GEN Exam 3 CHP 10-13
1/100
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
101 Terms
nucleic acids
the coding instructions for all living organisms
four important characteristics before DNA was discovered
1. the genetic material must contain complex information
2. genetic material must replicate faithfully
3. genetic material must encode the phenotype
4. genetic material must have the capacity to vary
the genetic material must contain complex information
- must be capable of storing large amounts of information and instructions for the traits and functions of organisms
genetic material must replicate faithfully
- genetic instructions must be copied with fidelity to be transmitted to progeny cells
genetic material must encode the phenotype
- must have the capacity to be expressed as a phenotype to code for traits
genetic material must have the capacity to vary
- genetic information must have the ability to vary, since different species and even individual members of the same species differ in their genetic makeup
Miescher
- biochemist who used white blood cells in pus that have large nuclei
- isolated DNA and protein, named it nuclein
- physical basis of heredity lies in the nucleus
- nuclein contained large amounts of phosphorus but no sulfur
- nuclein was not a protein
- nuclein is now DNA(deoxyribonucleic acid)
kossel
- found DNA contained four nitrogenous bases
- adenine, cytosine, guanine, and thymine
levene
- found DNA had large linked, repeating units called nucleotides
- each nucleotide contains a sugar, a phosphate, and a base
- incorrectly proposed that DNA was composed of repetitions of the same sequence of four nucleotides
- tetranucleotide hypothesis
