Bio 4 Exam Slideshow
1/117
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
118 Terms
Transcription
DNA to RNA
Nucleotides to nucleotides
Transcription
Translation
RNA to protein
What is the process of transcription
DNA must unwind (called transcription bubble), Template strand (RNA reads this), Non-template strand (coding strand)
Eukaryotic transcription-initiation
Promotors (TATA box), Transcription Factors
Transcription Elongation
RNA polymerase moves along the strand, mRNA transcript elongates (40 nucleotides per second)
What does ribosomes do in translation elongation?
Ribosomes synthesize polypeptides in the N-terminal to C-terminal direction
What does occurs in gene regulation for prokaryotes?
Three processing steps where regulation can occur
What does occurs in gene regulation for eukaryotes?
Four processing steps where regulation can occur
What does histone methylation do in histone modification?
Nucleosomes pack together tightly, transcription factors can’t bind to DNA, and genes are not expressed
Nucleosomes pack together loosely, transcription factors can bind to DNA, and genes are expressed
What does histone acetylation do in histone modification?
What does miRNAs do in eukaryotic post-transcriptional control?
Base pairs with mRNA, can target mRNA for degradation or, prevent initiation complex from forming
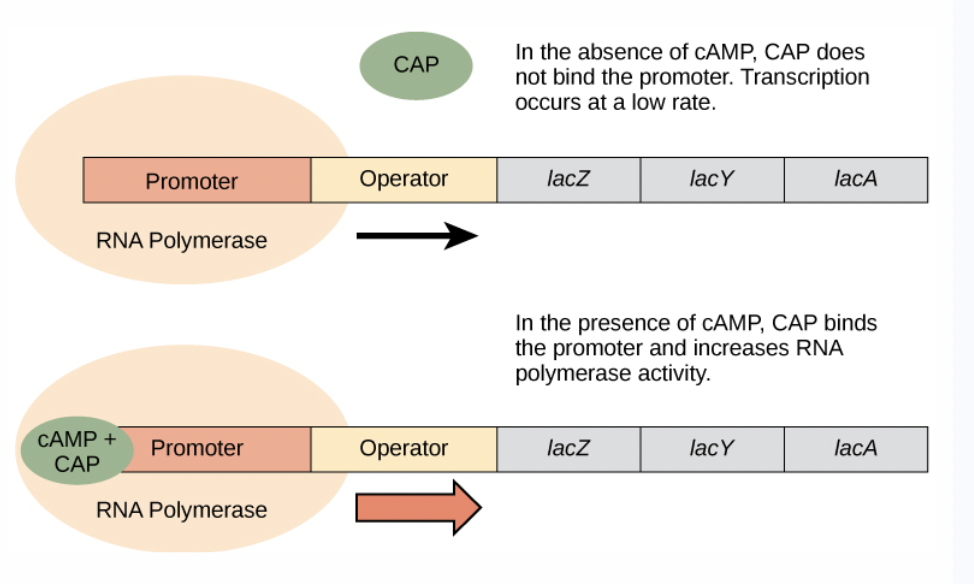
Is this positive or negative transcriptional control?
Positive regulation
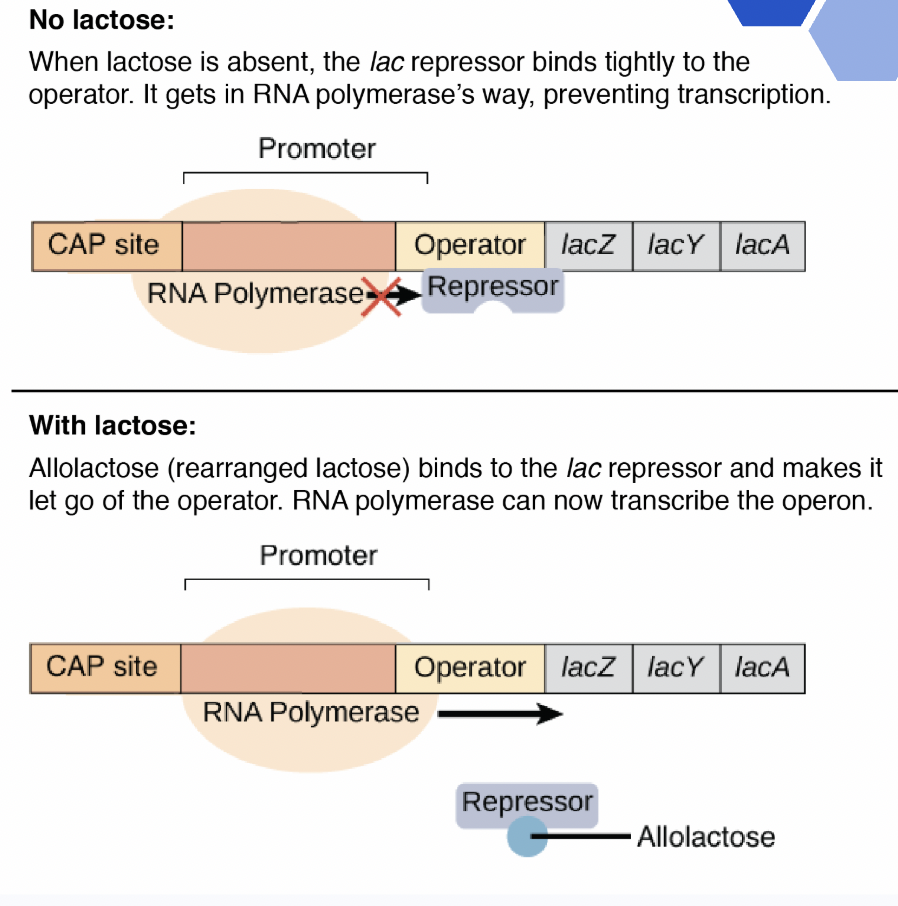
Is this positive or negative transcriptional control?
