Nucleotide Excision Repair
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
What is NER?
Removal of damaged base(s) as part of an oligonucleotide fragment - bulky adduct repair
SSBR and DSBR are non-bulky lesion repairs
What are the two major UV photoproducts? Which protein complexes recognise both damages?
Cyclobutane pyrimidine dimer (CPD) - recognised by XPC
TC (6-4) photoproduct (6-4PP) - recognised by DDB (1/2 complex)
How does UvrA2B2 and UvrC act in NER?
UvrA2B2 moves along DNA - damage surveillance
Binds bulky lesions - UvrB (helices) deposited at damaged site, opens DNA forming ‘bubble’ structure
UvrC (nuclease) nicks lesion and induces polymerase and ligase to repair lesion
What are the predispositions associated with Xeroderma Pigmentosum?
rare inherited disease which predisoses patient to:
pigmented lesions on skin exposed to sun
elevated skin cancer incidence
XP genes can be caused by mutations in several NER related genes
describe the clinical symptoms of XP, TTD and CS
XP: skin sensitivity to sunlight, acute sunburn, pigment abnormalities, multiple cancer predisposition, some neurological and intellectual disabilities
TTD: sulfur deficient, brittle hair, no cancer predisposition
CS: dwarfism, loss of adopts tissue, sunken eyes, premature ageing, retinal atrophy, no cancer predisposiiton
Symptoms are spectral based on severity of disorder
How did granule analysis confirm XP as causative of NER defects?
non-XP cells released granules (new DNA strands) when treated with UV, XP cells did not
granules = NER occurance
XP can be categorised by UDS level (unscheduled DNA synthesis)
XP-A has lowest UDS
CS has regular UDS
XP-C is the most prevelant in UK - however XP prevalence overall is low, potentially due to lack of UV
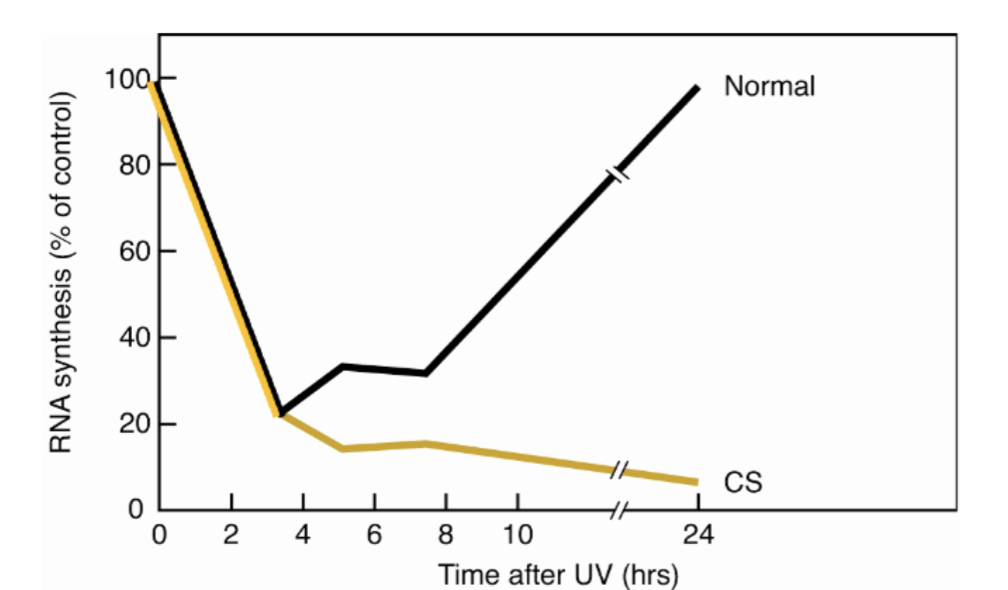
How is Cockayne Syndrome detected, what mutations cause CS?
UDS in CS is same a normal patient - so not differentiative
however RNA synthesis is very low in CS - suggests defect in transcription?
