Oxidative Phosphorylation
1/24
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
25 Terms
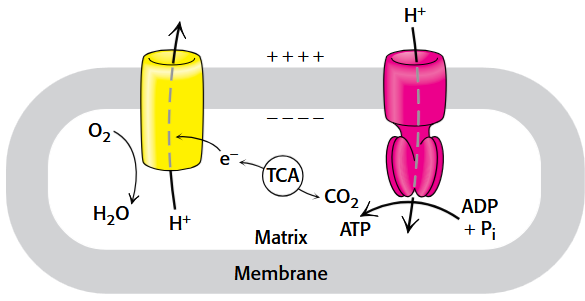
The Chemiosmotic Theory
The movement of protons down the concentration gradient of the mitochondria inner membrane (MIM) by three large protein complexes.
Energy from redox reactions or light is coupled to electron transfer that use this energy to translocate protons across the membrane.
Directional H+ pumping results in both a chemical gradient across the membrane, represented by ΔpH, and an electrical gradient due to the separation of charge, which can be measured as a membrane potential, Δψ.
ATP is produced.
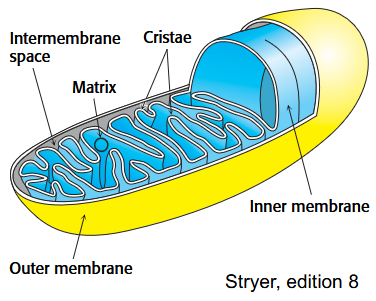
Why do mitochondria have threading inner membranes?
ratio of cell volume to mitochondrial membrane surface area
Cristae increase surface area of the inner mitochondrial membrane and create more sites for oxidative phosphorylation
estimated 14,000 m2 of inner mitochondrial membrane, which is the approximate equivalent of three football fields.
endosymbiotic theory
Once free-living bacteria, mitochondria were engulfed by eukaryotes
Outer vs Inner mitochondrial membranes
The outer membrane is permeable to most small molecules and ions (contains a transport protein called porin)
The inner membrane is impermeable to nearly all ions and polar molecules. Transporters shuttle ATP, and other metabolites across the inner mitochondrial membrane.
pH difference of mitochondria
intermembrane space = 6.8
matrix = 7.7
Δψ = 180 mV
Proton Uncoupling
uncouple redox energy from ATP synthesis → protons return to matrix via uncoupling protein/thermogenin → heat
ex. DNP, brown adipose
oligomycin
inhibit proton flow through ATP synthase complex → blocks synthase → NADH cannot be oxidised → cell death
Energy Yield of Aerobic Respiration
32 ATP per glucose molecule oxidized
redox potential in oxidative phosphorylation
the electron-transfer potential of NADH or FADH2 is converted into the phosphoryl-transfer potential of ATP
2 NADH + 2 H⁺ + 5 ADP + 5 Pi + O2 ⟶ 2 NAD⁺ + 5 ATP + 2 H2O
reducing agent
wants to donate electrons
negative reduction potential = oxidized form has lower affinity for electrons than does H2
NADH
oxidizing agent
ready to accept electrons
positive reduction potential = oxidized form of a substance has higher affinity for electrons than does H2.
O2
Standard Hydrogen Electrode
Measurement of Redox Potential with respect to
X- + H+ -> X + 1/2 H2
reduction potential of the H+ : H2 couple is defined to be 0 volts.
