Molecular Genetics Final Exam
1/493
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
494 Terms
What amino acid is best for alpha helices?
alanine
What amino acid is the worst for alpha helices?
proline
What does mRNA do
codes for the protein starting with the start codon and ending with the stop codon
What does tRNA do
brings the amino acid and recognizes specific codons
What do ribosome subunits do
binds the mRNA
What happens if there is a mutation in the first or second codon
changes the amino acid, but doesn’t change the chemical nature
What do ribosomes do
build proteins by catalyzing peptide bond formation
What are the 3 sites involved in translation
A, P, E
What happens in the A site in translation
charged tRNA is loaded
What happens in the P site in translation
protein is formed
What happens in the E site in translation
uncharged tRNA is released
What is covalently bound to the 3’ end of a tRNA
amino acid
What are the 8 steps to making a tRNA charged?
1. Introns are spliced out
2. 5’ leader sequence cleaved by RNAseP
3. 3’ trailer sequence cleaved by RNAseZ
4. CCA is added to the tail of the tRNA
5. tRNA exported out of the nucleus into the cytoplasm
6. ATP binds amino acid to tRNA synthase
7. tRNA loads onto tRNA synthase
8. Anticodon recognizes tRNA
What is the wobble model?
When tRNA binds to more than one codon
In small amino acids, how does tRNA synthase work for kinetic proofreading
tRNA synthase has a pocket that only fits the small amino acid
In large amino acids, how does tRNA synthase work for kinetic proofreading
tRNA synthase has a large pocket that can fit small and large amino acids, second pocket that only fits small amino acid and cleaves it
What are the 5 steps for translation elongation
1. tRNA is in the P site at the start codon
2. tRNA is loaded into the A site
3. Peptide bond forms between amino acid in P site and amino acid in A site
4. tRNA’s shift 1 site upstream
5. E site loads
Help the ribosome move alone or position tRNAs
elongation factors
What does EF-Tu do
binds GTP and loads/positions the incoming tRNA
What are the 6 steps for EF-Tu to bind
1. tRNA binds anticodon
2. EF-Tu spends GTP
3. GTP converted to GDP
4. tRNA is moved into the A site
5. Tu binds to Ts
6. GTP cleaves Ts and reloads Tu
What does EF-G do
binds GTP and moves everything down
mRNA is translated in a ____________ way
non-overlapping
What are the 6 steps for bacterial translation initiation
RBS has a Shine Dalgarno Sequence upstream of AUG
Scans along 30S subunit to find the AUG
30S recruited to shine dalgarno
Proteins stabilize sequence fMet-tRNA is recruited to AUG site
Large subunit is recruited
Bacterial operons express large mRNAs with multiple ORFs with different polycistrons
What are the 8 steps for eukaryotic translation initiation
1. Ternary complex forms
2. 43S pre-initiation complex forms in the cytoplasm
3. 5’ cap binding proteins recruit 43S to 5’ end of mRNA
4. 43S scans along the RNA using ATP
5. 43S reaches AUG
6. 43s stalls at AUG
7. 48S initiation complex forms
