Genetics exam #2
0.0(0)
Studied by 1 personCard Sorting
1/121
Earn XP
Description and Tags
Last updated 4:10 AM on 3/5/23
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
122 Terms
1
New cards
diphtheria facts
* sore throat, fever, coma and death
* epidemics in the 1920s killed 15,000 people in the US, mostly children
* Corynebacterium diphtheriae: toxin targets elongation factor 2 (EF2), stops translation
* we don’t have epidemics today due to vaccinations
* epidemics in the 1920s killed 15,000 people in the US, mostly children
* Corynebacterium diphtheriae: toxin targets elongation factor 2 (EF2), stops translation
* we don’t have epidemics today due to vaccinations
2
New cards
proteins
* synthesized from 20 amino acids
* composed of 1 or more polypeptides
* one string of amino acids linked together by peptide bonds
* amino acid structure
* composed of 1 or more polypeptides
* one string of amino acids linked together by peptide bonds
* amino acid structure
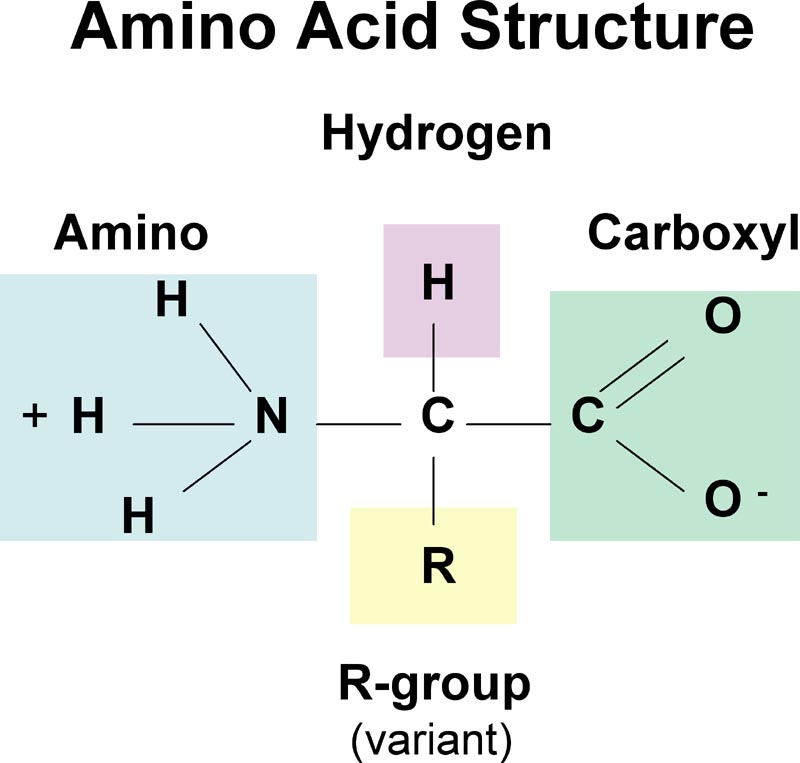
3
New cards
nonpolar, , aliphatic R groups
* Glycine (Gly, G)
* Alanine (Ala, A)
* Valine (Val, V)
* Leucine (Leu, L)
* Isoleucine (Ile, I)
* Proline (Pro, P)
* Alanine (Ala, A)
* Valine (Val, V)
* Leucine (Leu, L)
* Isoleucine (Ile, I)
* Proline (Pro, P)
4
New cards
Polar, uncharged R groups
* Serine (Ser, S)
* Threonine (Thr, T)
* Cysteine (Cys, C)
* Methionine (Met, M)
* Asparagine (Asn, N)
* Glutamine (Gln, Q)
* Threonine (Thr, T)
* Cysteine (Cys, C)
* Methionine (Met, M)
* Asparagine (Asn, N)
* Glutamine (Gln, Q)
5
New cards
Aromatic R groups
* Phenylalanine (Phe, F)
* Tyrosine (Try, Y)
* Tryptophan (Trp, W)
* Tyrosine (Try, Y)
* Tryptophan (Trp, W)
6
New cards
Positively charged R groups
* Lysine (Lys, K)
* Arginine (Arg, R)
* Histidine (His, H)
* Arginine (Arg, R)
* Histidine (His, H)
7
New cards
Negatively charged R groups
* Aspartate (Asp, D)
* Glutamate (Glu, E)
* Glutamate (Glu, E)
8
New cards
types of protein structures
primary, secondary, tertiary, quaternary
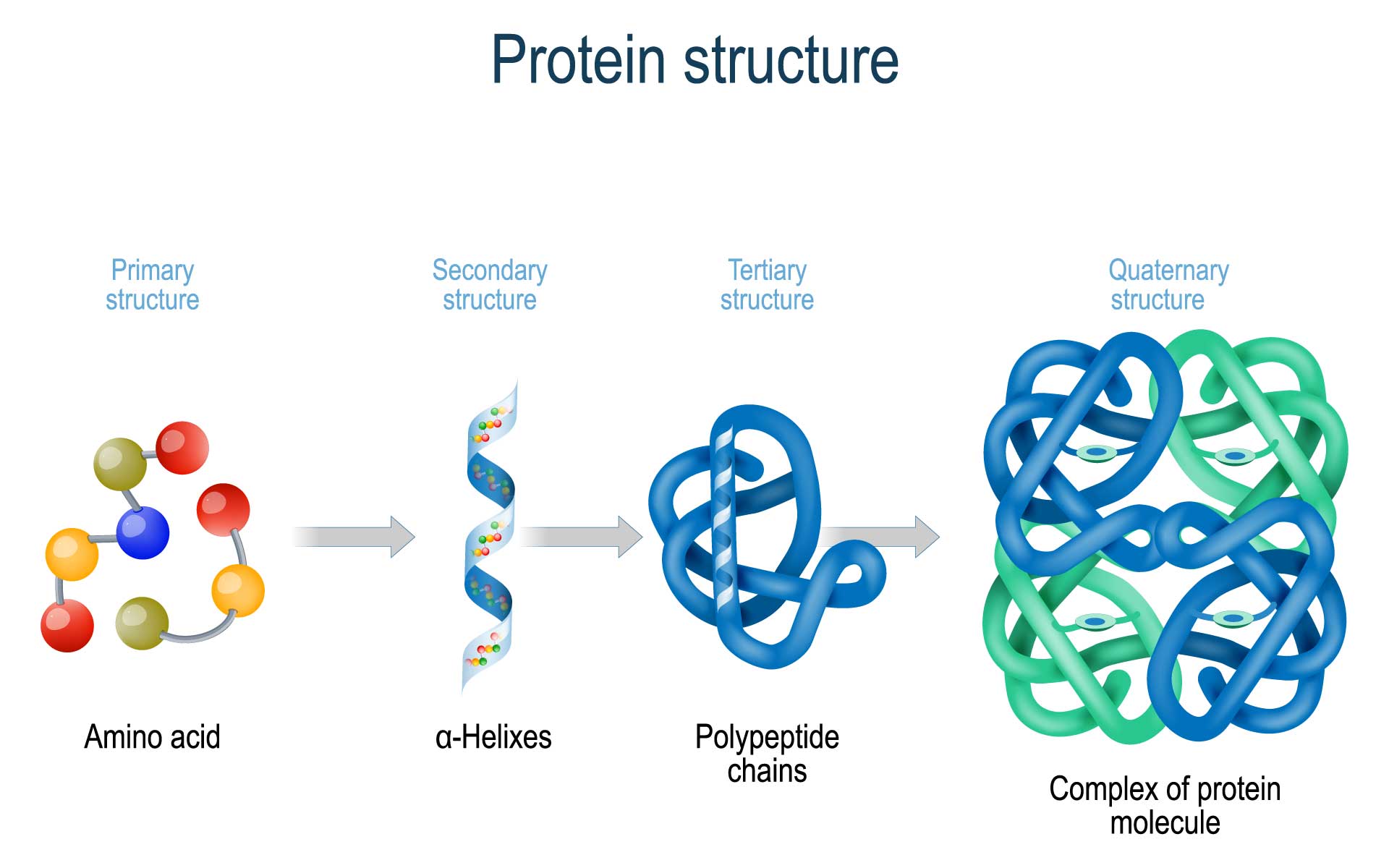
9
New cards
peptide bond
covalent bond formed between 2 amino acids
10
New cards
Francis Crick and James Watson
described the double-helix structure of DNA
11
New cards
genetic code is composed of nucleotide triplets
these three nucleotides are a codon, which specifies one amino acid
12
New cards
genetic code is non overlapping
each nucleotide belongs to one codon, which are read consecutively