chargoff rules
- disproved levene hypothesis
- studied base-composition of DNA
- the proportion of adenines equal that of thymines, and the proportion of guanines equals that of cytosines
- there is an equal proportion of purines(A&G) and pyrimidines(C&T)
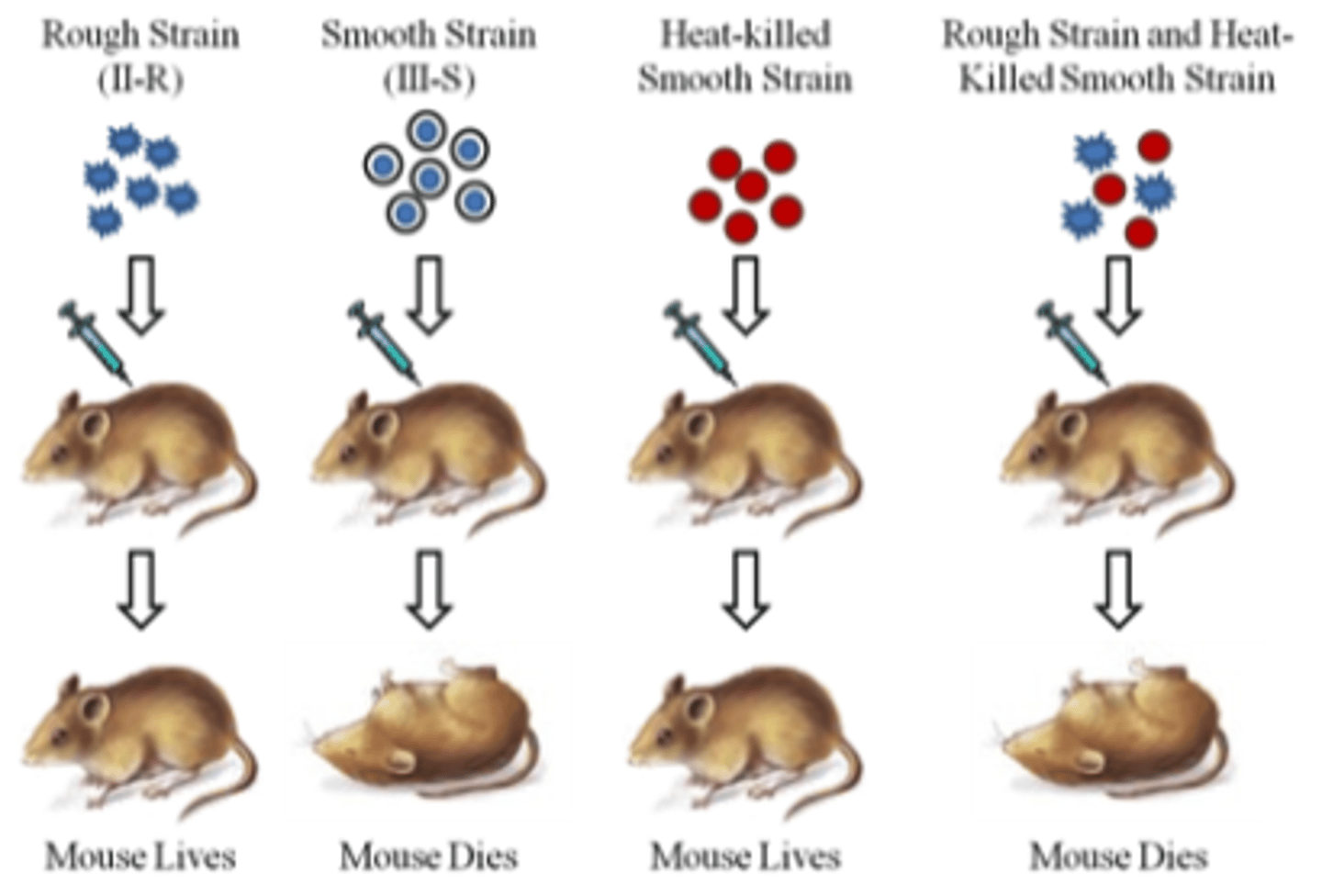
transformation principle
- initial step in identifying DNA as the source of genetic information
- first observed by Griffith, who used bacterium that caused pneumonia
- found smooth and rough coat
- some substance is capable of genetically altering bacteria
- a heritable change in a cell/organism brought about by exogenous DNA
- heat kills IIIS cells, but unknown substance survives and is capable of transforming IIR cells into the virulent IIIS cells
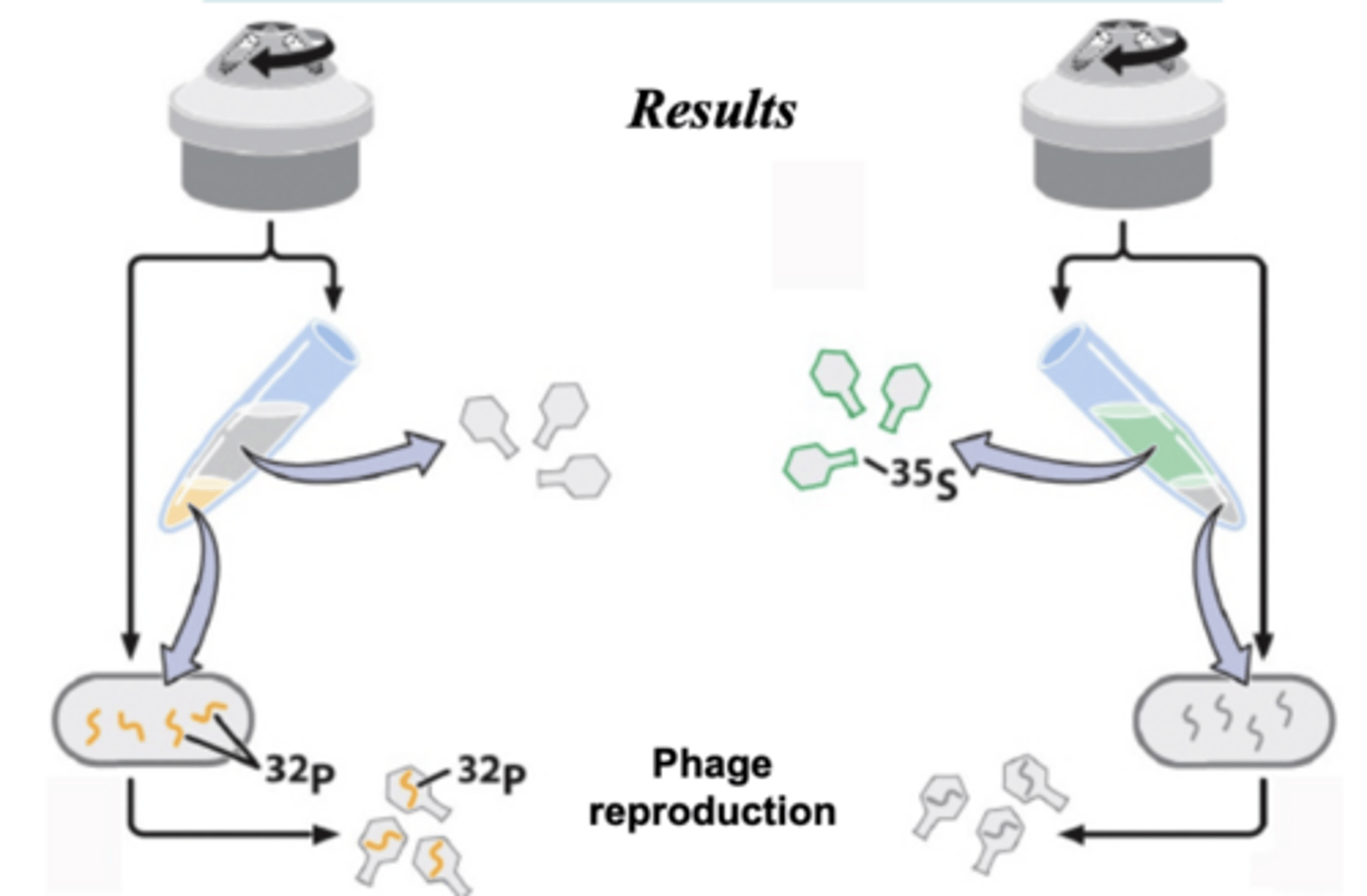
hershey-chase
- found DNA is not a protein but genetic material in bacteriophages
- DNA is the infectious agent in bacteriophages
- used radioactivity with phosphorus and sulfur
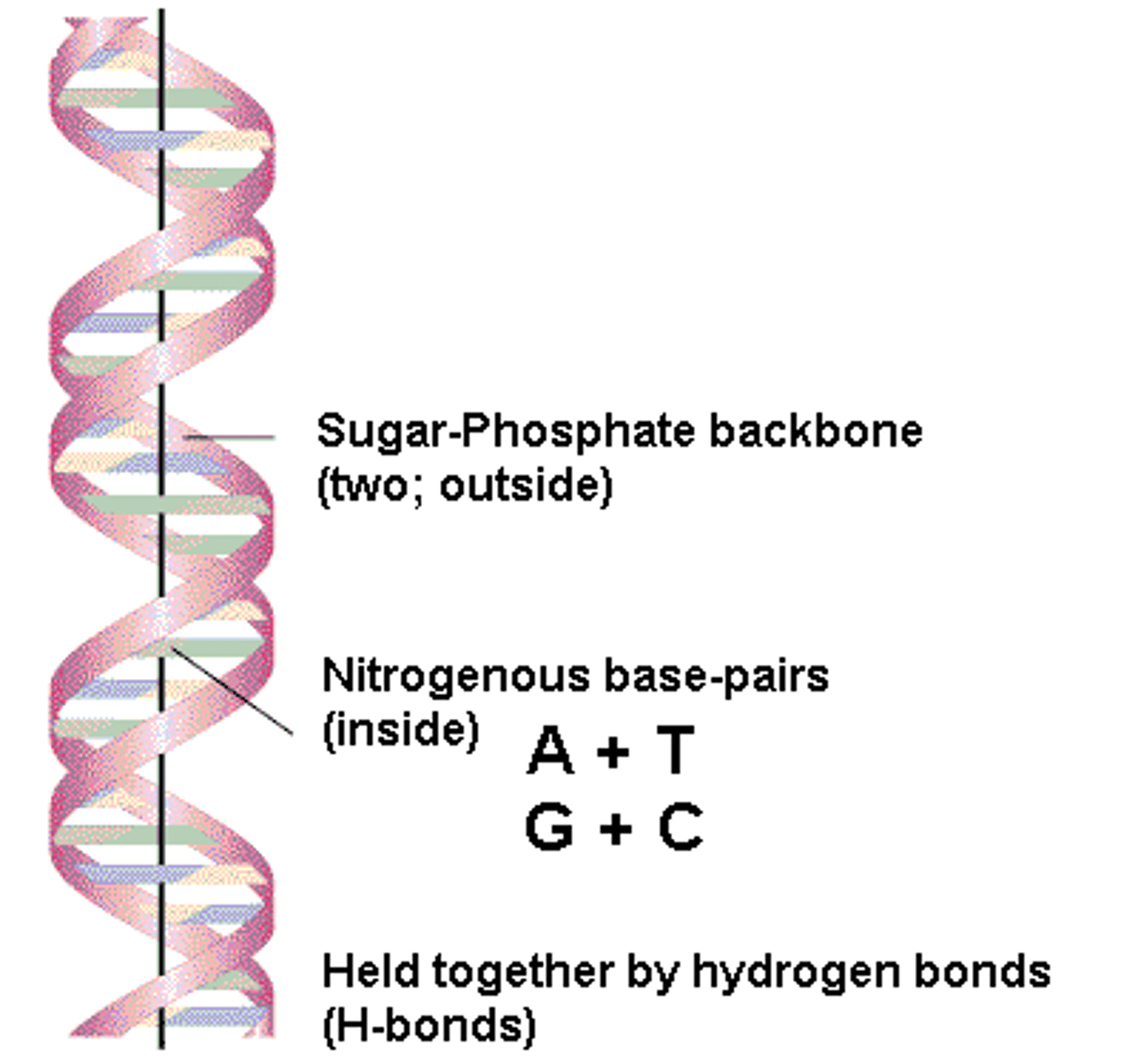
watson and crick
- proposed the double-helix model of DNA
- -deoxyribonucleic acid(DNA) is a double helix with antiparallel strands
- pentose sugars and phosphates backbone
- nitrogenous bases paired at the center by hydrogen bonds
- alternating monir and major grooves
- length of one whole turn on the helix is approx 10 base pairs
RNA
- genetic material in the virus of tobacco plants
forming double helix
- primary structure of DNA
- secondary structure of DNA
- tertiary structure of DNA
primary structure of DNA
- nucleotide structures are joined by phosphodiester linkages
seconday structure of DNA
- DNA's stable three-dimensional configuration is a double helix
