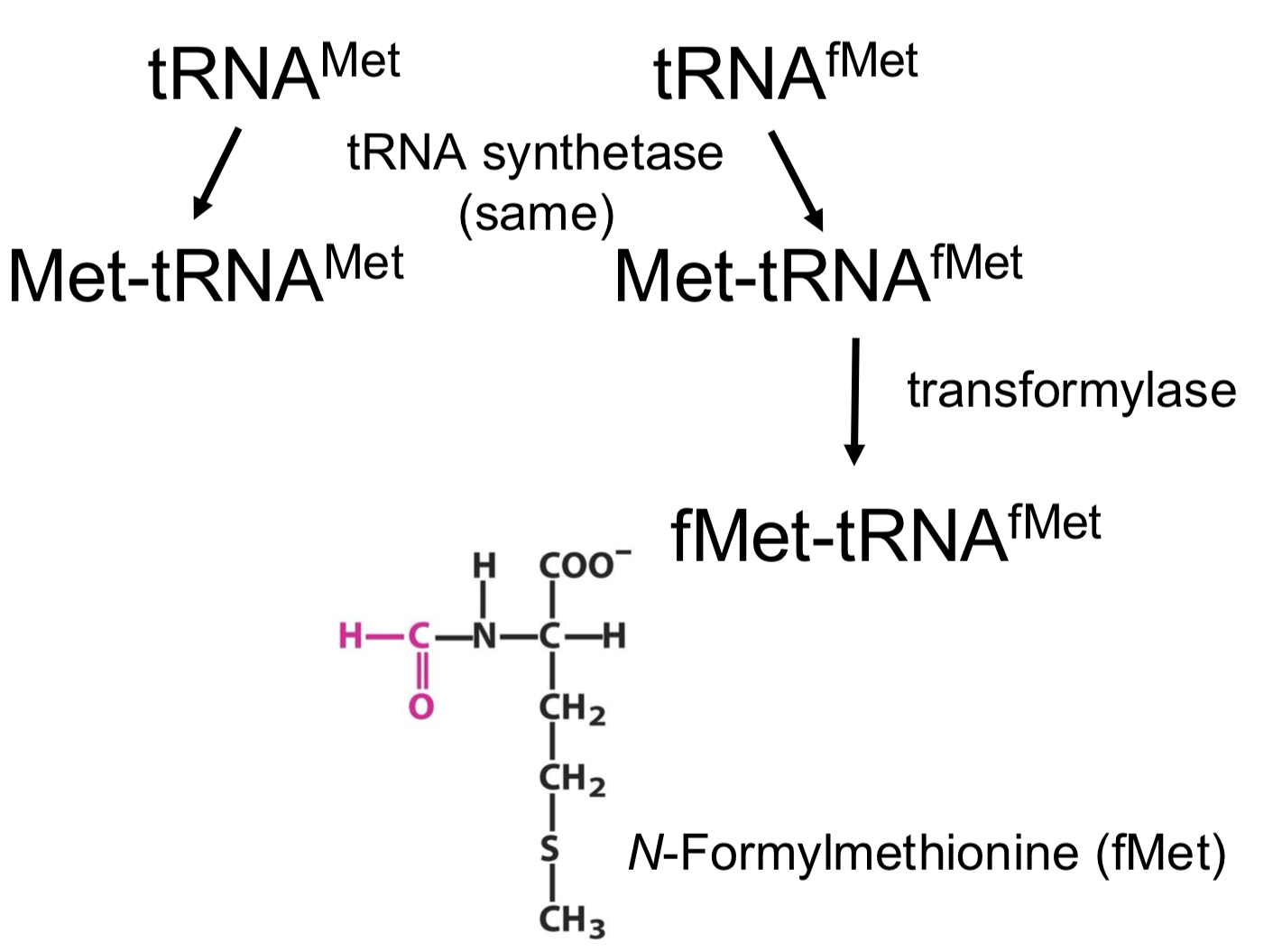
Two tRNAs for AUG codon (Met)
the tRNAfMet is a second tRNA for methionine
both tRNAMet (elongation) and tRNAfMet (initiation) have the same aminoacyl-tRNA synthetase
tRNAfMet recruits another enzyme that will modify the amino group of methionine (which then becomes the N-ter amino acid in the synthesized protein)
Initiation of Translation in Prokaryotes: Step 1
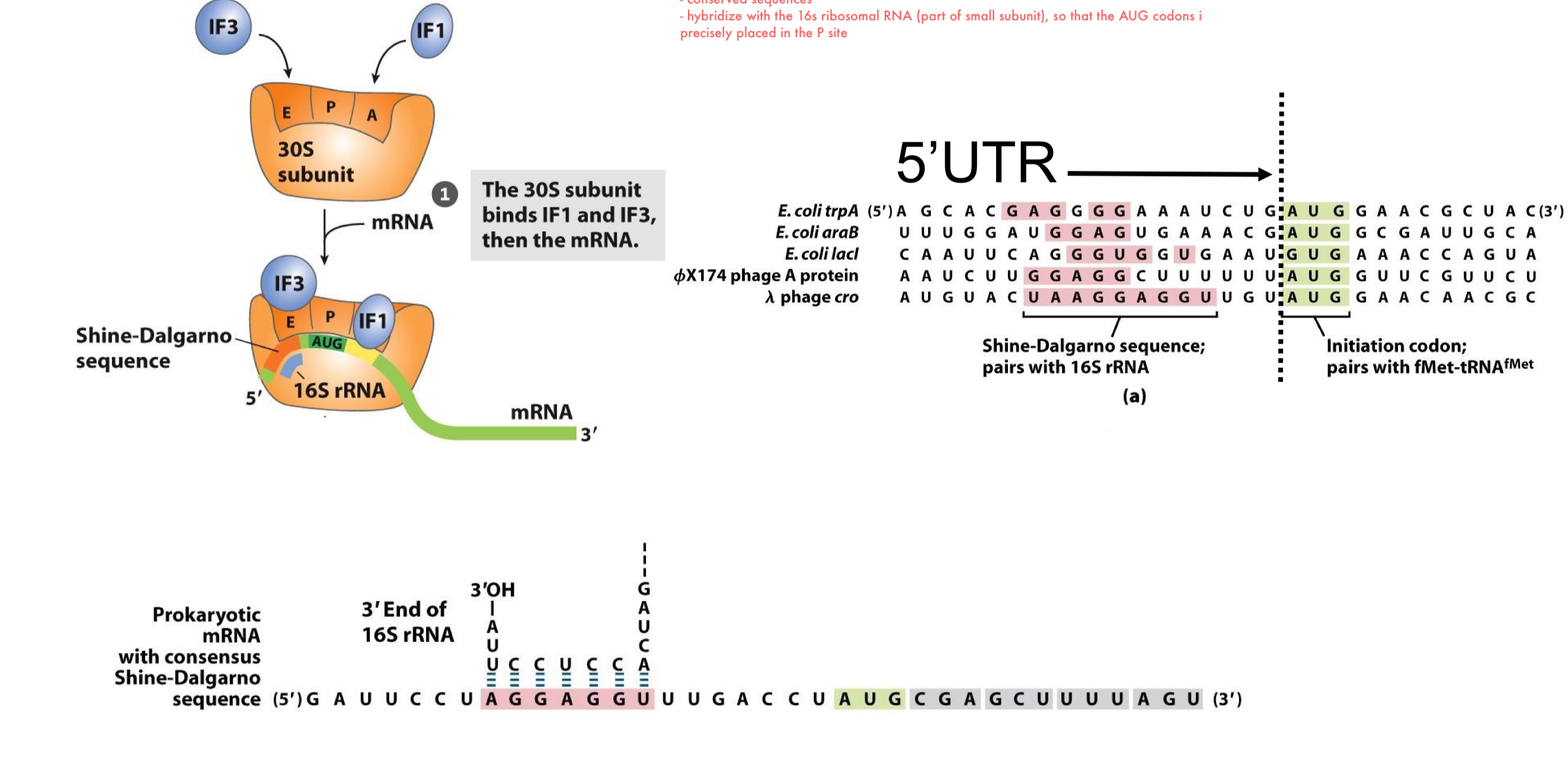
in the first step, a free 30S subunit will bind to IF1 & 3 followed by the binding to mRNA that is guided by the Shine-Dalgarno sequence that positions the first AUG codon in the P site of 30S
