Exam 4 Genetics Practice Problems
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
CH 4: Consider the set of crosses for hypothetical genes that control eye color and tail length in mice.
Diane is working with mice with recessive mutations for the genes that control light eye color (b) and short tail length (t). She knows that these genes display genetic linkage and are found on chromosome III.
In her work, she crosses a true‑breeding male with light eyes and a long tail (bbTT) and a true‑breeding female with dark eyes and a short tail (BBtt). She then crosses the resulting heterozygous progeny (BbTt) together in a dihybrid cross. The number of animals of each phenotype of this second cross is shown.
dark‑eyed,short‑tailed (𝐵𝐵𝑡𝑡)=22
dark‑eyed,long‑tailed (𝐵𝑏𝑇𝑡 or 𝐵𝐵𝑇𝑇)=55
light‑eyed,long‑tailed (𝑏𝑏𝑇𝑇)=26
light‑eyed,short‑tailed (𝑏𝑏𝑡𝑡)=2
What is the most likely explanation for why Diane sees light‑eyed, short‑tailed (bbtt) progeny in this cross?
Recombination
Gametes produced by independent assortment will lead to expected phenotypes where 25% of the offspring are dark‑eyed, short‑tailed (BBtt), 50% are dark‑eyed, long‑tailed (BbTt), and 25% of the offspring are light‑eyed, long‑tailed (bbTT)
In Diane's cross, she sees some progeny that are light‑eyed, short‑tailed (bbtt). These progeny are produced by recombination between homologous chromosomes.
This process generates new allele combinations along the same chromosome
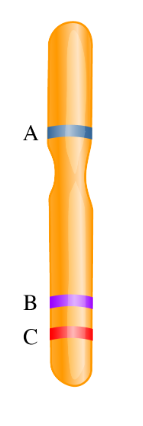

CH 4: The genetic map shows the location of three genes on a chromosome. Order the gene pairs based on their likelihood of recombining from most likely to least likely to recombine.
Most Likely: A + C, A + B, B + C: Least Likely
Genes that are located far apart on a chromosome are more likely to recombine than genes that are located in close proximity to one another
If two genes are very close together on a chromosome, the likelihood of the chromatid breaking in between those two genes is low, as compared to two genes that are far apart
CH 4: Arrange the events in chronological order for the physical process of meiotic recombination.
Prophase I of meiosis begins → Recombination is complete:
Enzymes resolve the Holliday junction
The 3’ overhang strand of DNA invades the double helix of the homologous chromosome
The duplexed strand of DNA serves as a primer for the synthesis of new DNA
Enzymes break both strands of DNA in one chromosome
Homologous chromosomes pair together
Enzymes degrade the 5’ strand of DNA
Homologous chromosomes pair together
Enzymes break both strands of DNA in one chromosome
Enzymes degrade the 5’ strand of DNA
The 3’ overhang strand of DNA invades the double helix of the homologous chromosome
The duplexed strand of DNA serves as a primer for the synthesis of new DNA
Enzymes resolve the Holliday junction
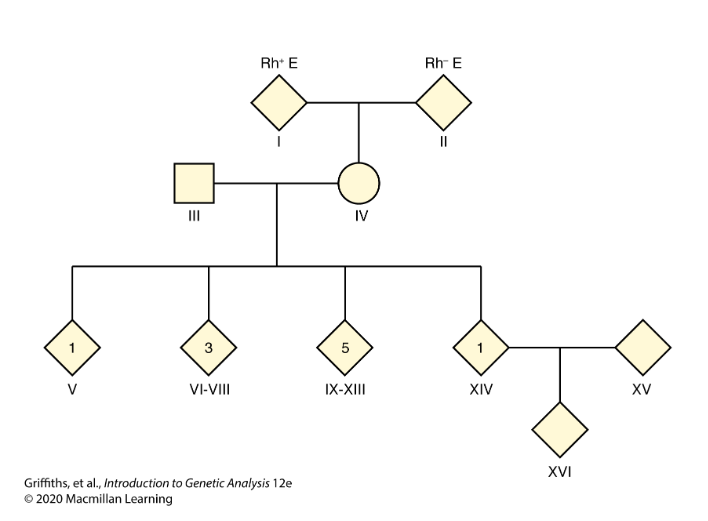
CH 4: The mother of a family with 10 children has blood type Rh+. She also has a very rare condition known as elliptocytosis (E) that causes red blood cells to be oval rather than round in shape, but that produces no adverse clinical effects. The father is Rh−, meaning that he lacks the Rh+ antigen. He has normal red blood cells (e).