13
New cards
genetic code is degenerate
all but two AA are specififed by more than one codon, usually 3rd base differs
ex: Arginine, CGU, CGC, CGA, CGG
ex: Arginine, CGU, CGC, CGA, CGG
14
New cards
wobble
a base in the 1st position of an anticodon can pair with any of several bases in position of 3 of codon
15
New cards
first position of tRNA
5’ end
16
New cards
third position of mRNA
3’
17
New cards
tRNA to mRNA bases
5’ to 3’
* A = U
* C = G
* G = C or U
* U = A or G
* I = A, U, or C
* A = U
* C = G
* G = C or U
* U = A or G
* I = A, U, or C
18
New cards
genetic code is ordered
codons for amino acids with similar chemical properties are closely related to each other
19
New cards
genetic code has start and stop codons
* start codon: AUG
* stop codons: UAA, UGA, UAG
* stop codons: UAA, UGA, UAG
20
New cards
genetic code is (almost) universal
same in viruses, bacteria, and humans
21
New cards
Nirenberg and Matthaei and Leder and Khorana
* the first to link specific coding sequences to specific amino acids
* laid cornerstone for the complete analysis of genetic code
* synthesized polypeptides in a cell free (in vitro) system for synthesizing proteins
* found means of producing synthetic mRNA to serve as templates for polypeptide synthesis in the cell-free system
* laid cornerstone for the complete analysis of genetic code
* synthesized polypeptides in a cell free (in vitro) system for synthesizing proteins
* found means of producing synthetic mRNA to serve as templates for polypeptide synthesis in the cell-free system
22
New cards
in-vitro translation
* synthesized polymers of RNA
* UUU-phenylalanine
* CCC-proline
* UUU-phenylalanine
* CCC-proline
23
New cards
in-vitro translation process
* uracil nucleotides w/ polynucleotide phosphorylase = Poly(U) homopolymer
* precipitate protein
* free amino acids separate from protein
* suction for further distillation
* precipitate protein
* free amino acids separate from protein
* suction for further distillation
24
New cards
4 parts of translation in prokaryotes
* tRNA charging
* initiation
* elongation
* termination
* initiation
* elongation
* termination
25
New cards
tRNA charging
aminoacyl-tRNA synthetase attaches correct amino acids to tRNA molecules (two step reaction, requires ATP)
26
New cards
initiation of translation
* IF1: binds to 30S subunit and prevents aminoacyl tRNA from binding to the A site