- sugar-phosphate linkages are on the outside of helix
- bases are stacked in the interior
- two polynucleotide strands run in opposite directions
- strands are held together by hydrogen bonds and stacking of force
nucleotides
- involved with primary structure of DNA
- the repeating units of DNA that have three parts: a sugar, a phosphate, and nitrogenous base
- in both DNA and RNA
- sugar + nitrogenous base + phosphate
ribose
- RNAs sugar
- has a hydroxyl group attached to 2' carbon
- ribonucleic acid
deoxyribose
- DNA's sugar
- has hydrogen atom at 2' carbon
- deoxyribonucleic acid
second component of nucleotide base
- two types: purine or pyrimidine
- both DNA and RNA have 2 purines
purine
- type of DNA bases
- double carbon and nitrogen aromatic ring structures
- adenine(A) and guanine(G) occur in both DNA and RNA
pyrimidine
- single carbon and nitrogen aromatic ring structures
- cytosine(C) occurs in both DNA and RNA
- thymine(T) occurs in DNA only
- uracil(U) occurs in RNA only
cytosine
- in both DNA and RNA
thymine
- only in DNA
uracil
- only in RNA
nucleoside
- a deoxyribose or a ribose sugar + nitrogenous base
phosphate group
- carry a negative charge that makes DNA acidic
- always bonded to the 5' atom of the sugar in a nucleotide
- DNA called deoxyribonucleotides
- RNA called ribonucleotides
polynucleotide strands
- has phosphodiester linkages
- a strand of nucleotides linked in this way
phosphodiester linkages
- link nucleotides in the polynucleotide chains
- bond between two nucleotides
- has oxygens(ORs) on both sides
- DNA and RNA
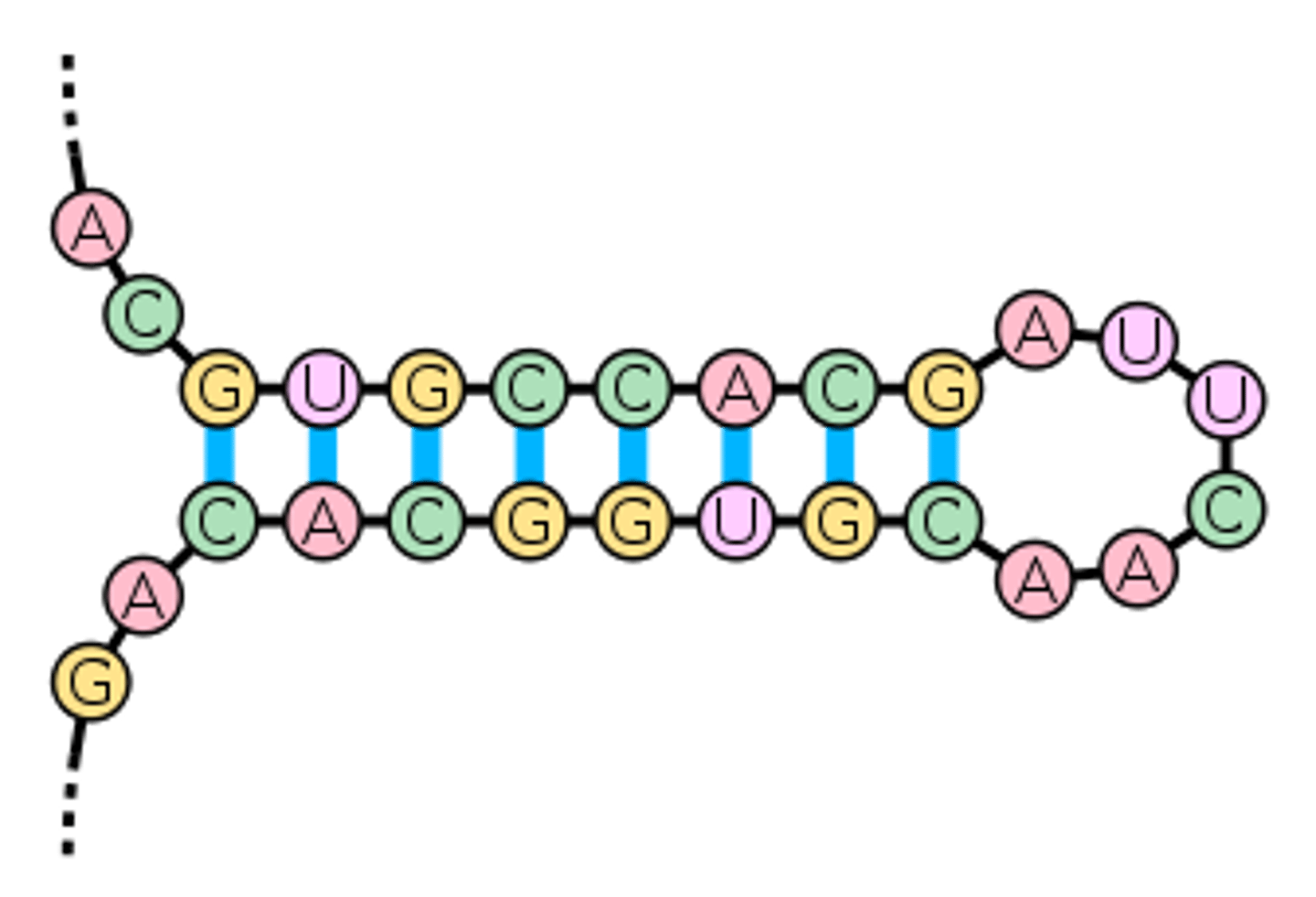
hairpin
- type of secondary structure
