Chemistry of Amino Acids
1/50
Earn XP
Description and Tags
L24 - Chemistry of Amino acids,L25 - Making and breaking biological outcomes
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
Why are proteins important in pharmacokinetics?
Drug metabolism is controlled by proteins
Proteins transport drugs in the body
Plasma protein binding affects distribution and half life
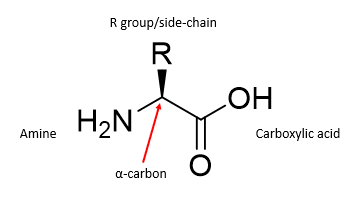
What are α-amino acids?
The monomers from which proteins are built
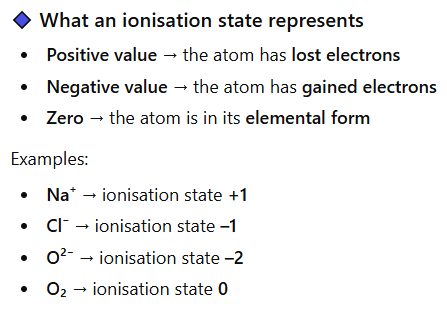
What does ionisation state mean?
The ionisation state describes the overall electrical charge a functional group or molecule has at a given pH (tells you how many e- an atom has lost, gained, or shared, compared to its neutral form).

What does pKa mean relative to 0?
The acid with the LOWER pKa is the STRONGER acid
pKa value less than 0 is a very strong acid
HCl has a pKa of -6
pKa value around 0 is a strong acid
pKa value higher than 0 is a weaker acid
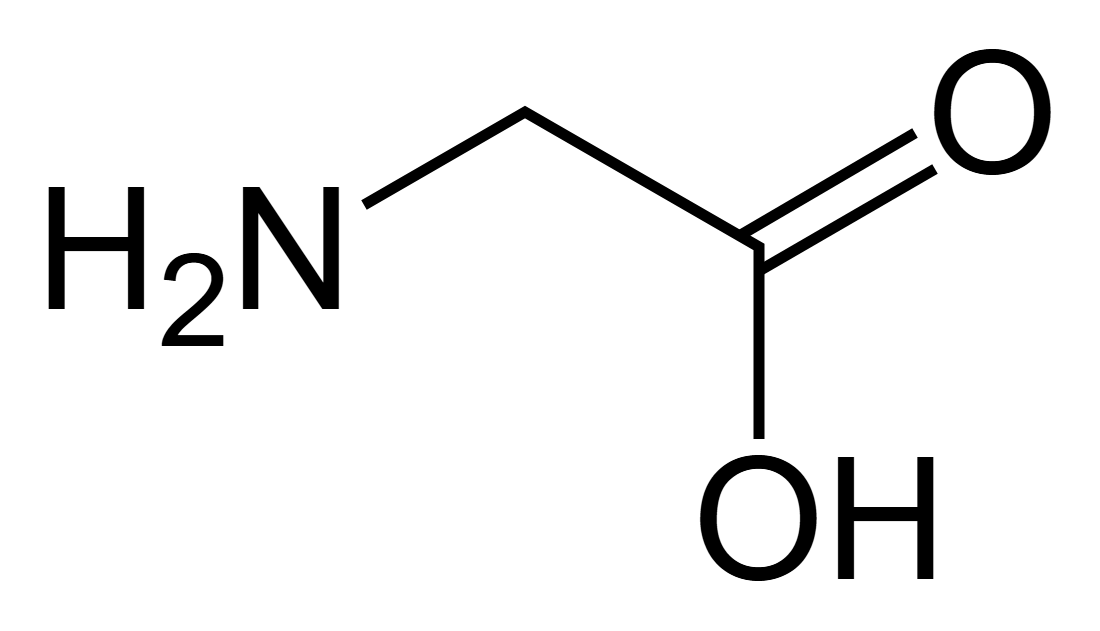
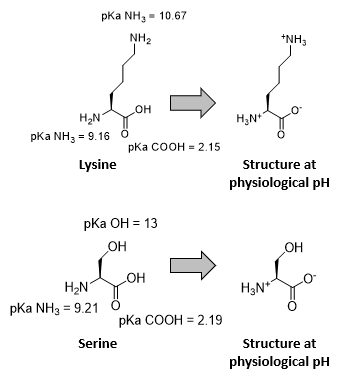
What are the ionisation states of glycine?
