1 enzymes
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
what type of catalysis is used by enzymes and why
dont use specific because [H3O+] and [HO-] are very low at physiological pH (approx 7_
but general is not a problem
how can enzymes act as general catalysts
bring reactants together via non-covalent interactions and can place catalytic functional groups in the right location to enable catalysis to occur

show the steps of an enzyme-catalysed reaction
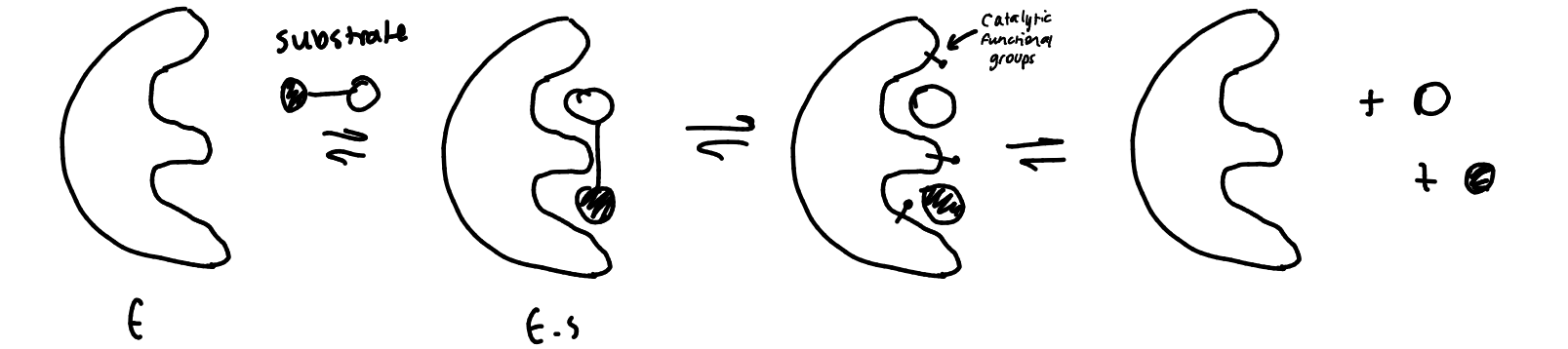
what makes RNA susceptible to hydrolysis
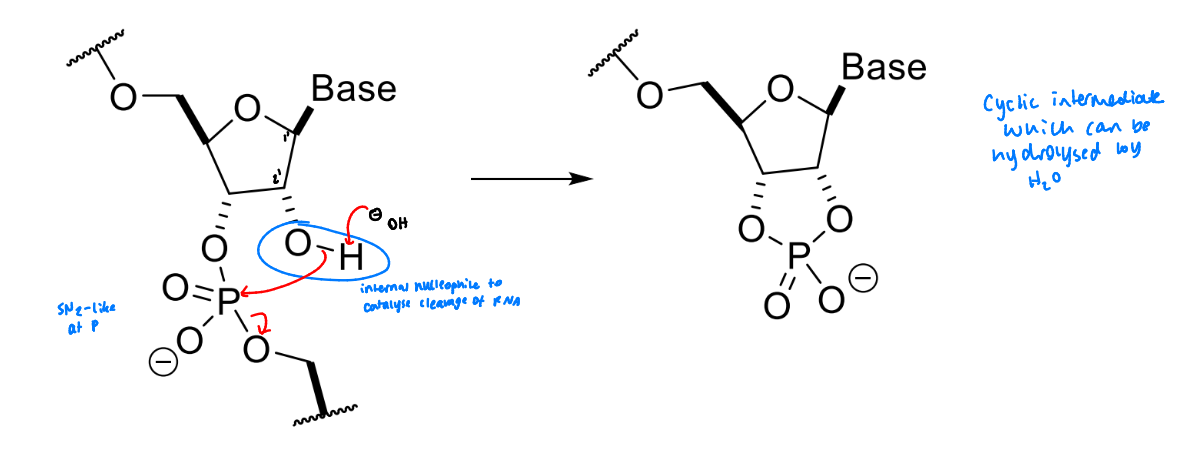
the presence of the 2’-OH group adjacent to the phosphate - the chain can be cleaved to form a cyclic product
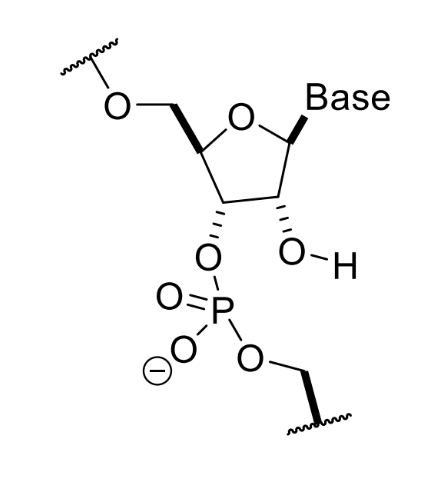
structure of RNA
show how RNA can be hydrolysed by bases
why is DNA not hydrolysed in the same way as RNA
it lacks the OH groups needed - the base would instead have to attack directly at the P
how is an RNase reaction catalysed
pKa?
catalysed by two histidines, pKa ~ 7
one is deprotonated and one is protonated at neutral pH
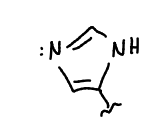
show deprotonated histidine
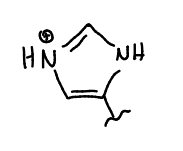
show protonated histidine
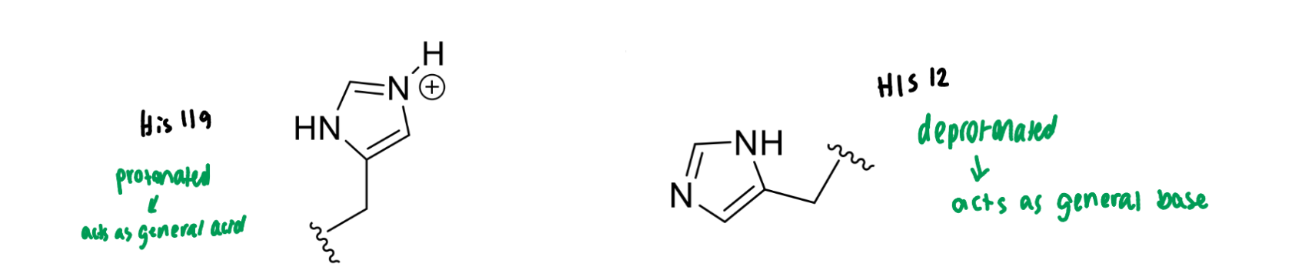
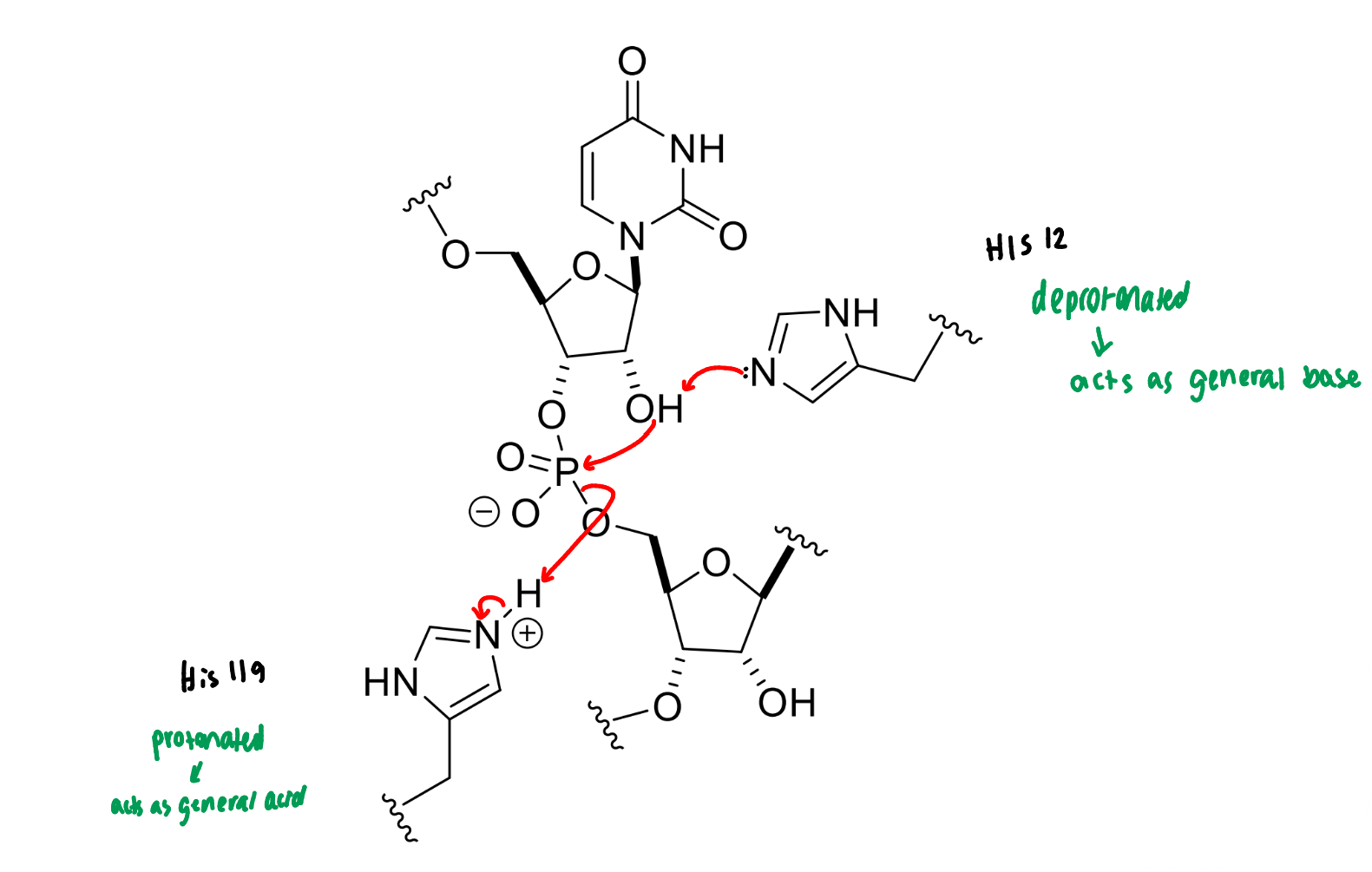
show the two histidines involved in the RNase reaction
what do they act as in the reaction?