Negative regulation
Operons allow bacteria to control when they turn on or turn off expression of genes
What do Operons allow bacteria to do?
What are operons controlled by?
All genes in the operon are controlled by the same regulatory element
What is the crucial step of a repressor binding?
Repressor binding to operator element is the crucial step of controlling operon function
What happens to the Lac I repressor protein when it binds to lactose, and how does this affect the transcription of the lac operon?
When bound to lactose, Lac I repressor protein is unable to bind to the lac operator, resulting in transcriptional induction of the lac operon when lactose is present
Transcriptional, post-transcriptional, translational, and post-translational levels
Control of gene expression in eukaryotic cells occurs at which level(s)?
What do Kinases do?
It adds phosphate groups
Remove them and changes the shape and activity of a protein
What do Phosphatases do?
In Post-Translational Control what does control protein folding do?
Cleave part of the amino acid sequence and proenzyme → enzyme
What happens to prokaryotes only in transcription and translation?
Transcription occurs in cytosol and transcription and translation can occur simultaneously
Transcription occurs in the nucleus, and RNA processing occurs before translation
What happens to eukaryotes only in transcription and translation?
It contains promoter regions, and additional proteins assist in the binding of RNA polymerase
What happens to prokaryotes and eukaryotes only in transcription and translation?
What happens to ribosome only in transcription and translation?
It catalyzes peptide bond formation, is made up of two subunits, and binds to start codon
What happens to mRNA only in transcription and translation?
It contains codons and is composed of nucleotides only
What happens to tRNA only in transcription and translation?
It contains anticodons, it binds to a start codon, and is composed of nucleotides only
DNA must unwind, a complicated process that helps regulate gene expression, called a transcription bubble
In transcription, what must DNA unwind?
It is a protein that regulates if a gene is transcribed or not
What is a transcription factor?
Some example of transcription factors are
A vitamin D receptor, retinoic acid receptor, and estrogen receptors
In transcription-elongation what does RNA polymerase do?
RNA polymerase moves along the strand and mRNA transcript elongates
How are splicing done in eukaryotic transcription-RNA processing?
They are done by things called small nuclear ribonuclear proteins (snPNPs). Exons are expressed, but introns are not
In translation, what does ribosomes do?
It has large and small subunits that dissociate when it’s not used
In translation, do ribosomes’ large subunits or small subunits bind first?
Small subunits binds first
Eukaryotic transcription termination
Not as well understood at prokaryotes, terminator equals particular sequences that signals the end of transcription
Eukaryotic transcription RNA processing
Alternative splicing, 5' capping, Poly A tail
5' capping
7 methylguanosine cap
Special codons
AUG (start), UAA, UAG, UGA (stop)
Translation basics
tRNA, transfer RNA, composed of RNA, anticodon is key
translation initiation
Requires initiation factors, small subunit and methionine's tRNA, cap binding proteins recognizes 5' cap, moves 5' to 3' til it gets to AUG, large subunit binds last
Translation elongation
3 compartments (A-aminoacyl site, P-peptidyl site, E-exit site), tRNA enters A (except the first methionine), peptide bond forms in P, E is for exiting
Translation termination
Stop codon signals the end, complex dissociates, protein folding
Protein folding
Some proteins need help folding, often a signal sequence is added as well
What are the components of an amino acid?