Standard Reduction Potentials of some reactions
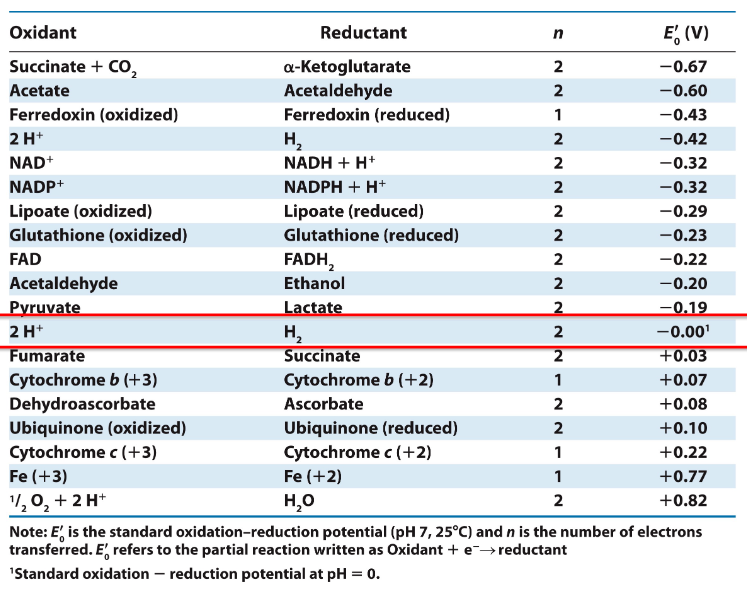
Change in Reduction Potential
∆Eo′ = Eo′(e- acceptor) – Eo′(e- donor)
Free Energy of Electron Transport
∆Go′ = –nF∆Eo′
n is the number of electrons transferred
F is the Faraday constant [96.48 kJ/mol•V (23.06 kcal/mol•V)
For negative DG need positive DE
E(acceptor) > E(donor)
reducing equivalents
1 mole of hydrogen atoms (one proton and one electron per H atom)
Flow of electrons in the Electron Transport Chain
a series of coupled oxidation–reduction (redox) reactions that transfer electrons from NADH and FADH2 to Oxygen
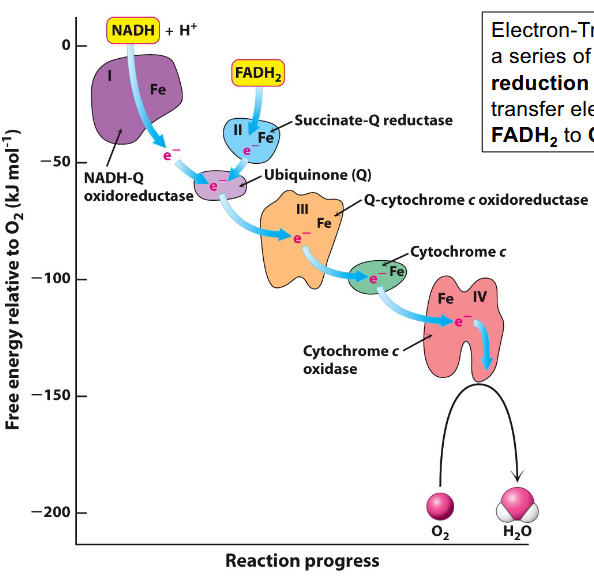
Protein Components of the Electron Transport System
Complex I NADH–ubiquinone oxidoreductase
Complex II Succinate dehydrogenase (TCA cycle)
Complex III Ubiquinone-cytochrome c oxidoreductase
Complex IV Cytochrome c oxidase
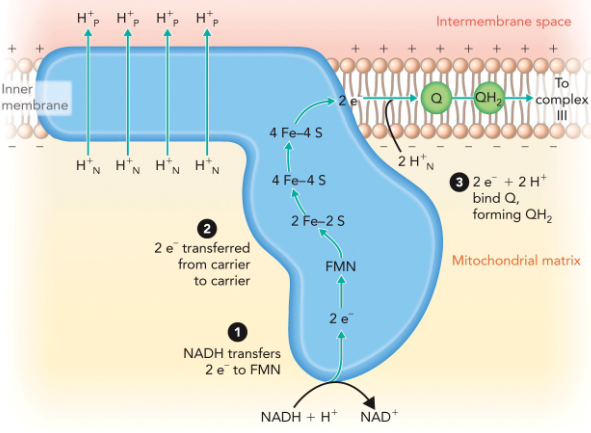
NADH–ubiquinone oxidoreductase
Oxidation of NADH in the matrix releases 2 e− (in the form of a hydride ion), which are transferred to FMN in a coupled redox reaction.
Electrons are then transferred from one carrier to another until they are donated in the last step to coenzyme Q
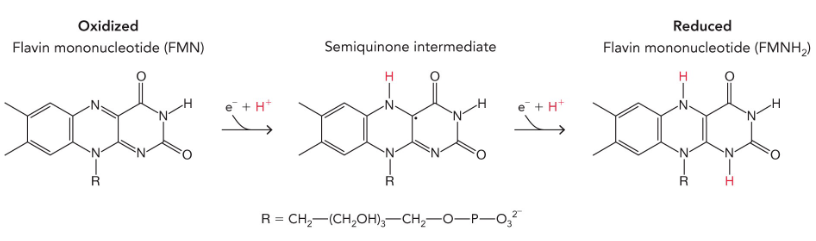
Flavin mononucleotides (FMN)
accepts one electron at a time to form semiquinone and reduced flavin mononucleotide
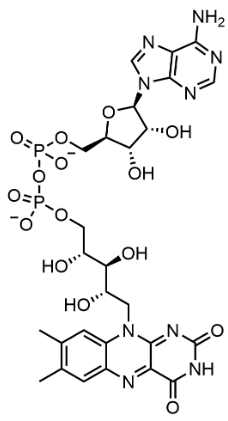
Flavin adenine dinucleotide (FAD)
adenine nucleotide (adenosine monophosphate) and the flavin mononucleotide (FMN) bridged together through their phosphate groups.
