Foundations of Biology Topic 2
1/59
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
60 Terms
energy
is the capacity to do work or cause change, where work occurs when a force acts over a distance
main types of energy in biology
Chemical energy → stored in bonds (e.g. ATP, glucose)
Electrical energy → charge differences across membranes
Heat (thermal energy) → molecular motion/temperature
Light energy → photons used in photosynthesis/vision
Mechanical energy → movement (muscles, cytoskeleton)
two broad categories of energy
Potential energy → stored energy (bonds, gradients, charges) → chemical bonds, ion gradients, ATP
Kinetic energy → energy of movement (heat, motion, particles) → muscle movement, diffusion, heat energy
thermodynamics
study of energy transformations and transfers in systems
open vs closed energy systems
Closed system: no matter/energy exchange (e.g. thermos)
Open system: exchanges energy + matter (e.g. organisms)
first law of thermodynamics
Energy cannot be created or destroyed, only transferred or transformed.
second law of thermodynamics
Every energy transfer increases entropy (disorder) because some energy is lost as heat
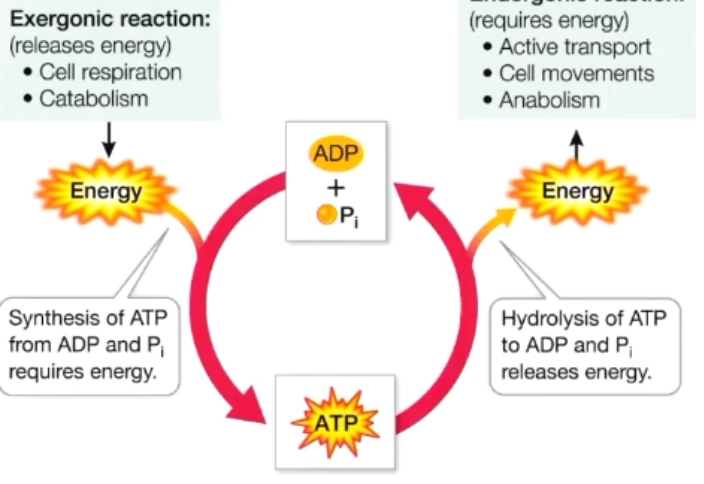
ATP
Adenosine triphosphate → a high-energy molecule used to transfer energy in cells.
energy rich → 3 negatively charged phosphate groups that repel each other
bonds are unstable → energy released when broken
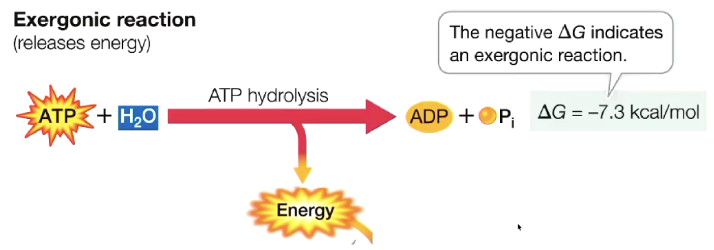
ATP hydrolysis
water added
ATP → ADP + Pi
releases energy (exergonic)
energy is used in the cell for work
phosphorylation
addition of phosphate group to a molecule, changes its shape and energy shape
Gibbs free energy
ΔG symbol, represents energy (useful energy)
exergonic and endergonic reactions
Exergonic: releases energy, spontaneous (-ΔG)
Endergonic: requires energy, non-spontaneous (+ΔG)
equilibrium in energy
state of maximum stability where no net energy change occursstate of maximum stability where no net energy change occurs
catabolic and anabolic reactions
Catabolic: breakdown large molecules into smaller ones, release energy (exergonic), -ΔG, usually hydrolysis and oxidation
Anabolic: build complex molecules from simple ones, uses energy (endergonic), +ΔG, usually condensation and reduction
are interconnected
oxidation
Loss of electrons
reduction
Gain of electrons
redox reactions
changes in electron transfer or charge of molecules
coupling of oxidation + reduction
transfer energy via electrons for work, driving processes like respiration and photosynthesis
enzymes
act as biological catalyst to increase reaction (metabolic and digestive) rate by lowering activation energy without being consumed
activation energy
the minimum energy required for reactants to reach the transition state and begin a reaction. lowering it means more molecules reach the transition state, more successful collisions per second thus faster reaction rate.
