Bio 3C Lab Quiz 3
1/166
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
167 Terms
what do restriction enzymes do
recognize and cut both strands of a specific sequence of DNA
what naturally makes restriction enzymes
Bacteria as an anti-viral response
what do we call viruses that infect bacteria
Bacteriophages
what is a recognized sequence is regards to restriction enzymes
a specific short DNA sequence that the enzyme can identify and bind to before cutting
recognized sequences are typically what?
palindromic/ an inverted repeat
EX: 5’C-C-G-C-G-G 3’
3’ G-G-C-G-C-C 5’
how can you tell if a cleavage product is sticky or blunt
sticky is if the cut is not the same place, blunt is if it is as the same place
in the photo top is blunt and bottom is sticky
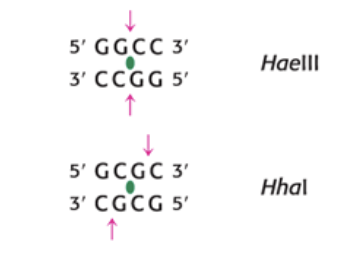

DNA fragments generated by resection enzyme digest can be separated by what
gel electrophoresis in agarose or polyacrylamide gels
Recombinant DNA general steps
cleave plasmid vector with a restriction enzyme (commonly EcoRI), cleave the insert (DNA fragment) with the restriction enzyme, anneal and fragments and rejoin using the enzyme DNA ligase
how can eukaryotic genes be expressed in bacteria
via reverse transcriptase
what is cDNA
complementary DNA made from mRNA via reverse transcriptase
only thing that naturally makes reverse transcriptase
retroviruses
example use of reverse transcriptase
the creation of proinsulin cDNA from proinsulin mRNA, the cDNA is then transcribed and inserted into a recombinant plasmid and bacteria are transformed
when making cDNA why do we use mRNA instead of DNA
because bacteria lack the machinery to splice introns out of the primary transcript which it made out of RNA nucleotides
Steps of reverse transcriptase
an Oligo(T) primer (linked thymidine) binds to the mRNA at the 3’ end, then reverse transcriptase creates the cDNA following the mRNA template, once the cDNA is complete the mRNA is hydrolyzed, then terminal transferase adds nucleotides to the 3’ end of the single strand cDNA which allows for DNA polymerase to come in and create the secondary complementary strand leading to a double-stranded cDNA
— comes hand in hand with —-
DNA analysis, sequencing DNA molecules
how is DNA sequencing done
the Sanger Dideoxy Method which is the controlled termination of replication
Sanger Dideoxy Method described, not modern
four tubes each with the same template DNA to be sequenced, primer, DNA polymerase, and a different labeled ddNTP (dideoxy nucleoside triphosphate). DNA polymerase will do its thing but randomly it will add a ddNTP and the reaction will stop. Each tube will have a variety of fragment in it that can then be isolated and examined via gel electrophoresis. these fragment correlate to the position of the nucleotides in the complementary strand
what are ddNTP’s
dideoxy nucleoside triphosphate, they are called a 2,2’,3,3’-dideoxy analog. They do not have an “OH” at carbon 3 so DNA polymerase cannot add onto it and replication will not continue
how are ddNTP’s labeled
either radiolabled or fluorescently
how do you read a Sanger Sequencing gel?
Bottom to top, bottom is the 5’ end and top is the 3’ end of the complementary stand. The Template strand is then the paring of that strand
How is modern Sanger Sequencing different?
each ddNTP has its own colored fluorescent label to it so everything can be done it one tube and on the gel we can organize the sequence based on the fluorescent color
PCR stands for what
polymerase chain reaction
PCR components
pair of primers that hybridize with the flaking sequences of the target DNA sequence, all four dNTP’s, and a heat stable DNA polymerase (often Taq)
what is the PCR cycle
strand separation at 95°C, hybridization of primer around 54°C, DNA synthesis at 72°C, and this all repeated about 30-40 times
in PCR during hybridization of primers why do parent duplexes (original DNA strands coming back together) not form
because the primers are present in large excess
How can the quantity of individual mRNA transcripts be determined?
