DNA Replication
1/193
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
194 Terms
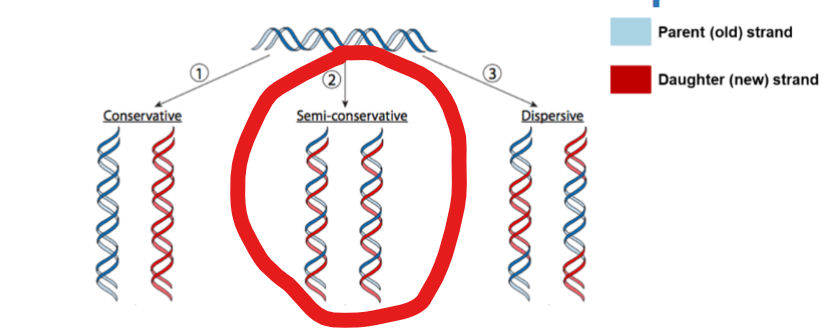
What is the mechanism of DNA Replication’
it is semi conservative
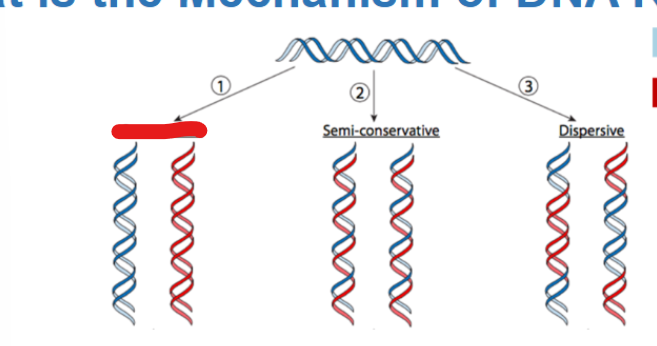
what type of replication yields an original intact DNA molecule and one entirely newly synthesized DNA
conservative replication
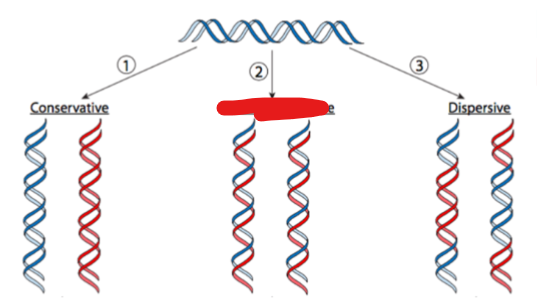
what type of replication yields 2 dna molecules, each with one parental and one newly synthesized strand
semi conservative
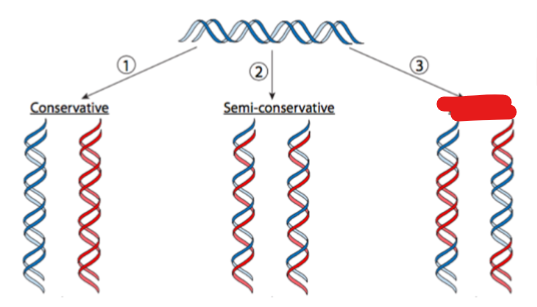
what replication yields 2 dna molecules that are hybrids ( or mixtures) of parental and newly syntehsIzed DNA
dispersive replication