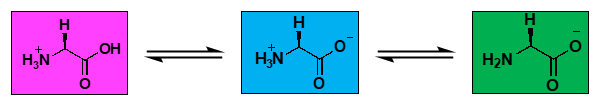
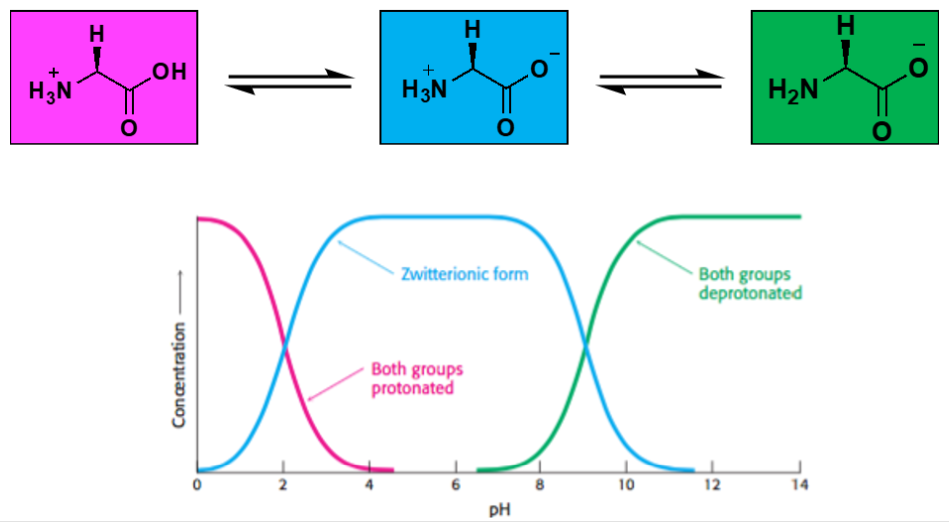
Explain this
It shows how protonation states change with pH.
Low pH: –NH₃⁺ / –COOH (net +1)
Mid pH: –NH₃⁺ / –COO⁻ (zwitterion, net 0)
High pH: –NH₂ / –COO⁻ (net –1)
Carboxyl group deprotonates first (lower pKa), amino group deprotonates last (higher pKa).
Define a zwitterion
A molecule with functional groups, of which at least one has a positive and one has a negative charge. The net charge of the molecule is zero.
What is the isoelectric point (pI)?
The pH at which a particular molecule carries no charge
What does ionisation of side chains mean?
Refers to whether the R group gains or loses H⁺ at a given pH
Each ionisable side chain has its own pKa, which determines the pH at which it gains or loses H+
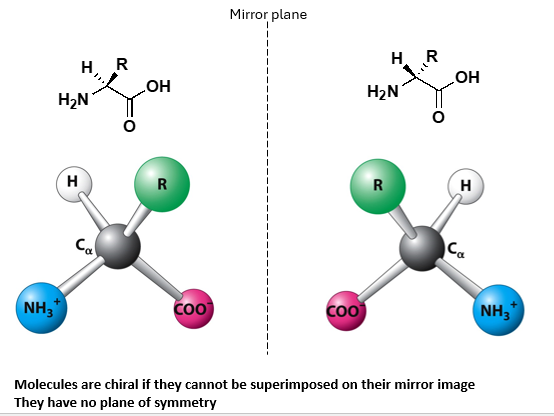
Are amino acids chiral molecules?
Yes
What is L/D nomenclature?
L/D nomenclature tells you whether the NH₂ is on the left (L) or right (D) in a Fischer projection.