8. 60S forms with the 48S into the 80S complex
What is the function of the kozak sequence
determines the AUG initiation
What happens when there is a strong kozak sequence
ribosome stalls and begins translation
What happens when there is a weak kozak sequence
AUG is skipped, translation does not start
genes that only produce 1 protein
monocistronic
What is an example of a monocistronic gene
mRNA
What does the release factor do in translation
cleaves the peptide chain off of the tRNA in the P site
What are the 8 steps to translation termination
1. RF1 binds
2. Polypeptide released
3. RF3 dissociates
4. RRF binds to the stop codon
5. RRF interacts with EFG
6. tRNA released in the E site
7. EFG does GDP hydrolysis
8. IF3 causes the ribosome to dissociate
What 3 components are required for translation
mRNA, tRNA, ribosome subunits
What is the component that actually carries out translation
ribosome
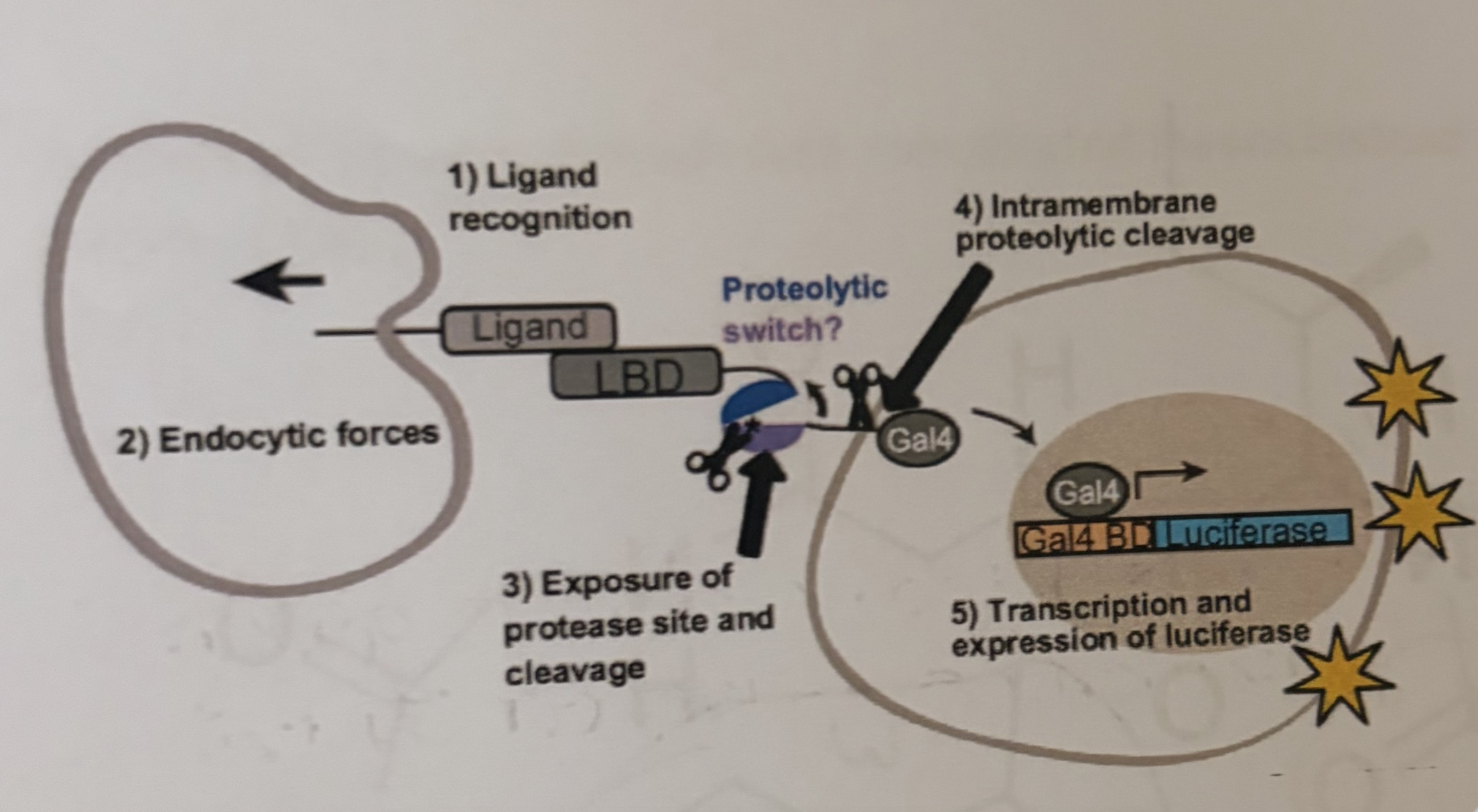
Give an example of a:
Enzyme
DNA Binding protein
Allosteric interaction
Active transport
Enzyme - Protease
DNA binding protein - Gal4
Allosteric interaction - exposure of protease site and cleavage
Active transport - Gal4 entering the nucleus
What type(s) of polymerase synthesize 3’ to 5’
None
Why is it okay that RNA polymerase has worse fidelity
the errors produced during transcription not inherited by offspring
In eukaryotes, how are the ends of chromosomes differentiated from double stranded breaks
Chromosome ends are protected by telomeres
What does DNA ligase do in DNA replication
seals breaks in the backbone by forming phosphodiester bonds
What does gyrase do in DNA replication
relieves supercoiling ahead of the replication fork by introducing negative supercoils
Which eukaryotic RNA polymerase produces the most RNA in the cell
RNA polymerase 1
How does RNA polymerase synthesis occur?
5’ to 3’
What does RNA polymerase 2 do?
Transcribes mRNA
What does RNA polymerase 1 do?
Encodes small RNAs that don’t need caps and are not meant to be translated (rRNA and micro RNA)
What does RNA polymerase 3 do?
Encodes for small RNAs and histones
What is an operon?
a series of genes that function together
What does RNA polymerase need to function?
DNA template, rNTPs, Mg²⁺/Mn²⁺, promoter, transcription factors
What are the 5 steps of transcription initiation through RNA polymerase
Polymerase and sigma factors bind promoter
DNA unwinds
Open complex forms
First rNTP pairs with template
Phosphodiester bond forms
How does RNA elongate
Polymerase moves 5’→3’, adds rNTPs complementary to template, DNA rewinds behind
How does transcription stop in bacteria
rho-dependent or rho-independent
How does transcription stop in eukaryotes
polyadenylation signal → RNA cleaved and released
what nucleotides does RNA polymerase use
ATP, GTP, CTP, UTP
How do mistakes occur by RNA polymerase
Wrong rNTP pairs with DNA template because RNA polymerase has no strong proofreading like DNA polymerase
How does RNA polymerase fix mistakes (2 ways)
Backtracking: Polymerase moves backward on the RNA.