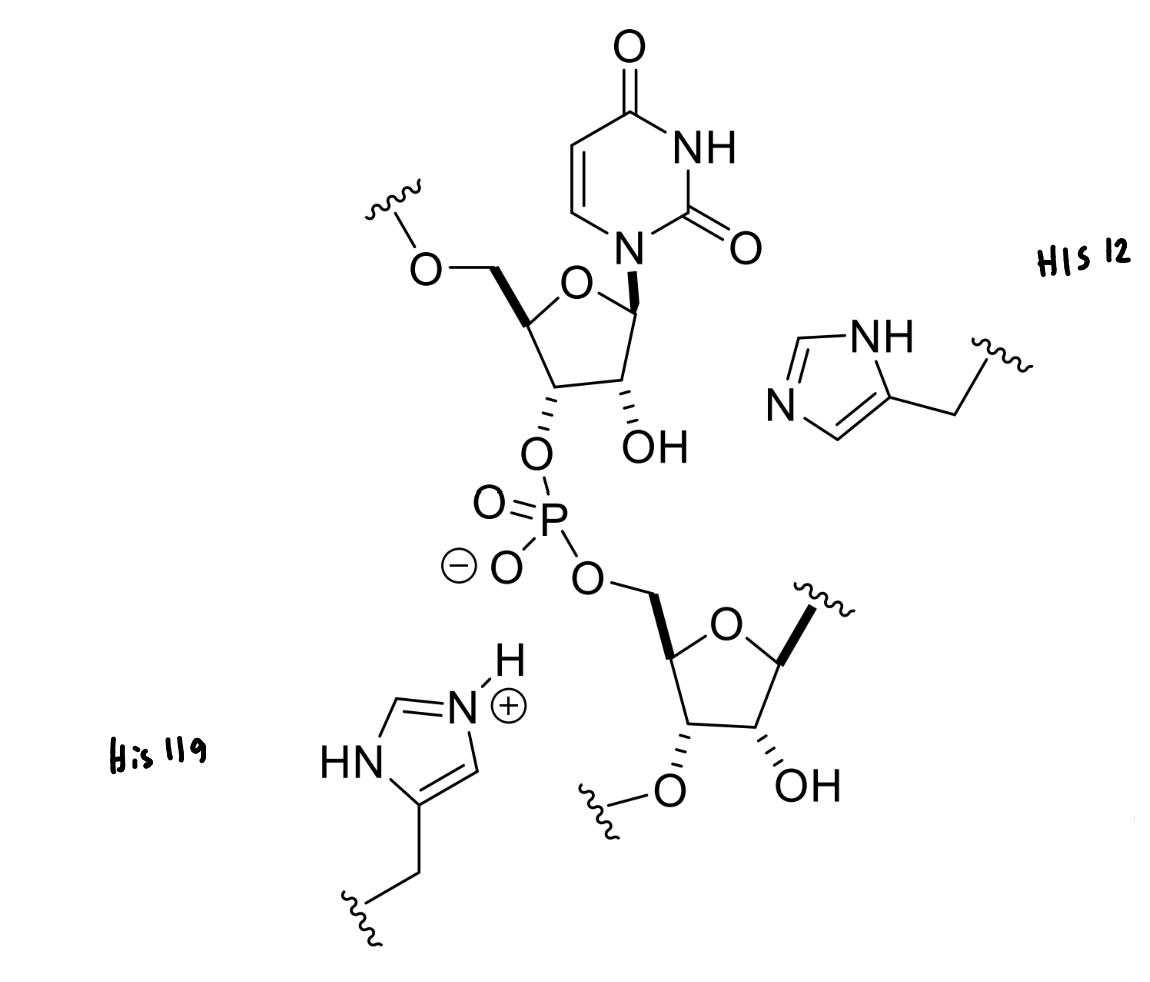
show the mechanism of RNase
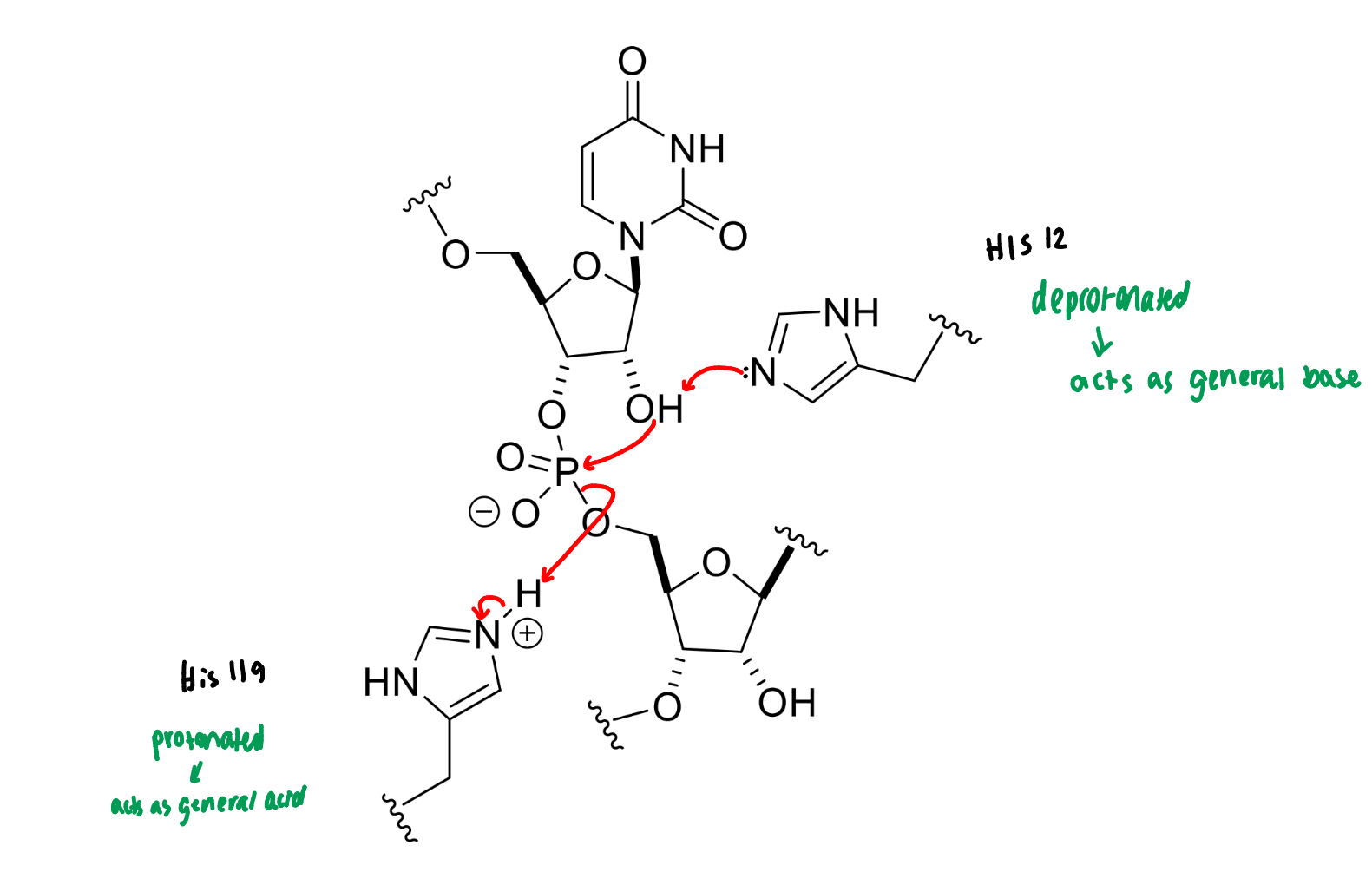
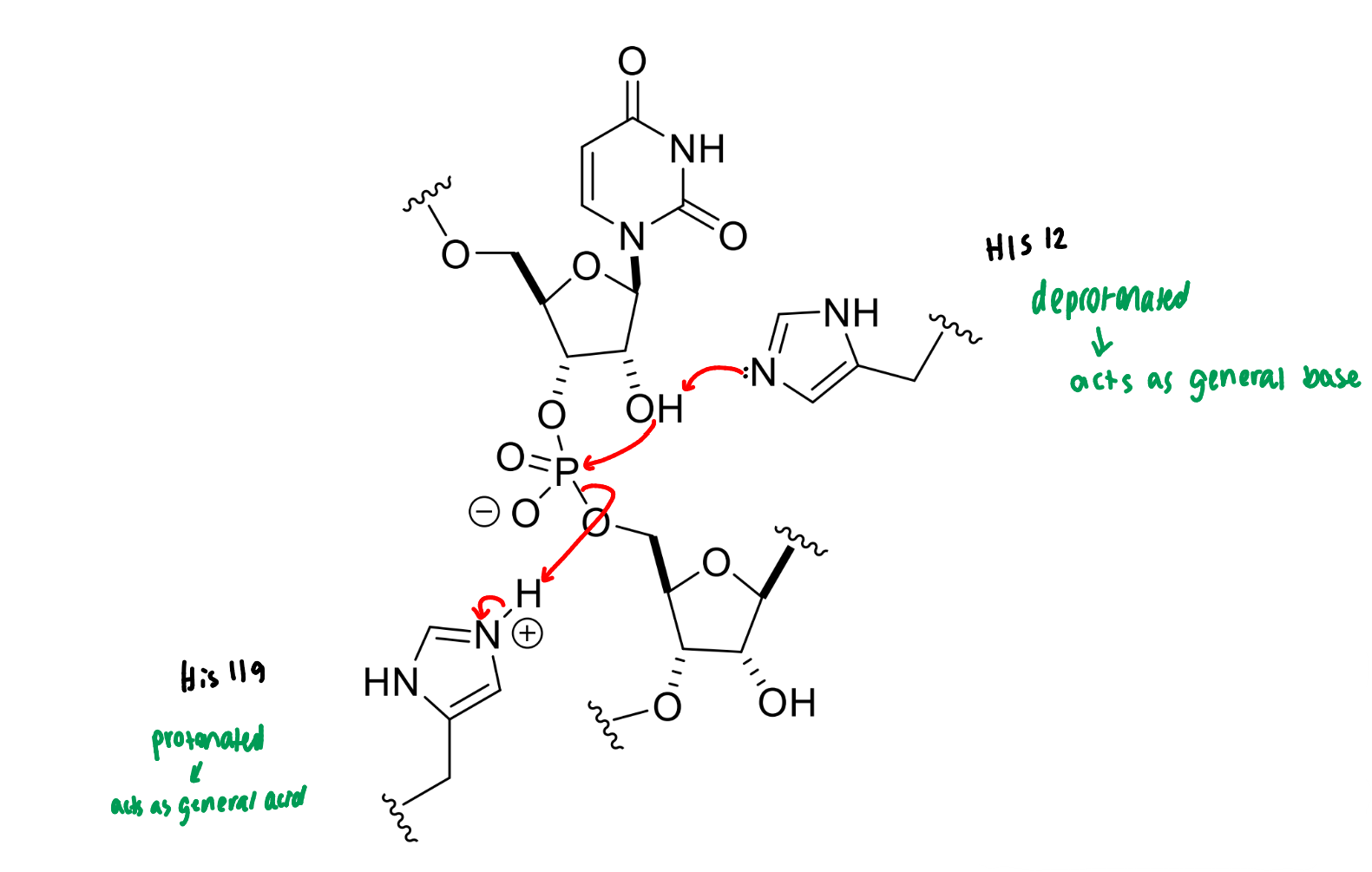
next step of RNase mechanism
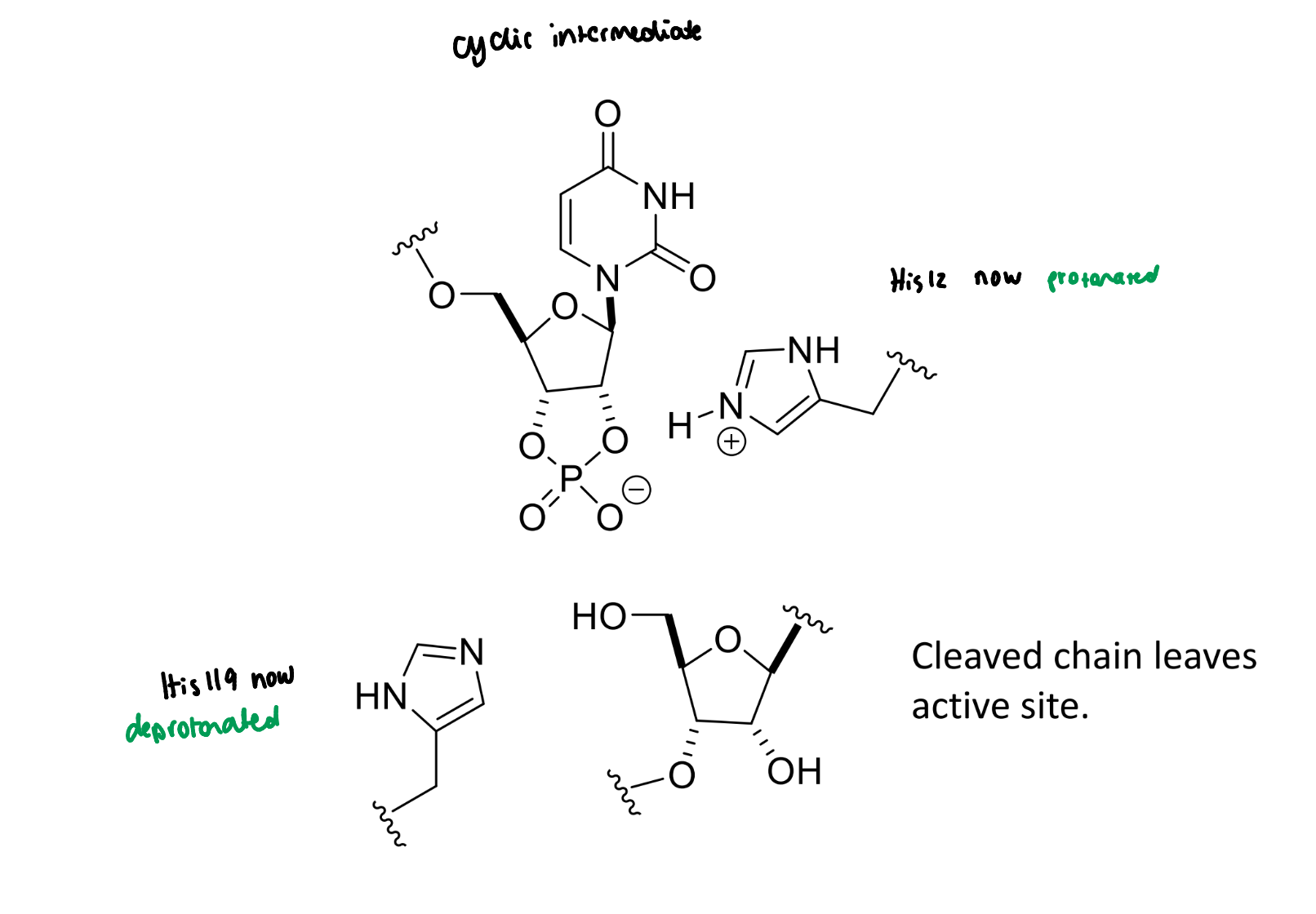
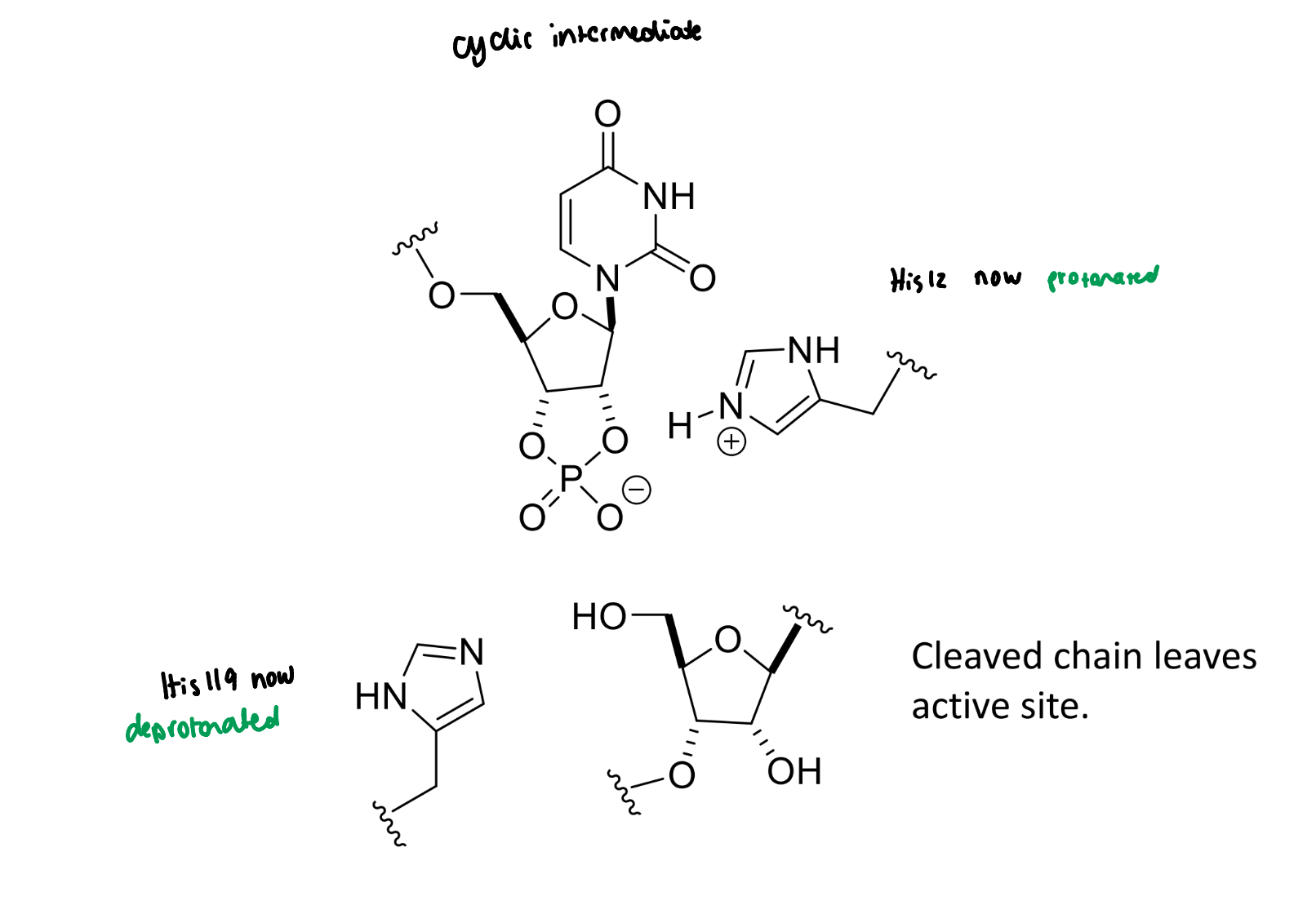
next step of RNase
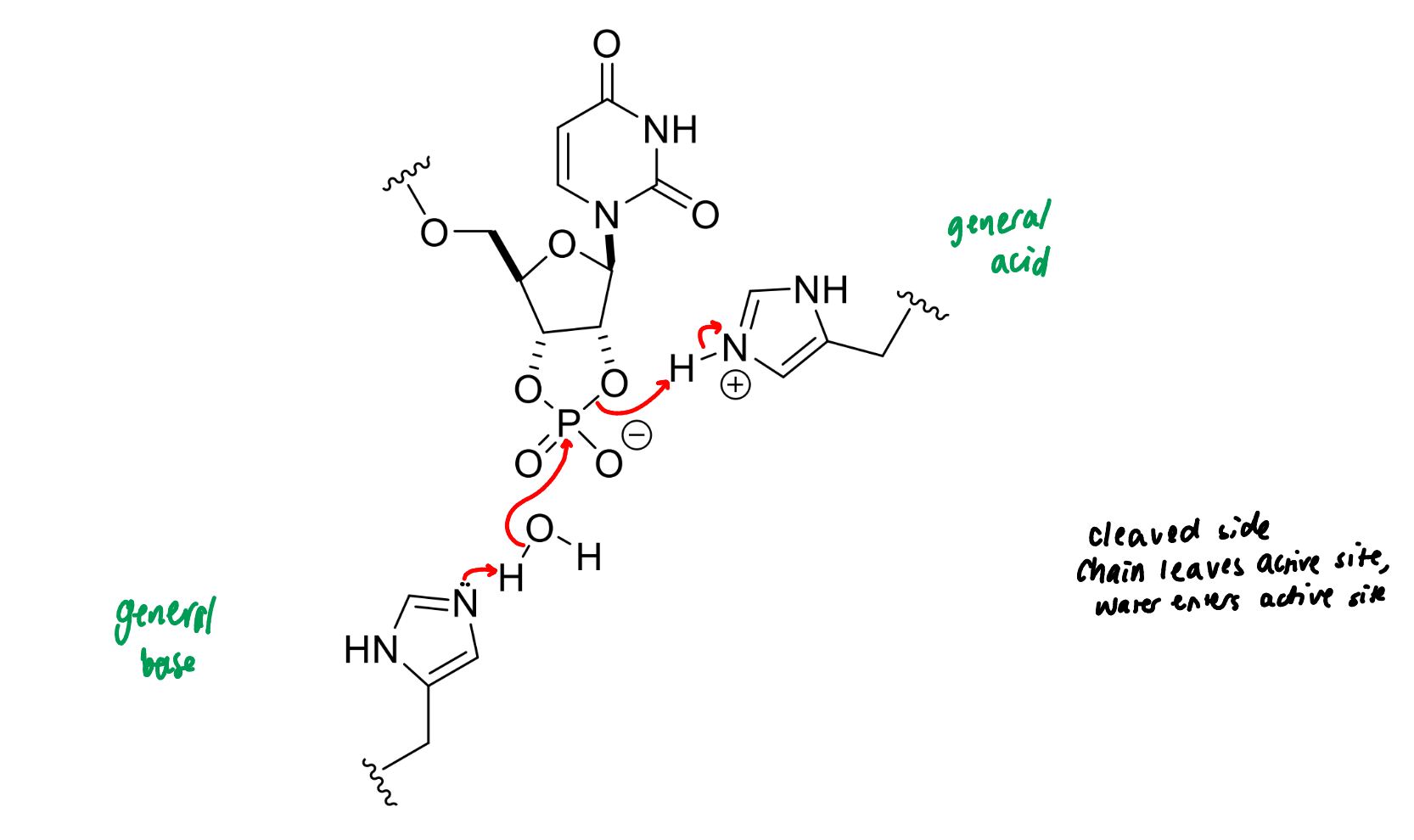
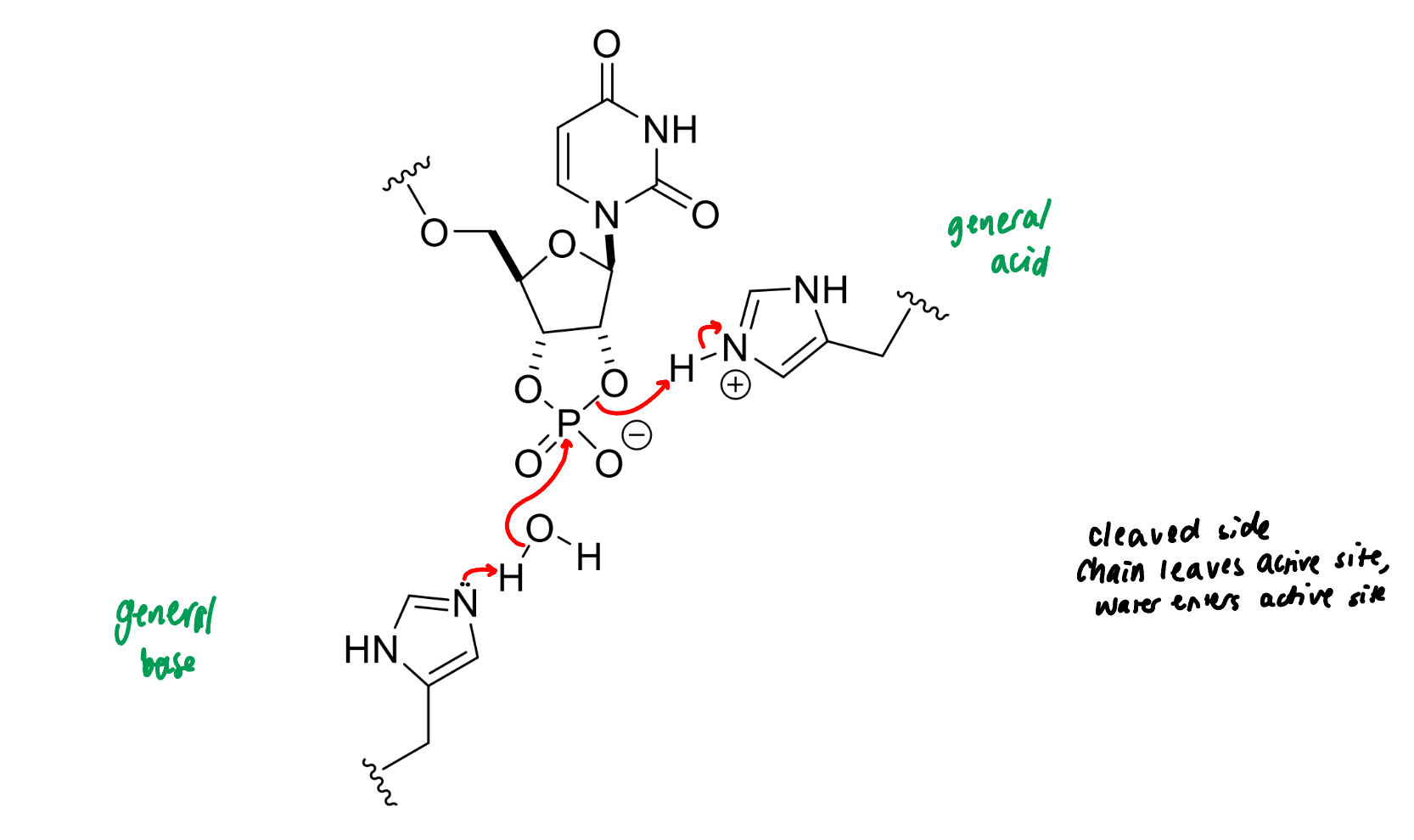
product of this stage of RNase mechanism
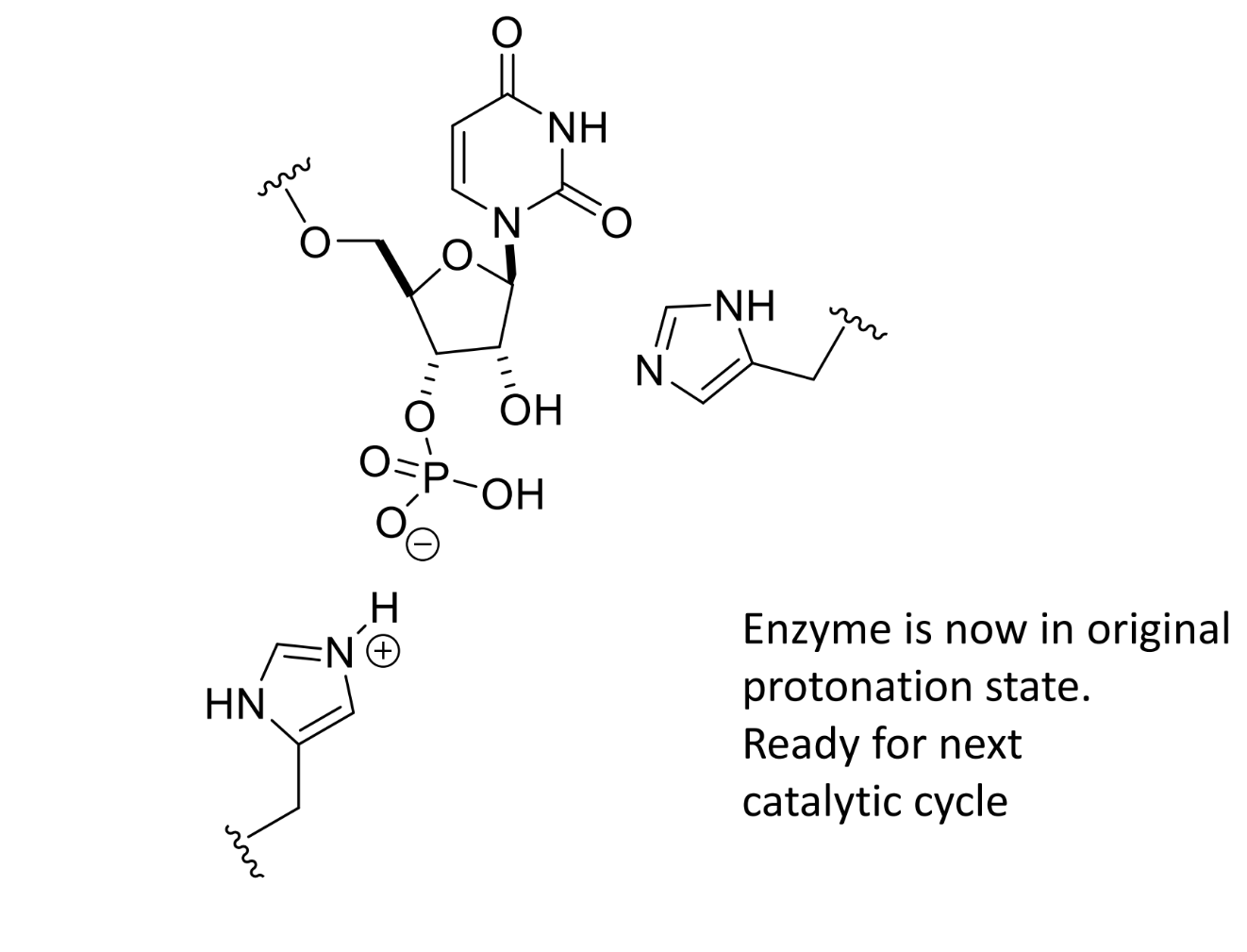
what is the RNase mechanism dependent on
dependent upon one histidine being protonated and one deprotonated
when is maximum activity for RNase and why
maximum activity at pH 6
below pH 6 both histidines are protonated