What is the reference molecule for L/D nomenclature?
Glyceraldehyde
Draw L-lysine and D-lysine

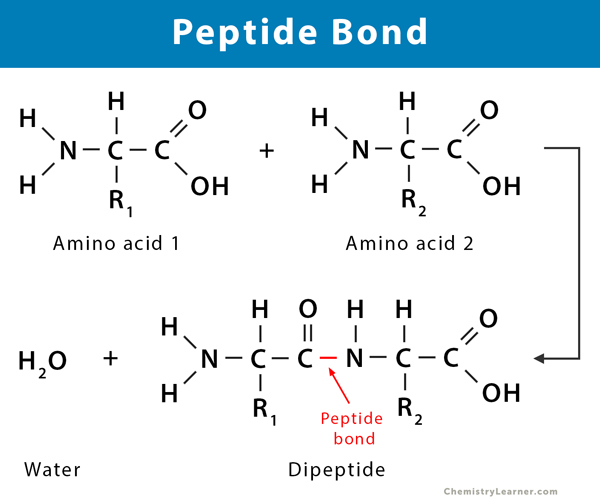
How are peptide bonds formed?
Carboxyl group of one amino acid reacts with the amino group of another amino acid
What is an amino acid unit in a peptide/polypeptide chain called?
A residue
What geometric isomers are possible for peptide bonds?
Cis / trans
Why do peptide bonds prefer trans conformation?
In the cis form, there are unfavourable steric clashes
What is hydrogen bonding?
The electrostatic attraction between a lone pair of electrons on an electronegative atom and a hydrogen atom bonded to an electronegative atom.
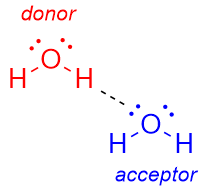
H bond donor Vs H bond acceptor
What is the hydrophobic effect in proteins?
Hydrophobic regions of a protein cluster together to minimise interactions with water.
Are proteins clustering together to minimise interactions with water an entropic effect or a formal interaction?
Entropic effect
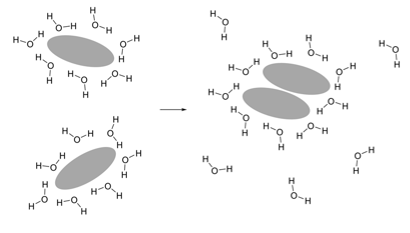
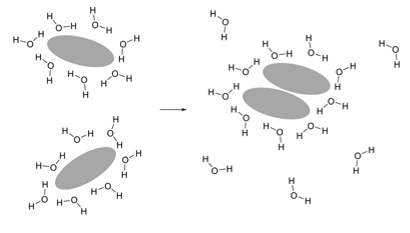
How does entropy increase with hydrophobic regions in proteins clustering together?
At first, it is ordered, has lower entropy, and this is disfavoured. There’s a limited number of conformations for water molecules to maintain hydrogen bond network
Then it is less ordered, higher entropy, and this is favoured. Reduction in size of hydrophobic water interface leads to fewer ordered water molecules
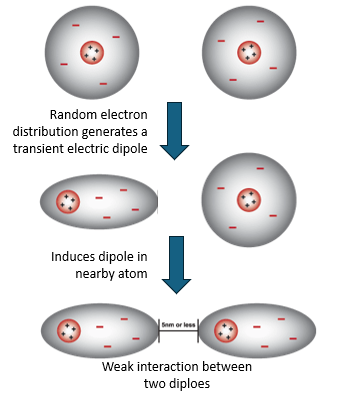
What are Van der Waals interactions?
The interaction between uncharged atoms held in proximity
Many weak interactions produce significant cumulative stability
Facilitates the dense packing of protein hydrophobic cores
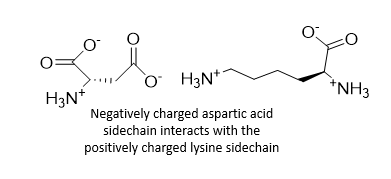
What are ionic interactions?