the Shine-Dalgarno sequence is upstream the AUG codon and will hybridize w/ 16S rRNA
IF3 prevents premature association of the large complex (50S)
IF1 sits in the A site, so that no undesired tRNA will associate to the complex
all mRNAs:
contain a Shine-Dalgarno sequence in the 5’UTR (upstream of initiation codons)
conserved sequences
Initiation of Translation in Prokaryotes: Step 1 FIGURE
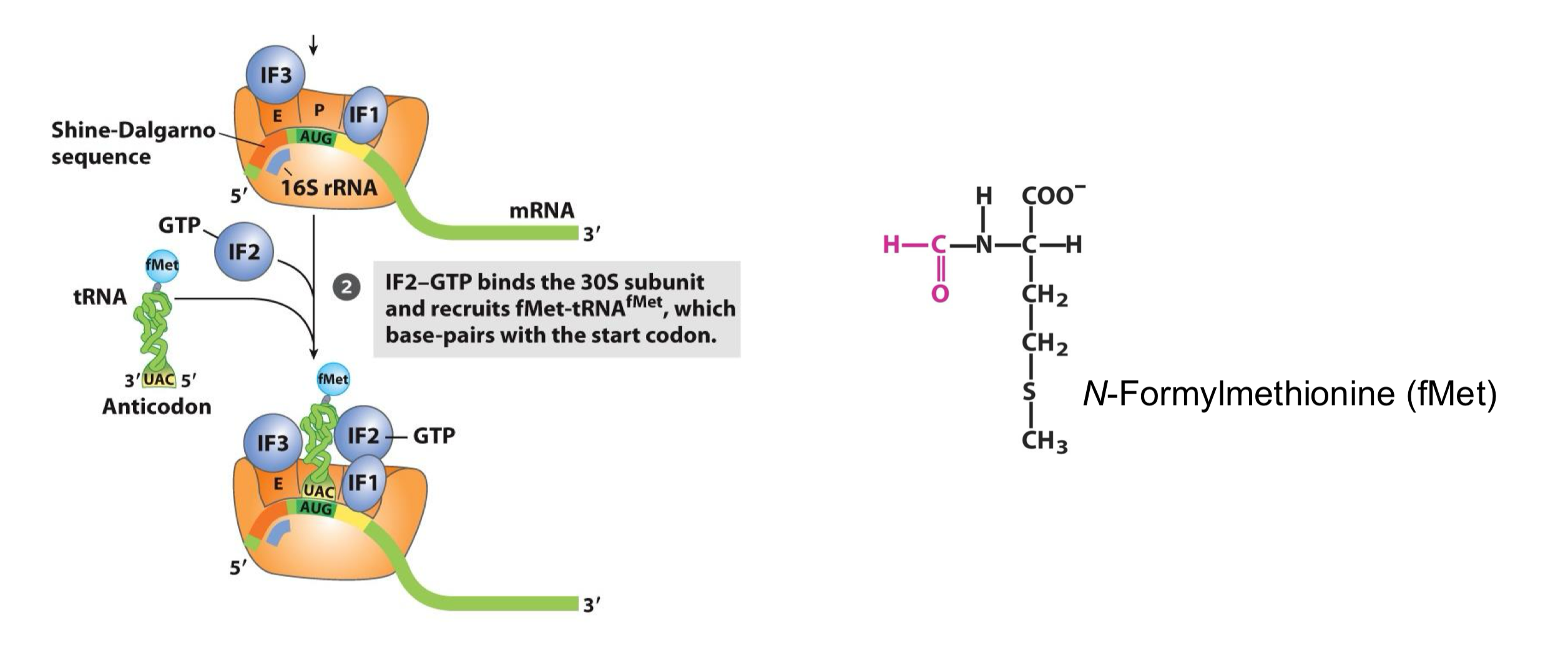
Initiation of Translation in Prokaryotes: Step 2
a GTP bound-IF2 mediates the recruitment of the initiation specific aminoacyl-tRNA in the P site, which is fMet-tRNA fMet
Initiation of Translation in Prokaryotes: Step 3
the large 50S subunit is recruited and following GTP hydrolysis, the three IFs are released