* IF2: binds f-met-tRNA to 30S-mRNA complex and hydrolyzes GTP
* IF3: binds to 30S subunit, preventing it from associating with the 50S subunit
* IF2: binds f-met-tRNA to 30S-mRNA complex and hydrolyzes GTP
* IF3: binds to 30S subunit, preventing it from associating with the 50S subunit
27
New cards
elongation of polypeptide
* EF-Tu: binds GTP, beings aminoacyl tRNA to the A site of the ribosome
* EF-Ts: generates active EF-Tu
* EF-G: stimulates translocation, GTP-dependent
* EF-Ts: generates active EF-Tu
* EF-G: stimulates translocation, GTP-dependent
28
New cards
termination of translation and release of polypeptide
* RF1: catalyzes release of the polypeptide chain from the tRNA and dissociation of the translocation complex; specific for UAA and UAG termination codons
* RF2: behaves like RF1, specific for UGA and UAA codons
* RF3: stimulates RF1 and RF2
* RF2: behaves like RF1, specific for UGA and UAA codons
* RF3: stimulates RF1 and RF2
29
New cards
translation components
* small subunit and large subunit in ribosome in GTP
* initiation factors
* elongation factors
* release factors
* anticodon and initiator tRNA
* many triplet codons
* initiation factors
* elongation factors
* release factors
* anticodon and initiator tRNA
* many triplet codons
30
New cards
initiation of translation
1. initiation factors bind to small subunits and attract mRNA
2. initiation complex: tRNA binds to AUG codon of mRNA in P site, forming initiation complex, IF3 is released
3. large subunit binds t complex; IF1 and IF2 are released. subsequent aminoacyl tRNA is poised to enter the A site
31
New cards
initiation
* must assemble components (ribosome, mRNA, fmet-tRNA, initiation factors, GTP)
* initiation factor 3 binds small ribosomal subunit (prevents large subunit from binding prematurely, IF1 assists)
* small subunit of ribosome binds to Shine Dalgarno sequence on mRNA
* IF2 forms complex with GTP
* fMet tRNA attaches to start codon on mRNA
* IF3 comes off, allowing large subunit to bind
* GTP hydrolyzes, IF1 and IF2 leave
* large ribosomal subunit binds
* tRNA-fMet is in P site
* initiation factor 3 binds small ribosomal subunit (prevents large subunit from binding prematurely, IF1 assists)
* small subunit of ribosome binds to Shine Dalgarno sequence on mRNA
* IF2 forms complex with GTP
* fMet tRNA attaches to start codon on mRNA
* IF3 comes off, allowing large subunit to bind
* GTP hydrolyzes, IF1 and IF2 leave
* large ribosomal subunit binds