through quantitative PCR aka real-time PCR
qPCR stands for what
quantitative polymerase chain reaction
what does qPCR monitor
DNA amplification as the reaction progresses
in qPCR how is the reaction progression monitored
by an increase in fluorescent signal which is created by SYBR Green 1 dye which binds to the DNA. It fluorescences brighter in dsDNA than in ssDNA, so there is an increase in fluorescence as the PCR reaction continues and dsDNA is formed
During/after the qPCR cycle what is made
a qPCR curve
phases of a qPCR curve
lag, exponential, linear, and plateau
what can you find from a qPCR curve
the CT which is the cycle threshold. This is the number of PCR cycles is takes for the fluorescence signals to cross a certain threshold and become detectable above background noise.
what does a low or high CT mean
A lower CT means there was more original template present so it reached the threshold faster, a higher CT means there was less original template present so it reached the threshold slower
what are DNA microarrays used for
to investigate the pattern and level of expression of all genes in a particular cell or tissue (which is the transcriptome)
How is a DNA microarray set up
mRNA is isolated and then converted to cDNA in the presence of fluorescent dNTPs (different colors for different samples), then you separate the cDNA into single strands and add it into the wells of a microarray, each well in the microarray holds a 20nt DNA sequence called an oligonucleotide, the cDNA will bind to any complementary sequences and anything unbound will be washed away. The remaining bound parts will hybridize and be visible after a scan due to their fluorescence. The spots will glow green, red, yellow, or not at all.
in a DNA microarray after the wells are scanned what do the different fluorescent colors signify
Green: decreased expression of a particular gene
Red: increased expression of a particular gene
Yellow: high expression in both samples
None: no hybridization/binding
what does CRISPR stand for
Clustered Regularly Interspaced Short Palindromic Repeats
in CRISPR describe Cas9 and guide RNA
Cas9-guide RNA complex is what binds to the complementary sequence of the target gene, then Cas9 cuts both DNA strands
when Cas9 cuts the DNA what happens to the broken strands
they are repaired by the cell. When they are repaired scientists can disable the target gene to study its normal function. Or the target gene with a mutation can be repaired by providing a normal copy of the gene
what is sgRNA
single guide RNA, an engineered form of guide RNA that forms a complex with Cas9, it is about 100nt in length and has two regions
What are the two regions of sgRNA
Guiding region: part of CRISPR RNA (crRNA in nature) that is typically a 20nt region complementary to the target DNA sequence and it is customized for each target of interest
Scaffold region: tracrRNA in nature, forms a multi-hairpin loop structure (scaffold) that binds tightly to a crevice in the Cas9 protein, this region is typically the same for all sgRNA
In order for the Cas9 complex to cut DNA what must occur
It first has to bind to a common sequence in the genome called a PAM (protospacer adjacent motif), then the sgRNA unwinds a part of the helix and binds to its complement, then Cas9 can cut the DNA
How can Cas9 be altered in CRIPSR to achieve alternate functions
Cas9 can have one or both cutting domains deactivated and an enzyme can be fused onto it. The Cas9 complex than acts as a transport for that enzyme to a specific DNA sequence. An example of this is fusing deaminase onto it which mutates specific DNA bases such as converting Cytidine into Thymidine
Cas9 can also be fully deactivated and then altered to promote gene transcription. This is done by adding proteins that activate transcription. This can be done by directly adding the proteins, by first adding peptides to which these proteins are added, or by recruiting these proteins to the gRNA
Blotting techniques? Names, what they look for, how they probe?
SNOW DROP
Southern Blot- looks for DNA and probes with DNA
Northern Blot- looks for RNA and probes with DNA or RNA
Western Blot- looks for proteins which binds to antibodies
CRISPR stands for
Clustered Regularly Interspaced Short Palindromic Repeats
where was CRISPR found
in bacteria as a defense mechanism
what are the two steps in gene editing
cutting the double-stranded DNA at a desired location and directing DNA repair to produce a desired sequence change
what happens to a bacterial cell when chromosomal DNA is cut
the cell will die unless the cut is repaired
How can cells repair double stranded breaks in DNA
Non-homologous end joining (NHEJ): Specific proteins reconnect the ends of the double stranded break back together. This process can randomly insert or delete one or more based which can lead to mutations that disrupt gene function or expression.