above pH 6 both histidines are deprotonated
what change may be made to the RNase reaction and what is the effect of this
changing either of His12 or His119 to alanine dramatically reduces the rate of reaction
show Ala12/Ala19

how can this be made to work with Ala119 instead
nitrophenolate anion is a good leaving group - doesn’t need to be protonated by His119
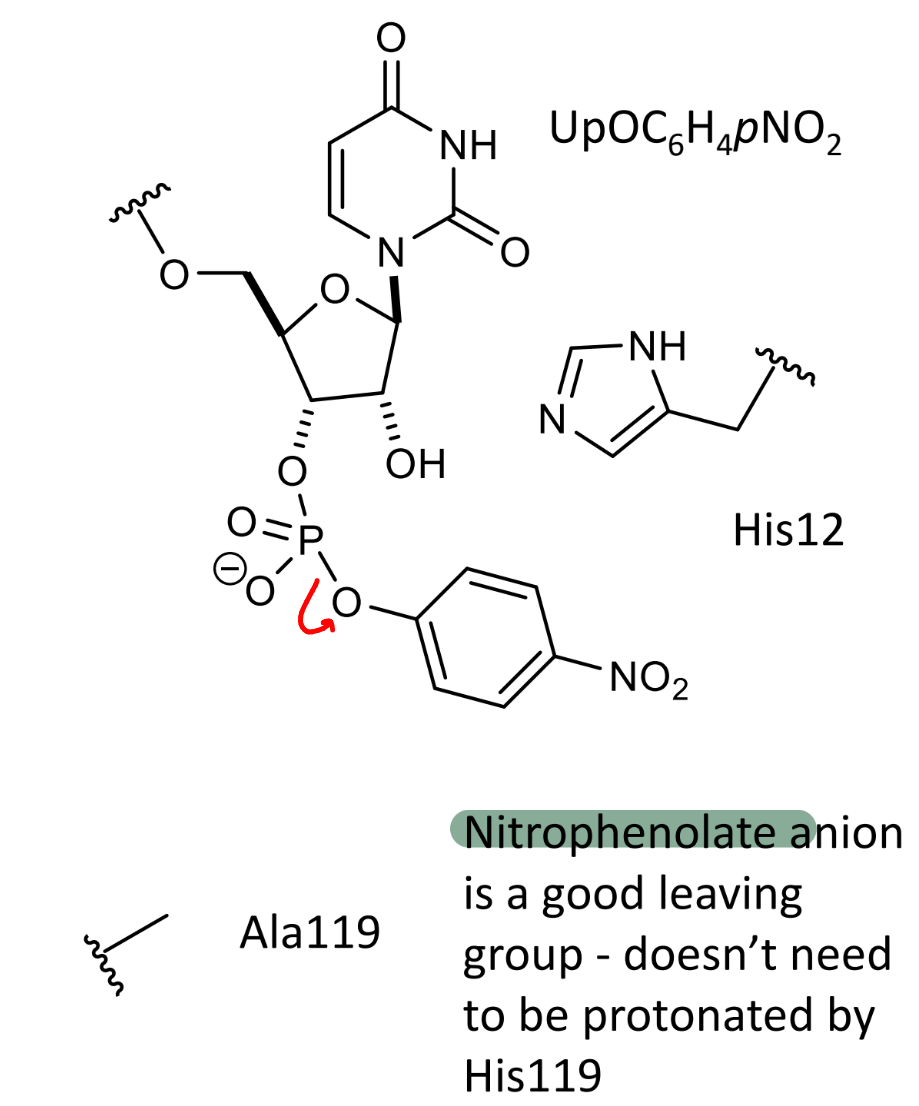
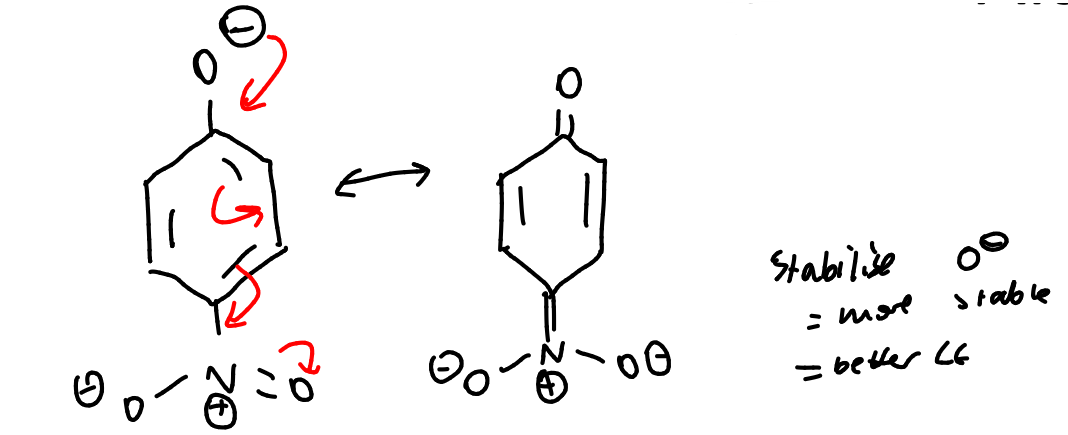
show the resonance forms of the nitrophenolate ion and explain why is is a good leaving group
why does nature want to cleave peptide/amide bonds
RNA is inherently unstable