* tRNA-fMet is in P site
32
New cards
three binding sites for ribosomal subunit
* E = exit
* P = peptidyl
* A = aminoacyl
* P = peptidyl
* A = aminoacyl
33
New cards
initiation: eukaryotes
* similar, but no shone Dalgarno
* 5’ cap is recognized
* slightly different initiation factors
* Poly A tail loops back to interact with proteins on the 5’ cap
* 5’ cap is recognized
* slightly different initiation factors
* Poly A tail loops back to interact with proteins on the 5’ cap
34
New cards
Elongation
* EF-Tu brings charged tRNA to A site
* requires GTP hydrolysis
* EF-Ts generate EF-Tu/GTP complex
* requires GTP hydrolysis
* EF-Ts generate EF-Tu/GTP complex
35
New cards
if codon and anticodon match:
1. peptide bond formed by peptidyl transferase (ribozyme)
2. translocation
1. ribosome moves 1 codon further
2. movement requires GTP hydrolysis and EF-G
3. uncharged tRNA is removed from E site
1. 2nd tRNA with growing peptide chain is in the P site, a site is empty
4. new charged tRNA enters A site, process repeats itself
36
New cards
termination
* ribosome moves until it gets to a stop codon
* released factors bind stop codons in A site
* GTP hydrolysis occurs to break polypeptide away from complex
* everything falls apart
* termination codon enters A site, RF1 or RF2 stimulates hydrolysis of the polypeptide from peptidyl tRNA
* ribosomal subunits dissociate and mRNA is released, polypeptide folds into native 3-D conformation of protein, charged tRNA released
* released factors bind stop codons in A site
* GTP hydrolysis occurs to break polypeptide away from complex
* everything falls apart
* termination codon enters A site, RF1 or RF2 stimulates hydrolysis of the polypeptide from peptidyl tRNA
* ribosomal subunits dissociate and mRNA is released, polypeptide folds into native 3-D conformation of protein, charged tRNA released
37
New cards
coupling of transcription and translation
only in prokaryotes (translation begins before transcription is complete)
38
New cards
polycistronic mRNA
* only in prokaryotes
* mRNA contains info for more than 1 gene
* each different gene has its own shine dalgarno and AUG
* often composed of genes for the same pathway
* mRNA contains info for more than 1 gene
* each different gene has its own shine dalgarno and AUG
* often composed of genes for the same pathway
39
New cards
why must antibiotics exploit differences between prokaryotic and eukaryotic cells?