Oppositely charged amino acid sidechains held in proximity will form electrostatic interactions
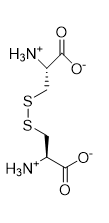
What are disulfide bonds?
Covalent bond linking together 2 cysteine residues
Not every cysteine in a protein forms a disulfide
What are Pi-interactions?
Non covalent interaction involving the electron density of aromatic rings
Phe, Tyr, and Trp residues in proteins
What type of Pi-interactions exist?
Pi stacking
Cation-Pi interactions
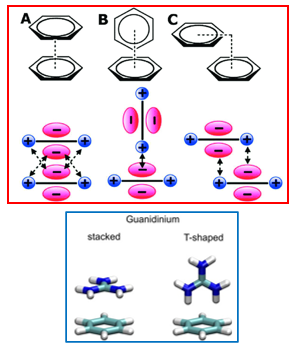
What is cation-pi interactions?
When a positively charged group interacts with the electron density of an aromatic ring
What does PTMs stand for?
Post-translational Modifications
What are PTMs?
Chemical changes which occur after a protein has been synthesised
The change in protein structure alters the function, stability, and / or cellular localisation
Are PTMs typically reversible?
Yes
What are a few examples of PTMs?
Protein function can be modulated by PTM such as:
Addition of small chemical groups like phosphorylation
Glycosylation, which is the addition of sugar molecules which can alter protein folding
How does TFEB activity and function get regulated in cells?
TFEB is a transcription factor that controls autophagy.
Its activity is regulated by cellular signalling, which determines whether it is active or inactive.
Post-translational modifications (like phosphorylation) change TFEB’s structure.
Structural changes alters where a protein is located in the cell, which affects its function.
Phosphorylation increases TFEB’s binding affinity, influencing its activity.
Protein dysfunction is a common cause of what?
disease
Other than PTMs, what else can change the physicochemical properties of proteins?
Mutations and environmental damage
What is “binding specificity” in amino acids?
A protein only binds molecules that fit its shape and match its chemical properties
How do amino acids in the binding site determine binding specificity?
The binding site has a specific shape
It also has favourable interactions (i.e. hydrogen bonds, ionic interactions, hydrophobic contacts)
Only molecules that complement these features can bind effectively.
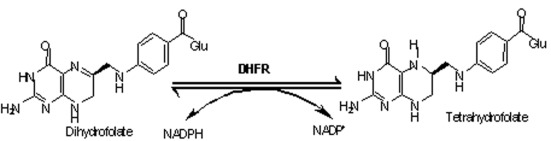
DHFR reduces dihydrofolate to tetrahydrofolate using NADPH.
How is the substrate binding stabilised?
Hydrogen bond network (enzyme–substrate–water)
π-stacking with an active-site phenylalanine
Ionic interaction between substrate carboxylate and arginine
What is protein dysfunction?
Proteins must fold correctly and maintain chemical properties to perform biological function
Protein dysfunction leads to disease, since it changes stability, structure, activity, binding, and / or abundance
What 3 ways do mutations affect proteins?
Loss of function
Gain of function
Altered expression
How can mutations lead to a loss of function?
Changes in amino acids lead to a loss of catalytic activity, reduced binding, or structural destabilisation
How can mutations lead to a gain in function?
Changes in amino acids lead to new interactions or aggregation
How do mutations lead to altered expressions?
Too much or too little protein activity alters the biochemical pathways
Name the factors of the environment that damages proteins
Oxidation
UV
Reactive metabolites
pH changes
Environmental damage (oxidation, UV, reactive metabolites, pH changes) do what to proteins?
Produces altered protein chemistry
What is the structure of haemoglobin?