DNA polymerases with a numeral are found where?
within prokaryotes

DNA polymerases with greek letters are loacted where?
eukaryotes
with DNA polymerase what are the substrates for it
dNTPS
what does DNA polymerase do?
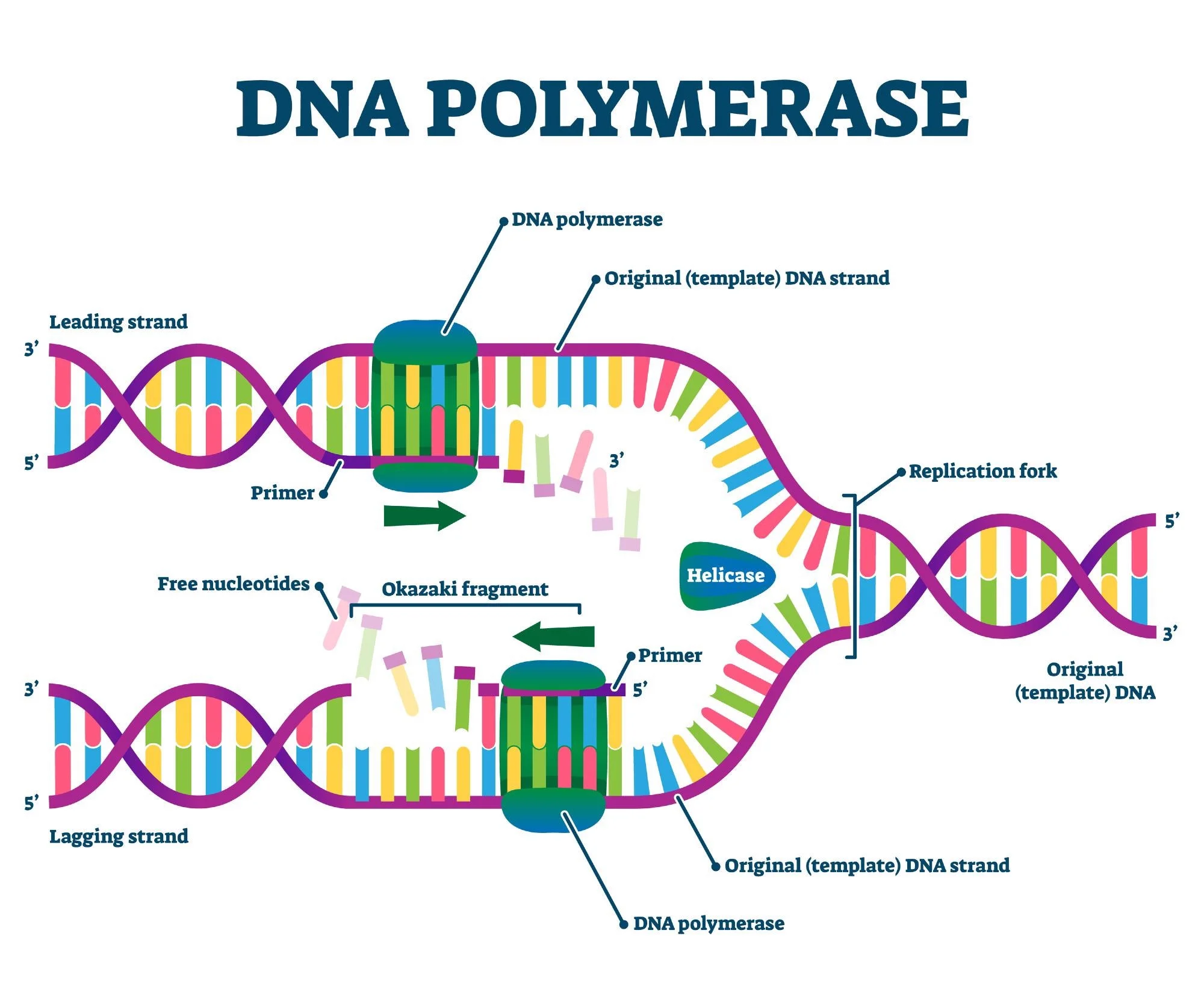
it catalyzes the extension of DNA strand one dNMP at a time
What direction does the dna polymerase syntehsize in
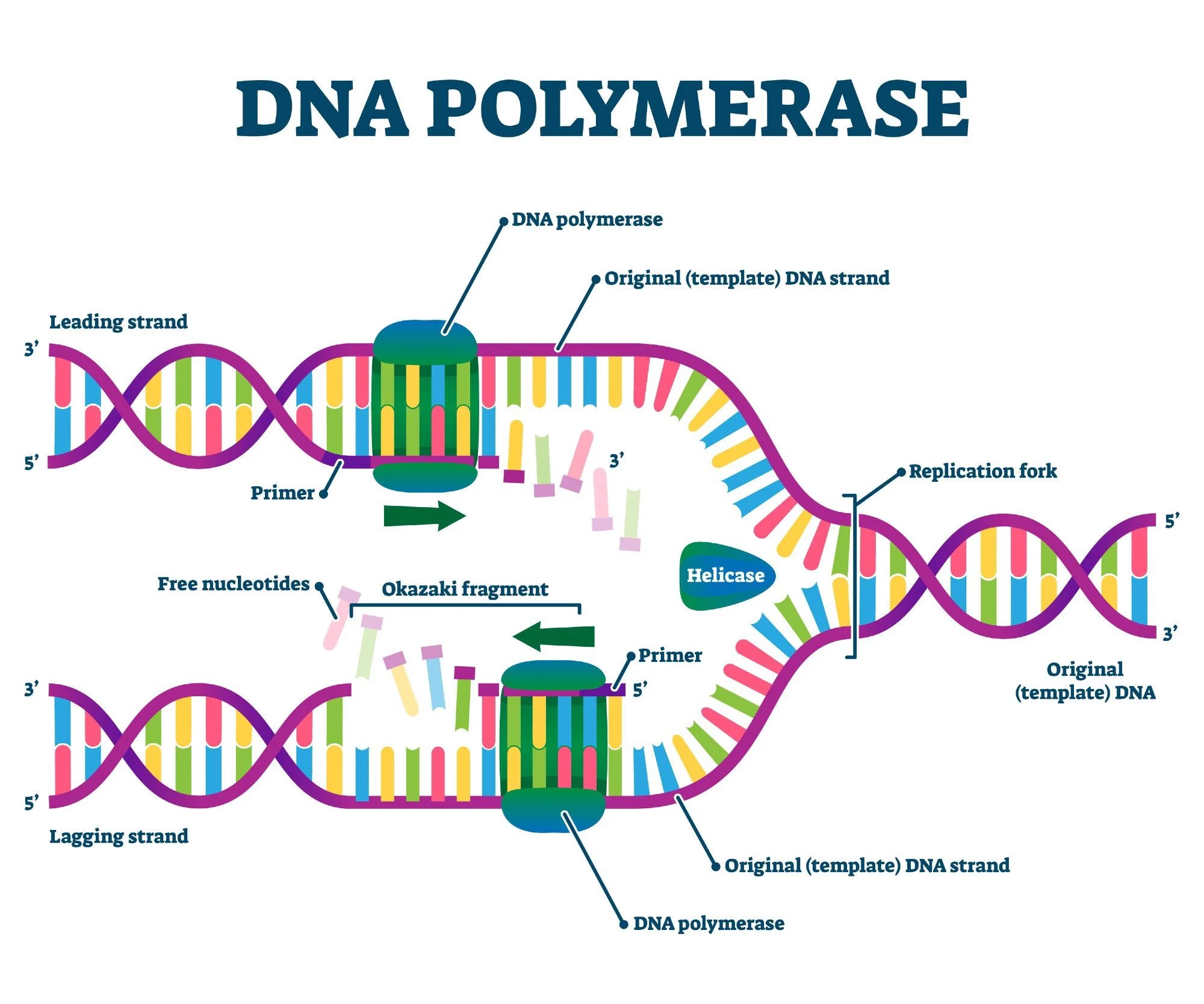
5’ to 3’ direction
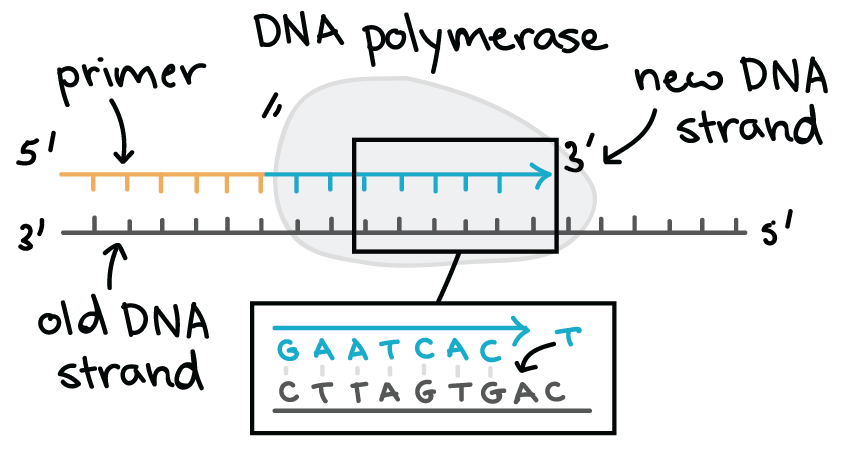
what does the dna polymerase require to work?
a template and DNA/RNA primer
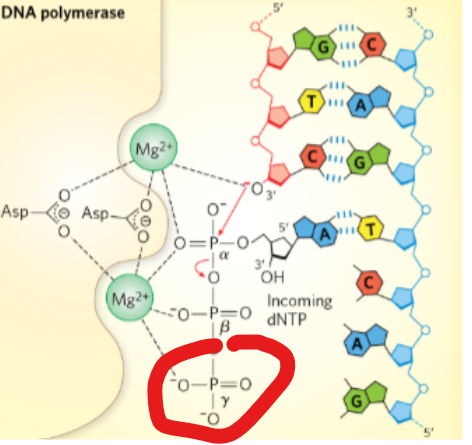
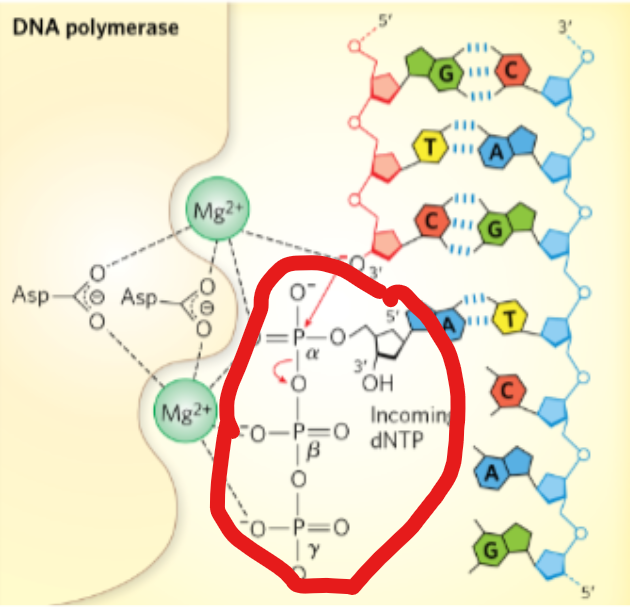
During a DNA synthesis reaction, what type of transfer occurs?