Homology Directed Repair (HDR): Enzymes patch the break using donor template DNA. Researchers design the donor template DNA which can include a desired sequence added onto both ends called a homology arm. These homology arms match the sequence upstream and downstream of the cut. A complementary DNA strand in created during repair.
What did we personally use CRISPR-Cas9 to do in lab
cut bacterial chromosomal DNA at a specific location within the lacZ gene, then take advantage of the cell’s HDR to create a desired change in the lacZ gene. This change was due to the donor template DNA have a stop codon in it to disrupt gene function.
In the CRISPR lab what was the E. coli used for
The E. coli strain was HB101-pBRKan (often called pBRkan), this strain held the Cas9 enzyme and it carried a plasmid containing a kanamycin resistance gene, a naturally functional lacZ gene, and a gene for HDR that was controlled by an arabinose-inducible promoter (so only with arabinose present will bacterial takes be able to take up donor template DNA to preform HDR and fix the double strand breaks)
What is Cas9
a protein nuclease that uses an RNA guide to cut a specific sequence of DNA
what was our Cas9 in the CRISPR lab engineered to cut
only at the lacZ gene
what is the lacZ gene
gene that codes for the enzyme β-galactosidase (β-gal) which is part of the lac operon. It allows for E. coli to break down lactose, specifically it can also hydrolyze a sugar known as X-gal which when hydrolyzed leads to a blue pigment. Through this we can use blue-screening to look for blue colonies and determine is the lacZ gene is functional
what does Cas9 need in order to cut
a guide RNA that will bring it to the specific target DNA sequence
in the CRISPR lab how did we transform the E. coli
We had two different plasmids with different contents but both had a spectinomycin (SPT) resistance gene
pLZ Donor (pD): has the donor template DNA that can be used by HDR which has a stop codon to stop lacZ’s function
pLZ Donor Guide (pDG): has the donor template DNA and a sequence that codes for the sgRNA, once the bacteria has this plasmid it can transcribe the sgRNA gene then the sgRNA can guide Cas9 to cut at the lacZ gene
in the CRISPR lab what two plasmids are present and what is in each
HB101-pBRKan plasmid with kanamycin resistance, lacZ gene, and Cas9 gene.
pLZDonor or pLZ Donor Guide with spectinomycin resistance, donor template DNA and/or not sgRNA gene)
In the CRISPR lab when picking out colonies from the starter plates, what type of starter plates did we have and what was in them
Starter Plate IX: which has IPTG (for the I) to allow expression of the lac operon (lacZ transcription), and X-gal (for the X) which is the chromogenic substrate of β-galactosidase and results in blue colonies. No arabinose so no HDR
Starter Plate IX/ARA: which has IPTG (for the I) to allow expression of the lac operon (lacZ transcription), and X-gal (for the X) which is the chromogenic substrate of β-galactosidase and results in mostly blue colonies. Arabinose allows for activation of HDR
what is IPTG
Isopropyl β-D-thiogalactopyranoside, which is an analog of allolactose and used to induce expression of the lac operon by binding to the repressor protein so that DNA Polymerase can bind to the promoter and the lacZ gene is able to be transcribed
In the CRISPR lab what do these plates under these conditions look like?
E. coli + pLZDonor (pD) on IPTG/X-gal Starter Plate (IX)
E. coli + pLZDonorGuide (pDG) on IPTG/X-gal Starter Plate (IX)
E. coli + pLZDonor on IPTG/X-gal/Arabinose Starter Plate (IX/ARA)
E. coli + pLZDonorGuide on IPTG/X-gal/Arabinose Starter Plate (IX/ARA)
Blue colonies, no cutting of the lacZ gene
Cutting of the lacZ gene, but no arabinose so no repair, so no colonies (chromosomal DNA when cut and not repaired dies)
Blue colonies, no cutting of the lacZ gene
White colonies, lacZ gene is cut but donor template DNA is inserted and results in the stopping of the lacZ gene so no β-galactosidase can hydrolyze X-gal and make blue pigment
CRIPSR. Once we have the transformed bacteria and want to grow them what do we plate them on and why?