Cleavage of RNA: The misincorporated nucleotide is cut out.
How are transcripts modified by RNA polymerase?
5’ capping, 3’ poly-a tail, splicing
Why is a 5’ cap important?
RNA stability, translation, and transporting RNA out of the nucleus
Why is a 3’ poly-a tail important
mRNA stability
What does RNA splicing do?
Determines which codon is being used
4 steps to operon function
RNA pol binds promoter
Repressor binds operator
Activator binds DNA
Structural genes code proteins
In the lac operon, regulation of _____ is done by the control of ____________
Initiation, RNA polymerase binding
In the trp operon, regulation of ________ is done by _________
elongation/termination, attenuation
What does attenuation cause
premature termination of transcription under certain conditions
a sequence consisting of two identical or highly similar inverted repeats which are either adjacent to one another or separated by a spacer region
Palindrome
What is an example of a palindrome in bacteria
Transcription factor binding sites
In the lac operon, what level of gene expression is present when there is low glucose, high cAMP, no lactose
None
In the lac operon, what level of gene expression is present when there is : high glucose, low cAMP, present lactose
Low
In the lac operon, what level of gene expression is present when there is low glucose, high cAMP, present lactose
High
How is bacterial initiation regulated?
Sigma factors, activators, and repressors
How do repressors work in bacteria?
Bind to the operator and prevent RNA polymerase from initiating transcription
What inactivates repressors in bacteria
Inducers
How do activators work in bacteria?
Bind to DNA near the promoter and help RNA polymerase bind
What do activators in bacteria need to function
Co-activator or ligand
What is the default state of a promoter in bacteria?
On
How is eukaryotic initiation regulated? (5 things)
Promoter
Activators/Repressors
Mediator
Chromatin
Coactivators/Corepressors
What is the promoter in eukaryotic transcription and what does it do
TATA box ; recruits RNA Pol II
What do activators/repressors do in eukaryotic transcription
Bind enhancers/silencers to boost or block transcription
What does the mediator do in eukaryotic transcription
Bridges TFs to RNA Pol II and integrates signals
What does chromatin do in eukaryotic transcription
Histone modifications and nucleosome remodeling control accessibility
What do coactivators/corepressors do in eukaryotic transcription
Modify chromatin or stabilize polymerase
What do activators do in eukaryotes
Bind enhancers to help RNA Pol II initiate transcription
What do repressors do in eukaryotes
Bind silencers to block transcription
What is the default state of a promoter in eukaryotes
Off
How is the beginning of the transcript modified?
5’ capping
Describe Rho-independent transcript termination in bacteria (3 steps)
RNA forms a hairpin loop and poly-U sequence
Hairpin destabilizes RNA-DNA hybrid
Polymerase releases RNA
Describe Rho-dependent transcript termination in bacteria (5 steps)
Rho protein binds rut site on RNA
Moves along RNA in the 3’ direction
RNA polymerase reaches the termination site
Stem-loop causes RNA polymerase to pause
Helicase separates RNA from DNA
How is the end of the transcript modified (5 steps)
polyA signal recognized
RNA cut downstream of signal
PolyA polymerase adds A’s
Poly-A binding proteins bind tail
Rat1/hXrn2 degrades leftover RNA
Short single-stranded DNA or RNA that bind to specific targets with high affinity and specificity
Aptamers
Aptamer tied to a regulatory structure; Regulate gene expression when there are environmental or cellular changes
Riboswitches
Regulatory region of 100,000 base pairs upstream of the promoter; Lots of space that influences if the region is on or off
INK/ARF locus
bind activators and transcription factors to start transcription
Enhancers
How does splicing occur mechanistically? (7 steps)
2’OH attacks 5’ splice site
Exon 1 released
2’OH forms covalent bond with 5’ end of exon
3’OH attacks 3’ splice site
Exon 1 and exon 2 link
Intron lariat released
Spliced RNA processed
Why does splicing occur? (3 reasons)
Removes non-coding introns
Joins exons together to produce a continuous open reading frame
Allows alternative splicing
What is alternative splicing?
A process where different combinations of exons are joined from a single pre-mRNA
Describe the 2 types of alternative splicing
Minor spliceosome - recognizes AU5’ and AC 3’
Self-splicing introns - introns that splice themselves with only RNA, no proteins involved
How does alternative splicing affect transcripts?
allows a single pre-mRNA to produce multiple mature mRNA isoforms
removed pieces of RNA
intron
kept and translated during RNA processing
exon
large complex that does the splicing
spliceosome
splice donor
5’ splice site
splice acceptor
3’ splice site
proteins coded from the same gene but are spliced differently
isoforms