active site
specific region where the substrate binds and the reaction occurs, involving specific amino acid R groups.
active side models
Lock and key: rigid and fixed active site shape, perfect fit
Induced fit: active site changes shape to fit substrate (more accurate model)
steps is enzyme catalysis
substrate binds to active site
enzyme stabilises transition state
activation energy is lowered
products are released, enzyme unchanged
substrate concentration on enzyme activity
rate increases as substrate increases
at saturation point all active sites are full → maximum rate (Vmax)
temperature on enzyme activity
Low temp → slow activity
Optimal temp (~37°C in humans) → max activity
High temp → denaturation → loss of function
pH on enzyme activity
Low temp → slow activity
Optimal temp (~37°C in humans) → max activity
High temp → denaturation → loss of function
cofactors
Non-protein helpers required for enzyme function
can be inorganic ions or organic molecules
coenzymes
non protein organic molecules, transfer chemicals from active site of an enzyme to active site of another enzyme
prosthetic groups
permanently bind to active site to transfer chemicals from active site of enzyme to some other substance
inhibitors
molecules that reduce enzyme activity by blocking or altering active sites
Competitive: compete with substrate
Non-competitive: bind elsewhere, change enzyme shape
Irreversible: permanently inactivate enzyme
how to measure enzyme reaction rate
Decrease in substrate concentration
Increase in product concentration
Change per unit time (kinetic measurement)
feedback inhibition
A regulatory mechanism where the end product of a metabolic pathway inhibits an enzyme earlier in the pathway, slowing or stopping further production.
importance:
prevent overproduction of products
conserves energy and resources
how does feedback inhibition work
final product accumulates
binds to an enzyme early in the pathway, often allosterically
enzyme activity decreases → pathway slows or stops
feedback activation
when a product or molecule activates an enzyme, increasing pathway activity and directs metabolic flow.
how can biological pathways be regulated by feedback + example
Products from one pathway can:
Inhibit their own pathway
Activate other pathways
Helps balance metabolic needs
example:
Phosphofructokinase (glycolysis enzyme):
Activated by AMP (low energy)
Inhibited by ATP and citrate (high energy)
adjusts respiration rate based on energy demand
allosteric regulation
Regulation where a molecule binds to a site other than the active site (allosteric site), causing a shape change that alters enzyme activity.
allosteric enzyme
An enzyme with:
Multiple subunits
Multiple binding sites
Ability to switch between active and inactive forms
allosteric activators
Bind to allosteric site
Stabilise active form
Increase enzyme activity
allosteric inhibitors
Bind to allosteric site
Stabilise inactive form
decrease enzyme activity
metabolism
the sum of all biochemical reactions in a cell, often organised into metabolic pathways where products of one reaction become substrates for the next
metabolic pathways
reactions occur in stepwise sequences
each step is catalysed by a specific enzyme
pathways are regulated at key steps
catabolism and anabolism
Catabolism: breakdown of large molecules to smaller ones, usually hydrolysis and oxidation, releasing energy (exergonic, -ΔG)
Anabolism: Synthesis of complex molecules from simpler ones, usually condensation and reduction, requiring energy (endergonic, +ΔG).
they are interconnected
lipid metabolism
catabolism: energy production
lipolysis: triglycerides → glycerol + fatty acids, occurs in adipose tissue
fatty acids bind to coenzyme A → fatty acyl-Co, tranport to mitochondria for energy production or lipid synthesis
β-oxidation (mitochondrial matrix): fatty acids broken down into acetyl-CoA (TCA cycle), NADH, FADH2 (ETC)
anabolism: energy storage
lipogenesis: when energy is abundant, acetyl-CoA from excess glucose or amino acids → fatty acids → triglycerides stored in adipose tissue
gluconeogensis
production of glucose within a cell, it is almost a reversal of glycolysis. making new glucose from a non carb source.