On plates with spectinomycin so that only bacteria that as taken in the pD or pDG plasmid will survive and grow
Why did the starter plates have kanamycin on them?
to ensure that only E. coli of the HB101-pBRKan strain with the pBRKan plasmid would grow
if you grew HB101-pBRKan on a plate with arabinose and then moved them what would happen
if grown on arabinose they will keep the enzymes needed for HDR even if moved, but their daughter cells will not produce those HDR enzymes unless exposed to arabinose
We can use CRISPR for two diseases, what and how?
Sickle-cell anemia and transfusion dependent beta-thalassemia which both use Exa-cel. Exa-cel stands for exagamglogene autotemcel. This cuts out the BCL11A gene, which prevents the production of fetal hemoglobin, and it edits the enhancer region. Once the BCL11A edited gene is put back in it undergoes less transcription so the transcription of fetal hemoglobin can occur. As a result there is more fetal hemoglobin with two alpha and two gamma subunits instead of two alpha and two beta subunits as in adult hemoglobin.
what does ELISA stand for
Enzyme Linked Immunosorbent Assay
what is immunology
the study of the immune system
what are the two main branches of the immune system
Innate Defenses: Nonspecific and have a variety of factors such as normal microbiota, mucous membranes, phagocytic cells, fever, and antimicrobial substances
Adaptive Defenses: Specific to each pathogen and involve the production of antibodies. Antibodies location and attache themselves to their target and facilitate the destruction and removal of pathogens, they are highly specific to their targets/antigens
How are animals involved with antibodies
Animals can be injected with a specific antigen and their body will make antibodies against that specific injected antigen which is located in their serum. We can harvest and isolate that antibody.
What are primary antibodies
Antibodies that recognize and bind to antigens
What type[s] of antibodies does ELISA use
primary and secondary antibodies
How can we produce secondary antibodies
Same idea as primary. We inject animals with the human primary antibody, they will make antibodies against the primary antibody which we can then harvest and isolate
What are secondary antibodies
antibodies that bind to the primary antibody which is already attached to the antigen. They are often enzyme-linked
what is another term for an enzyme linked secondary antibody
a labeled secondary antibody
In our ELISA lab was the enzyme that the secondary antibody was linked to and its substate
the enzyme is HRP, horseradish peroxidase. And the substate is TMB, 3,3’,5',5’-tetramethylbenzidine
What is HRP and what is its substrate/ does to it
horseradish peroxidase, the enzyme linked to the secondary enzyme in the ELISA lab. its chromogenic substrate is TMB: 3,3’,5,5’-tetrametylbenzidine, which gets oxidized and forms a blue product
what does TMB stand for/do
3,3’,5’,5’-tetramethylbenzidine, it is the substrate for the enzyme linked to the secondary antibody, it is light sensitive
what type of ELISA did we use and what does that mean
we used an indirect ELISA. This means we start with antigen bound to the wells, then the primary antibody is added and binds to the antigen, then the secondary enzyme-linked antibody is added which binds constant chain of the already bound primary antibody, substrate is added and then colored product is formed
What is direct ELISA
sandwich ELISA. When primary antibody is first bound to the well then antigen is added which binds to the primary antibody, then the enzyme-linked antibody specific for the antigen is added and binds, substrate is added and colored product is formed
What is the point of a blocking buffer in ELISA
used to stop any non-specific binding by using an irrelevant protein or by using a non-ionic detergent
in our ELISA lab what blocking buffer did we use, where do we find this?