in contrast, an amide bond is essentially indefinitely stable in physiological conditions
how can the hydrolysis of an amide bond be performed
using enzymes called proteases
where are the important roles of proteases (4)
digestion, cell signalling, blood clotting, inflammation
type(s) of catalyst used by serine protease
acid/base and nucleophilic
what are the 3 residues that serine protease relies on
what is this called?
Ser
His
Asp
Catalytic triad
pKa of serine
approx 14
show serine protease’s catalytic triad and how it links to the backbone
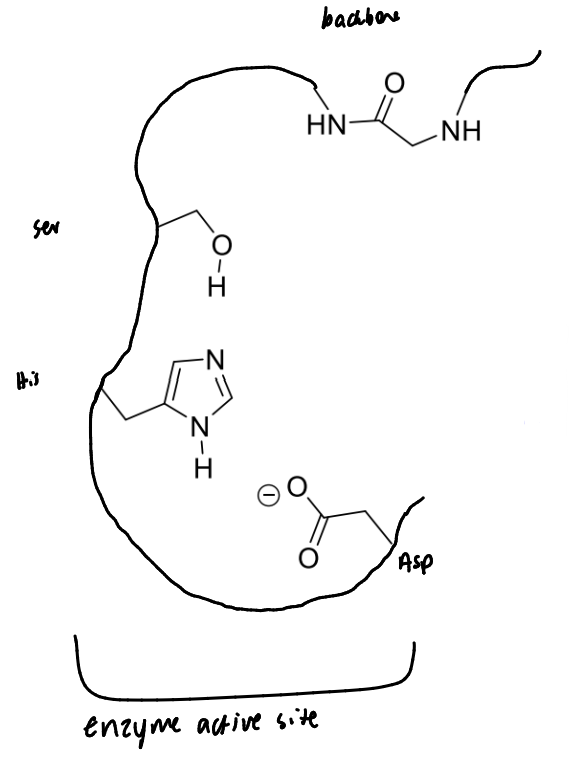
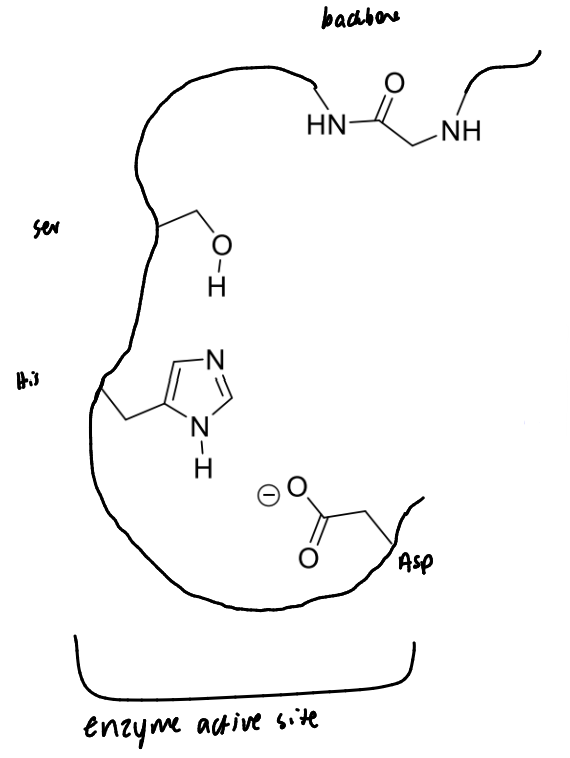
show the first step of serine protease’s mechanism with a peptide bond