phosphoryl group transfer
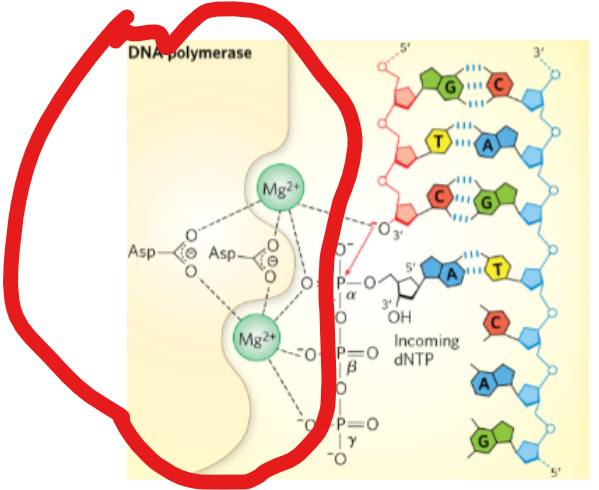
During a DNA synthesis reaction,what is required at the active site in the DNA polymerase enzyme?
Mg²+
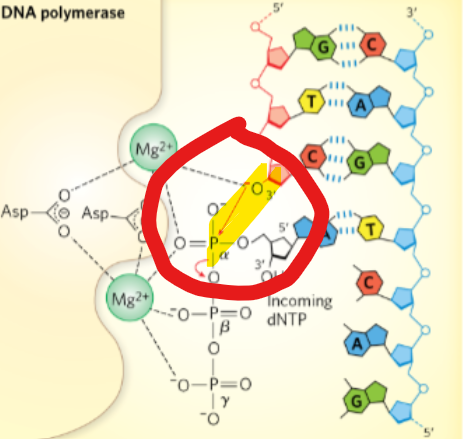
During a DNA synthesis reaction: ________ of the 3’ nucleotide attacks _______ of incoming dNTP
3’OH; α phosphate
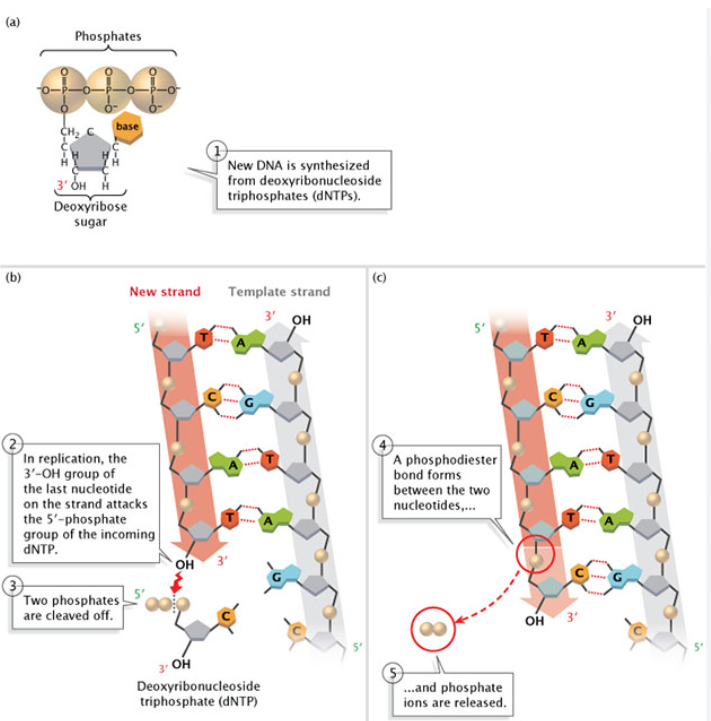
During a DNA synthesis reaction,what is released?
it releases PPi (pyrophosphate)
what does an error in replication do?
it can introduce a mutation into the genome
An error in replication can introduce a mutation into the genome.
What will come from the mutation?
it will be permenant and inherited by subsequent daughter cells
What is the average E. coli mutation
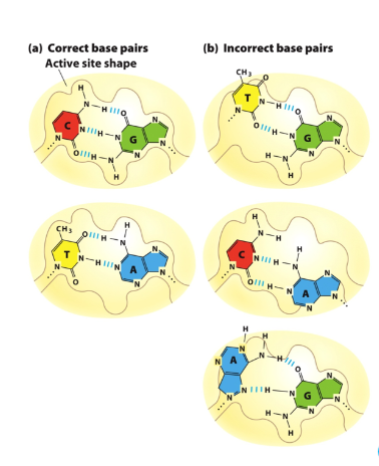
What does the DNA Polymerase active site restrict base pairing to ?
watson-crick-franklin bp
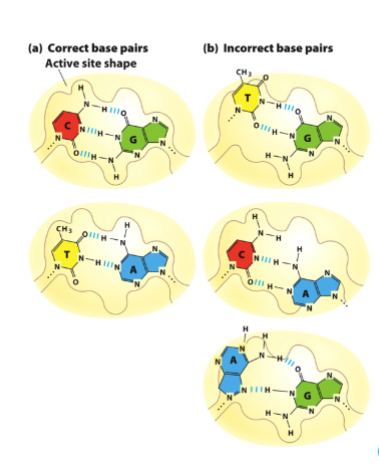
DNA Pol active site restricts base pairing to Watson-Crick-Franklin bp
What does this mean?
that before a nucleotide is even added, the enzyme essentially “checks” whether the incoming base can form the correct hydrogen bonding pattern and fit geometrically with the template strand
DNA polymerase has an active site that is highly specific in shape and chemistry, which only properly accommodates correct Watson–Crick–Franklin base pairs (A–T and G–C). This means that before a nucleotide is even added, the enzyme essentially “checks” whether the incoming base can form the correct hydrogen bonding pattern and fit geometrically with the template strand
What is this known as?
presynthetic error control
DNA polymerase has an active site that is highly specific in shape and chemistry, which only properly accommodates correct Watson–Crick–Franklin base pairs (A–T and G–C). This means that before a nucleotide is even added, the enzyme essentially “checks” whether the incoming base can form the correct hydrogen bonding pattern and fit geometrically with the template strand—this is known as presynthetic error control. Because incorrect bases do not align properly in the active site, they are usually rejected before incorporation.