- forms when sequences of nucleotides on the same strand are inverted
- has a region of pairs bases(the stem) and intervening unpairs bases(a loop)
H-DNA
- three-stranded structure
- DNA unwinds and a single polynucleotide strand pairs with double stranded DNA
DNA methylation
- methyle groups are added by specific enzymes to certain positions on nitrogenous bases
central dogma
- molecular bio proposed that information flows in a one-way direction from DNA to RNA to protein
conservative replication
- entire double-stranded DNA molecule serves as a template for a whole new molecule of DNA
- the original DNA molecule is fully conserved during replication
dispersive replication
- both nucleotide trands break down into fragments that serve as templates for synthesis
- DNA fragments somehow reassemble into two complete DNA molecule
- original DNA is not conserved
semiconservative replication
- intermediate between two models
- the two nucleotide strands unwind and each serve as a template for new DNA molecule
meselson and stahl
- equilibrium density gradient centrifuge proved DNA replicates semi-conservatively
- DNA migrates to the point within the gradient where its own density matches the sencsity of CsCl
- used E. coli in a medium of 15N(heavy) and 14N(light)
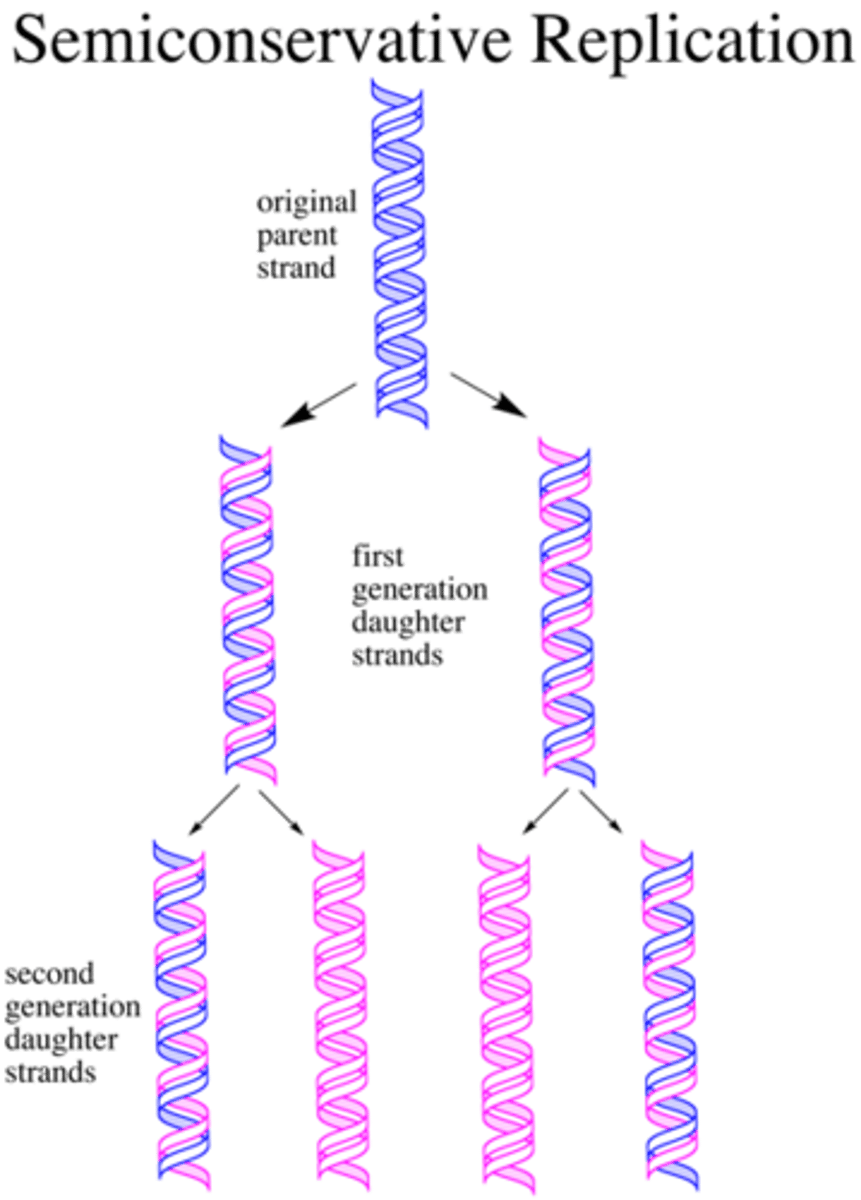
modes of replication
- replicon:a segment of DNA that undergoes replication
- origin of replication: each replicon has an origin
- replication starts at the origin and continues until the entire replicon has been replicated
- theta replication
- rolling-circle replication
- linear eukaryotic replication
theta replication
- takes place in circular DNA(like e. coli)