describe the roles of the groups involved
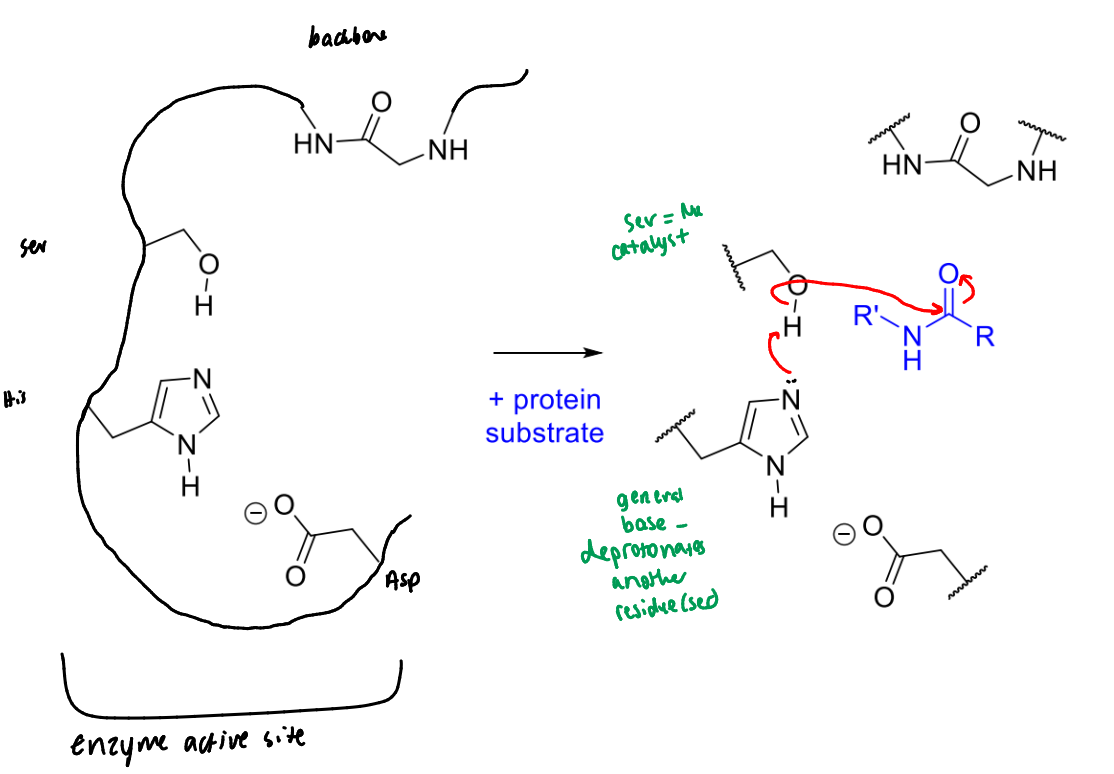
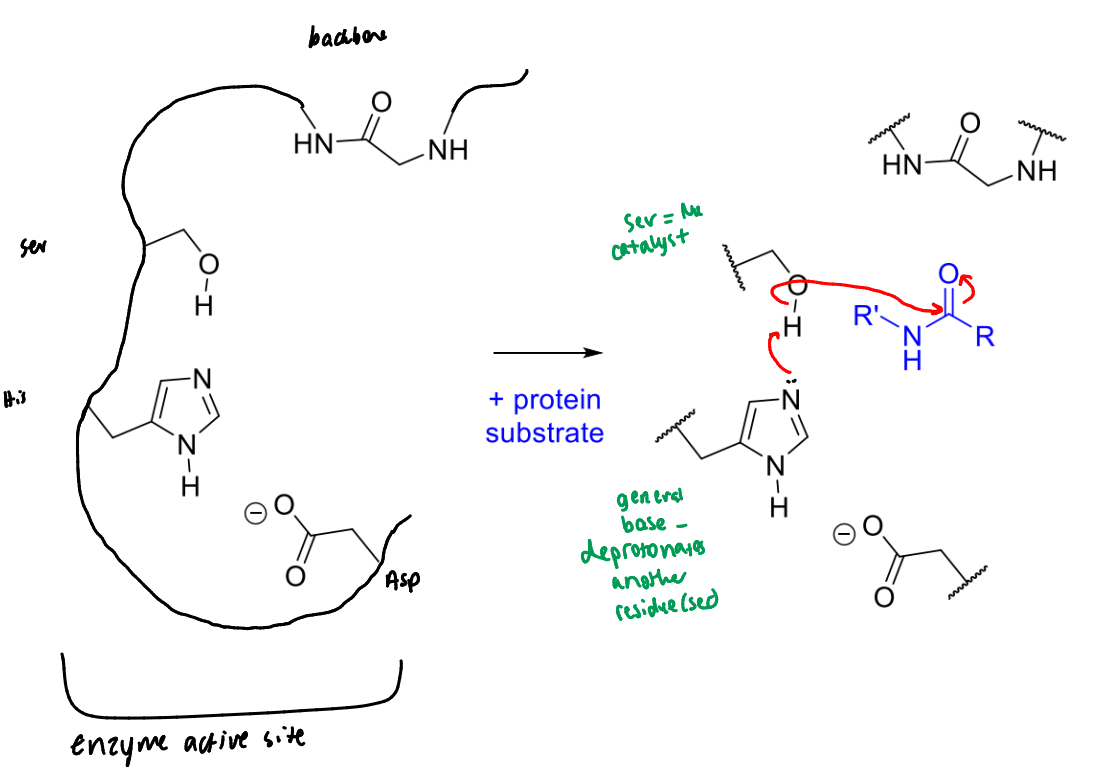
what is the product of this step of serine protease’s mechanism
explain any charges and relevent intermolecular forces
His protonated - general acid
Asp helps stabilise His positive charge
tetrahedral intermediate - active site has H bonding to help stabilise negative charge
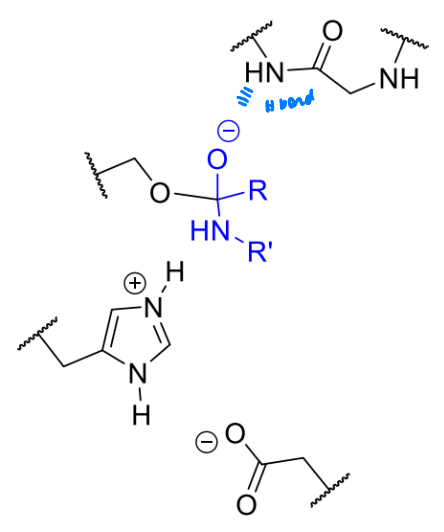
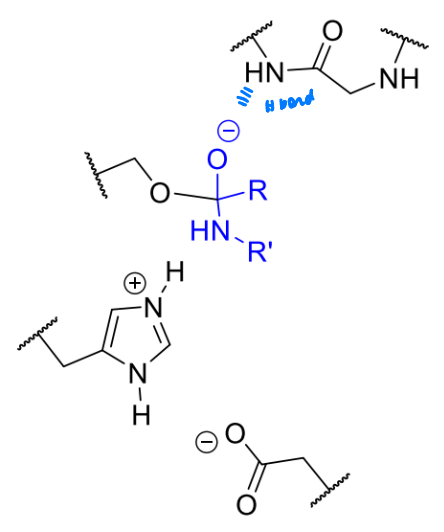
what is the next stage in the serine protease mechanism
comment on the leaving group?
NH not good leaving group but protonate with His as it leaves
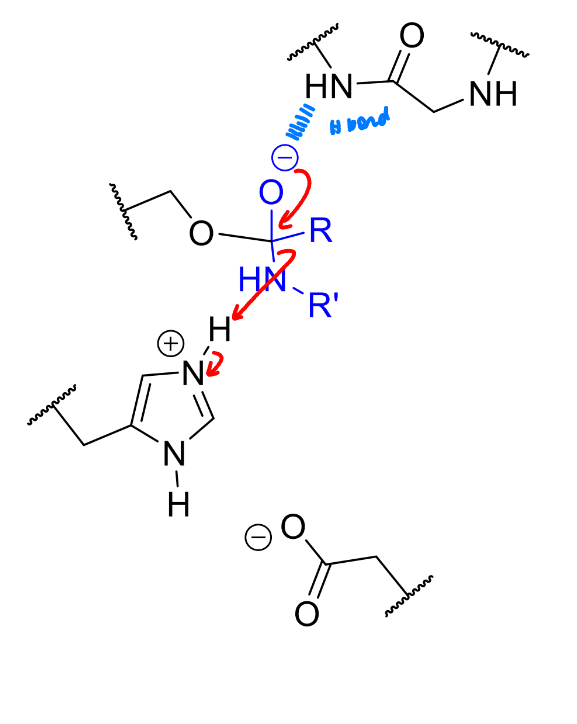