70S complex is formed with mRNA and fMet tRNA
the complex with the mRNA and fMet-tRNAfMet in the P site can now proceed to the elongation
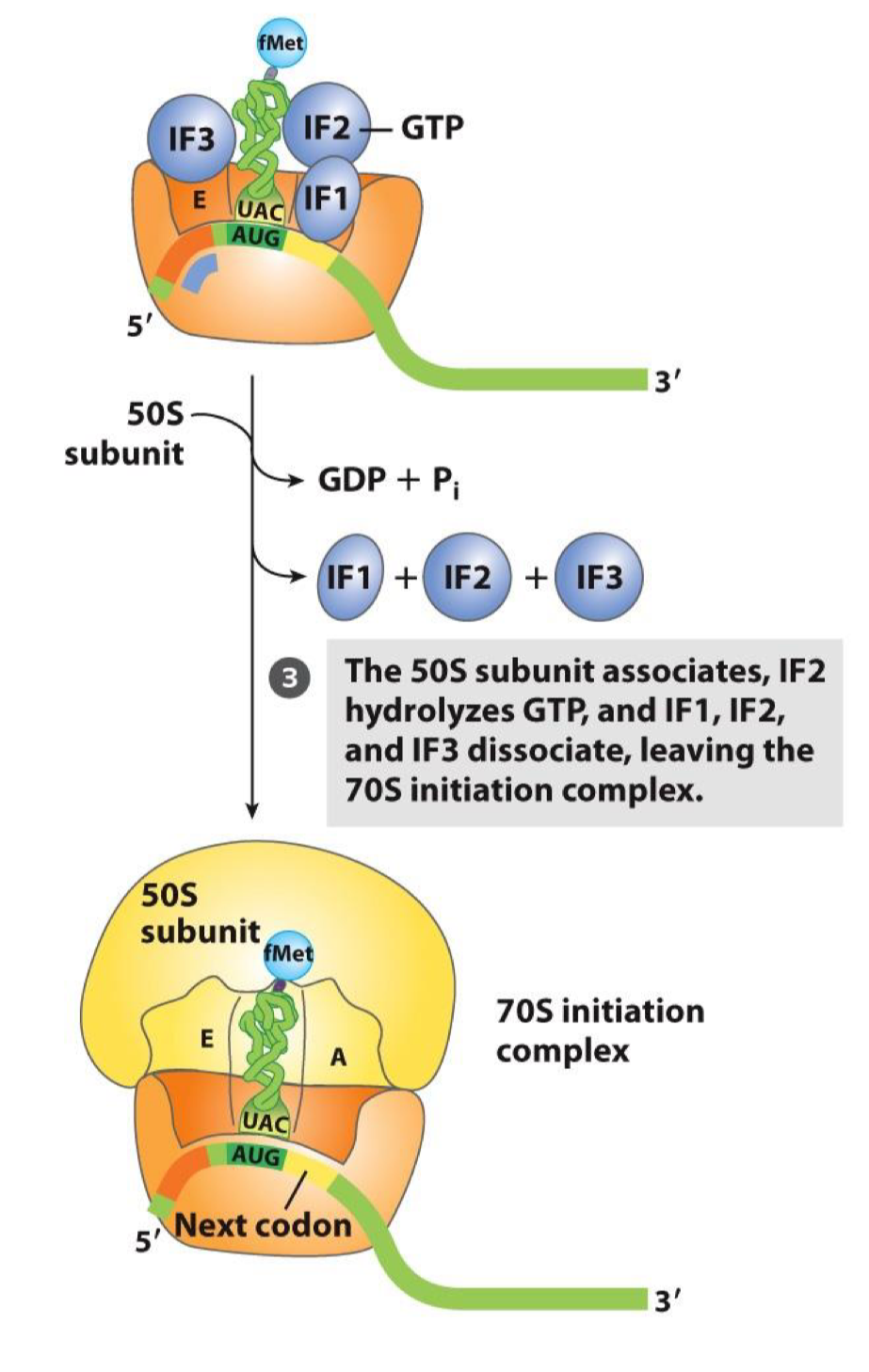
Elongation of Translation: Step 1
Recruitment of a charged AA-tRNA in the A site (based on the codon-anticodon hybridization)
the charged AA-tRNA first binds to Ef-Tu that is associated to GTP (1)
EF-Tu mediates entry into A site
tRNA don’t freely enter the ribosome, they need elongation factors
AA-tRNA-Tu (GTP) enters in the A site (2)
following GTP hydrolysis, Tu(GDP) is released (an exchange factor will then promote association to GTP for a new cycle) (3)