- generates intermediate structure that looks like theta
- double stranded DNA unwinds at the origin making single nucleotide strands
- replication bubble: unwinding of double helix that generates a loop
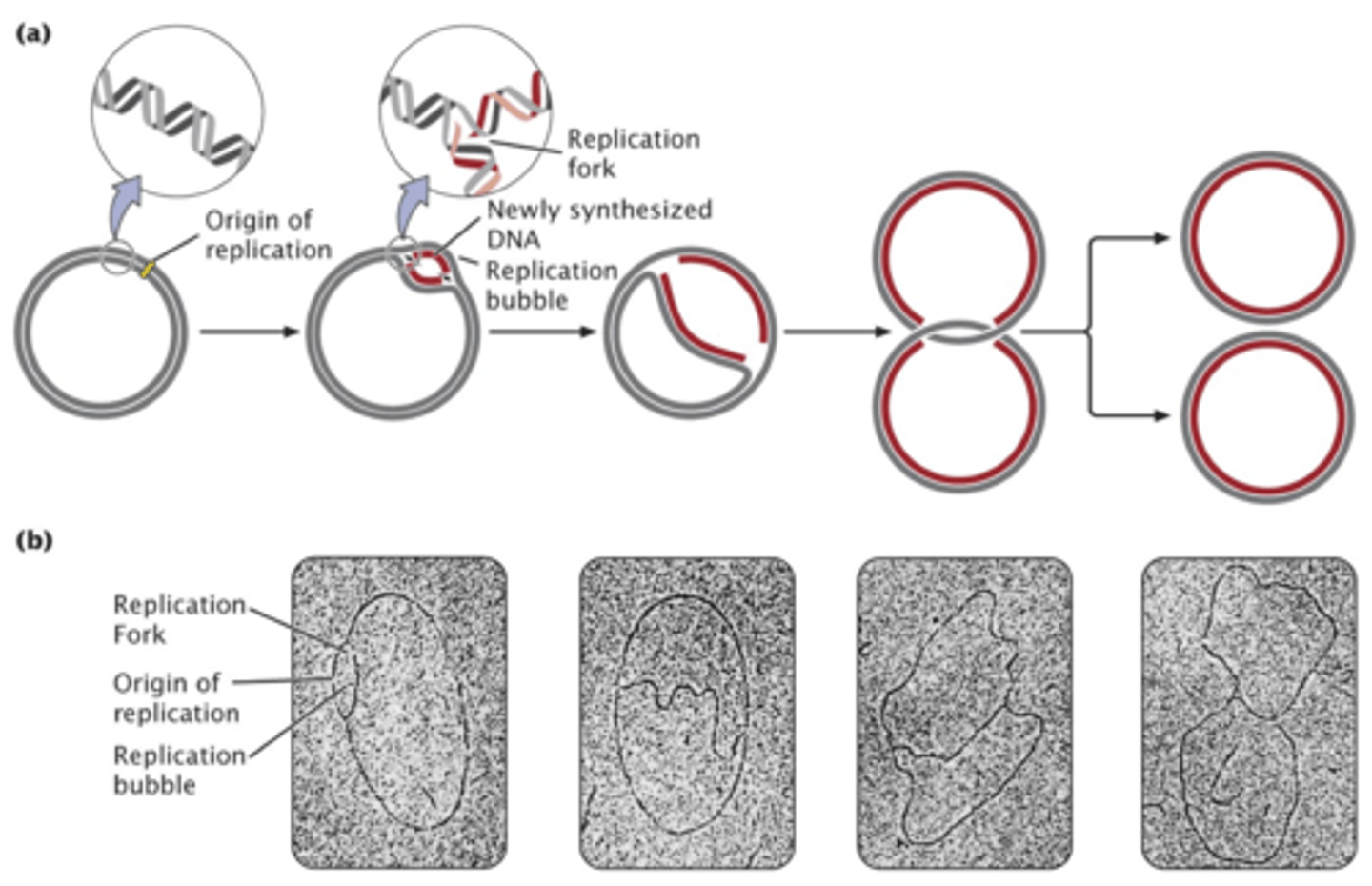
replication fork
- point of unwinding where two strands separate from the double stranded DNA helix
bidirectional replication
- two replication forks, one at each end of bubble, proceed outward and unwinds + replicates DNA

rolling circle replication
- initiated by a break in one of the nucleotide strands then new nucleotides are added
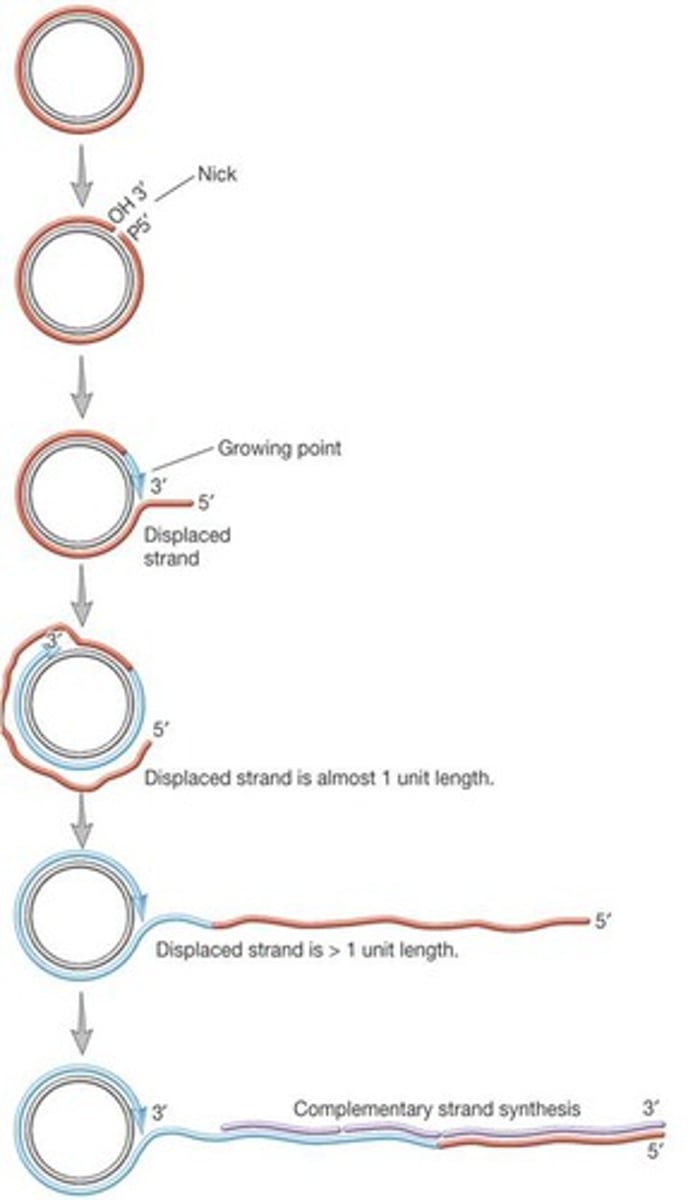
linear eukaryotic replication
- too much DNA so it unwinds and produces a replication bubble
- replications occurs on both strands at each end of the bubble with replication forks spreading outward
- eventually run into each other and fuse, making one long new DNA
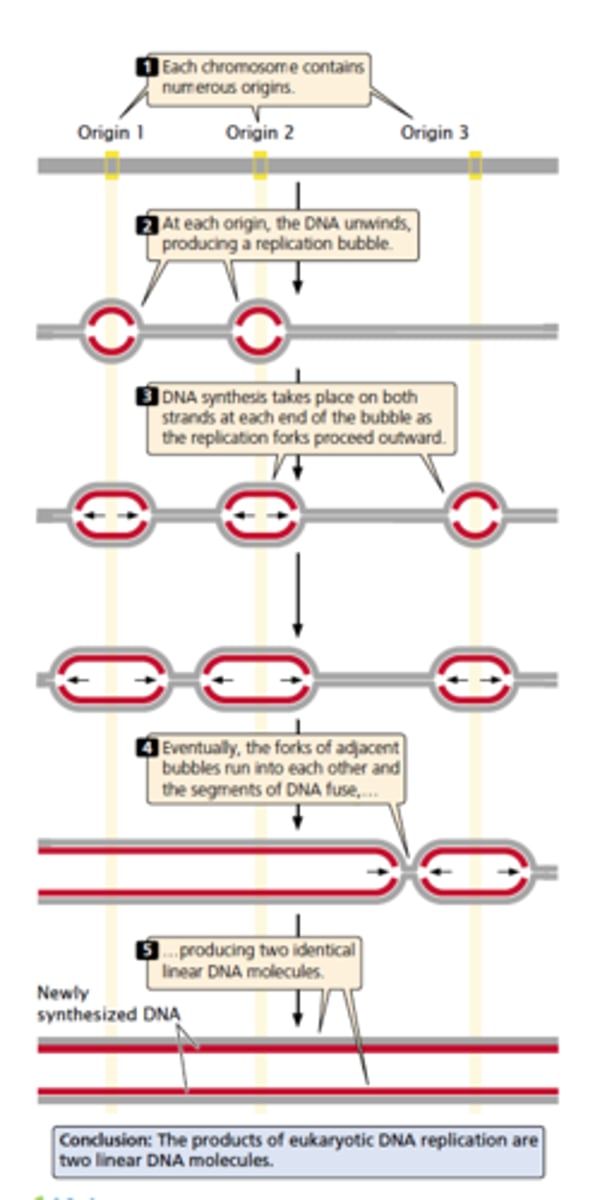
requirements of replication
1. a template consisting of single stranded DNA
2. raw materials to be assembled into new nucleotide strand
3. enzymes and other proteins that read the template and assemble substrates into DNA
griffith
- discovered bacterial transformation in mice
- proposed four properties of the unknown genetic material
properties of unknown genetic material
- capable of replication
- capable of storing information
- complex enough to generate thousands of gene products
- any changes in genetic material(mutations) should directly lead to changes in phenotypes
transformation
- a heritable change in a cell or organism brought about by exogenous DNA
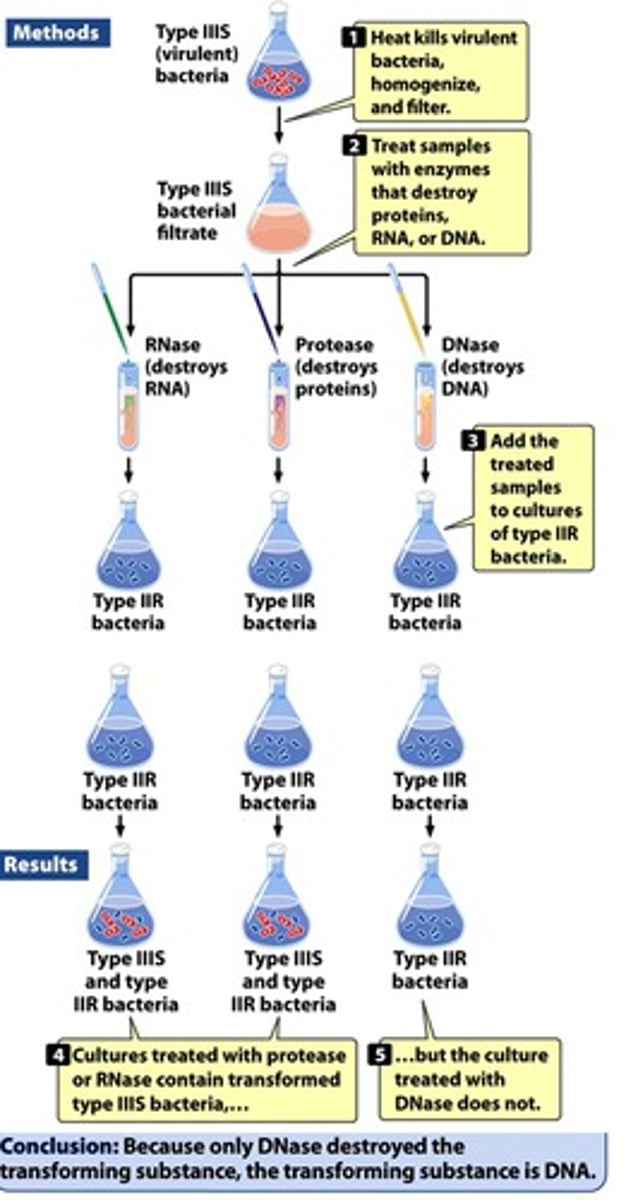
Avery, MacLeod, and McCarty experiments