One child is Rh+e, four children are Rh+E, and five children are Rh−e. The mother’s parents are Rh+E and Rh−e.
One of the four children expressing Rh+E marries someone who is Rh+e, and they have an Rh+E child.
Is the pedigree in agreement with the hypothesis that the Rh+ allele is dominant and the Rh− allele is recessive?
Yes, the pedigree supports the hypothesis
Mother is Rh+, but she has Rh− children → she must be Rr (carrier)
Father is Rh− → must be rr
So you expect both Rh+ and Rh− children, which matches:
5 Rh− children
5 Rh+ children (1 Rh+e + 4 Rh+E)
Rh− individuals only appear when both alleles are recessive (rr)
Rh+ appears even when only one dominant allele is present (Rr)
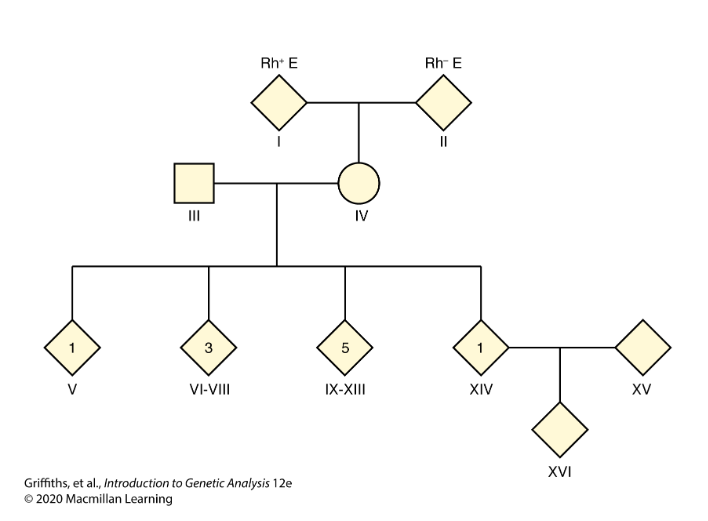
CH 4: The mother of a family with 10 children has blood type Rh+. She also has a very rare condition known as elliptocytosis (E) that causes red blood cells to be oval rather than round in shape, but that produces no adverse clinical effects. The father is Rh−, meaning that he lacks the Rh+ antigen. He has normal red blood cells (e).
One child is Rh+e, four children are Rh+E, and five children are Rh−e. The mother’s parents are Rh+E and Rh−e.
One of the four children expressing Rh+E marries someone who is Rh+e, and they have an Rh+E child.
What is the mechanism of transmission of elliptocytosis?
