Genetics Final Exam Objectives
1/106
Earn XP
Description and Tags
Exam on 4/28/2026! Covers lectures 1-19 :)
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
107 Terms
(16) Understand basic features of bacteria as well as advantages for using them in genetic studies
Features
Unicellular microbes
Shapes: spheres, rods, spirals
Found throughout human body (1:1)
Replication of genome and division by binary fission
Lab Advantages
Rapid reproduction
Haploid genome allows all mutations to be expressed directly
Small genomes
(16) Know the difference between auxotrophic and prototrophic bacteria and minimal versus complete media
Prototrophic bacteria: (wild-type) can synthesize essential molecules for growth and replication
Auxotrophic bacteria: (mutant) cannot synthesize essential molecules, requires molecules be added to media
Minimal media: Only required by prototrophic bacteria
Complete media: Contain all substances required by all bacteria, including auxotrophic bacteria
(16) Describe the techniques: bacterial culture in liquid broth, spread plate, streak to isolation, & replica plating
Spread plate: Bacteria grown in liquid media and spread onto agar plate with triangle
Streak to isolation: Bacteria grown in liquid media and spread on agar plate with loop, separating individual colonies
Replica plating: Using a primary plate of bacteria to print secondary plates on media that selects for various phenotypes (leu-)
(16) Know the structure of a plasmid and it’s three main components
Plasmids: Small circular DNA molecules capable of replicating independently of bacterial chromosome
They contain: ORI (Origin of replication), Selection marker (ex. antibiotic resistance), and Cloning site (synthesized)
(16) Explain E. coli F factor, its role in exchanging DNA between bacteria
F factor = Fertility factor. Controls mating and exchange between E. coli
F factor contains and ORI, genes for replicaion (green), genes for transfer (blue), and insertion sequences (IS2, IS3)
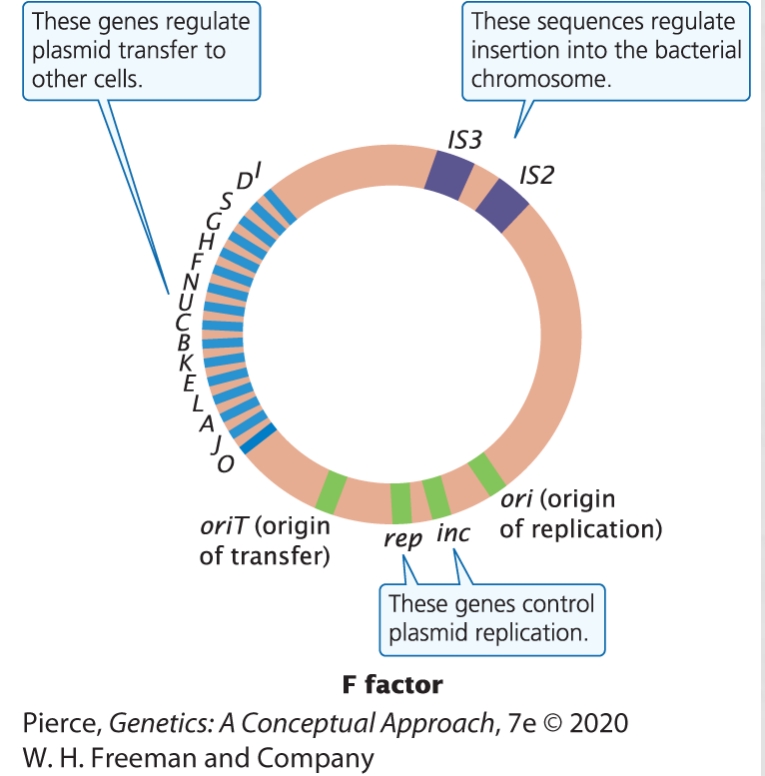
(16) Define episome
Freely replicating plasmids that integrate into host chromosome
(16) Describe the experiment Lederberg and Tatum designed and their conclusions
Demonstrated that bacteria exchange genetic material through conjugation
Two auxotrophic E. coli strains were combined and plated, and new strains could finally grow in minimal media
(16) Define plasmid
Small circular DNA molecules capable of replicating independently of bacterial chromosome
(16) Define conjugation
Direct transfer of DNA from one bacterium to another
(16) Define Transformation
Bacterium takes up free DNA
(16) Transduction
Bacterial viruses take DNA from one bacterium to another
(16) Explain the process of conjugation, between F+ & F- cells, Hfr & F- cells
Conjugation depends on a donor cell having a F factor.