Quaternary structure
4 subunits
Each subunit contains a haem group that binds oxygen
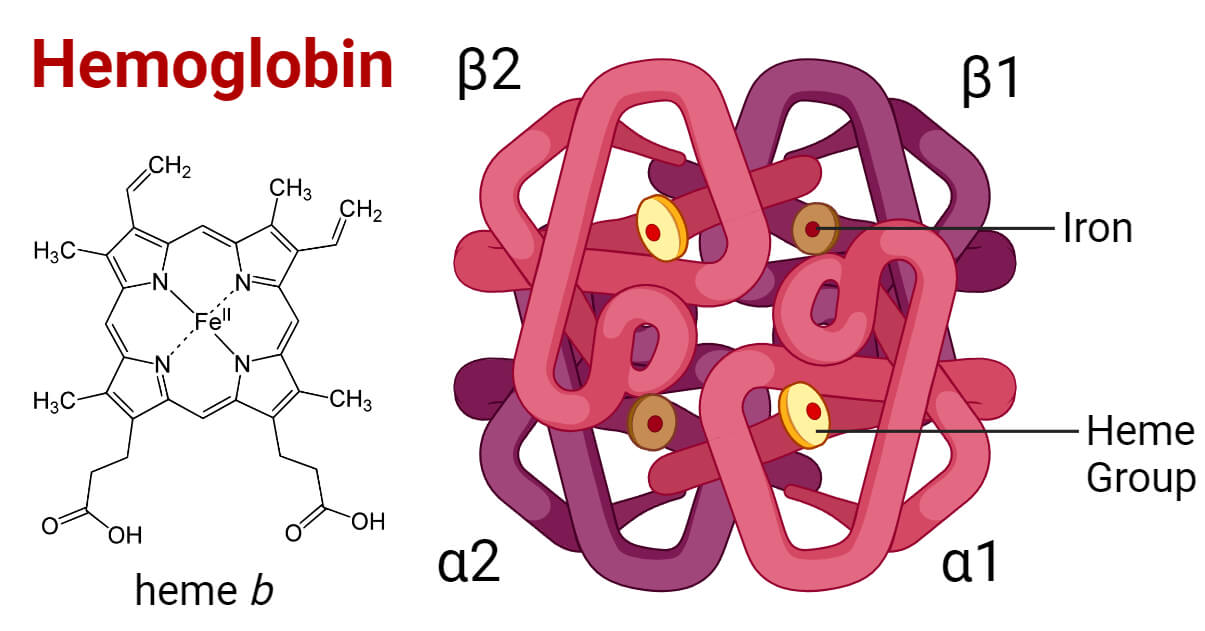
What is the function of haemoglobin?
It is an oxygen transport protein located in red blood cells
How does the mutation in sickle cell disease affect haemoglobin and red blood cells?
Sickle cell disease is caused by a single amino acid substitution, where glutamic acid is replaced by valine (E6V).
The substitution creates a hydrophobic patch on the surface of haemoglobin
The hydrophobic patch causes abnormal protein–protein interactions
As a result, haemoglobin aggregates and forms polymeric fibres
These fibres distort the red blood cells into a sickle cell shape, blocking capillaries and reducing oxygen delivery
Why do drugs act on proteins?
Drugs act on proteins in order to:
Antagonise overactive proteins
Agonise underactive proteins
Degrade overexpressed proteins
Prevent/reverse misfolding
Rebalance signalling pathways
How can somatostatin biology be used for targeted cancer therapy?
Somatostatin is an inhibitory peptide hormone that signals through specific somatostatin receptors
These receptors are overexpressed in some neuroendocrine tumours
A somatostatin-like peptide can be used to target tumour cells
When this peptide is linked to a radionuclide, it delivers localised radiation, allowing selective cancer cell killing whilst sparing the tissue
Explain how DPP-4 inhibitors improve glucose control in type 2 diabetes.
GLP-1 suppresses glucagon (this is good)
DPP-4 degrades GLP-1
Therefore, DPP-4 inhibitors prevent GLP-1 breakdown, prolonging GLP-1 action
This increases insulin release and improves glucose homeostasis
Demonstrates targeting a regulatory enzyme rather than the primary defect