PCB3063 EXAM THREE
1/94
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
95 Terms
Cancer
uncontrolled cell growth and spread of abnormal cells
Cyclin-dependent kinases & Ras-signaling pathway
two mechanisms involved in the regulation of the cell cycle
Protooncogenes
stimulate cell division
mutations cause increased cell growth (gain-of-function)
Tumor Suppessor Genes
stop the cell cycle, DNA repair, or induce apoptosis
mutations cause damaged cells to continue growing and dividing
Basal Lamina and Attachment Proteins
mutations allow cells to break away from their original tissue
DNA repair systems
cells accumulate more mutations due to lack of repairs
Cell identity
cancer cells can have an abnormal appearance and invade other tissue
Benign Tumors
form noncancerous, single multicellular mass
due to mutations in TS genes and proto-oncogenes
Malignant Tumors
benign cells break away and starts multiplying
invade other tissues and can cause life-threatening problems
Hyperplasia
increase in cell number
Dysplasia
abnormal cells
Metastasis
cell migration, breaking free from tissue
Malignancy
cell invasiveness
Driver Mutations
gives a growth advantage to a tumor cell
Passenger Mutations
no direct contribution to the cancer phenotype and is acquired over time during cell division
XP
involved in nucleotide excision repair and mutations lead to skin cancer
FAP
genetic predisposition to colon cancer due to:
mutation in APC gene
tumor-suppressor controlling cell-cell contact and growth inhibition
mutation in MUTYH gene
gene involved in DNA repair and mutations can cause loss of heterozygosity
pRB
phosphorylated to obtain transcription factor E2F
E2F activated transcription of genes required for the cell cycle like cyclins
Hardy-Weinberg Assumptions
no natural selection
random mating
no new alleles from mutations
infinitely large population
no migration
Hardy-Weinberg for two alleles
p = dominant allele & q = recessive allele
p + q = 1
p2 + 2pq + q2 = 1
Hardy-Weinberg for multiple alleles
p + q + r = 1
p2 + q2 + r2 +2pq + 2pr + 2qr = 1
Hardy-Weinberg for X-linked alleles
frequency of allele = frequency of males expressing the X-linked gene
female carriers = 2pq
females affected = q2
Hardy-Weinberg with Fitness
find total fitness
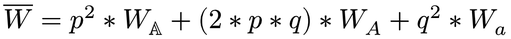
find new values
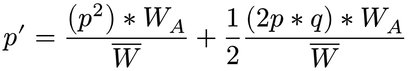
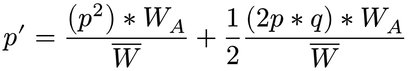
Directional Selection
selection favors one genotype related to a phenotypic extreme
shift in population mean
Stabilizing Selection
both phenotypic extremes are selected against and intermediate types are favored
mean stays the same but variance decreases
Disruptive selection
both phenotypic extremes are selected for and intermediates are selected against
increasingly bimodal population
Migration effect on allele frequency
pi’ = (1 - m)pi + mpm
Positive Assortive Mating
similar genotypes are more likely to mate than dissimilar ones
Negative Assortive Mating
dissimilar genotypes are more likely to mate than similar ones
Inbreeding
mating individuals are related, removing heterozygosity
Change in Allele Frequency
s = 1 - Waa
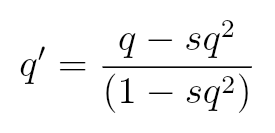
for lethal recessive alleles
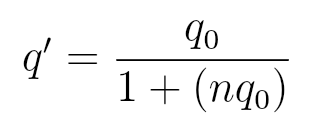
Meristic Traits
polygenic traits in which the phenotype is counted in integers
ex. number of appendages
Threshold Traits
polygenic but have a small number of discrete phenotypic classes, often influenced by environmental factors
Estimating number of polygenes
n = number of polygenes
any amount of polygenes: (1/4)n = 1/(individuals expressing one extreme)
low number of polygenes: 2n + 1
Variance

Covariance
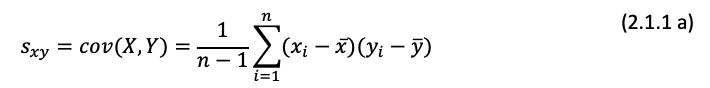
Correlation Coefficient
r = covxy/sxsy
Heritability
proportion of total phenotypic variation in a population due to genetic factors
Broad-sense Heritability
H2 = Vg/Vp
Phenotypic Variance
VP = VG + VE + VGxE
Narrow-sense Heritability
h2 = VA/VP
h2 = (M2 - M)/(M1 - M)
h2 = R/S
Genetic Variance
genetic variance = additive variance + dominance variance + interactive variance
VG = VA + VD + VI
Quantitative Trait Loci
multiple genes contributing to a quantitative trait
Log Of Odds (LOD)
odds = [prob(data: QTL)] / [prob(data: no QTL)]
LOD ≥ 3 are considered significant