Dominant inheritance
Trait appears in every generation (no skipping)
The affected person usually has an affected parent
Males and females are affected equally
About half the children of an affected × normal parent show the trait
The trait shows even when only one copy of the allele (E) is present
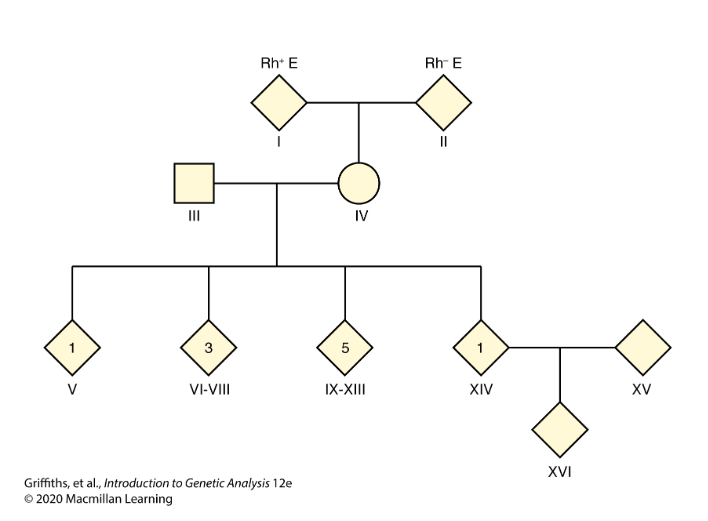
CH 4: The mother of a family with 10 children has blood type Rh+. She also has a very rare condition known as elliptocytosis (E) that causes red blood cells to be oval rather than round in shape, but that produces no adverse clinical effects. The father is Rh−, meaning that he lacks the Rh+ antigen. He has normal red blood cells (e).
One child is Rh+e, four children are Rh+E, and five children are Rh−e. The mother’s parents are Rh+E and Rh−e.
One of the four children expressing Rh+E marries someone who is Rh+e, and they have an Rh+E child.
Could the genes governing the E and Rh phenotypes be on the same chromosome? If so, estimate the map distance between them.
Yes, the Rh+e child represents a recombinant if the genes are linked. The distance between loci is 10 map units.
You are told:
4 children = Rh+E → parental type
5 children = Rh−e → parental type
1 child = Rh+e → this is unusual (recombinant)
Calculate RF:
Recombination frequency = 1/10 = 0.10 = 10%
CH 4: A plant of genotype 𝐴𝐵/𝑎𝑏 is testcrossed with 𝑎𝑏/𝑎𝑏. If the two loci are 10 map units apart, what percent of progeny are expected to inherit the genotype 𝐴𝐵/𝑎𝑏?
45% of the progeny are expected to be AB/ab
Parental 90% total
Recombinant 10% total
% of progeny = 90/2 = 45%
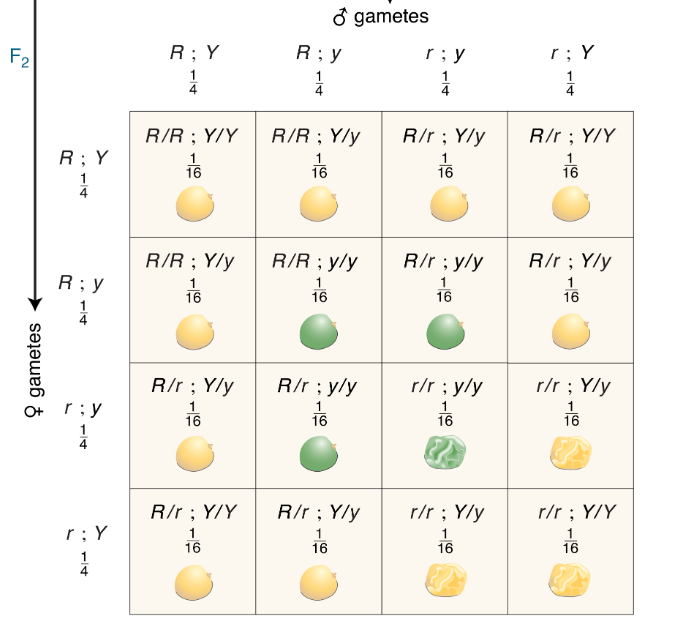
CH 3: How many different genotypes are shown in the 16 squares of the grid?