protein metabolism reasons
3 key points:
there is no storage form of proteins, all proteins consumed is turned into carbs or fat and the nitrogen from the amino group must be eliminated through the urea cycle
essential amino acids must be consumed daily for proteins to be made
proteins are always degraded, therefore there is a constant need for quality protein to maintain the product’s structure
How are proteins catabolised?
amino acids undergo:
Transamination (transfer NH₂ group to a keto acid)
Deamination (remove NH₂ → ammonium ion is removed)
Carbon skeleton enters glycolysis or citric acid cycle
urea cycle
takes 2 metabolic waste products, ammonium ions and carbon dioxide and produces urea, a relatively harmless soluble compound that is excreted in urine
why are proteins not a preferred energy source
no storage form
requires nitrogen removal (urea cycle)
complex, slower
how are proteins synthesised
Amino acids formed via amination (ammonium ion used to form an amino group) and transamination (amino group transferred to keto acid)
Assembled into proteins
aerobic cellular respiration
metabolic process in which cells break down glucose using oxygen to produce energy in the form of ATP, releasing carbon dioxide and water as waste
C6H12O6(glucose)+6O2(oxygen)→6CO2(carbondioxide)+6H2O(water)+ATP(energy)
glycolysis: in cytoplasm, break down glucose to pyruvate and makes 2 tap
TCA cycle: in mitochondrial matrix, producing carbon dioxide and transfer electrons
ETC: in inner mitochondria membrane, produce majority of ATP through oxidative phosphorylation
glycolysis
in cytosol
10 step pathways
inputs: glucose, ATP, NAD+, ADP + Pi
steps 1-5: (energy investment) use ATP to phosphorylate glucose
steps 6-10: (energy pass off): produces ATP and NADH
outputs: pyruvate, 2 ATP (net), NADH
regulation of glycolysis
key enzyme in glycolysis: phosphofructokinase
reaction in pathway: fructose-6-phosphate → fructose-1,6-bisphosphate
Regulation:
Activated by: AMP, ADP → when low energy → ↑ glycolysis
Inhibited by: ATP, citrate → when high energy → ↓ glycolysis
TCA cycle/citric acid/krebs cycle
inputs: acetyl CoA, NAD+, FAD, ADP + Pi, H2O
outsputs: CO2, NADH, FADH2, ATP
cycle spins twice per glucose, making 2 ATP
TCA cycle regulation
High NADH/FADH₂ inhibits enzymes
Slows cycle when energy is abundant
energy generation in TCA cycle
Energy (electrons and H+)captured as NADH and FADH₂ (electron carriers)
Small ATP via substrate-level phosphorylation
Electron carriers go to electron transport chain → major ATP production
Electron transport chain
inner mitochondrial membrane
inputs: NADH, FADH2 (electron carriers), O2 (final electron accepter)
NADH and FADH₂ are oxidised → release high-energy electrons
Electrons pass through a series of protein carriers
Energy released is used to pump H⁺ (protons) from matrix → intermembrane space
process done through chemiosmosis
ATP Synthesis:
H⁺ flows back into matrix via ATP synthase
Drives ADP + Pi → ATP
👉 This is oxidative phosphorylation
outputs: 30-32 net ATP, water, NAD+, FAD
chemiososis
from a proton gradient
Creates electrochemical gradient (high H⁺ outside, low inside)
This stores potential energy
anaerobic cellular respiration
without O2, no ETC
cells use: glycolysis and fermentation (lactic acid and alcoholic)
fermentation regenerates NAD+ for glycolysis
lactic acid fermentation
pyruvate → lactate
in muscle cells
2 atp
alcoholic fermentation
pyruvate → ethanol + CO2
in yeast and plants cells
2 atp