the AA-tRNA will be ideally repositioned in a stop called accommodation to allow the next step of elongation
Elongation of Translation: Step 1 FIGURE
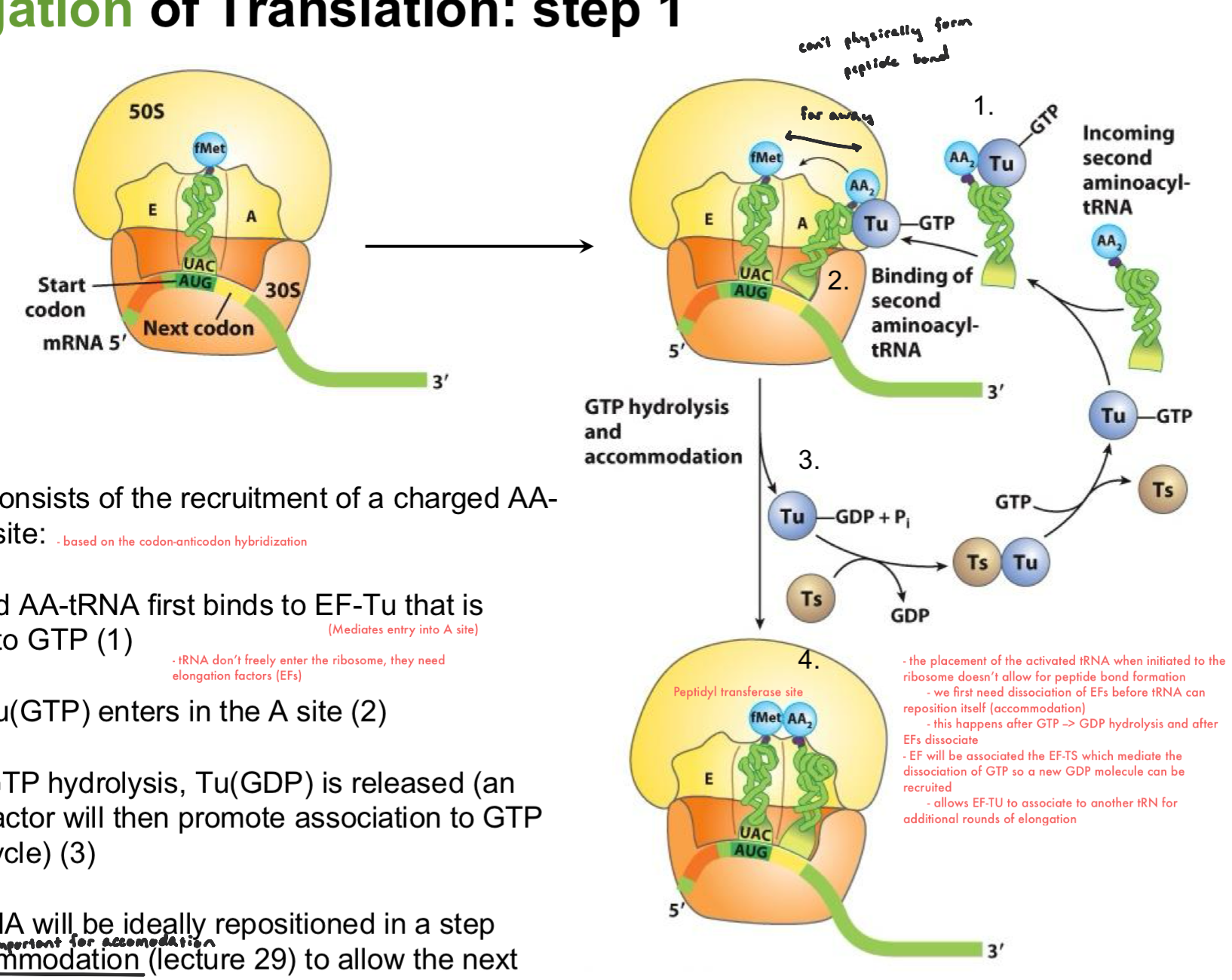
Elongation of Translation: Step 2
Peptide bond formation occurs in the second step:
the free amine group of the incoming AA (in A site) attacks the carboxyl group of the previous AA (in P site) (1)
following the formation of the transient tetrahedral, the first ester bond is cleaved and the preceding AA dissociates from the tRNA in the P-site
the amine group of the incoming AA forms a peptide bond w/ the preceding AA.
The elongated polypeptide in P-site remains associated via its carboxyl end to the incoming tRNA in the A-site.