However, this system is not perfect .. what does DNA polymerase do?
it still inserts the wrong nucleotide- but the fidelity of DNA replication remains extremely high because additional proofreading and repair mechanism act after this step to correct most mistakes
When does a mutation occur during replication
rarely
What is the big picture of the accuracy or replication
Even though DNA polymerase isn’t perfect, the combination of multiple correction systems makes DNA replication so accurate that mutations are rare events, not common ones—only appearing after thousands of replication cycles.
what digests polynucleotide chains
nucleases
which nuclease is specific for DNA
DNase
which nuclease is specific for RNA
RNase
what nucelase breaks a phosphodiester at one end of a polynucleotide chain
exonuclease
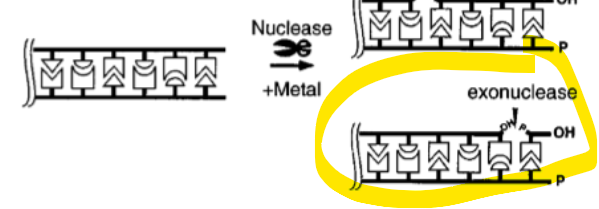
what direction does exonucelase work
5’→ 3’ OR 3’→5’
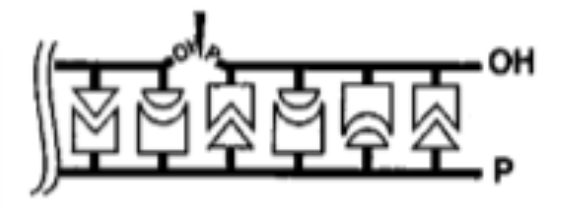
which type of nuclease breaks a phosphodiester bond within a polynucleotide chain
endonuclease
what two forms can an endocnuclease be
sequence independent to sequence specific
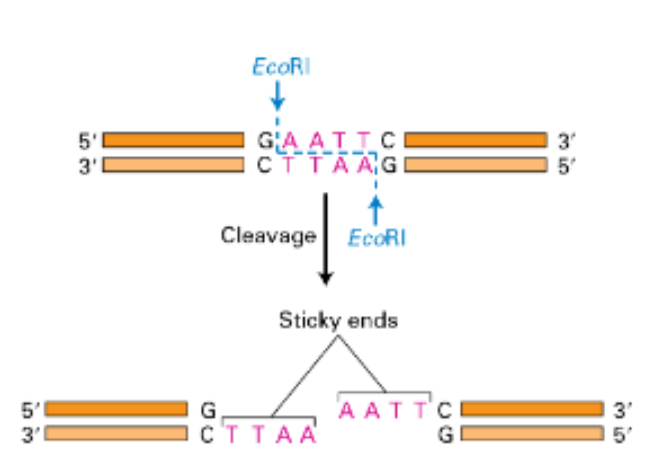
what type of breaks can endonuclease do
single strand (nick)
double strand break
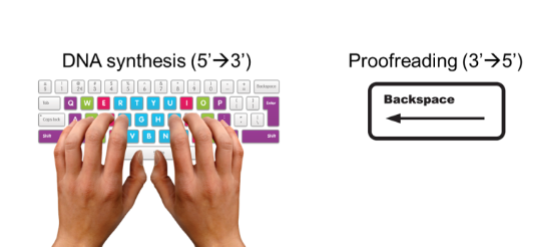
What is a critical mechanism that ensures the extremely high accuracy of DNA replication by correcting mistakes immediately after they occur.
Proofreading
High-fidelity DNA polymerases have two distinct active sites, each with a different function
What are the active sites
a catalytic site for DNA synthesis
a 3’→5’ exonuclease site for removing mis-incorporated nucletides
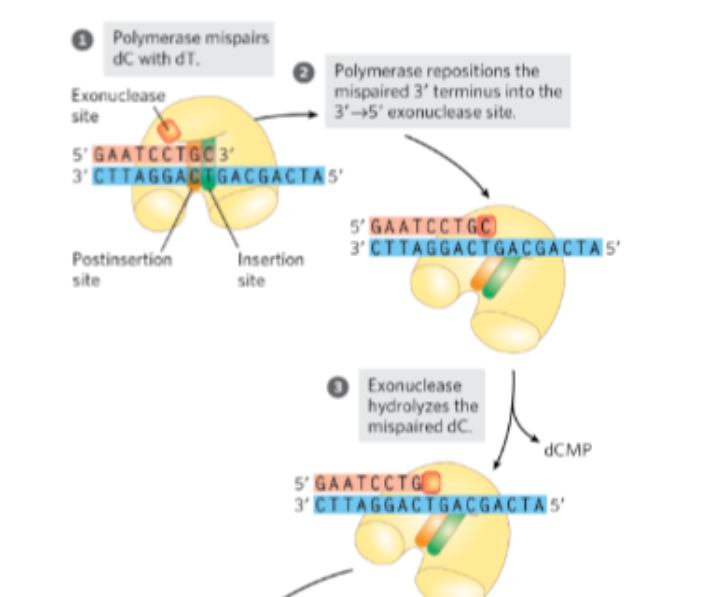
The DNA strand is then shifted to the second active site, which has 3’ → 5’ exonuclease activity, meaning it removes nucleotides from the end of the strand
What does this site do?
excises the incorrectly paired nucleotide, effectively “backing up” the synthesis.
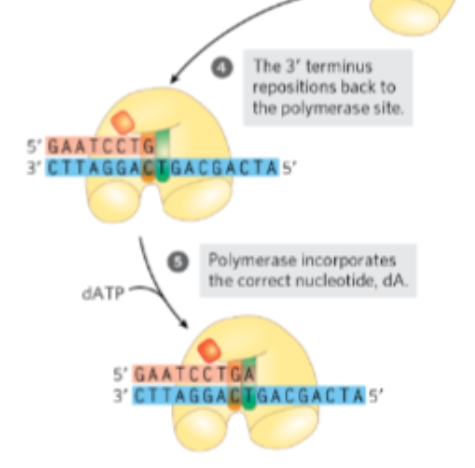
The DNA strand is then shifted to the second active site, which has 3’ → 5’ exonuclease activity, meaning it removes nucleotides from the end of the strand. This site excises the incorrectly paired nucleotide, effectively “backing up” the synthesis.