- large quantities of IIIS bacterial cultures in liquid media were centrifuged, collected, heat-killed, lysed, and homogenized
- enzymes and extractions with detergents and organic solvents eliminated carbohydrates, lipids, and proteins, resulting in a solution that had a nitrogen-phosphorus ratio consistent with a nucleic acid that could transform IIR cells into IIIS cells
- mixtures with the enzymatic destruction of proteins, RNA, or DNA
- resulting mixtures were added to cultures of IIR cells
- because only DNase destroyed the transforming substance, the transforming substance is DNA
- IIIS DNA can transform IIR cells into IIIS cells
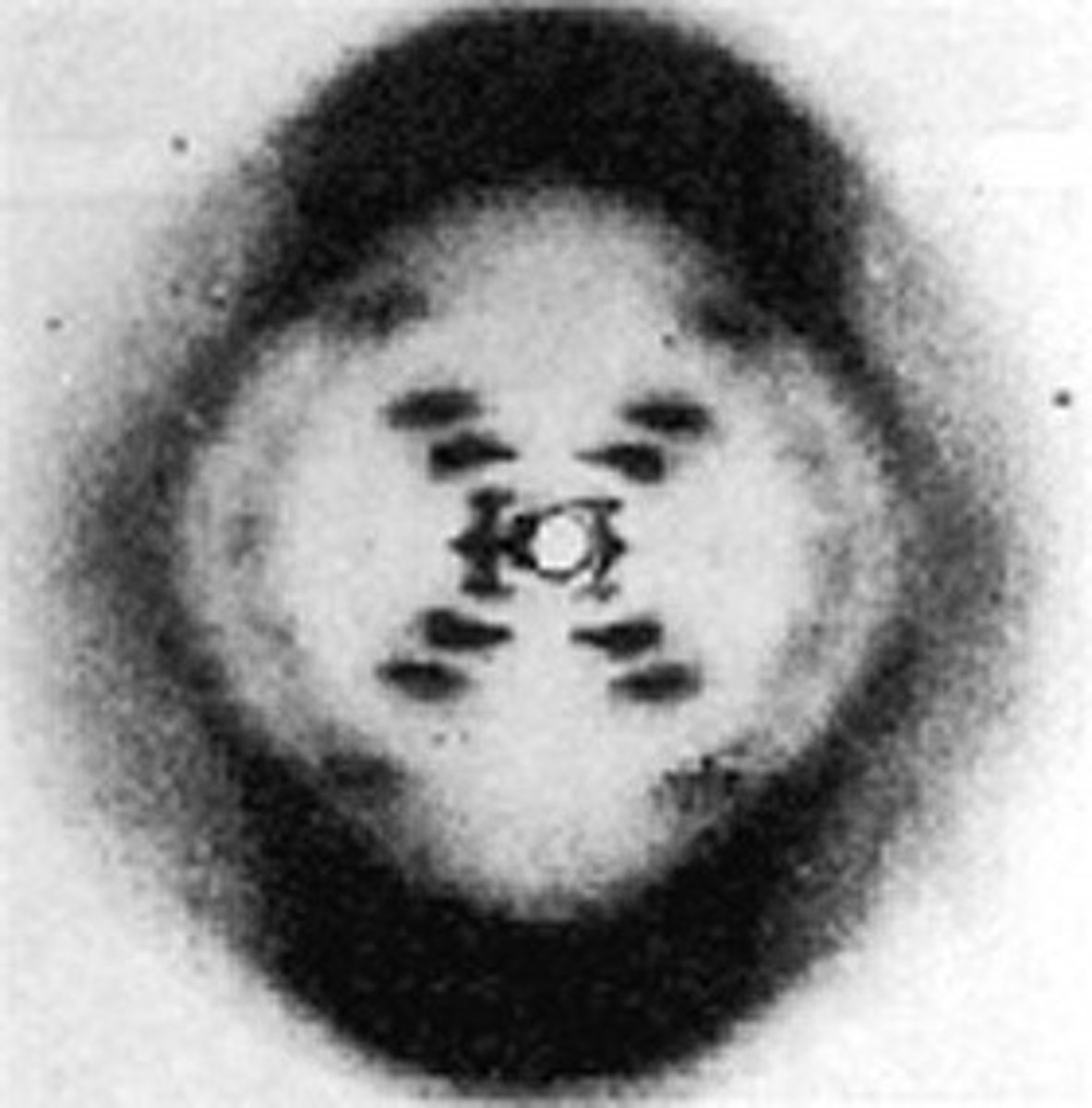
franklin and wilkins
- conducted the x-ray crystallography diffraction of DNA fibers
- diffraction was consistent with a helical structure, possible a double helix, with a sugar-phosphate backbone outside
ribose and deoxyribose pentose sugars
- numbering of the carbons in the pentose sugars: 1' to 5'
- ribose present in RNA
- deoxyribose lacks the 2'-OH group and is present in DNA
deoxyribonucleosides nomenclature
- deoxyadenosine
- deoxycytidine
- deoxyguanosine
- deoxythymidine
ribonucleosides nomenclature
- adenosine
- cytidine
- guanosine
- uridine
UMP nomenclature
uridine monophosphate
CDP nomenclature
cytidine diphosphate
dTDP nomenclature
deoxythymidine diphosphate
ATP nomenclature
adenosine triphosphate
cyclic nucleotide
- cAMP: cyclic adenosine monophosphate
structure of DNA
- double-stranded
- antiparallel
- opposing 5'->3' polarities of the two strands joined by hydrogen bonds
watson-crick base pairs
- two hydrogen bonds form between adenine and thinine in DNA
- adenine and uracil in RNA