The cell is not F+ until the entire F factor is transferred. F factor moves through sex-pilus
Hfr cell: a cell in which f-factor has been integrated into the bacterial chromosome. If f-facor excises from Hfr chromosome, it is called F’ (F prime)
Process is called sexduction: producing partial diploids called merozygotes = cells with 2 copies of some genes
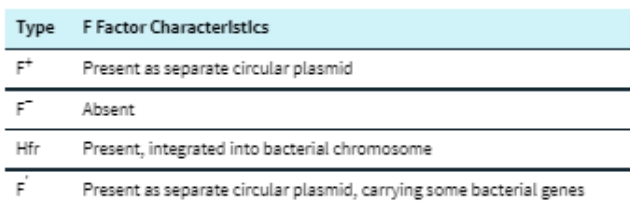

(16) Characteristics of E. coli cells with different types of F-factor
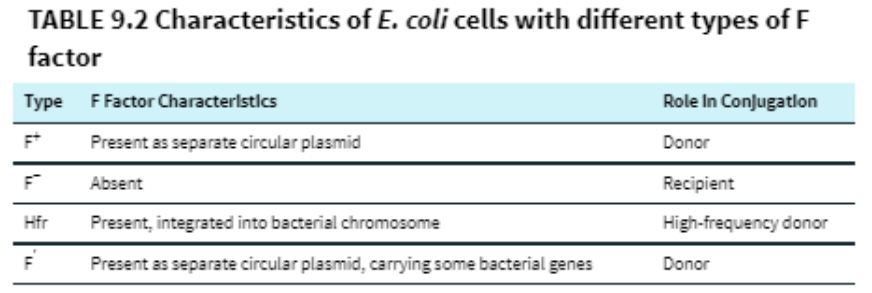


(16) Results of conjugation between E. coli cells with different F factor
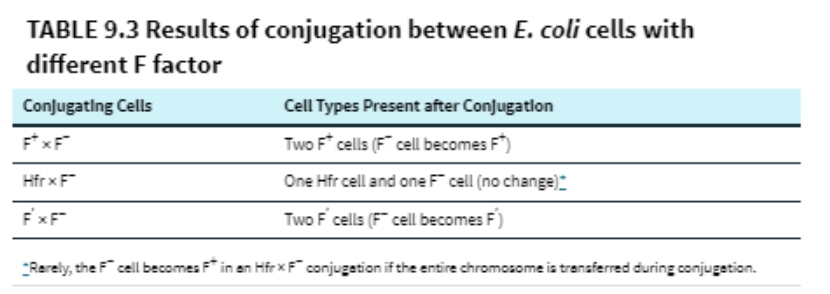
(16) Describe how conjugation can be used to map bacterial genes
Time required for certain genes to be transferred tells about their location in reference to F factor. As more time passes, genes that enter the F- cell are father from the F-factor at the 5’ end.
Unit of distance: Minute
Conjugation can be used to map bacterial genes by mixing Hfr and F- cells and interrupting conjugation at many intervals
(16) Explain how DNA is obtained by transduction and transformation
Transformation: occurs when cells take up DNA from the medium and recombination integrates DNA into the chromosome. Cells that can take up DNA are competent.
Transduction: Bacterial viruses carries DNA from one bacteria to another. Usually occurs between bacteria of same or closely related species
(16) Virus
A particle consisting of RNA or DNA and protein that must infect a living cell to reproduce
(16) Bacteriophage
A virus that specifically infects bacteria. Virulent phages kill host by cell lysis
(16) Describe the difference between the lytic or lysogenic cycle of a phage or virus
Lytic: Virus replicates —> cell bursts
Lysogenic: Viral DNA integrates into host genome (dormant)
(16) Describe the three major components of a virus and its basic life cycle
Three major components
Nucleocapsid (nucleic acid, protein core, icosahedral shaped)
Viral tegument (protein layer)
Outer envelope (lipids, proteins, glycoproteins)
Basic life cycle
Attachment
Un-coating
Replication (Integration, txn, tsn)
Assembly and release (budding and/or lysis)
(16) Describe what is unique about a retrovirus and it’s life cycle
Retrovirus: RNA viruses can convert RNA genome to DNA to insert into host genome
Reverse transcriptase: synthesizing DNA from RNA or DNA template
Life cycle
Virus attachment
Viral core enters cytoplasm
RNA —> DNA via reverse transcriptase
DNA integrates in host genome
T cell activation begins transcription of viral genes. RNA transported to cytoplasm
RNA and proteins are packaged into new viral particles, which acquire a new envelope, and they bud from the cell
(16) What are co-transformation and co-transduction and what do they indicate?
Fragments that co-transform/transduct are closer together on the chromosome
(17) Explain why we need to be able to turn gene expression on and off
We need to be able to turn gene expression on and off to save energy (human cells only express cytokines in the presence of pathogens)
(17) What are the 6 major levels of control of gene expression?