CS defined by defective transcription coupled repair - all NER is global (up to 24hrs)
Mutation in CS-A or CS-B leads to CS - epistatic
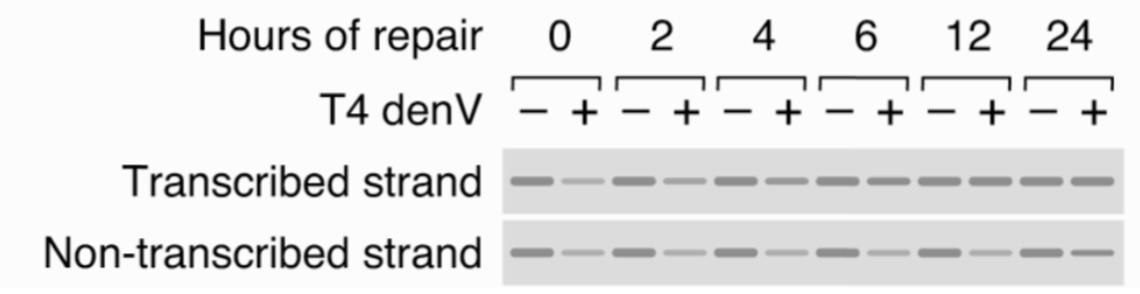
How does CS repair bulky lesions in comparison to WT?
WT cells repair lesions in transcribed strands - non-transcribed strand lesions do not affect protein-level
T4 denV endonuclease cuts DNA
TCR repairs DNA required for trasncription at a faster rate than GG-NER - crucial
CS cells are defective in all TCR, gene is effectively inactive until repaired by GG-NER
XP-C is opposite, only carry out TCR-NER, defective in GG-NER
How is a lesion recognised in GG-NER?
Detected by one damaged base in 1 million undamaged bases.
CPD and 6-4PP are recognised by DDB heterodimer (DDB1/2 - DDB2 is XP-E)
CPD is poorly recognised - better recognised by XP-C
DDB1 breaks helical structure of damaged site
DDB is part of a ubiquity ligase complex:
ubiqutinates chromatid proteins
recruits XP-C for CPD recognition
self ubiqutinates for self degradation
Describe the three significant subunits of the TFIIH complex and their importance in normal transcription and NER.
TFIIH is a basal transcription factor - subunits include XP-B, XP-D and TTD-A
Helicase in NER and normal transcription