watson-crick base pairs
- three hydrogen bonds form between guanine and cytosine
- high G-C content tends to have higher density and higher melting point
chargaffs rules about base-composition of DNA
- the proportion of adenines equals that of thymines, and the proportion of guanines equals that of cytosines
- there is equal proportion of purines(A&G) and pyrimidines(C&T)
- the proportion of C+G does not necessarily equal that of A+T
nucleic acid DNA summary
- backbone: phosphate and deoxyribose sugar
- nitrogenous purine bases: adenine and guanine
- nitrogenous pyrimidine bases: cytosine and thymine
nucleic acid RNA summary
- backbone: phosphate and ribose sugar
- nitrogenous purine bases: adenine and guanine
- nitrogenous pyrimidine bases: cytosine and uracil
intramolecular hydrogen bonds in transfer RNAs
- most RNAs are single-stranded
- intramolecular hydrogen bonds can form that make endless 3D structures
nucleosome
- the basic unit of organization of DNA in the eukaryotic nuclear chromosomes
- single nucleosome is nine proteins and DNA
- core of either histone molecules
organization or DNA in eukaryotic nucleus
- chromosomes are always present in the nucleus but only visible during cell division
- nucleosome
- two turns of DNA wrapped around a cluster of eight histone proteins(the nucleosome core particle) and held by a ninth histone(histone 1)
- adjacent nucleosomes pack together to form the 30-nm chromatin fibers
- higher order levels of organization result in the condensation of the entire chromosome into the visible chromosome, beginning at prophase of mitosis or meiosis
isolation and purification of DNA
- isolation of DNA follows procedures similar to Avery, MacLeod, and McCarty