9
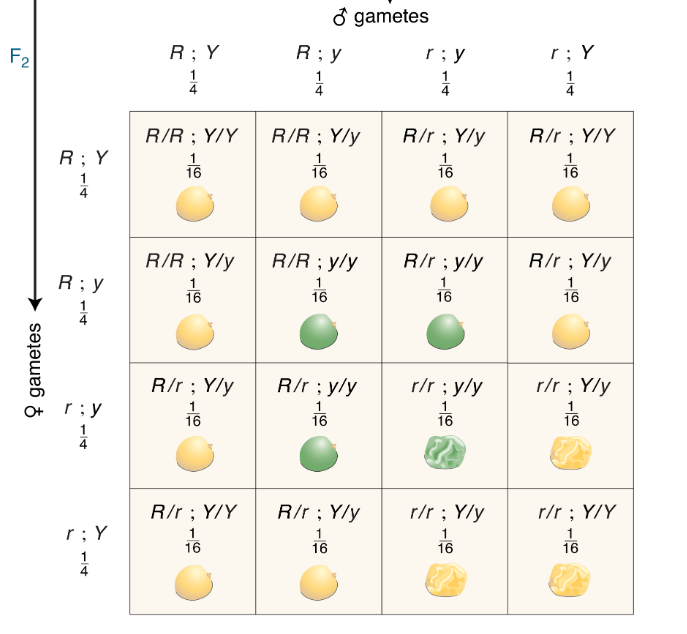
CH 3: Can you devise a simple formula for the calculation of the number of progeny genotypes in dihybrid, trihybrid, and so forth crosses? Use 𝑛 for the number of genes.
3n
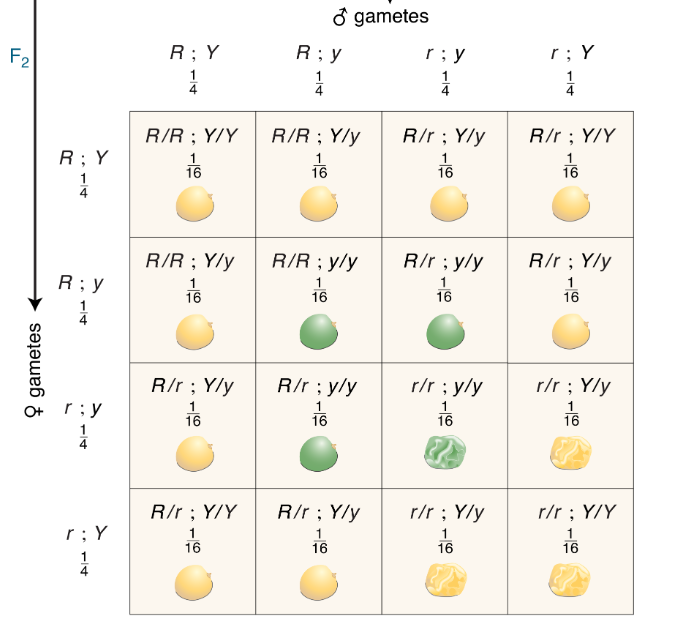
CH 3: Can you devise a simple formula for the calculation of the number of progeny phenotypes in dihybrid, trihybrid, and so forth crosses? Again, use 𝑛 for the number of genes.
2n
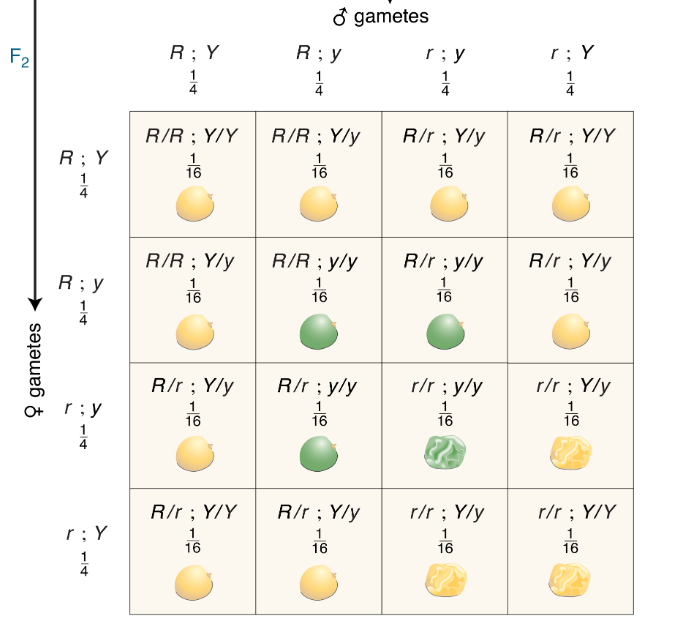
CH 3: Mendel predicted that, within all but one of the phenotypic classes in the Punnett square, there should be several different genotypes. In particular, he performed many crosses to identify the underlying genotypes of the round, yellow phenotype.