XP-B: opens promoter sites via helices activity, not essential for NER but involved
XP-D: not essential for transcription, but essential for NER
TTD-A: not essential but involved in NER
XPB and XPD are helicases in opposing polarities
TFIIH forms ‘bubble’ structure in NER - defective TFIIH associated with TTD
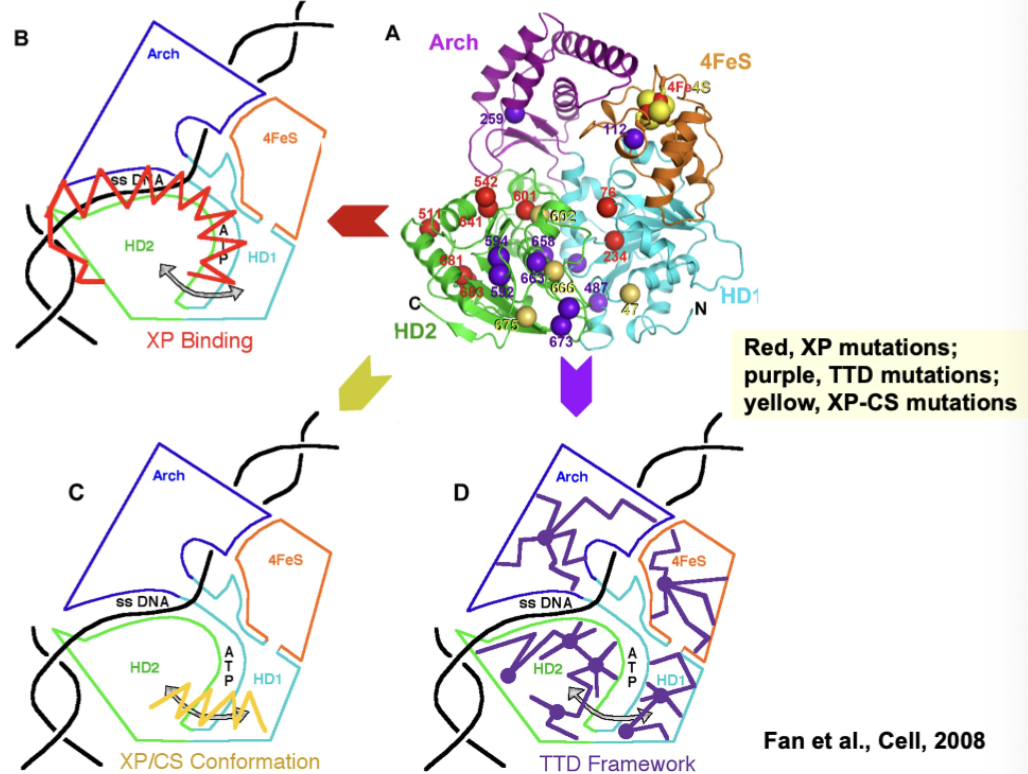
How does each X-ray crystallography mapping common mutations for each syndrome suggest differences in phenotype?
XP mutations in ability to bind damage DNA suggest GG-NER defect
TTD mutations in core of protein suggest protein misfolding and TRANSCRIPTIONAL defect
XP-CS mutations appear to be a combination - defects in both?
What is the role of XP-A in NER? Is it an essential gene?
XP-A is a zinc finger protein which binds to UV-irradiated DNA, homologous to Rad14 in yeast
binding increased by RPA
verifies damage and postioning in NER - previously thought to recognise damage
XPA/XPC KO cells sensitive to UV, gene is important but not essential
Which nucleases are involved in NER? Are they essential genes?
XPG nuclease cuts from 3’ to damage, ERCC1 and XPF cuts from 5’ to damage
These genes are essential in KO mice, implies another crucial function
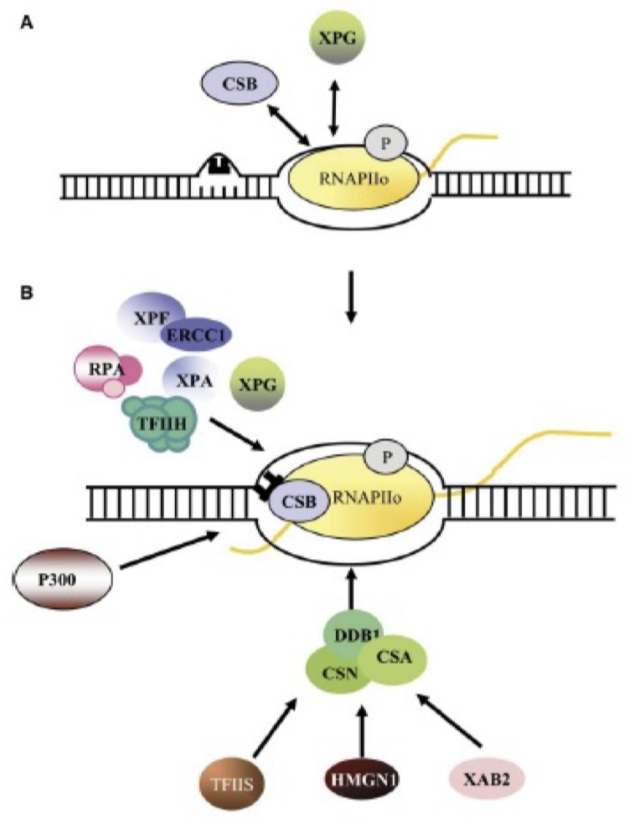
How was the sequential protein activation in NER determined?