center carbon, carboxyl group, amino group, hydrogen atom, R group (side chain)
The central dogma refined
Gene (dna) -> pre-mRNA -> mature RNA -> polypeptide -> functional protein
transcription bubbles
the dna must be opened up and unwound and this is formed
Promoter
Where transcription actually starts
transcription factors
bind to promoter sequence, direct RNA polymerase to begin transcription
terminator
directs the end of transcription, proteins interact with the terminator sequence and cause dissociation of RNA polymerase
5' UTR
untranslated region on the 5' end
3' UTR
untranslated region on the 3' end
direct binding to DNA
estrogen (E2) enters the plasma membrane and binds to nuclear ERS (nERs) then translocates into the nucleus, directly induces transcriptional changes in estrogen-responsive genes
binding to transcription factors
E2-ER complex can interact with transcription factore
anticodon
sequence found at the end of tRNAs that complement a codon
aminoacyl site
receives incoming charged tRNA
peptidyl site
formation of peptide bonds occurs
exit site
uncharged tRNA leaves the ribosome
initiation complex
mRNA + small subunit + Met tRNA, cap binding protein recognizes 5' cap, moves5' to 3' until it gets to AUG, large subunit binds last
Stop codon
signals the end, complex dissociates
release factor
causes the ribosome to disassemble and the synthesized polypeptide is release
bacteria
almost exclusively transcription
eukaryotes
transcription, RNA processing, translation, post-translational
bacteria transcription control
most of bacteria control of regulation is focused on transcription (if it is transcribed it is usually translated and then active), operons
eukaryotes control of transcription
transcription factors, methylation, miRNA
methylation
addition of a methyl group
5' Cap, poly A tail
slow exonucleases, can change rate of degredation
eukaryotes post translational
regulation by adding to proteins (phosphate), kinases add phosphate to proteins, phosphatases remove a phosphate to a protein
acetyl groups
similar to phosphate, addition or removal activates or deactivates
ubiquitin
added to protein, can do many things (degradation, move to different area, alter activity, prevent interactions)
prokaryotes gene regulation
three processing steps where regulation can occur, most occurs on the transcriptional level
Eukaryotes gene regulation
four processing steps where regulation can occur, regulation is distributed across levels, transcriptional regulation is very important though
transcriptional control methods
regulation by environment and cell signaling, epigenetic regulations, depends on transcription factors and regulatory elements
vitamin D receptor
works with vitamin D and the retinoic acid receptor
CpG islands
stable methylation of cytosine, preserved through cell division
chromosome packaging
single human somatic cell can span up to 6.5 ft when untwisted and stretched out, chromosome packaging is highly organized
histone acetylation
adding an acetyl group to histones, nucleosomes pack together loosely, transcription factors can bind to DNA, genes are expressed
miRNAs
micro RNAs, small, non coding, RNA molecules, base pairs with mRNA, can target mRNA for degradation or prevent initiation complex from forming
positive regulation activator
enhances polymerase binding, increasing transcription
negative regulation repressor
blocks transcription
phosphorylation
kinases add phosphate groups, phosphates remove them, changed the shape and activity of a protein, can either activate or deactivate a protein.
What are the types of mutations?
Silent, missense, nonsense, and frameshift
silent mutation
does not change amino acid sequence, usually happens when the mutation is in the 3rd codon
missense mutation
changes the amino acid, it might change the protein or not, depends on the amino acid changed, sickle cell anemia is the result of missense
nonsense mutation
AA -> stop codon, produces truncated protein, usually not good, some types of cystic fibrosis
frameshift mutation
insertion or deletion not divisible by 3, alerts the reading from, changes every AA following the mutation, usually very bad
wild type mutant
most common/consensus/original variant of a gene
spontaneous mutation
results of abnormality in DNA replication or crossing over/anaphase
induced mutation
caused by something environmental, chemical (caused by some kind of chemical, nitrous acid, base analogs), physical (something from the environment, radiation)
germaline
mutation in gametes, passed onto next generation
somatic
occurring elsewhere in the body, not passed on
DNA repair
nucleotide excision, Uvr proteins, proofreading and repair mechanism, removes large section of DNA
nucleotide excision
thymine dimers, modified bases, mismatch + missing bases
proto-oncogene
when mutated becomes oncogene, expressed at high levels, control cell growth and cell division, gain function mutations in proto-oncogenes lead unregulated growth
tumor supressor gene
when functioning prevent cancer
EGFR pathway
controls growth and cell division in some tissues Ras -> Raf -> Mek -> Erk, Ras is a proto-oncogene, missense mutation changes it to an oncogene
P53
Guardian of the genome, half of all cancers connect to p53, controls ability of cell to pass in to G1, can activate apoptosis, tumor supressor
EGF Receptor
part of the signaling pathway with RAS, when activated signals go through cell division, usually responds to epidermal growth factor, but can get stuch on
G1, first gap
cell grows, preparing for DNA replication
S, synthesis
DNA replication