With a testcross, complete heterozygosity will yield a 1:1:1:1 phenotypic ratio. Homozygosity at one locus will yield a 1:1 phenotypic ratio, while homozygosity at both loci will yield only one phenotypic class.
With selfing, complete heterozygosity will yield a 9:3:3:1 phenotypic ratio. Homozygosity at one locus will yield a 3:1 phenotypic ratio, while homozygosity at both loci will yield only one phenotypic class.
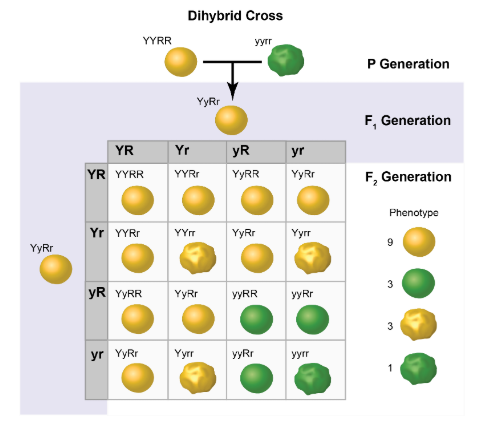
CH 3: During his experiments with pea plants, Gregor Mendel crossed plants that were heterozygous for two traits with one another in order to determine laws of inheritance. The Punnett square shows the results of this cross.
Allele pairs do not influence one another during segregation for gamete formation.
A dihybrid cross between heterozygous individuals will result in exactly nine genotypes.
CH 3: In pea plants, the genes for seed color and seed shape are located on different chromosomes and assort independently of one another. Consider that Y=yellow, y=green, R=round, and r=wrinkled. According to the law of independent assortment, what are all the possible phenotypes for offspring of a self‑pollinated pea plant with a genotype of YyRr?
Yellow round seeds, yellow wrinkled seeds, green round seeds, and green wrinkled seeds
CH 3: Suppose Pablo breeds flowers and wants to optimize production of offspring with both short stems and white flowers, which are coded for by two genes with the recessive alleles t and p, respectively. In flowers, T codes for tall stems and P codes for purple flowers. Pablo crosses two heterozygotes that produce 656 offspring.
How many of these 656 offspring are predicted to have both short stems and white flowers?
1/16 ttpp → 656(1/16) = 41
CH 3: Suppose a diploid slime mold is completely heterozygous at all 12 of its chromosomes (2𝑛=12). How many different combinations of gametes can be produced by this slime mold, assuming no homologous recombination between chromosomes?
26 = 64
CH 3: Suppose a person is heterozygous for heterochromia, an autosomal dominant condition that causes two different‑colored eyes in an individual, produced 25 offspring with a person who does not have heterochromia. Of their children, 16 had heterochromia and 9 did not. Calculate the chi‑square value for this observation.
X2= ∑ (O-E)2/E
Expected heterochromatic offspring = 25 × 0.50 = 12.5
Heterochromatic: 𝑂−𝐸 = 16 − 12.5 = 3.52 = 12.25
Normal: 𝑂−𝐸 = 9 − 12.5 = −3.52 = 12.25
Heterochromatic: 12.25/12.5 = 0.98
X2= ∑ 0.98 + 0.98 = 1.96
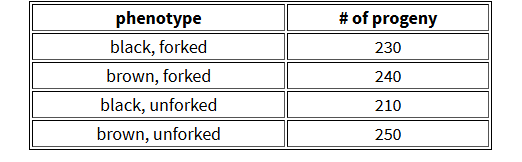
CH 3: A presumed dihybrid in Drosophila, B/b;F/f, is testcrossed with b/b;f/f (B=black body; b=brown body; F=forked bristles; f=unforked bristles). The results of the cross are outlined in the table. Assume that the genes assort independently.