Why switch to FAD; why not just stick with NAD
FAD has a m ore positive reduction potential than NAD+ and is a stronger oxidizing agent.
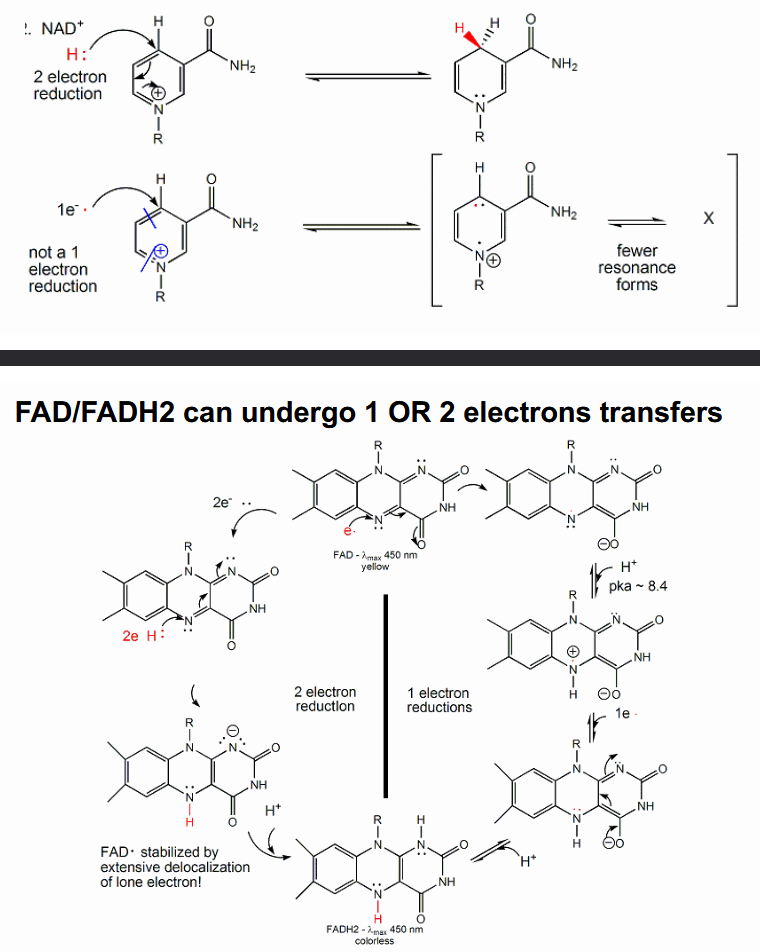
All NAD+/NADH reactions in the cell involve 2 electron hydride transfers; FAD/FADH2 can undergo 1 OR 2 electrons transfers (resonance)
Flavins can be held tightly by the enzymes so the cell is not generating a diffusible reactive species
Succinate dehydrogenase
FADH2 → CoQ
Coenzyme Q/ubiquinone Reduction
ferry the electrons from one complex to the next.
hydrophobic quinone that diffuses rapidly within the inner mitochondrial membrane.
fully reduced form holds its protons more tightly → electron-transfer reactions are coupled to proton binding and release, a property that is key to transmembrane proton transport
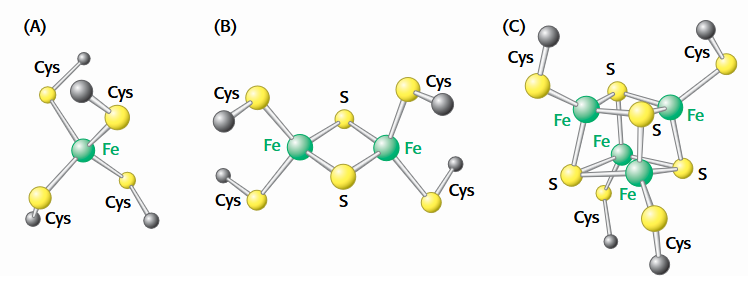
Iron-Sulfur complexes in ETC
(A) A single iron ion bound by four cysteine residues.
(B) 2Fe-2S cluster with iron ions bridged by sulfide ions.
(C) 4Fe-4S cluster.