* prokaryotic cells have exploitable differences from eukaryotic cells
* antibiotics like penicillin inhibit bacterial cell wall production with minimal effects on animal cells
* antibiotics like tetracycline inhibit prokaryotic ribosomes but have less effect in eukaryotes
* antibiotics like penicillin inhibit bacterial cell wall production with minimal effects on animal cells
* antibiotics like tetracycline inhibit prokaryotic ribosomes but have less effect in eukaryotes
40
New cards
puromycin
* looks like 3’ end of aminocyl tRNA
* binds A site, peptidyl transferase creates a bond between peptide and puromycin
* no more elongation, peptide released
* binds A site, peptidyl transferase creates a bond between peptide and puromycin
* no more elongation, peptide released
41
New cards
streptomycin
* binds to small ribosome subunits
* blocks initiation of translation
* blocks initiation of translation
42
New cards
tetracycline
prevents aninoacyl tRNA from binding to A site
43
New cards
chloramphenicol
* binds large subunit
* blocks peptidyl transferase activity
* blocks peptidyl transferase activity
44
New cards
gene regulation in prokaryotic cells
* no differentiation
* growth and survival
* growth and survival
45
New cards
genes are regulated primarily by:
* transcriptional control
* RNA stability
* translational control
* posttranslational control (protein function)
* RNA stability
* translational control
* posttranslational control (protein function)
46
New cards
inducible systems
synthesis of enzymes occurs only when substrate is present
47
New cards
repressible systems
synthesis of enzymes stops when product is not needed, detect excess of end products
48
New cards
inducible enzymes
made in response to a particular stimulus
49
New cards
constitutive enzymes
made continuously
50
New cards
lactose operon of E. coli
* sequence of adjacent genes under the transcriptional control of the same promoter and operator
* E.coli grows best using glucose as a carbon sources
* will grow on lactose
* E.coli grows best using glucose as a carbon sources
* will grow on lactose
51
New cards
additional enzymes required to grow E.coli on lactose
* normally transcribed at a very low rate
* induced in the presence of allolactose
* induced in the presence of allolactose
52
New cards
LacZ operon
* B-galactosidase
* splits lactose into glucose and galactose
* converts lactose into allolactose
* splits lactose into glucose and galactose
* converts lactose into allolactose
53
New cards
lacY operon
* lactose permease
* active transport of lactose into cell
* active transport of lactose into cell
54
New cards
LacA operon
* transacetylase
* function not well understood
* function not well understood
55
New cards
LacZ, Y, and A are what?