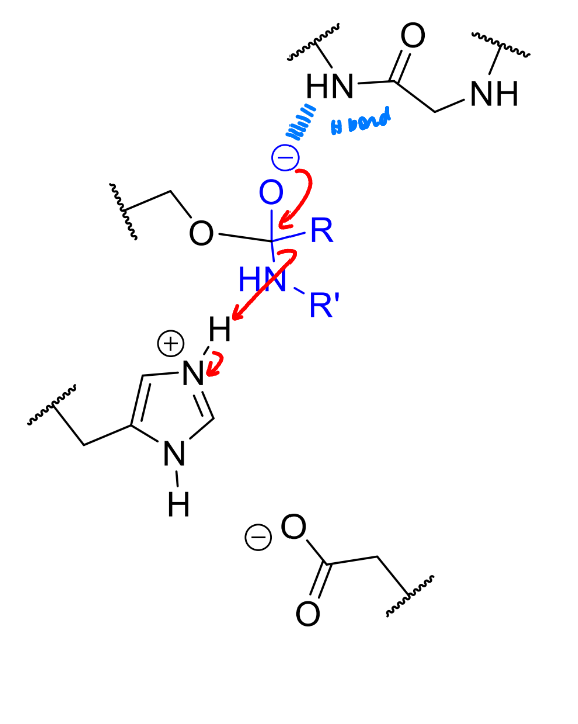
product of this stage of the serine protease mechanism
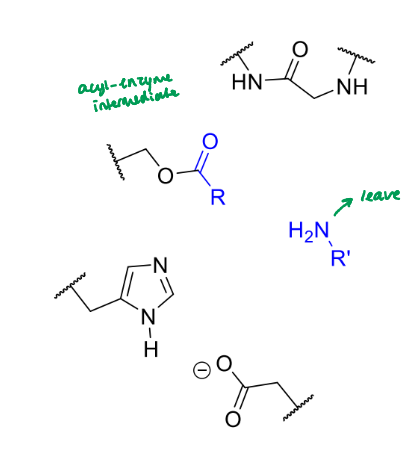
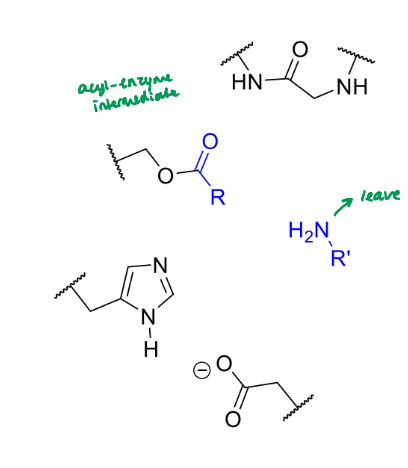
what happens next in the serine protease mechanism
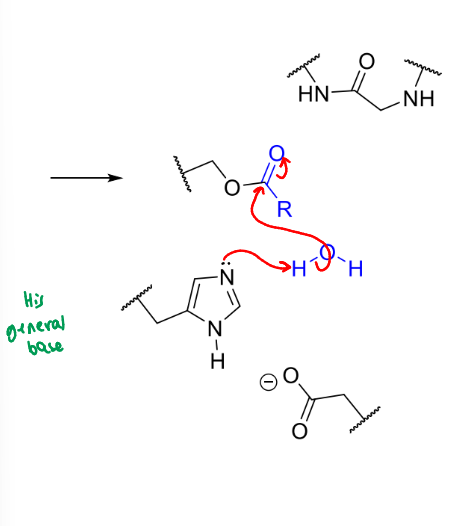
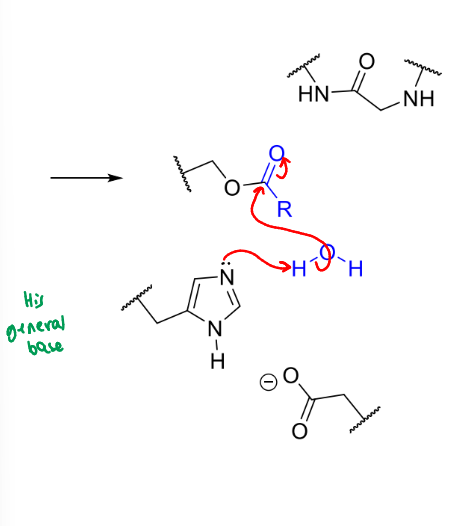
what is the product of this stage of serine protease
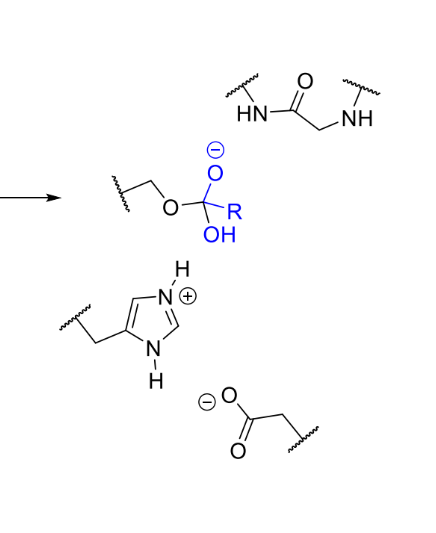
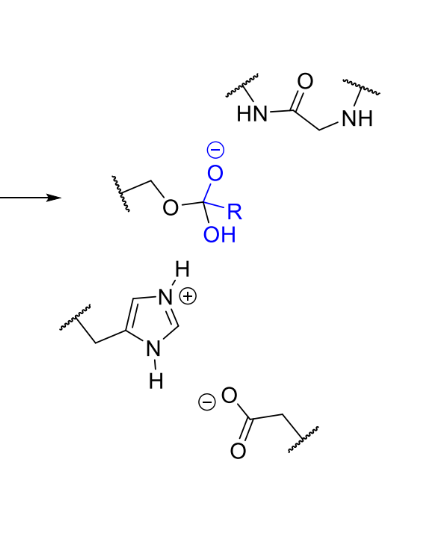
what happens next in serine protease mechanism
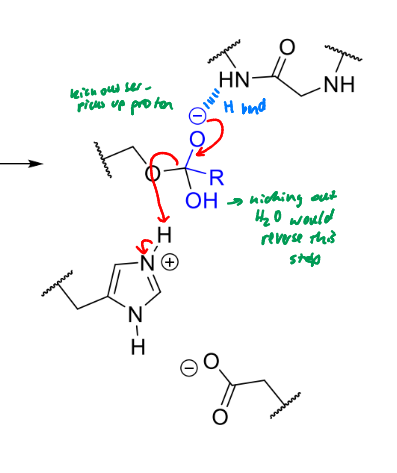
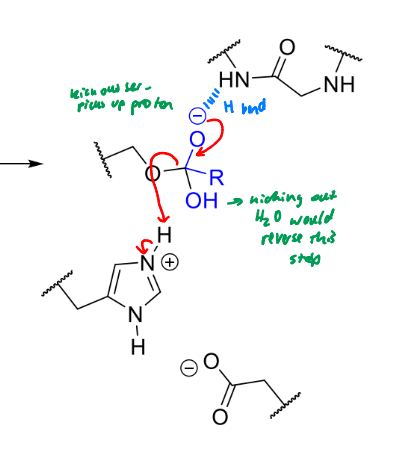
product of this stage of serine protease mechanism
enzyme in original state, product leaves