What happens after removal
the corrected strand is repositioned back into the catalytic site, and DNA synthesis resumes with the correct base. This proofreading step dramatically reduces the error rate of DNA replication
Truie or False: DNA synthesis is bidirectional
true
What sequence does DNA synthesis begin at
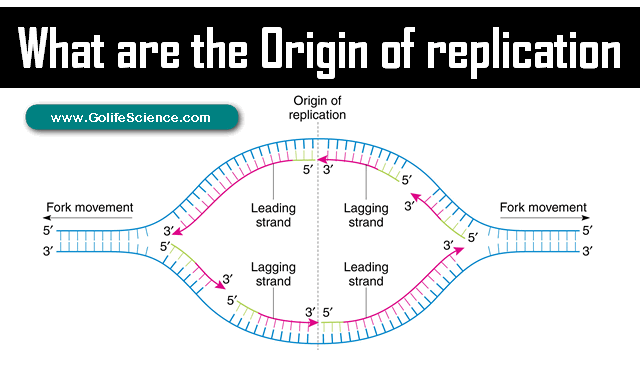
origin of replication
what strands seperate in dna synthesis
Parent strands
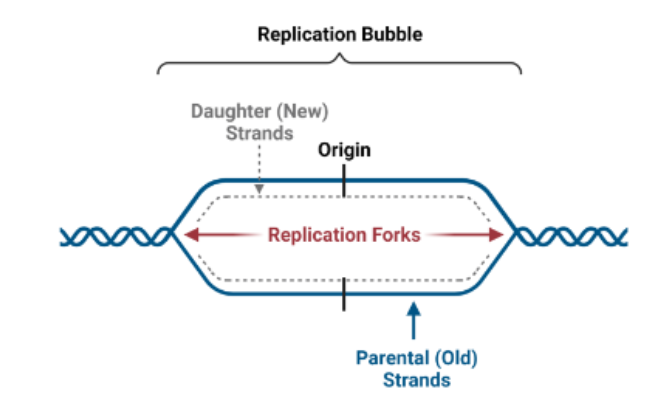
what does it mean for dna synthesis to be bidirectional
DNA is synthesized in both directions starting at the origin
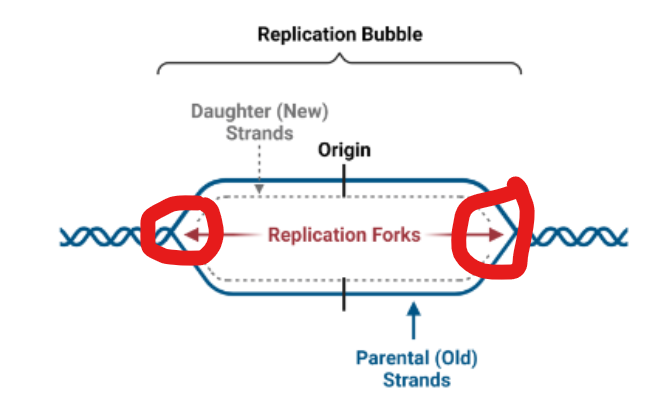
DNA is synthesized by DNA Pol at sites called
replication forks
what happens at a replication fork
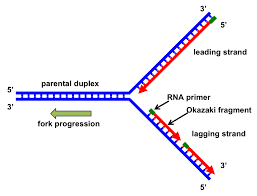
parent DNA is being used as a template for replication by DNA polymerase
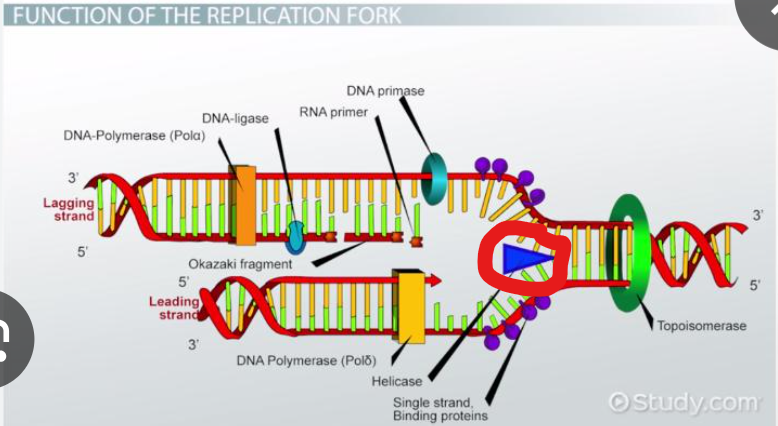
What is parent DNA first unwound by
helicase
Parent DNA is first unwound by helicase. What does it use?
ATP hydrolysis
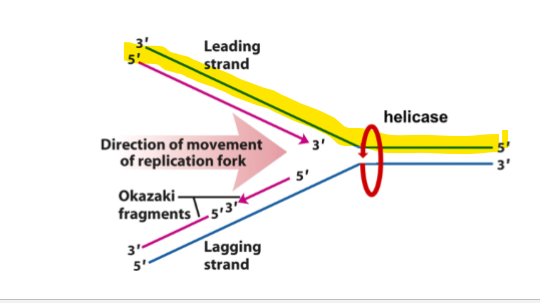
which strand is has DNA synthesis occurs continuously 5’→3’
leading strand
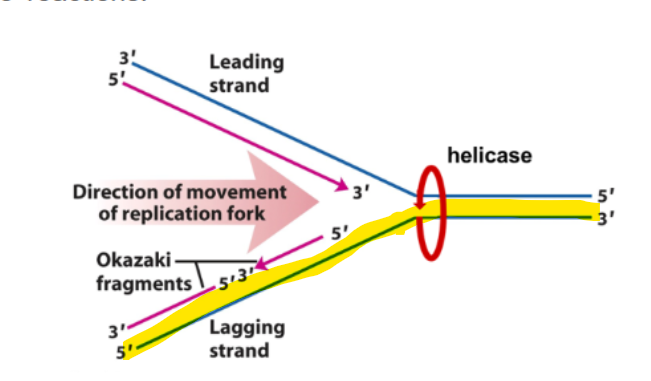
what strand happens in DNA synthesis occuring discontinuosly
Lagging strand
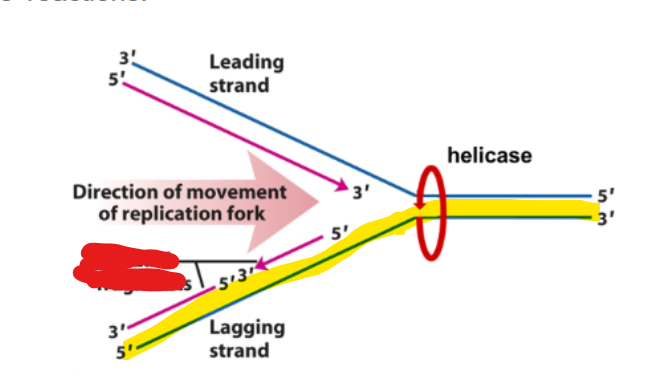
DNa synthesis occuring discontinously via the lagging strand is in what type of fragments as a series of 5’→3’ reactions
okazaki fragments
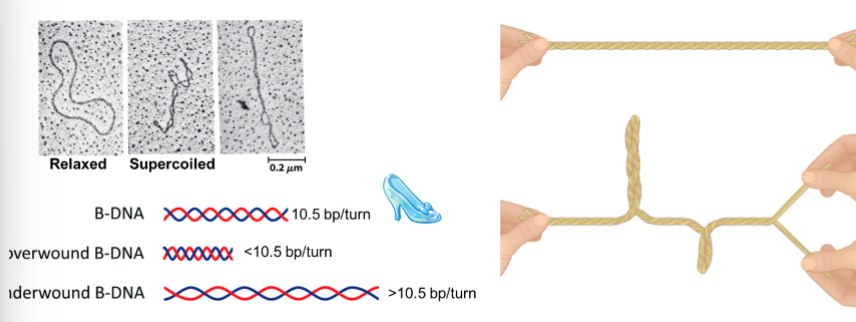
Under-winding or over-winding DNA causes torsional stress, what does this result in
the formation of supercoils
What are the enzymes that add/remove supercoils in DNA by cutting phosphdiester bonds in one or both strands, unwrap the helic, then reseal the strands
topoisomerase
Topoisomerases are the enzymes that add/remove supercoils in DNA by cutting what type of bonds in one or both strands , unwrapping the helix, and then resealing strands
phosphodiester bonds
Supercoiling can be helpful in what type of packaging
genome
Supercoiling can be helpful in genome packaging
what is used in bacteria that introduces negative supercoils to compavct the genome and also removes positive supercoils in front of replication forks
Topoisomerase II (DNA gyrase)
what are antibiotics that target bacterial DNA gyrase
fluoroquinolones
True or false: Fluoroquinolones blocks DNA gyrase (toposiomerase II) ability to reseal DNA
True
What are fluoroquinolones used to treat
to treat urinary tract infections, respiratory infections, and gastrointestinal infections
What type of trait does fluoroquinolones use that is selctive for bacterial enzymes
selective toxicity
Where does bacterial DNA replication initiation occur at?