DNA methylation
reversible addition of methyl groups to DNA, blocking transcription factors by blocking the major groove
CpG islands
clusters of CpG dinucleotide sites in promoter and upstream sequences
genes with adjacent methylated regions are transcriptionally silenced
heterochromatin methylation
dinucleotides adjacent to genes but are in repetitive DNA sequences like the centromere, maintaining chromosome stability by preventing translocations and related abnormalities
demethylation
passive - failure to methylate new DNA strands
active - removal of methyl groups independent of DNA replication
histone modifications
covalent modification of amino acids of the N-terminal through acetylation, methylation, and phosphorylation
histone acetylation
relaxes the grip of histones of DNA, allowing transcription to occur
short ncRNAs
act as repressors of gene expression
piRNA interact with proteins to form RNA-protein complexes that participate in epigenetic gene-silencing in germ cells
Antisense lncRNA
lncRNA genes partially overlap protein-coding genes and transcribe in opposite directions, reducing transcription
Intronic lncRNAs
lncRNA genes are located in introns and transcription does not overlap adjacent exons
Bidirectional lncRNAs
lncRNA genes use protein-coding genes’ promoter but is transcribed in opposite directions
Intergenic lncRNAs
lncRNAs are discrete transcription units located outside protein-coding genes
Monoallelic expression
only the parental or maternal allele is transcribed and the other is transcriptionally inactive
occurs in genomic imprinting, autosomal genes, and random inactivation of an X chromosome
Genomic Imprinting
expressed in a parent-of-origin pattern
genomic region escapes demethylation and remethylation during gamete formation making the alleles stay transcriptionally silent
usually found in clusters on the same chromosome
Epimutations
mutations in imprinted genes that affect adjacent genes due to DNA sequence changes or coordinately controlled imprinted genes
random inactivation of an X chromosome
happens in cells of female mammals
Xist lncRNA turns inactivated chromosome into a Barr body
Tsix lncRNA prevents the active chromosome from being silenced and represses the Xist lncRNA
Monoallelic expression (MAE) of autosomal genes
scattered throughout the genome
causes expression of both alleles (biallelic), only the maternal or paternal allele, or expression of neither allele
genomic hypomethylation
occurs in all cancer cells allowing for unrestricted transcription of many gene sets
selective hypermethylation
occurs in certain regions of the genome in cancer cells to silence certain genes like tumor-suppressors
alternative splicing
creates different spliceforms/variation which different protein variant
isoforms
different variants of mRNA caused by alternative splicing
cassette exons
most common type of alternative splicing in mammals
exons can be excluded by joining the 3’ end of an upstream exon to the 5’ end of a downstream exon
alternative splice site
type of alternative splicing where an exon near an intron gets spliced as well
alternative promoters
type of alternative splicing where there is more than one site where transcription is initiated
CT/CGRP gene
in thyroid cells, mature mRNA contains only the first four exons which is processed into a calcitonin (CT) peptide that regulated blood calcium levels
in neurons, exon 4 is alternatively spliced out, processing a peptide hormone, CGRP, that stimulates the dilation of blood vessels
proteome
number of proteins in an organism
Dscam gene
gene in fruit flies that codes for a protein that guides axon growth
alternative splicing leads to different tags on neurons, allowing for correct wiring
splicing enhancers and silencers
cis-acting sequences which promote or inhibit the splicing of nearby splice sites
SR proteins
bind to enhancers and activate splicing by recruiting spliceosome components
hnRNPs (heterogeneous nuclear ribonucleoproteins)
class of proteins that bind to splicing silencers and inhibit splicing
RBPs (RNA-binding proteins)
class of proteins that bind to RNA sequences or structures and can bind or hide splice sites to promote the use of alternative sites
Sex lethal (Sxl) gene
regulatory gene that encodes an RNA-binding protein