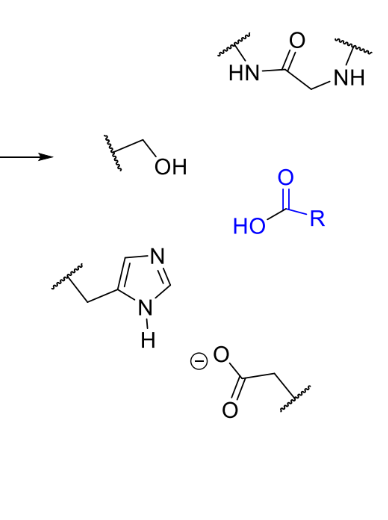
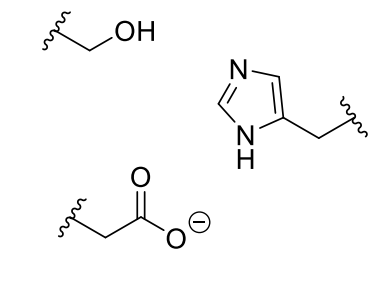
how is it clear this is the reaction mechanism?
if any of the three amino acids of the enzyme are replaced, there is a large drop off in enzyme activity
shows it is essential
what makes enzyme activity pH-dependent
necessity of charges (or lack of charges) on certain amino acid side chains
pKa of Asp
4
pKa of His
7
pKa of Ser
14
what are the three serine proteases that are chiefly responsible for digesting dietary protein in small intestines
trypsin
chymotrypsin
elastase
how do trypsin, chymotrypsin and elastase cleave
cleave amide bonds adjacent to specific amino acid residues