polycistronic (deciphers various proteins and is an attribute of many prokaryotic bacterial and chloroplast mRNAS, carried information for the synthesis of more than one protein)
56
New cards
Lac operon
* grow E.coli in glucose
* basal level of txn
* 3 molecules of B-gal per cell
* switch to lactose
* induced
* 3000 molecules of B-gal per cell
* Lac I
* produced constitutively
* produces laci repressor
* basal level of txn
* 3 molecules of B-gal per cell
* switch to lactose
* induced
* 3000 molecules of B-gal per cell
* Lac I
* produced constitutively
* produces laci repressor
57
New cards
the component of the wild-type lac operan and the response in the absence and presence of lactose
repressor gene, promoter, operator gene, leader, structural genes Z + Y + A
58
New cards
absence of lactose
* Lacl protein binds lac operator
* RNA pol can not bind promoter, cannot initiate txn
* Lac operon is off
* NOTE: low level of txn reflects the fact that repressor does not bind irreversibly
* RNA pol can not bind promoter, cannot initiate txn
* Lac operon is off
* NOTE: low level of txn reflects the fact that repressor does not bind irreversibly
59
New cards
presence of lactose
* B-gal converts some lactose to allolactose (inducer)
* allolactose binds repressor (allosteric shift)
* repressor dissociates from DNA
* RNA pol now initiates txn
* allolactose binds repressor (allosteric shift)
* repressor dissociates from DNA
* RNA pol now initiates txn
60
New cards
the lac operon in the absence of lactose
* lacl protein binds the lac operator
* if RNA polymerase cannot bind promoter, then transcription cannot be initiated
* this means the lac operon is OFF
* low level of transcription reflects the fact that repressor does not bind irreversibly
* if RNA polymerase cannot bind promoter, then transcription cannot be initiated
* this means the lac operon is OFF
* low level of transcription reflects the fact that repressor does not bind irreversibly
61
New cards
the lac operon in the presence of lactose
* B-galactose converts some lactose into allolactose (inducer)
* allolactose binds repressor (allosteric shift)
* repressor dissociates from DNA
* RNA polymerase now initiates transcription
* allolactose binds repressor (allosteric shift)
* repressor dissociates from DNA
* RNA polymerase now initiates transcription
62
New cards
mutations in lac operon system
* promoter of lac operon:
* so RNA pol cannot bind
* different results in glucose and lactose
* operator
* delete operator sequence
* Lacl
* delete gene for lac I
* or over-express lac I
* so RNA pol cannot bind
* different results in glucose and lactose
* operator
* delete operator sequence
* Lacl
* delete gene for lac I
* or over-express lac I
63
New cards
catabolite repression
* if both glucose and lactose are present, glucose will be used
64
New cards
catabolite repression with glucose
* adenyl cyclase inhibited by glucose
* not as much as CAP-cAMP
* not as much as CAP-cAMP
65
New cards
catabolite reaction with no glucose
* cAMP made by adenyl cyclase
* CAP binds cAMP
* CAP-cAMP binds DNA at CAP site (upstream of lac promoter)
* DNA bends, RNA poly binds more strongly
* CAP binds cAMP
* CAP-cAMP binds DNA at CAP site (upstream of lac promoter)
* DNA bends, RNA poly binds more strongly
66
New cards
catabolite repression with absent glucose
* glucose absent
* CAP (catabolite activating protein) + cAMP
* =
* promoter region (includes CAP-binding site and polymerase site)
* structural genes (transcription and translation occur)
* CAP (catabolite activating protein) + cAMP
* =
* promoter region (includes CAP-binding site and polymerase site)
* structural genes (transcription and translation occur)
67
New cards
cAMP facts
as cAMP levels increase, cAMP binds to CAP, causing an allosteric transition
68
New cards
catabolite repression with present glucose
* glucose
* CAP means cAMP levels decrease
* =
* CAP cannot bind efficiently
* RNA polymerase seldom binds
* transcription and translation diminished
* CAP means cAMP levels decrease
* =
* CAP cannot bind efficiently
* RNA polymerase seldom binds
* transcription and translation diminished
69
New cards
tryptophan operon
* repressible system
* turned off when end product builds up
* 5 structural genes A-E, promoter, operator, 162 bp leader region
* turned off when end product builds up
* 5 structural genes A-E, promoter, operator, 162 bp leader region
70
New cards
trpR
repressor located away from operon
71
New cards
components of repressible operon
* promoter, operator, leader, and attenuator
* repressor gene before regulatory region and structural genes in trp operon
* repressor gene before regulatory region and structural genes in trp operon
72
New cards
repressor proteins have what?