Transcription initiation
mRNA stability (change half life of mRNA)
Alternative splicing (eukaryotes only)
Translation initiation
Posttranslational modification: phosphorylation, acetylation, proteolytic cleavage
Protein stability: environmental conditions can trigger a protein to be targeted for degradation
(17) Explain the basic components of an operon
Operon: transcriptional unit including: promoter + operator + structural genes
Regulator/Regulatory gene: DNA sequence encoding products that affect the operon function, but are not part of the operon (transcribed and translated to produce a regulator protein that will bind to operator to repress or activate)
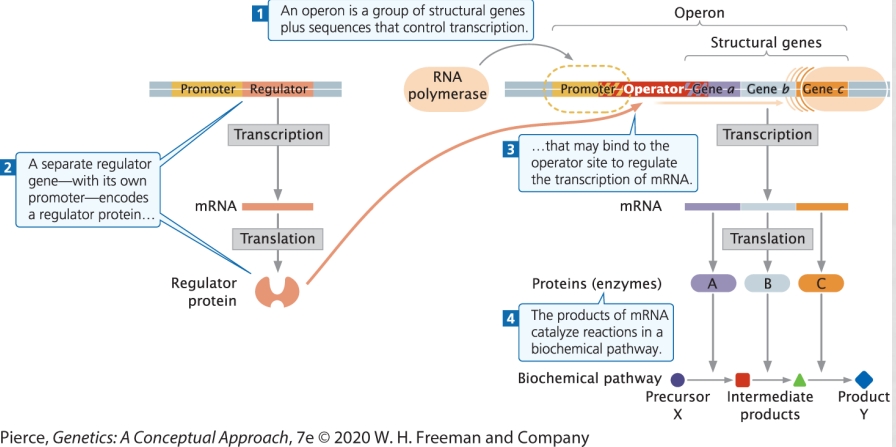
(17) Describe the roll of an effector molecule in an inducible versus repressive operon
In inducible operons, effector molecules turn transcription ON
In repressible operon, effector moleciles turn transcription OFF
(17) Know the difference between positive and negative control of operons
Negative control: regulatory protein is a repressor (OFF switch)
Positive control: regulatory protein is an activator (ON switch)
(17) Know the difference between inducible and repressible operons
Inducible operon: transcription is usually off and needs to be turned ON
Repressible operon: transcription is normally on and needs to be turned OFF
(17) Be able to describe negative inducible, negative repressible, positive repressible, and positive inducible operons
Negative inducible: Repressor bound to operator, blocking POL from binding promoter —> no TXN
Negative repressible: Repressor inactive, POL can bind promoter = TXN
Positive inducible: Activator inactive, POL can’t bind = TXN off
Positive repressible: Regulatory protein binds DNA = TXN on
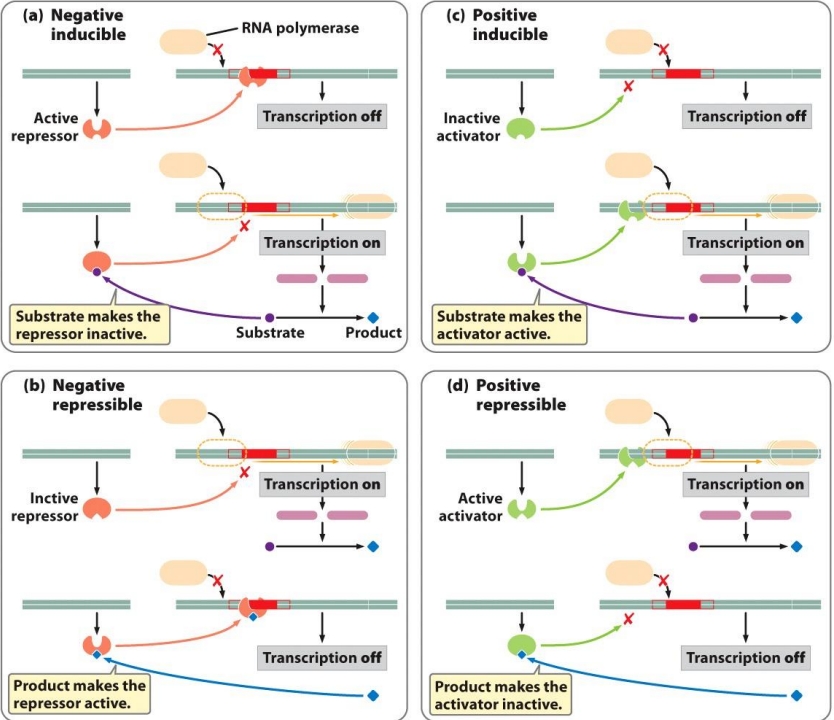
(17) Know the components of the lac operon and be able to identify or draw the operon structure
The lac operon of E. coli is a negative inducible operon.