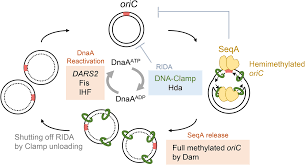
oriC
When does bacterial replication occur?
only once per round of cellular division
To make sure that replication in prokaryotes occurs only once per roound of cellular deivision- What is the most regulated step in bacterial dna replication
initiation
What is a unique 245 bp sequence in prokaryotes?
the origin of replication “oriC”
What are R1-5 and I1-3 binding sites for?
Used for DnaA proteins to intiate replication
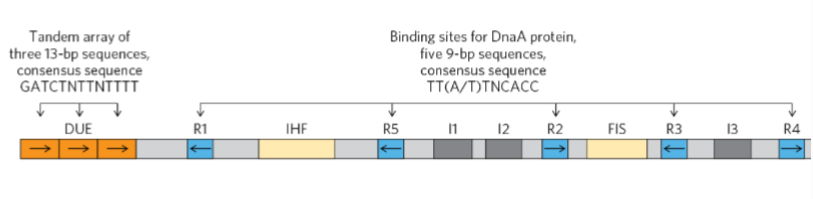
What is an AT-rich segement where strand seperation occurs for bacterial dna replication during initiation
DNA unwinding element (DUE)
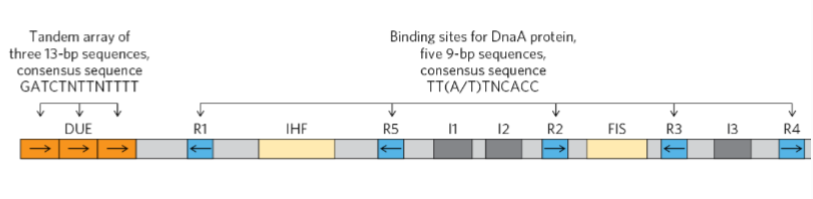
What are binding sites for proteins called replication initiation factors
IHF and FIS
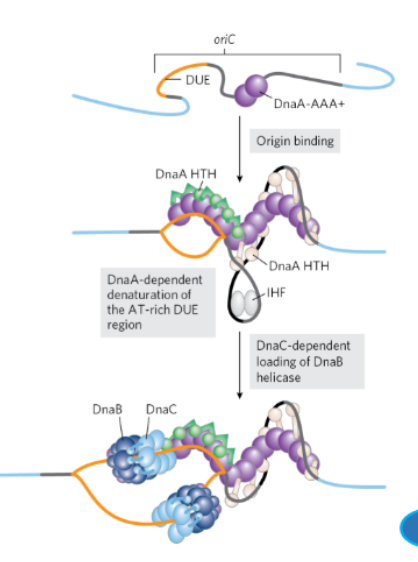
What are the steps of E. coli DNA Replication Initiation
DnaA binds ATP to become active
DNaC binds ATP and loads a DNaB helicase at both ends of the replication bubble
DNA polymerase and additional proteins are added to both DnaB helicases
Hydrolysis of the ATP bound to DnaA
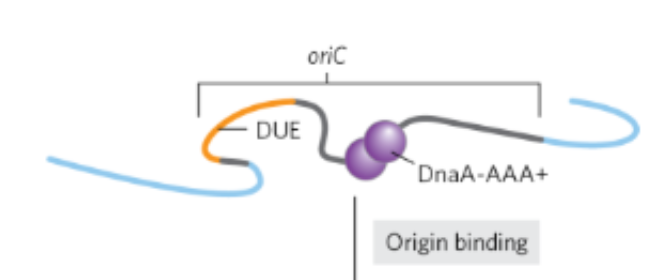
In the first step of E.Coli DNA replication initiation what happens
DnaA binds ATP to become active
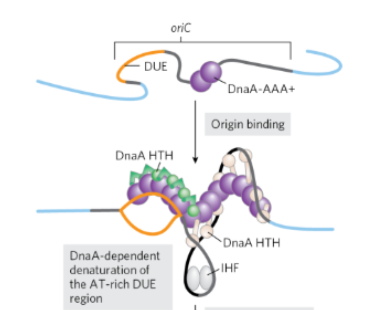
What happens once DnaA binds ATP to become active
DnaA protein binds to oriC→ psotive supercoil→ DNA denaturation at the DUE
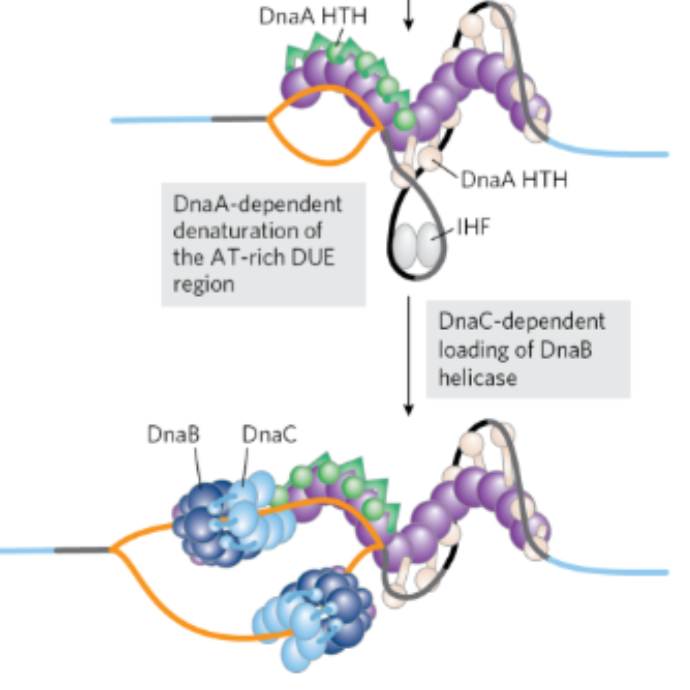
What happens once DnaA binds ATP to become active allowing for the DnaA proteins bind to oriC for a postive supercoil and dentauration at the DUE
DnaC binds ATP and loads a DnaB helicase at both ends of the replication bubble
WHat happens once DnaC binds ATP and loads a DNaB helicase at both ends of the replication bubble
helicase leads the replication fork seperating DNA strands 5’→3’ using ATP and DnaC dissociates
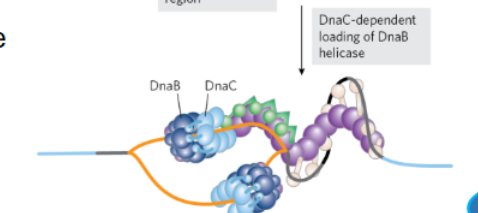
_________ and additional proteins are added to both DnaB helicases since a pre-existing single stranded DNA and primer is now available ( which is required for this enzyme)
DNA polymerases
What happens after DNA polymerase and additional proteins are added to both DnaB helicases
Hydrolysis of the ATP bound to DnaA (topoisomerase II)
What happens after hydrolysis of the ATP bound to DnaA
DnaA dissociates and is very slow to release ADP allowing for strict regulation
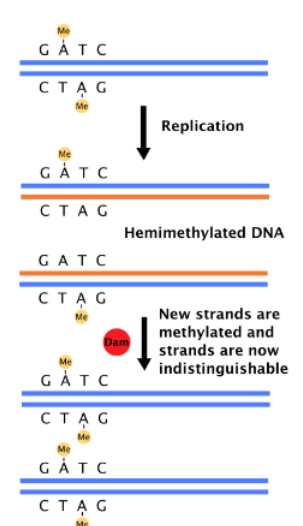
What regulates initiation
DNA Methylation
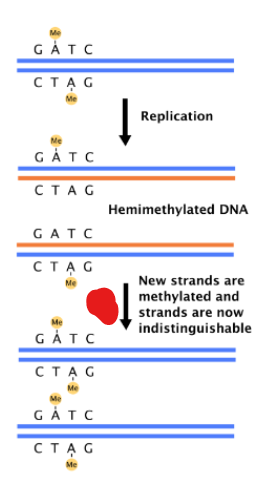
What is oriC DNA methylated by ?