The uncharged (deacetylated) tRNA remains associated to P-site, but shifts toward the E site
Elongation of Translation: Step 2 FIGURE
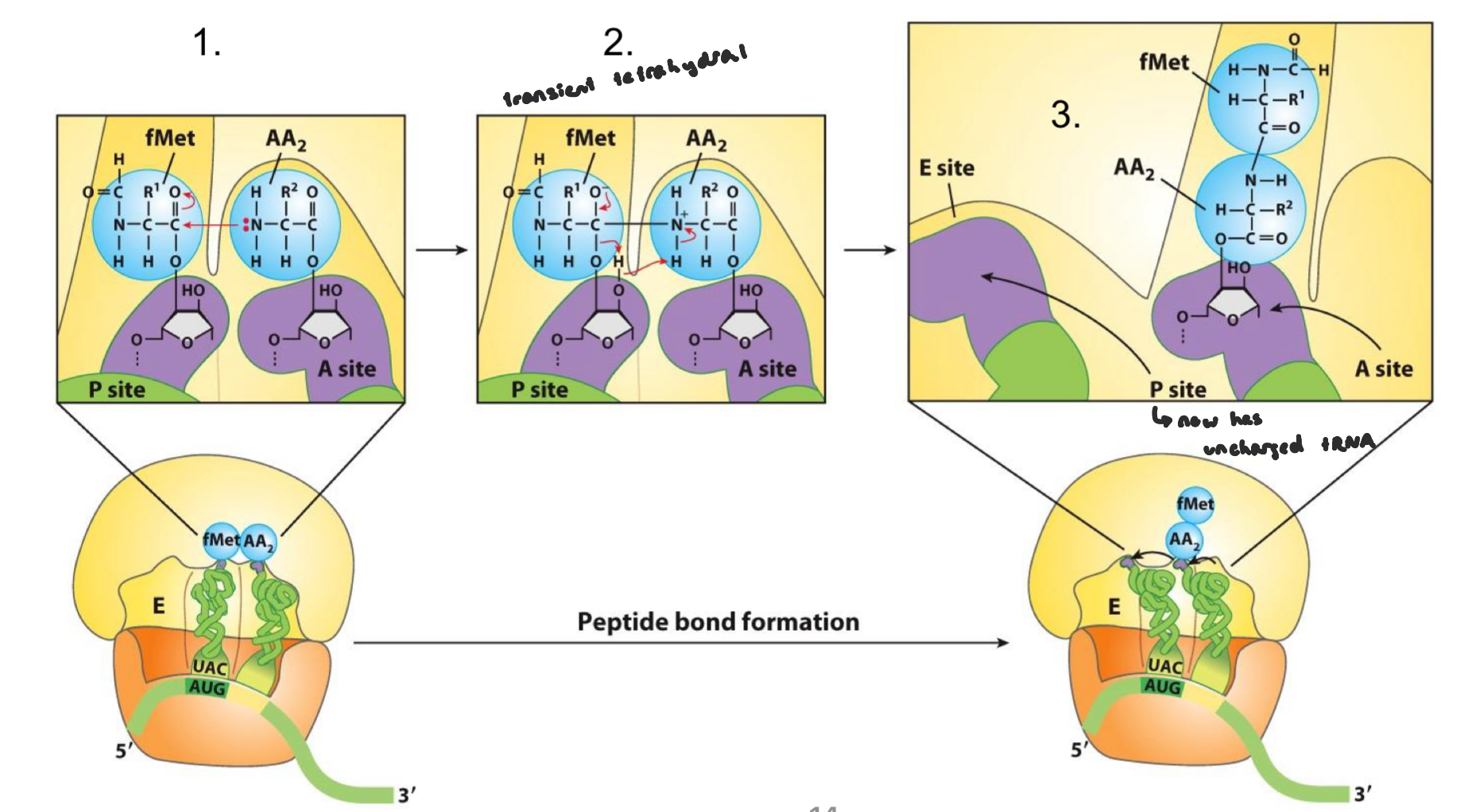
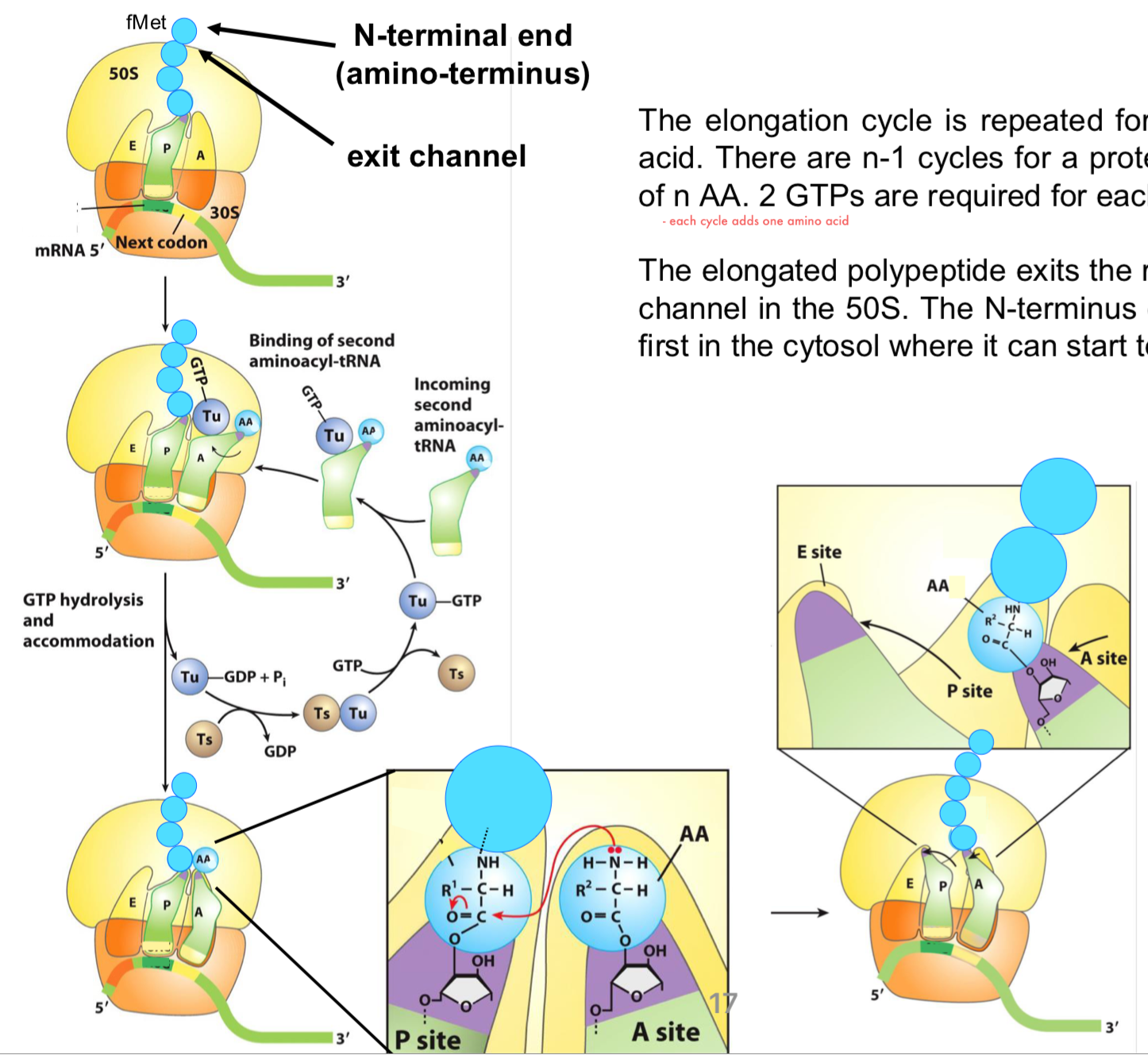
Elongation of Translation: Step 3 Part 1
translocation of the ribosome that moves toward the 3’ end of mRNA to position itself for the next round of elongation
EF-G associated to GTP binds to the A-site as its structure mimics EF-Tu-tRNA (1)
when EF-G associated in the A site, it will push the ribosome so that the first and second tRNA transition to the E and P sites
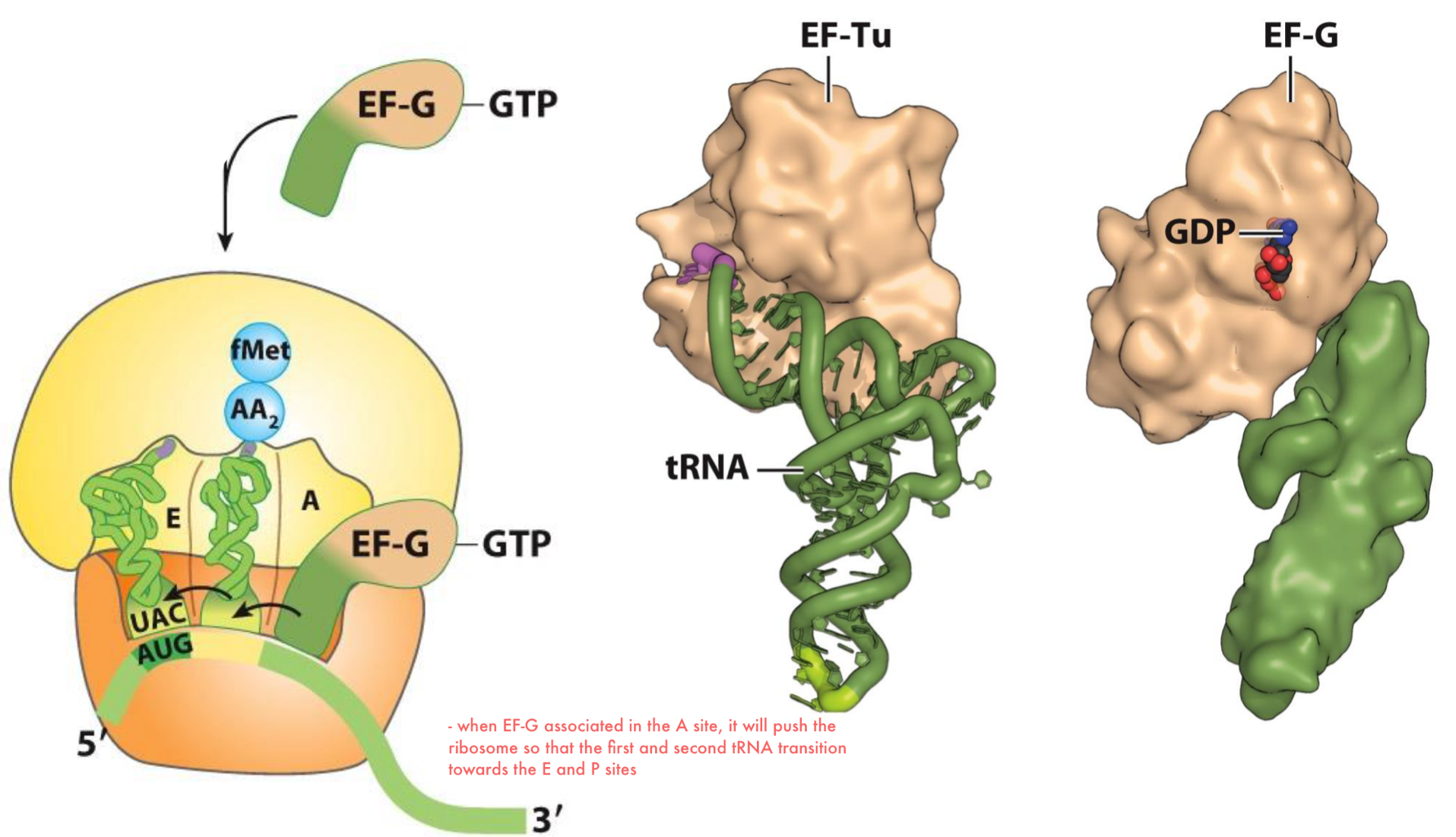