Inducer: allolactose
LacI: gene that encodes repressor protein
LacP: operon promoter (binds POL)
LacO: operon operator (binds repressor)
LacZ: Encodes b-galactosidase (converts lactose —> glucose and galactose)
LacY: Encodes permease (transports lactose across membrane)
LacA: Encodes transacetylase (function unknown)
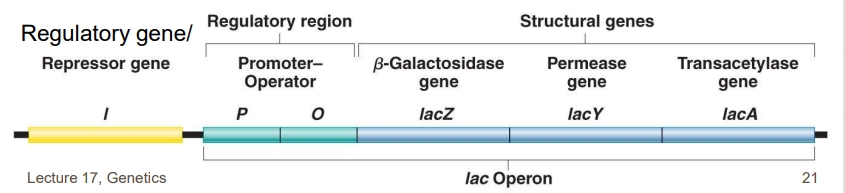
(17) Understand how the lac operon functions in the presence/ absence of lactose and high/ low glucose
In absence of lactose: lacI produces repressor, which binds operator = no TXN
In presence of lactose: lactose converted to allolactose, which binds to active repressor, inhibiting repressor from binding the operator = TXN
In presence of lactose AND glucose: lac operon turned OFF
To express lac operon, you must have 1. presence of lactose and 2. absence of glucose
In high glucose, less cAMP available to bind CAP, and absence of CAP/cAMP at promoter = less TXN
In low glucose, more cAMP is available to bind CAP, CAP/cAMP binds to promoter = more TXN
(17) Understand the function of the trp operon in the presence and absence of tryptophan
The trp operon of E. coli is a negative repressible operon. The binding activator is trp, allows repressor to bind operator, blocking promoter = no TXN
In the presence of trp: trp binds to repressor and makes it active = no TXN
In the absence of trp: trp repressor is inactive, it cannot bind to operator = TXN ON
(17) Gene expression
The steps through which a gene influences phenotype
(17) Structural genes
Genes that encode proteins
(17) Regulatory genes
Encoding products that interact with other sequences and affect the transcription and translation of these sequences
(17) Regulatory elements
DNA sequences that are not transcribed but play a role in regulating other nucleotide sequences
(17) Constitutive expression
Continuously expressed under normal cellular conditions
(17) Regulator protein
Controls transcription
(17) Operon
Transcriptional unit including promoter + operator + structural genes
(17) How do amino acids in DNA-binding proteins interact with DNA?
By forming hydrogen bonds with DNA base
(18) Describe Recombinant DNA technology (genetic engineering)
Set of molecular techniques for locating, isolating, altering, combining and studying DNA segments. Produces recombinant DNA (DNA that has been recombined)
(18) Cut & paste DNA: explain how restriction enzymes cut DNA & how they are used with genes of interest and plasmids
Restriction enzymes: recognize and make DS cuts in DNA at specific NT sequences. Can produce cohesive or blunt ends.
Cohesive ends: fragments with short, single stranded overhangs
Blunt ends: even length ends from both single strands
Two fragments cut with the same restriction enzyme will recombine. Cut gene of interest and cut plasmid to insert gene into plasmid.
(18) Visualize DNA: Explain how gel electrophoresis is used, explain how a southern blot is used
Gel electrophoresis: technique used to separate DNA fragment by size (gel stained with ethidium bromide, fluoresces under UV light)
Southern blotting: Process by which DNA is transferred form a gel to a solid support such as a nitrocellulose or nylon fiber (buffer moves up through paper and transfers DNA to membrane)
(18) Amplify DNA: What are the three basic steps of gene cloning, describe the entire process of gene cloning (including the specific example using the lacZ & amp genes described)
Three basic steps
Create plasmid vectors (insert foreign DNA into plasmid using restriction enzymes)
Transformation of host cells with plasmids
Screening cells for recombinant plasmids (selectable markers used to confirm whether cells have been transformed or not)
Gene cloning process
Gene of interest inserted into plasmid containing ampR and partial lacZ gene (recombinant plasmid also contains gene of interest). Bacteria that will receive the plasmids are deficient in lacZ (lacZ-) and amp sensitive
Bacteria are transformed, they can:
Not take up plasmid (killed by ampicillin)
Take up plasmid without inserted gene (gain amp resistance, can make b-galactosidase and cleave x-gal, colonies blue)
Take up plasmid with inserted gene (gain amp resistance, gene insertion disrupts partial lacZ gene so cannot make b-galactosidase, colonies white)
(18) Give an example of how genetic engineering is used in tobacco plants
BT toxin. Lethal to insects, but does not harm other plants/animals. BT toxin FNA fragmented and ligated to neo.