sex determinism in fruit flies
transcription factors encoded by genes on the X chromosome allow for the transcription of the Sxl (sex lethal) gene
lower concentration of transcription factors in males does not activate transcription
SXL protein binds to tra (transformer) gene and uses alternative splicing to create a fully functional SR protein
translation of male mRNA leads to a nonfunctional protein
male and female spliceoforms create different DSX (double sex) isoforms
DSX-F proteins repress transcription of genes that control male sexual development
DSX-M proteins activate transcription of genes that control male sexual development and repress the transcription of genes that control female sexual development
exoribonucleases
enzymes that degrade RNA via the removal of terminal nucleotides
deadenylation-dependent decay
process initiated by deadenylases, enzymes that shorten the poly-A tail
if it shortens thedeadeny poly-A tail to less than 30 nucleotides, mRNA will be degraded
Exoribonuclease destroys the mRNA in a 3’ to 5’ manner
OR
decapping enzymes remove the 5’ cap, allowing XRN1 to destroy mRNA in the 5’ to 3’ direction
deadenylation-independent decay
endoribonucleases that cleave mRNA internally
P (processing) bodies
mRNAs accumulated in cytoplasmic complexes due to not being actively translated
Adenosine-uridine rich element (ARE)
cis-acting sequences that regulate mRNA stability
ARE-containing mRNAs encode proteins that promote cellular multiplication
TTP (tristetrapolin)
recruits decay machinery to promote mRNA degradation
RNAi (RNA interference)
mechanism by which ncRNA molecules guide the posttranscriptional silencing of mRNAs in a sequence-specific moments
microRNA (miRNA)
regulatory ncRNA that causes translational downregulation of mRNA due to complementary base pairing
if the match is perfect, the mRNA is degraded
if the match is partial, translation is blocked
small interfering RNAs (siRNAs)
RNA derived from double-stranded RNA caused by virus infection
Dicer enzyme cleaves and evicts one of the two strands as a siRNA guide to recruit RISC to a complementary mRNA
RISC cleaves mRNA in the middle of the siRNA-mRNA complementary region
cleaved mRNA lacking a cap or poly-A tail is degraded by exoribonucleases
microRNA formation
primary miRNAs are created from miRNA genes
Dorsha enzymes removes noncomplementary 5’ and 3’ ends to produce pre-miRNAs and hairpins are exported to the cytoplasm
cleaved by Dicer to produce mature double-stranded miRNAs
further processing by RISC to become single stranded
long noncoding RNAs (lncRNAs)
linked to diverse regulatory functions with chromatin modification, altering patterns of gene expressions, and alternative splicing
can function as competing endogenous RNAs, which act as decoys binding to miRNAs, leading to more cell differentiation
circular RNA (circRNA) compete for miRNA binding
CPE (cytoplasmic polyadenylation element) sequence
CPEB protein recognizes mRNAs not being translated
PARN shortens the poly-A tail
shortened tail is less bound to PABP
Makin binds to a cap binding protein (eIF4E), blocking its interaction with eIF4G which is important for translation initiation
reversal of CPE-containing mRNAs
CPEB is phosphorylated by kinases
PARN is released
cytoplasmic poly-A polymerase lengthens the poly-A tail
poly-A tail is bound by additional PABPs which displace Maskin allowing for translation initiation
ZPB1 (zip code binding protein 1)
blocks transition initiation by preventing the association of the large subunit of the ribosome until mRNA is in the correct location
Phosphatases
enzymes that remove phosphate groups from proteins
Ubiquitin
a protein that targets others for degradation
covalently attaches to target protein’s lysine side chain through ubiquination
proteasome recognizes tagged proteins, unwinds them, removes ubiquitin tags, breaks proteins into small peptides
Ubiquitin ligases
enzymes that bind to specific proteins and catalyze the processive addition of ubiquitin residues
p53
activates transcription of genes that encode proteins that stop the cell cycle and promote DNA repair
bound to Mdm2 and tagged for degradation in normal cells, but Mdm2 is phosphorylated when the cell senses DNA damage