Elongation of Translation: Step 3 Part 2
following the release of EF-G (GDP), the A-site can accept the next charged AA-tRNA (2)
the uncharged tRNA dissociates from the E site (3)
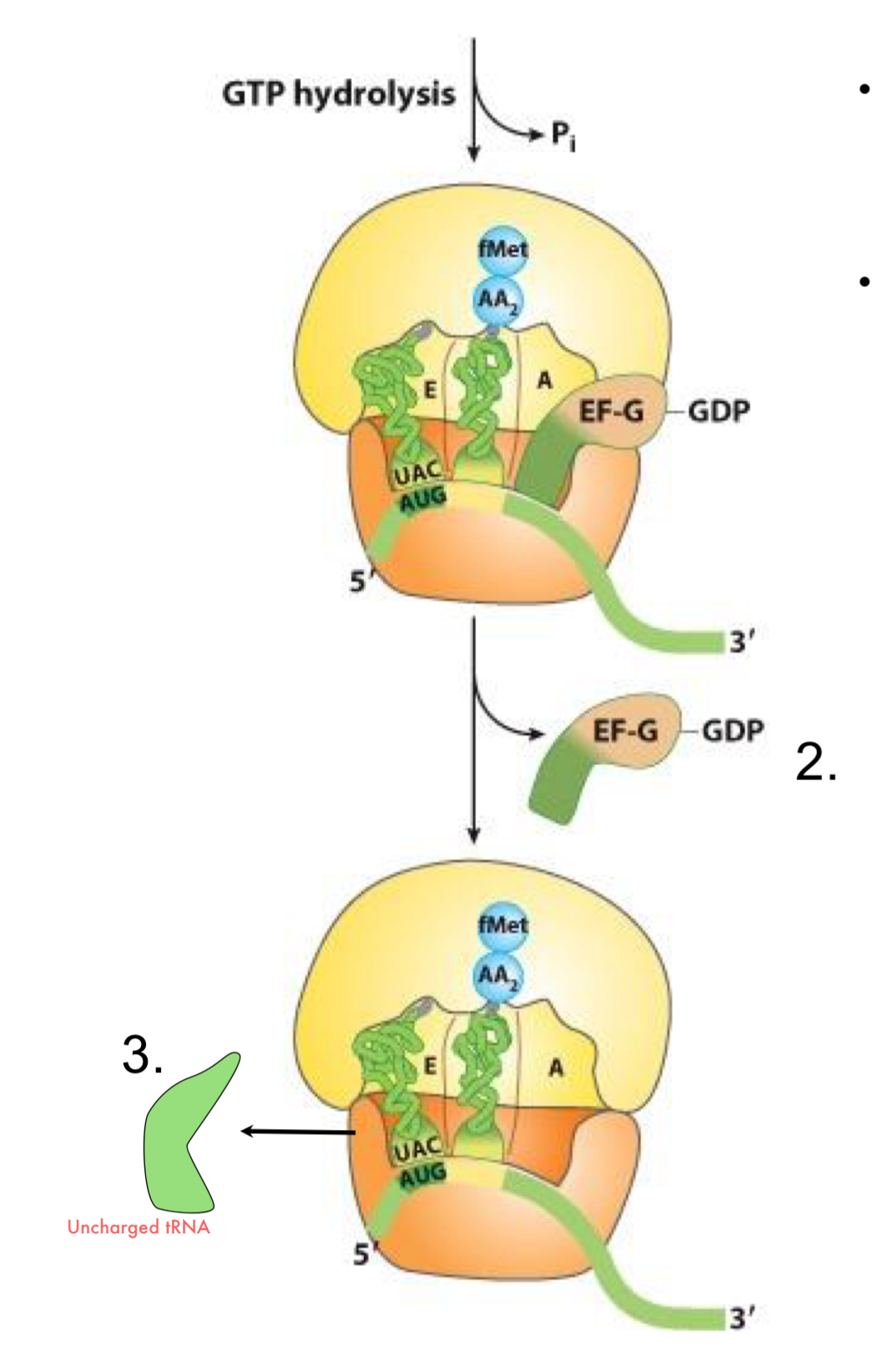
Elongation of Translation (n-1 cycles)
the elongation cycle is repeated for each added amino acid
there are n-1 cycles for a protein of a given length of n AA
2 GTPs are required for each cycle
the elongated polypeptide exits the ribosome via the exit channel in the 50S. The N-ter of the polypeptide is first in the cytosol where it can start to fold
Termination of Translation
signaled by the presence of one of three stop codons