Fragments cut with restriction enzymes and inserted in “expression vectors” (plasmids that contain everything needed to transcribe and translate the gene)
A. tumefaciens transformed and selection with neomycin
Tobacco plants infected with recombinant A. tumefaciens
DNA transferred to plant genome and plants can produce BT toxin!
(18) Describe the three steps of PCR, what is special about Taq Polymerase
Denaturation: Separates DNA strands so templates are single stranded
Annealing: Allows primers to anneal but not renaturation
Extension: Allows DNA to be replicated
Taq polymerase is special because it is a stable DNA polymerase at high temperature.
(18) Find genes of interest: What are the steps to create a gene library? What are the steps in screening a gene library?
Steps to making gene library
Bacteria are transformed with the plasmids, creating a genomic library. A genomic library is a collection of clones containing all the DNA fragments from one source
DNA isolated from cells
DNA cut by cocktail of restriction enzymes
Fragments are ligated into different plasmids (can contain overlapping fragments of genes)
This collection of plasmids should contain all of the fragments of DNA
Screening gene libraries
Bacteria are plated on agar
Some colonies adhere to nitrocellulose fiber
Filter treated to disrupt cells, denature DNA and fix to filter
Probe hybridizes to DNA of interest
Excess probe washed off, filter exposed to visualize DNA of interest and determine corresponding bacteria on agar
(18) Analyze Gene function: what’s the difference between a transgenic mouse and knockout mouse?
Transgenic mouse: organism permanently altered by the addition of a DNA sequence to its genome (carries a transgene)
Knckout mouse: mouse in which a normal gene has been disabled (it’s non-functional)
(18) Probe
DNA or RNA with a base sequence complementary to a sequence in the gene of interest
(19) What caused the decline in population of isolated Bighorn Sheep in the 1900s?
They were bred in isolation for 60 years, limited gene pool caused decreased genetic variation
(19) Explain the relationship between genetic variation, the gene pool, and survival as a species
Genetic variation comes from the gene pool and is the basis of evolution. More variation = better ability to adapt
(19) Know how to mathematically calculate phenotypic frequencies
Ex. In a sample of a population of flowers where flower color shows incomplete dominance, there are 30 red, 20 white, and 50 pink.
Phenotypic frequency: portion of individuals in a population that have a given phenotype
f(red) = # red individuals / N = 30 / 100 = 0.3
(19) Know how to mathematically calculate genotypic frequencies
Ex. In a sample of a population of flowers where flower color shows incomplete dominance, there are 30 red, 20 white, and 50 pink.
red are R1R1
white are R2 R2
pink are R2R1
Genotypic frequency: the portion of individuals in a population that have a given genotype
f(R1R1) = # R1R1 individuals / N = 30 / 100 = 0.3
(19) Know how to mathematically calculate allelic frequencies
Ex. In a sample of a population of flowers where flower color shows incomplete dominance, there are 30 red, 20 white, and 50 pink.
red are R1R1
white are R2 R2
pink are R2R1
Allele frequency: the fraction, out of all copies of a given gene, that are of a given allele
Measuring allelic frequency
Count # copies of a given allele in a population
Divide this # by total # of copies
f(R1) = # copies of R1 allele / tot # copies of allele at the locus 2N
110/200 = 0.55
(19) Know how to mathematically calculate allelic frequencies of multiple (three) alleles

(19) Understand how Hardy-Weinberg relates allele frequency to genotype frequency in a population
p²: homozygous dominant allele pair frequency (AA)
q²: homozygous recessive allele pair frequency (aa)
2pq: heterozygous allele frequency (Aa)
Hardy-weinberg equation: p²+2pq+q²=1
(19) Describe the assumptions made by the Hardy-Weinberg Law
Population is large (so no genetic drift changes allele frequencies)
Random mating
The gene is not under selection (none of the genotypes have a greater chance of survival)
Gene pool is isolated (no new alleles)
Mutation is not introducing new alleles or changing allele freq.