Dam methylase
What does Dam mean ?
Dna Adenine Methylation
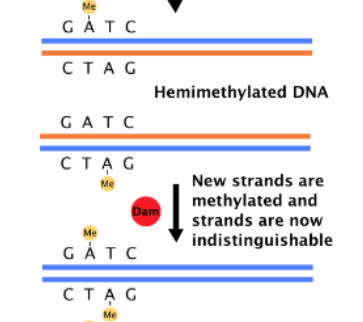
What does Dam methylate
it methylates N^6 position of A within (5’) GATC sequence
What location has a lot of methylated N^6 position of A within (5’) GATC sequence
the origin
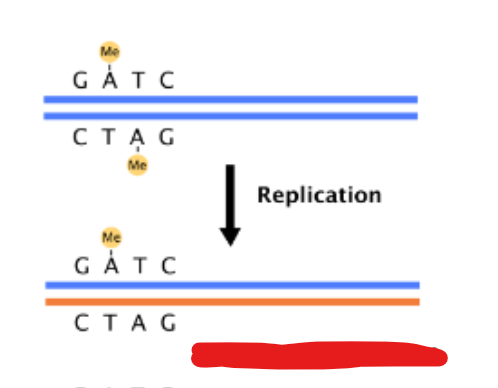
What happens after DNA replication
DNA is hemimethylated
What does hemimethylated oriC associated and sequestered with ?
plasma membrane
When can replication begin again
only after the DNA is fully methylated
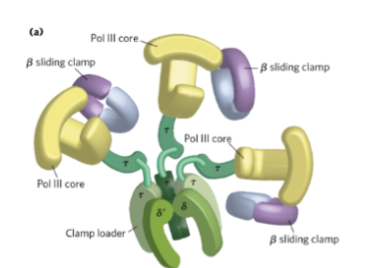
What catalyzes DNA synthesis
Core polymerase (Pol III)
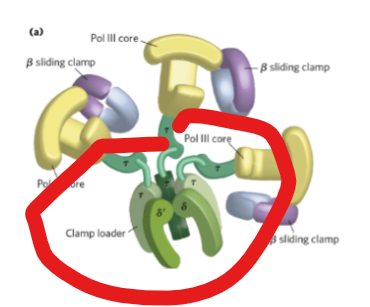
What serves as:
a scaffold for DNA polymerase III complex
to to assemble the β clamp onto DNA using ATP.
coordinates the replication fork by interacting with DnaB helicase through τ subunits.
the clamp loader
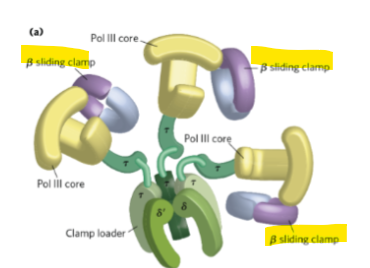
What is the β clamp (sliding clamp) on the Polymerase III holoenzyme ?
it tethers the core polymerase to DNA
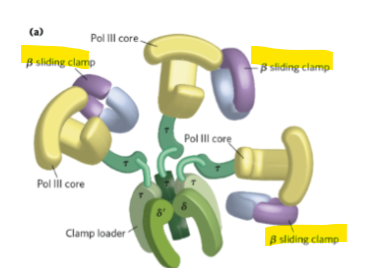
What does the β clamp (sliding clamp) decrease ?
it decreases the polymerase dissociation from DNA to increase processivity
What generates primers for Pol III
Primase
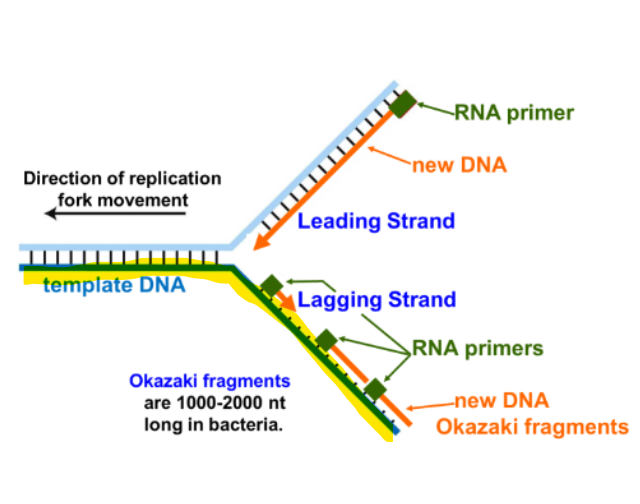
What does DNA polymerase III require
primed DNA template (3’-OH)
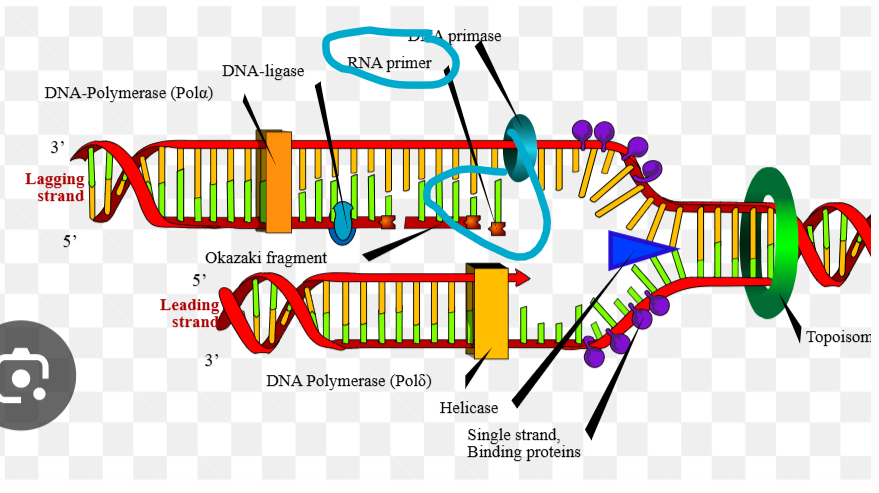
What is an RNA polymerase
a primase
True or False: Primase is DNA template-dependent
true
True or false: primase is is primer independent?