X2= ∑ (O-E)2/E
230 + 240 + 210 + 250 / 4 = 233
(230 − 233)2 + (240 − 233)2 + (210 − 233)2 + (250 − 233)2 / 233 = 3.75
CH 18: In humans, red blood cells have a number of proteins embedded in the cell membrane. One type of protein, the Rh factor, is controlled by a single gene and is either present or missing from the red blood cells. If present, the individual has the Rh+ phenotype. If missing, the individual has the Rh− phenotype. Rh+ is the dominant to Rh−. Suppose that, in the Welsh population, the frequency of the Rh− phenotype is 0.04.
Using the Hardy–Weinberg equations, calculate the frequency of the Rh+ allele to at least two decimal places.
p2 + 2pq + q2 = 1
p = 0.004 → √0.04 = 0.2
1 - 0.2 = 0.8
You are studying three genes: the K gene, the P gene, and the R gene. If they were written with the following nomenclature, which of these conclusions would be true?
R/r ; KP/kp
The K and P genes are encoded on different chromosomes (;)
The product rule states that the probability of independent events occurring together
Is calculated by multiplying their individual probabilities
We are growing a new crop plant with high yield in our greenhouse. You propose an experiment to create a more sustainable plant combining our high-yielding line and a stress-tolerant variety from Australia. Of the four traits you are pursuing (good flavor, high yield, drought tolerance, pathogen resistance), each parent line has some of the desired traits:
UMSL Parent: F/f ; Y/Y; dr/dr; RES/res Australian Parent: F/f; y/y; DR/dr; RES/res
You want to combine these traits into a single line. If you crossed these two lines and grew the progeny in our space, how many would you expect that were f/f ; Y/- ; DR/- ; RES/ - ?
f/f = 1/4
Y/- = 1 (since Y/Y × y/y → all Yy)
DR/- = 1/2
RES/- = 3/4
Multiply:
¼ x 1 × ½ x ¾ = 3/32
In a testcross for two traits that show independent assortment, the recombination frequency (RF) will be:
About 50%
A dihybrid cross occurs between a true breeding red, long-stamen plant with a true breeding, white, short-stamen plant. This F1 plant is self-fertilized to produce an F2 population that has 300 red, long-stamen plants and 100 white, short-stamen plants. What can you conclude?
(300:100) → 3:1 Independent assortment is occuring
The smaller the distance between linked genes, the _____ the chance of crossovers in the region between the genes, and the _____ the proportion of recombinants that would be produced.
lower; lower
Which of the following statements about epistasis are true:
Epistasis is the genetic term to describe when the allele of one gene eliminates or masks the expression of the allele of another gene.
Epistasis can be seen when two genes act in the same pathway, for instance a biochemical pathway or developmental pathway.
Epistasis can be quantified as a 9:3:3:1 genotypic ratio for a dihybrid trait.
Choices a AND b are both correct.
Choices a AND c are both correct.
Choices a AND b are both correct
A testcross is done with a heterozygous F1 individual to examine the possible linkage between the N gene (smooth leaves vs notched leaves) and the L gene (green leaves vs. purple leaves):
Parent 1: N/N ∙ l/l with smooth purple leaves
Parent 2: n/n ∙ L/L with notched green leaves
F1 testcross: N/n ∙ L/l x n/n ∙ l/l
The progeny show:
39 smooth green leaves
58 smooth purple leaves
55 notched green leaves
40 notched purple leaves
Determine the recombination frequency for these two genes. Based on this support, would you conclude these traits are linked or not linked?
Analyze the data using the χ2 test (an unmarked chart would be provided). Be sure to include your hypothesis and whether you would accept or reject the hypothesis. Based on this support, would you conclude these traits are linked or not linked?