a non-ionic detergent called Tween-20, found in our wash buffer
in our ELISA lab what kid did we use to test for what
we used a kit from Bio-Rad to determine if mock parents have HIV viral proteins in their blood
what do actual HIV texts look for
anti-HIV antibodies since virion numbers will be low in the serum but antibody levels will be high
in our ELISA lab what was in the wash buffer and what did it do
Has 10% Tween-20 which is an non-ionic surfactant that removes anything unbound while also binding to the plastic (polystyrene) wells and saturating biding sites as to avoid nonspecific binding
in our ELISA labs what are the wells made out of and why
Polystyrene which binds to proteins by hydrophobic interactions
in ELISAs why is it important to not have non-specific binding
if antibody is bound non-specifically to the well the that means it will be present even if antigen isn’t present which can lead to a false positive result
in ELISAs why do we wash between each step
to remove anything that is unbound and prevent previous components from reacting in later steps and possibly causing a false positive
What happens in an ELISA when you add a primary antibody if the antigen is and isn’t present
if the antigen is present and you add primary antibody, the antibody will bind specifically to the antigen creating an antigen-antibody complex that will not wash off
if the antigen is not present and you add primary antibody there will be no binding and no creation of an antigen-antibody complex so the primary antibody will be washed away
What are three other immunoassays other than ELISA
radioimmunoassays, western blot, lateral flow assays
In order for an ELISA to determine if there is a positive result what do we need to add
a chromogenic substrate in the end to produce a colored positive result
in the ELISA lab what antigen, primary antibody, and secondary antibody did we use exactly
Antigen: chicken gamma-globulin (purified egg yolks)
Primary antibody: polyclonal rabbit antibody raised against chicken gamma-globulin
Secondary antibody made from goats immunized with rabbit IgG
In ELISA what can lead to false positives or negatives
False positive can result from recent vaccinations. False negatives can result from recent immunosuppression, or administering the test too soon after infection with HIV
Antibody structure?
Heavy inside chain and light outside chain, variable (top four section) region and constant (bottom eight section) region, FC and Fab region. The part of the antibody that binds to and recognizes the antibody is called the epitope.
Antigen defintion
a substance that causes the body to produce specific antibodies or sensitized T cells
what are immunoglobulins
globular proteins, specialized proteins called plasma cells in the immune system
what is valence in terms of antibodies
the total number of antigen binding sites on the antibody
How are B cells activated
Antigen binds to receptor, B cells take in the antigen, B cells then present pieces of the antigen on the surface via MCH Class II molecules, Helper T cells then binds to the MCH Class II molecules, Helper T cells release cytokines, those cytokines then fully activate the B cells which then differentiate into plasma or memory cells
what are the classes of immunoglobulins, what determines that class
IgG, IgM, IgA, IgD,IgE. This is determined by the amino acid sequence of the constant region of the heavy chains
what are the protective mechanisms of binding antibodies to antigens
Agglutination: Antibodies bind to multiple antigens at once and clump pathogens together to make them easier for immune cells to detect and remove since they are grouped and not spaced out
Opsonization: Antibodies coat a pathogen and act like a tag that allows for immune cells (like macrophages) to recognize the antibody and phagocytose the pathogen more easily
Neutralization: Antibodies bind to viruses or toxins and block them from entering or damaging cells. They inactive the threat without destroying it.
Antibody-dependent cell-mediated cytotoxicity: Antibodies attach to infected or abnormal cells, then immune cells (like natural killer cells) recognize those antibodies and kill the targeted cell
Activation of Complement: Antibodies trigger the complement system which is a group of proteins in the blood that leads to cell lysis, increased inflammation, and increased opsonization
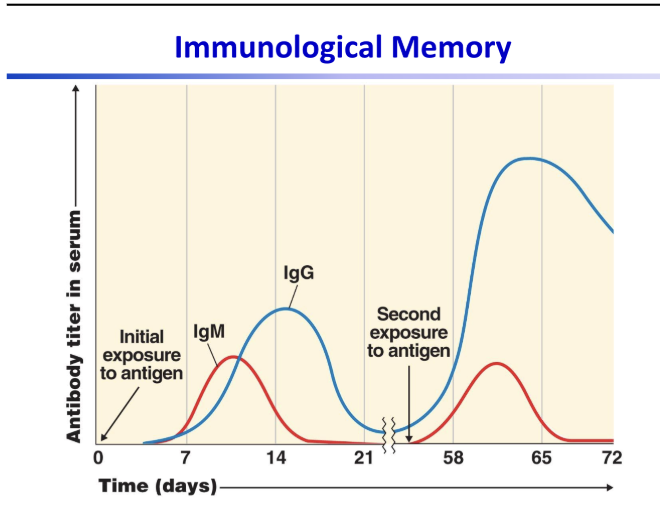
understand the idea of this graph

the first exposure takes longer to produce antibodies and makes less of them. the second exposure creates antibodies way faster and produces way more of them.
what are common blocking agents besides the Tween-20 used in our ELISA
non-fat dry milk, gelatin, and bovine serum albumin