(19) Know how to mathematically calculate genotypic and allelic frequencies in populations that are in Hardy-Weinberg

(19) Know why the sum of all frequencies of genotypes at one locus, or of alleles for one gene, equals one
Because they represent the sum of all possible outcomes
(19) Genetic drift
Change in frequency of a gene variant (allele) in a population due to random sampling of organisms
(19) Population genetics
Study of genetic makeup and inheritance of genes in a population
(19) Population
Group of individuals who can breed
(19) Gene pool
All of the genetic information within a Mendelian population
(1) List the Key Characteristics of Genetic material
Genetic material must contain complex information
Genetic material must replicate faithfully
Genetic material must encode phenotype
Genetic material must have capacity to vary
Genetic material is carried in DNA or RNA
(1) Describe the 3 qualities of genetic material
Stores information
Makes information available to cell
Is accurately replicated and evenly distributed (to daughter cells during cell division, to gametes during cell reproduction)
(1) Describe the features of a double helix & be able to identify the components of DNA structure
Features
3D configuration
Phosphodiester bond backbone
Hydrogen bond and base pairing
Antiparallel strands
Complementary strands
Hydrogen bonds between bases
Major grooves and minor grooves
(1) Describe the difference between coding and non-coding genes
Coding genes: Make proteins
Non-coding genes: Functional RNA (transcribed but not translated into protein) —> tRNA, rRNA, snRNA, etc.
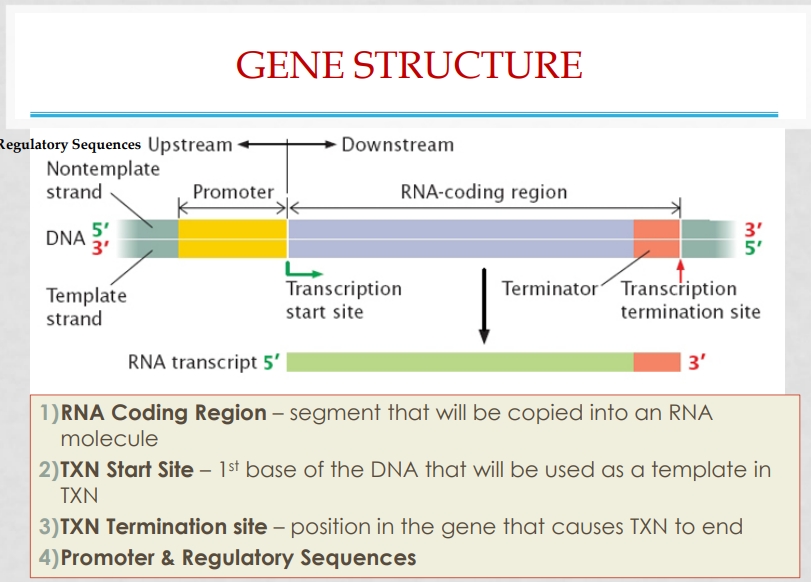
(1) Draw and describe the basic structure of a gene
(1) Genome
Complete set of genetic instructions for any organism
(1) Gene
Sequence of DNA that contains instructions for particular trait
(2a) Describe differences between prokaryotes & eukaryotes & viruses
Prokaryotes: No membrane-bound organelles, no nucleus, genome is usually one circular DNA molecules
Eukaryotes: Genome is multiple linear DNA molecules
Viruses: Outer protein coat surrounding nucleic acid(
(2a) Explain how a prokaryotic cell reproduces
Simple division: separation of replicated circular chromosome
Origins of replication segregate to opposite sides
Cell divides
(2a) Know the difference between chromosomes, chromatids, and chromatin
Chromosome: structure consisting of DNA and associated proteins that carries and transmits genetic information (each circular or linear strand of DNA)
Chromatin: Form of the chromosome where DNA has been condensed with proteins to fit into the EUK nucleus
Chromatid: Further condensed form of DNA, referring to one of the two strands of the same chromosome after replication, which remain attached to each other and are called “sister chromatids”
(2a) List differences between viral genomes & prokaryotic genomes
Viral genomes
dsDNA, ssDNA, dsRNA, or ssRNA containing minimal genes to produce new viruses
Prokaryotic genomes
Circular DNA molecule, one chromosome containing all genes, little non-coding DNA
Contain plasmids (small, non-essential, independently replicating circular DNA separate from chromosome. Easily transferred between prokaryotes.)