39 + 58 + 55 + 40 = 192 total
Parentals (highest):
58 smooth purple
55 notched green → 113
Recombinants:
39 + 40 = 79
RF:
79/192 = 41 → Linked
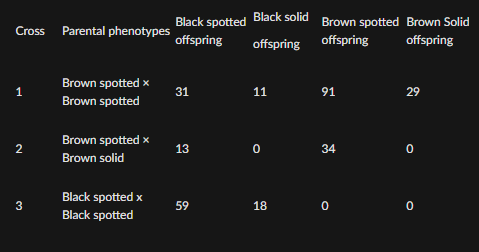
In alpacas,
brown fur coloration is dominant over black
spotted fur is dominant over solid coloration
Assume that these effects are caused by two independently assorting genes. For each cross below, determine the genotypes of the parents. Use the symbols F and f for the brown and black fur-color alleles and the symbols S and s for the spotted and solid alleles.
Cross 1: Brown spotted × Brown spotted
Offspring:
all four phenotypes appear:
black spotted
black solid
brown spotted
brown solid
To get the black offspring (ff)
BOTH parents must carry f
To get solid (ss) offspring:
BOTH parents must carry s
So both parents must be heterozygous for both genes → FfSs × FfSs
Cross 2: Brown spotted × Brown solid
Offspring:
Black spotted = 13
Black solid = 0
Brown spotted = 34
Brown solid = 0
NO ss offspring at all
Parents → FfSs × Ffss
Cross 3: Black spotted × Black spotted
Offspring:
black spotted = 59
black solid = 18
brown spotted = 0
brown solid = 0
Key observations:
NO brown offspring → no F allele present
→ both parents must be ff
spotted : solid ≈ 3:1 (59:18 ≈ 3:1)
So both parents are heterozygous for S gene
Parents → ffSs × ffSs
Another scientist studying alpaca traits comments that the gene influencing coarse hair (C) shows a recombination frequency of 12% with the F gene that influences fur coloration.
What does this finding reveal about these two genes in terms of their physical relationship?
What does this finding suggest in terms of their genetic inheritance?
How was the recombination frequency determined?
Genes are linked on the same chromosomes and close together
Shows Genetic linkage and will be inherited together.
RF = (recombinant offspring ÷ total offspring) × 100

In a jungle plant, the hypothesized pigmentation pathway for leaves is shown below. The first reaction is catalyzed by an enzyme encoded by the YEL gene and the second reaction is catalyzed by an enzyme encoded by the GRE gene.
Two mutants were identified:
A pale mutant with no coloration, hypothesized to be defective in step 1
A yellow mutant, hypothesized to be defective in step 2
If the two genes are independently assorting, how would you write the genotype for these two genes in a wild-type line?
Write the genotype for these two genes in each mutant background. Because these single mutants were selected in the lab, each mutant is known to be homozygous dominant for the other gene.
You cross the pale mutant with the yellow mutant. Describe the genotype and phenotype of the F1 plant.
Define the resulting genotypes if you cross two F1 individuals. For full credit, be sure to specifically match the ratio with the corresponding genotypes.
Define the resulting phenotypes if you cross two F1 individuals. For full credit, be sure to specifically match the ratio with the corresponding phenotypes.
Wild type: YEL/YEL ; GRE/GRE (both functional enzymes)
Genotype for these two genes in each mutant:
Pale mutant (step 1 broken):
yel/yel ; GRE/GRE
Yellow mutant (step 2 broken):
YEL/YEL ; gre/gre
yel/yel ; GRE/GRE × YEL/YEL ; gre/gre → YEL/yel ; GRE/gre → phenotype: wild-type green
F2 (F1 × F1 genotypes):
Classic dihybrid ratio:
9 YEL_ GRE_ (green)
3 YEL_ gre/gre (yellow)
3 yel/yel GRE_ (pale intermediate block step 1)
1 yel/yel gre/gre (no pigment)
F2 phenotypes:
9 green
3 yellow
3 pale
1 colorless