presence of a stop codon in the A-site mediates the recruitment of a release factor (RF) that contributes to the hydrolysis of the peptidyl-tRNA bond to release the polypeptide from the ribosome and into the cytosol
the complex remains associated to mRNA util it can recruit ribosomal recycling factors IF3, EF-G and EFs bound to GTP
When the ribosome recruits ribosome recycling factor (RRF), along with IF3 and EF-G (bound to GTP), hydrolysis of one final GTP causes the ribosome to split into its 30S and 50S subunits. At the same time, the mRNA, the deacylated (uncharged) tRNA, and the associated factors dissociate
two ribosomal subunits, RF, RRF, EF-G (GDP) and the last uncharged tRNA dissociate
the recycled components can participate to other rounds of translation
IF3 stays associated to the small subunit
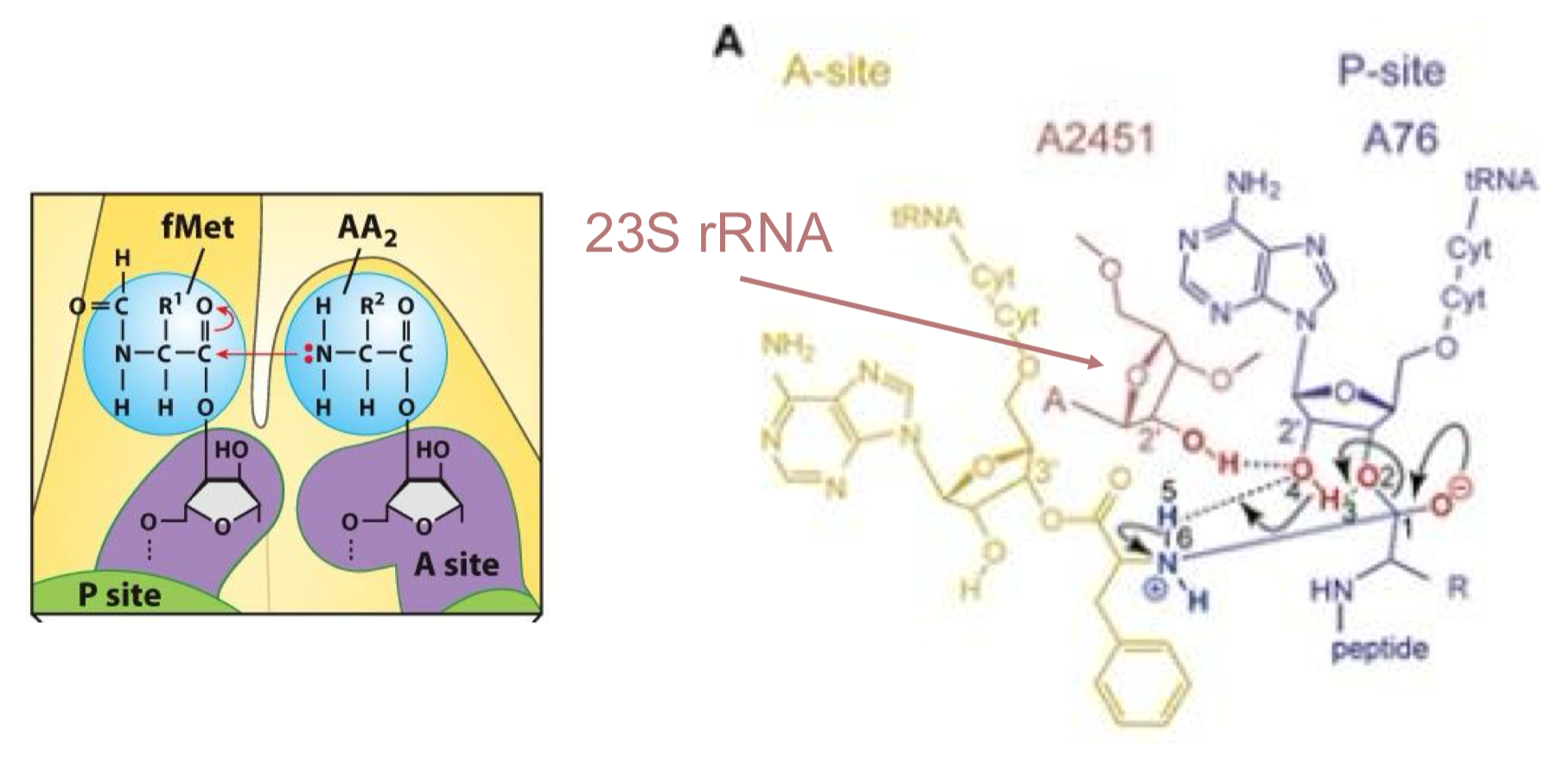
Translation is an RNA-driven Process
The ribosome is a ribozyme as the 23S rRNA (in the 50S subunit) catalyzes peptide bond formation with the help of a 2’OH, while there is no protein within 18ºC A of the active site
meaning proteins aren’t involved in catalysis
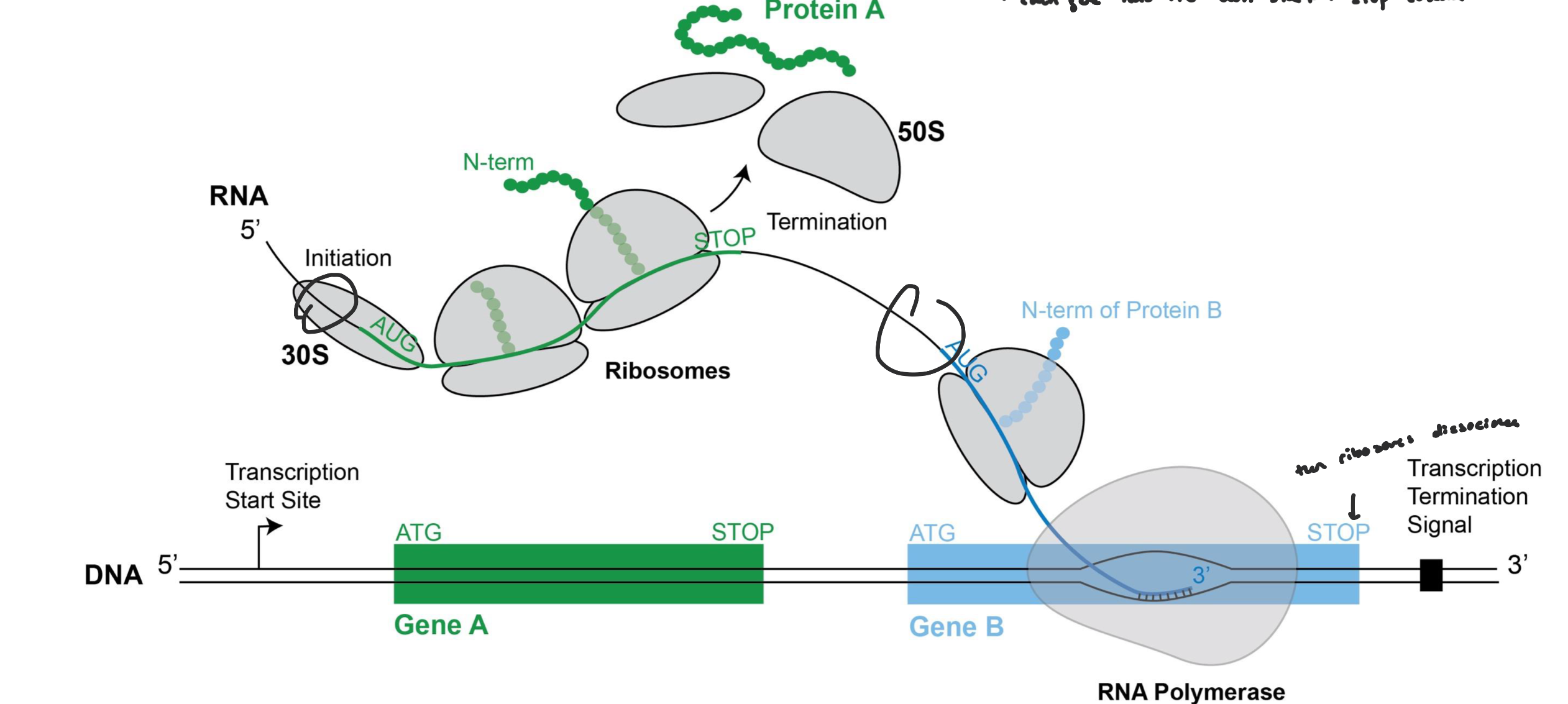
Operon
an operon is a region of DNA where multiple genes are transcribed together on the same mRNA (often genes that work in the same pathway)
that mRNA will translate multiple proteins
each protein has its own start and stop codon sites