(2a) Human Genome: Know size, estimated # of genes, and percent encoding protein
3.2 million bp
Estimated 20k genes
Only ~1-2% encodes proteins
(2a) Describe chromosome structure in prokaryotes versus eukaryotes
Prokaryotes: Single circular DNA molecule
Eukaryotes: Linear DNA molecules inside membrane-bound nucleus, organized with histone proteins, forming chromatin. Have telomeres and centromeres
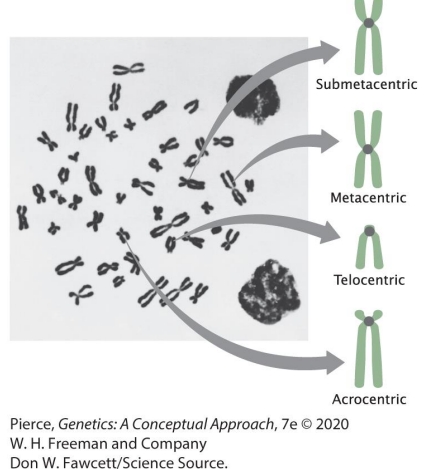
(2a) Describe human chromosomes: number, structure, homologous pairs versus sister chromatids, 4 types
22 pairs (autosomes) + sex chromosomes
Homologous pairs: 1 from mom, 1 from dad. During replication, each of these forms an identical copy (sister chromatid)
Telomere: protective ends
Centromere: location of kinetochore assembly, attachment point for spindle MTs
Kinetochores: forms before cell division, MT attach here
4 types: Metacentric, submetacentric, acrocentric, telocentric
(2a) Know the phases of the cell cycle, what occurs in each phase, and checkpoints
Interphase
G1: Growth; proteins necessary for rep. synthesized
G1/S CHECKPOINT
S: DNA synthesis
G2: biochemical prep for cell division
G2/M CHECKPOINT
M Phase
Mitosis: separation of sister chromatids
Cytokinesis: separation of cytoplasm
(2b) Describe: semi-conservative DNA replication
When DNA is copied, each new DNA molecule contains one original (parental) strand and one newly synthesized strand.
(2b) Explain the entire process of DNA replication knowing the function of: primase, primers, nucleotides, helicase, gyrase, DNA polymerase, ssBPs, ligase
Initiation: initiator protein binds to the ORI and causes a short piece of DNA to unwind
Unwinding
Helicase: breaks h bonds between bases. binds to lagging strand and moves 5’ —> 3’
Single stranded binding protein (ssBP): stabilizes ssDNA
Gyrase (topoisomerase): controls DNA supercoiling
Primase (RNA polymerase): synthesizes short RNA primers that bind DNA
Primer: a group of RNA nucleotides created by primase, with a 3’ -OH group to which a new dNTP can be added
Elongation
DNA polymerase: elongates poly NT strand by polymerization
DNA ligase: seals breaks in DNA sugar phosphate backbone
(3) Know all the stages of the cell cycle and their purpose (individual slides & summary in Table 2.1)
G0 Phase: Stable, nondividing period
Interphase
G1 phase: growth and development of the cell; G1/S checkpoint
S: synthesis of DNA
G2 phase: prep for division; G2/M checkpoint
M phase
Prophase: chromosomes condense and mitotic spindle forms
Prometaphase: nuclear membrane disintegrates, and spindle microtubules anchor to knietochores
Metaphase: chromosomes align on metaphase plate, spindle assembly checkpoint
Anaphase: sister chromatids separate, becoming individual chromosomes that migrate toward spindle poles
Telophase: chromosomes arrive at spindle poles, the nuclear membrane re-forms, and condensed chromosomes relax
Cytokinesis: cytoplasm divides, cell wall forms in plant cells
(3) Describe the differences between Mitosis & Meiosis
Mitosis
Part of cell cycle
Genetically identical progeny
Diploid
One division
Division = chromatids
Meiosis
Part of life cycle
Genetically diverse progeny
Haploid
Two divisions
Division 1 = homologs, division 2 = chromatids
(3) Explain the difference between Meiosis I & Meiosis II
Meiosis I: separation of homologous chromosome pairs, and reduction of chromosome number by half
Meiosis II: Separation of sister chromatids, also known as equational division
(3) Understand crossing over & genetic recombination
Crossing over: Non-sister chromatids cross over, causing recombination. Cells inherit chromosomes with different combos of alleles on each. Crossing over between homologs causes recombination, the creation of new alleles (versions of a gene) on a chromatid
Random separation of homologs: Cells inherit different combos (paternal vs maternal) of chromosomes. Recomb. of alleles can occur due to random separation of homologous chromosomes
(3) Sister chromatids
Identical copies of same chromosome
(3) Homologous pair
Two of the same chromosome #, one from mom one from dad
(4) Know the structure of RNA & the differences between RNA & DNA
RNA structure: ribose sugar, U instead of T, has -OH on 2’ carbon