UV-irradiate cell fluorescence analysis
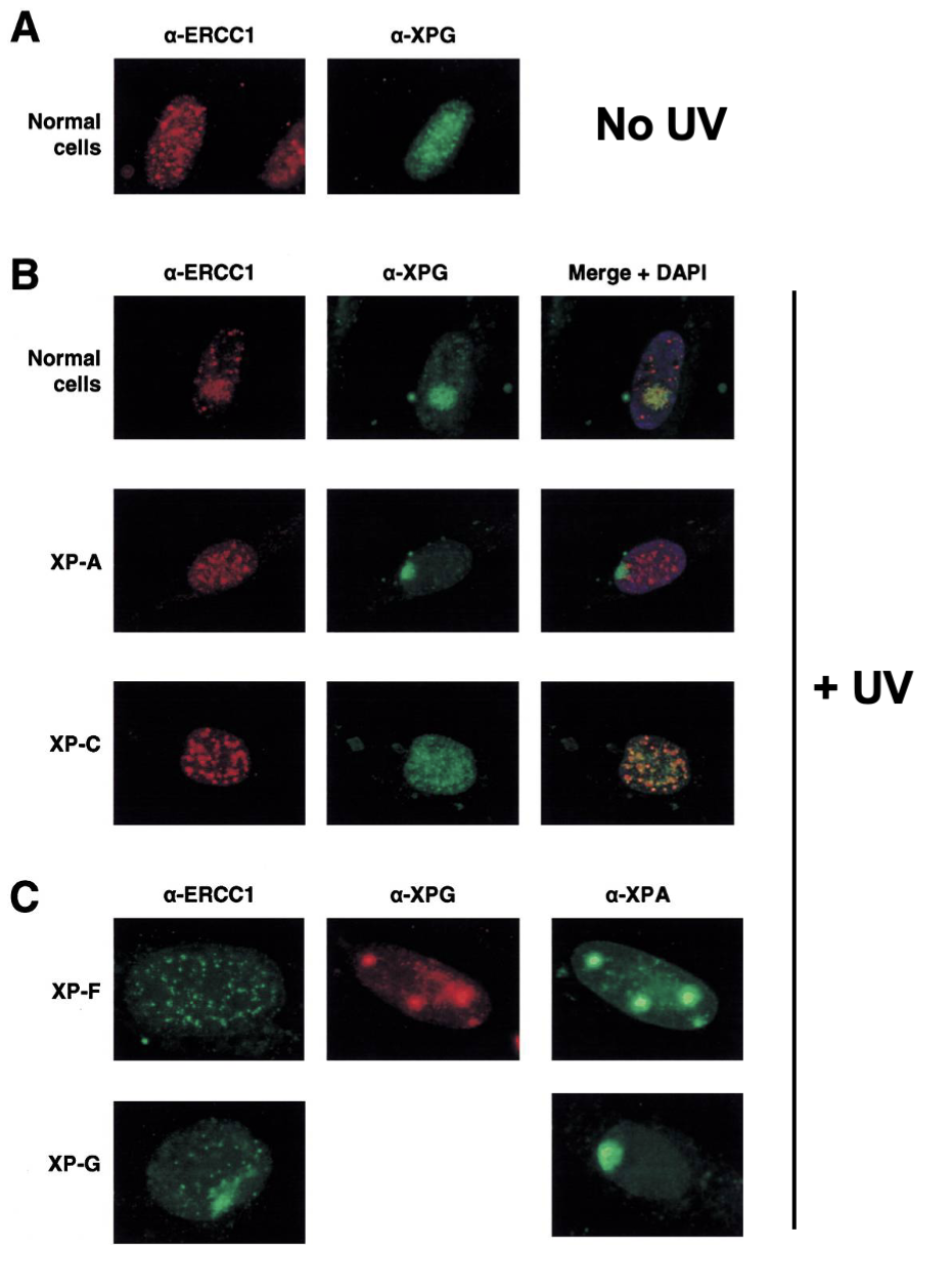
What is the difference in TCR, compared to GG-NER?
Recognition proteins and mechanism different
Blocked RNAPII signals lesion recognition for TC-NER
recruitment of remodellers by CS-A/B proteins enable RNAPII to be pushed back and NER proteins can recruit to and bind damage.
further process is same as GG-NER