tryptophan binding site
73
New cards
with tryptophan abundant:
* tryptophan binds repressor, repressor binds operator, txn reduced 70x
74
New cards
without tryptophan
* repressor inactive
* txn occurs (attenuation possible)
* txn occurs (attenuation possible)
75
New cards
attenuation
* involves leader region between operator and 1st structural gene
* mRNA region termed leader transcript
* four subregions 1-4
* can form stem loops
* includes tryptophan codons
* mRNA region termed leader transcript
* four subregions 1-4
* can form stem loops
* includes tryptophan codons
76
New cards
in the attenuation model, tryptophan operon is regulated by:
changing direction
77
New cards
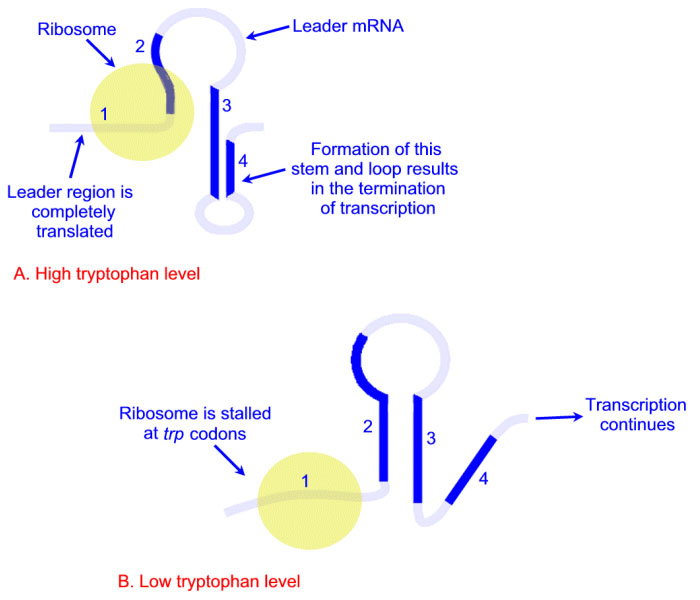
attenuation with tryptophan
* txn, tln proceed up leader peptide gene
* ribosome overlaps region 2
* 3-4 terminator stems forms
* txn terminated
* ribosome overlaps region 2
* 3-4 terminator stems forms
* txn terminated
78
New cards
attenuation without tryptophan
* ribosome pauses at trp codon
* 2-3 loop forms
* so 3-4 does not
* txn proceeds
* 2-3 loop forms
* so 3-4 does not
* txn proceeds
79
New cards
ribosome stals on tryptophan codons
allows formation of 2-3 stem loop = transcription proceeds
80
New cards
ribosome moves into 1-2 region
prevents formation of 2-3 step loop, transcription terminates and translation initiated
81
New cards
high levels of tryptophan
* complete leader peptide
* ribosome
* trp codons 1 & 2 on line
* 3 & 4 on loop
* stem loop: transcription termination signal
* ribosome
* trp codons 1 & 2 on line
* 3 & 4 on loop
* stem loop: transcription termination signal
82
New cards
low levels of tryptophan
* incomplete leader peptide
* ribosome stalls
* alternate stem loops: transcription continues
* trp codons 1 & 4 on line
* codons 2 & 3 on loop
* trp regulated genes at the end
* ribosome stalls
* alternate stem loops: transcription continues
* trp codons 1 & 4 on line
* codons 2 & 3 on loop
* trp regulated genes at the end
83
New cards
attenuation: general starvation
* no ribosomes
* 1-2 and 304 formed
* txn terminated
* repression occurs
* 1-2 and 304 formed
* txn terminated
* repression occurs
84
New cards
more complicated regulation in eukaryotic cells
areas of control:
* chromatin structure
* transcription activators/enhancers/silencers
* RNA processing/degredation
* RNA interference
* translation/protein modification
* epigenetics
* chromatin structure
* transcription activators/enhancers/silencers
* RNA processing/degredation
* RNA interference
* translation/protein modification
* epigenetics
85
New cards
chromatin structure: location of nucleosomes
may occlude binding of transcription factors to promoter, chromatin remodeling proteins may be required
86
New cards
chromatin structure: histone acetylation
destabilize nucleosome structure
87
New cards
chromatin structure: histone methylation
may activate or silence, depends on specific amino acid method
88
New cards
in diagrams, acetylation does what?