(4) Describe the difference between coding and non-coding RNA
Coding RNA: mRNA (carries code for proteins), tRNA (incorp. amino acids in polypeptide chain), rRNA (struc. and func. components of ribosome)
Non-coding RNA: pre-mRNA, snRNA, crRNA…
(4) Know the components of a transcription unit and the structure of a gene (slide 11,34,35)
TXN unit: sequence NT in DNA that has info to encode RNA molecule
Includes: promoter, RNA coding sequence, terminator
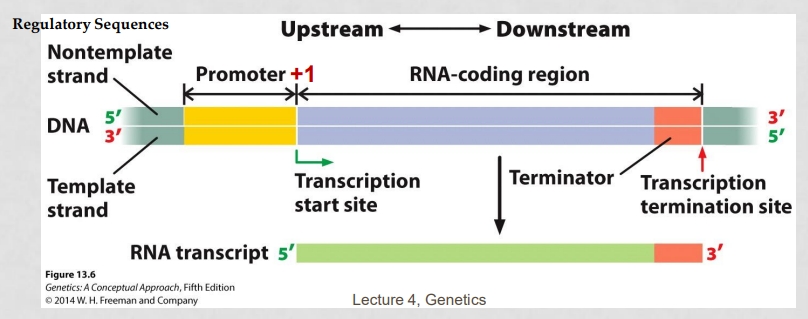
(4) Explain how RNA is produced from a template strand
RNA polymerase reads template strand (3’ —> 5’), builds RNA 5’ —> 3’, RNA sequence matches coding strand (except U replaces T)
(4) Know the process of TXN in prokaryotes: Initiation ( consensus sequences, transcription bubble, sigma factor, holoenzyme, unwinding, etc), Elongation (TXN bubble, pausing, accuracy, addition of NTs), Termination (Rho dependent vs. independent)
Initiation: TXN apparatus assembles on the promoter and begins synthesis of RNA
Promoter recognition: -10 consensus sequence of Pribnow box = TATAAT. -35 consensus sequence = TTGACA
Formation of transcription bubble
Creation of the first bonds between rNTPs
Escape of the TXN apparatus from the promoter
Elongation: DNA is threaded through RNA POL, where it is unwound and new NTs are added to the 3’ end of the growing RNA strand
TXN bubble: RNA continuously synthesized
TXN pausing: POL slides back on DNA to prevent TSN
TXN accuracy: RNA POL proofreads
Termination: recognition of the end of the TXN unit and the separation of the RNA molecule from DNA template
Rho- dependent termination: uses rho factor
Rho-independent termination: hairpin structure formed by inverted repeats, followed by string of uracils
(4) Explain the following processes: 5’ Capping, Polyadenylation
5’ capping: a nucleotide with 7-methylguanine 5’-5’ bond is attached to the 5’ end of the RNA
Polyadenylation: addition of poly(A) tail, adenine NTs added to the 3’ end of mRNA for stability, ribosome attachment, export
(5) Know the features of the genetic code
Triplet: 3 bases make codon
Degenerate: more than 1 codon per amino acid
No gaps: continuous
Written 5’-3’ as in mRNA
(5) Explain reading frame versus open reading frame
Reading frame: way NT seq. is read in groups of 3 NT in TSN
ORF: begins w/ start (AUG), ends w/ stop (UAA, UAG, UGA)
(5) Explain the process of translation, the event that occur during Initiation, Elongation, & Termination and the factors involved
Initiation: IF3 binds to 30S to prevent 50S binding (so mRNA can bind). Shine delgarno consensus sequence (AGGAGG) must be present upstream the AUG start codon. mRNA shine delgarno sequence binds 16S in small subunit, positioning the initiation codon.
Elongation: XXXX
Termination: There is no tRNA for the stop codon. Release factors bind mRNA and disassemble the complex
(5) Explain charging of tRNA, including the structure of the tRNA, the role of the aminoacyl transferase, & the molecular interactions that occur
tRNA: 1 RNA molecule, folded into cloverlead, held by hydrogen bonds, carries AAs to ribosome to create a polypeptide
tRNA charging: attaching an amino acid to tRNA. AA needs to bind the 3’ adenine of the tRNA. AA carboxyl group (COO-) binds to the carbon of the tRNA adenine.
Aminoacyl-transferease: guide specific AA to bind.
(6) Describe the benefit of studying mutations
Reveal gene function
Explain genetic disease
(6) Categories of Mutations: What are the four types/ classes of mutations
Germline: in cells that make gametes
Somatic: in cells that dont make gametes
Mosaic: Patches of mutated cells (occurs in embryo prior to cell diff.)
De novo: mutation that appears for 1st time
(6) Know the difference between frame shift and in-frame
Frameshift mutation: changes reading frame, can drastically affect phenotype (not a set of 3)
In-frame indels: does not change reading frame (sets of 3)
(6) List and describe the types of Protein coding sequence mutations: Silent, Missense, Nonsense
Silent (synonymous): Base sub. in NT seq of codon but encodes same AA (no AA change)
Non-synonymous: Base sub, but alters protein encoded
Missense: NT sub in codon —> changes AA
Nonsense: NT sub in codon —> makes STOP codon