where does the selectivity of trypsin, chymotrypsin and elastase come from?
a binding pocket next to the active site
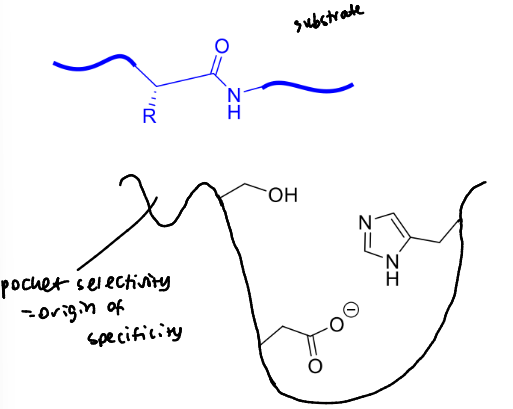
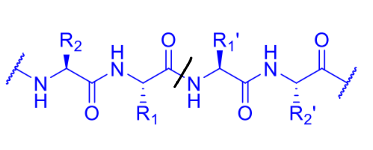
label this to describe the residues on either side of the cleavage site as well as where the binding pockets are
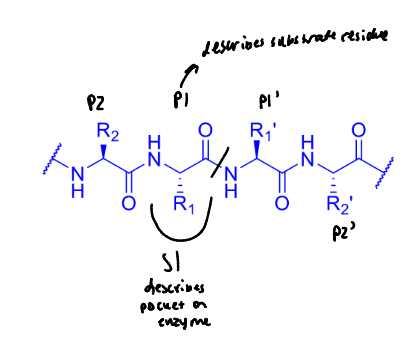
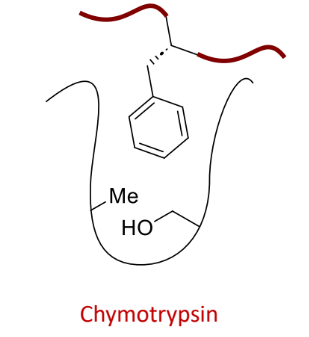
what does this protease bind?
large, aromatic side chains
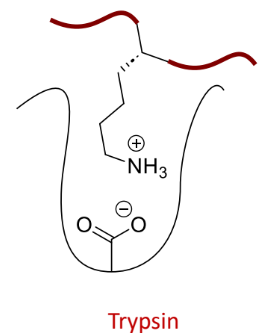
what does this protease bind
positively-charged side chains
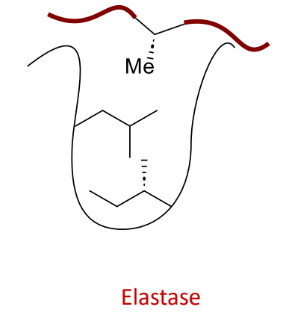
what does this protease bind
small side chains