true
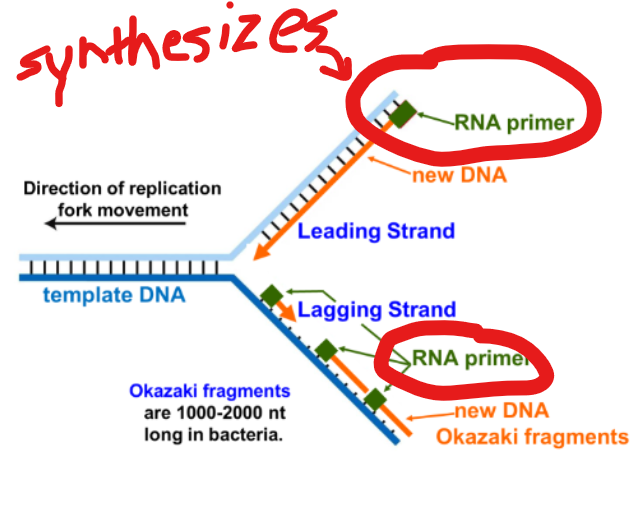
What does primase do?
it synthesizes a <9nt RNA primer at the beginning of the leading strand and each okazaki fragment
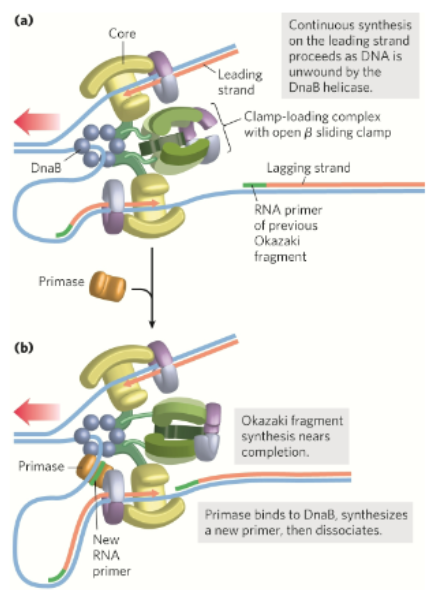
What is the basic lagging strand synthesis
DnaB helicase travels along the lagging template strand in the 5’→3’ direction and unwinds the DNA
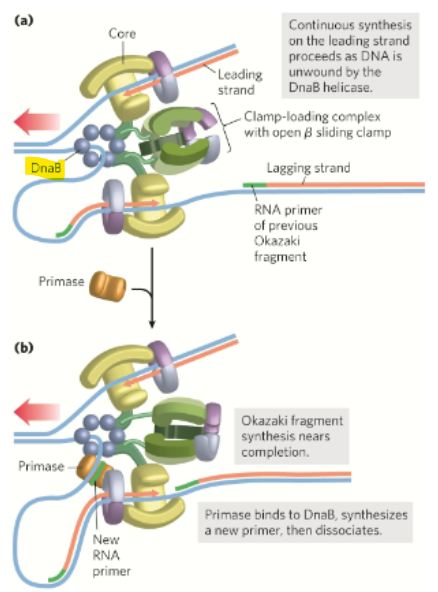
What travels along the lagging template strand in the 5’→3’ diorection and unwinds the DNA
DnaB helicase
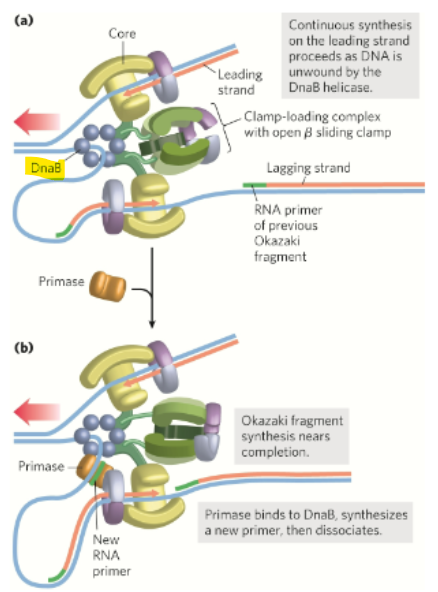
What does DnaB helicase travel along in the 5’→3’ direction to unwind the DNA
the lagging template strand
As DnaB helicase travels along the lagging template strand in the 5′→3′ direction and unwinds the DNA
what binds the single stranded DNA
single strand DNA binding protein (SSB)
As DnaB helicase travels along the lagging template strand in the 5′→3′ direction and unwinds the DNA
what occasionally associates with DnaB helicase and synthesizes a short RNA primer
DnaG primase
As DnaB helicase travels along the lagging template strand in the 5′→3′ direction and unwinds the DNA
where does primase function?
at the replication fork just as core polymerase nears completion of an Okazaki fragment
What happens after:
As DnaB helicase travels along the lagging template strand in the 5′→3′ direction and unwinds the DNA
A new β clamp is loaded onto the lagging strand at each new RNA primer by the clamp loader
Durin glagging strand synthesis what happens after:
A new β clamp is loaded onto the lagging strand at each new RNA primer by the clamp loader
A new β sliding clamp is loaded onto the lagging strand at each new RNA primer by the clamp loader
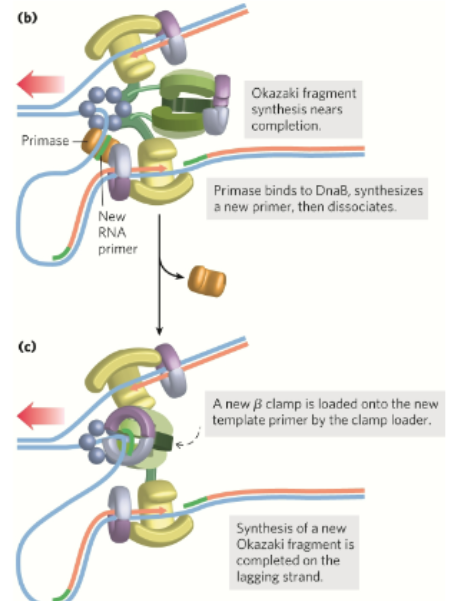
In lagging strand synthesis part 2 , what happens after:
A new β sliding clamp is loaded onto the lagging strand at each new RNA primer by the clamp loader
clamp loader binds ATP then sliding clamp
The clamp loiader opens the sliding clmap at one subunit interface
ATP hydrolysis closes the clamp around the DNA, and the clamp loader dissociates
In lagging strand synthesis part 2
after a new β sliding clamp is loaded onto the lagging strand at each new RNA primer by the clamp loader what closes the clamp around the DNA and the clamp loader dissociates
ATP hydrolysis