loosens positively charged tail
89
New cards
DNA methylation
* on CpG islands (5’ CG 3’)
* Cpg islands often found at promoter sites
* cytosine methylated on carbon 5 (methyl group may sit at major groove)
* methylation associated with inactive genes
* methylation status is heritable
* Cpg islands often found at promoter sites
* cytosine methylated on carbon 5 (methyl group may sit at major groove)
* methylation associated with inactive genes
* methylation status is heritable
90
New cards
transcription
* mediated by several factors
* proteins bind to enhancers and/or promoters
* several transcription factors needed
* tissue specific enhancers
* response elements coordinate regulation
* Prokaryotes just need sigma factor and RNA polymerase
* proteins bind to enhancers and/or promoters
* several transcription factors needed
* tissue specific enhancers
* response elements coordinate regulation
* Prokaryotes just need sigma factor and RNA polymerase
91
New cards
mRNA processing and stability
* RNA must be processed to mature mRNA
* polymerase A splicing
* how stable is the transcript? will it last months or minutes?
* polymerase A splicing
* how stable is the transcript? will it last months or minutes?
92
New cards
translation produces large T antigen while translation produces a…
small t antigen
93
New cards
translational control
* regulation of mRNA localization and localized translational control are governed by cis-regulatory sequences on the mRNA and trans-acting RNA-binding proteins
94
New cards
protein modification and degradation
* must undergo this process before translation, uses RNA-stabilizing proteins which protect it from degradation
* most important part is addition of stabilizing and signaling factors at the 5’ and 3’ ends of the molecule, and the removal of intervening sequences that don’t specify the appropriate amino acids
* most important part is addition of stabilizing and signaling factors at the 5’ and 3’ ends of the molecule, and the removal of intervening sequences that don’t specify the appropriate amino acids
95
New cards
epigenetics
* changes in gene expression NOT related to changes in DNA sequence
* alterations in chromatin structure
* methylation patterns in DNA
* DNA methylation associated with histone deacetylation
* regulation by noncoding RNA
* alterations in chromatin structure
* methylation patterns in DNA
* DNA methylation associated with histone deacetylation
* regulation by noncoding RNA
96
New cards
epigenetics example “good mother rats”
offspring that received more grooming/licking have a different pattern of DNA methylation (genes involved in stress response are expressed
97
New cards
DNA methylation
* on CpG islands
* 5’ CG 3’
* cytosine is methylated on carbon 5
* methyl group may sit at major groove
* methylation associated with inactive genes
* methylation status is heritable
* 5’ CG 3’
* cytosine is methylated on carbon 5
* methyl group may sit at major groove
* methylation associated with inactive genes
* methylation status is heritable
98
New cards
99
New cards
genomic imprinting
gene expression depends on the parent of origin
100
New cards
Prader-Willi and Angelman syndromes
Prader-Willi
* small, poor sexual development, below average IQ
* develop huge appetites and become obese
Angelman
* uncontrolled laughter, muscle movement, seizures
* developmental delays
Deletion on chromosome 15
* PW: delete father’s copy
* Angelman: delete mother’s copy
* small, poor sexual development, below average IQ
* develop huge appetites and become obese
Angelman
* uncontrolled laughter, muscle movement, seizures
* developmental delays
Deletion on chromosome 15
* PW: delete father’s copy
* Angelman: delete mother’s copy