- precipitation of DNA by the addition of ethanol
- followed by the redissolving of the precipitate is the last step in the purification of DNA
exonucleases
- enzymes that remove nucleotides one by one from the ends(exo) of a polynucleotide chain
endonucleases
- enzymes that cleave phosphodiester bonds within(endo) a polynucleotide chain
restriction endonuclease
- enzyme that recognizes a specific base pair sequence in DNA
- restriction site with a palindrome sequence and makes a cut at that site
restriction fragments
- smaller linear DNA fragments with either complementary single-strand cohesive tails(sticky ends) or blunt ends
nucleases
- exonucleases
- endonucleases
- restriction endonuclease with a restriction site
- restriction digestion of DNA results in restriction fragments
electrophesis
- method for separating fragments of nucleic acids according to their sizes
- the molecules migrate through a semisolid medium in an electric field toward the (+) end due to the negative charges in the phosphate groups
- smaller molecules migrate faster
preparation of an agarose gel for DNA electrophoresis
- tray with gel is placed in an electrophoresis chamber that is filled with buffer solution to cover gel
- samples of DNA are loaded into the wells
- electric field is applied: the cathode(-) is placed at the end of the gel, the anode(+) at the opposite end
ethidium bromide staining of separated nucleic acids
- the fluorescent dye becomes nested between the nitrogenous bases in nucleic acids and can be excited by ultraviolet light
DNA sequences
- presented by showing the sequence of nucleotides of only one of the two strands
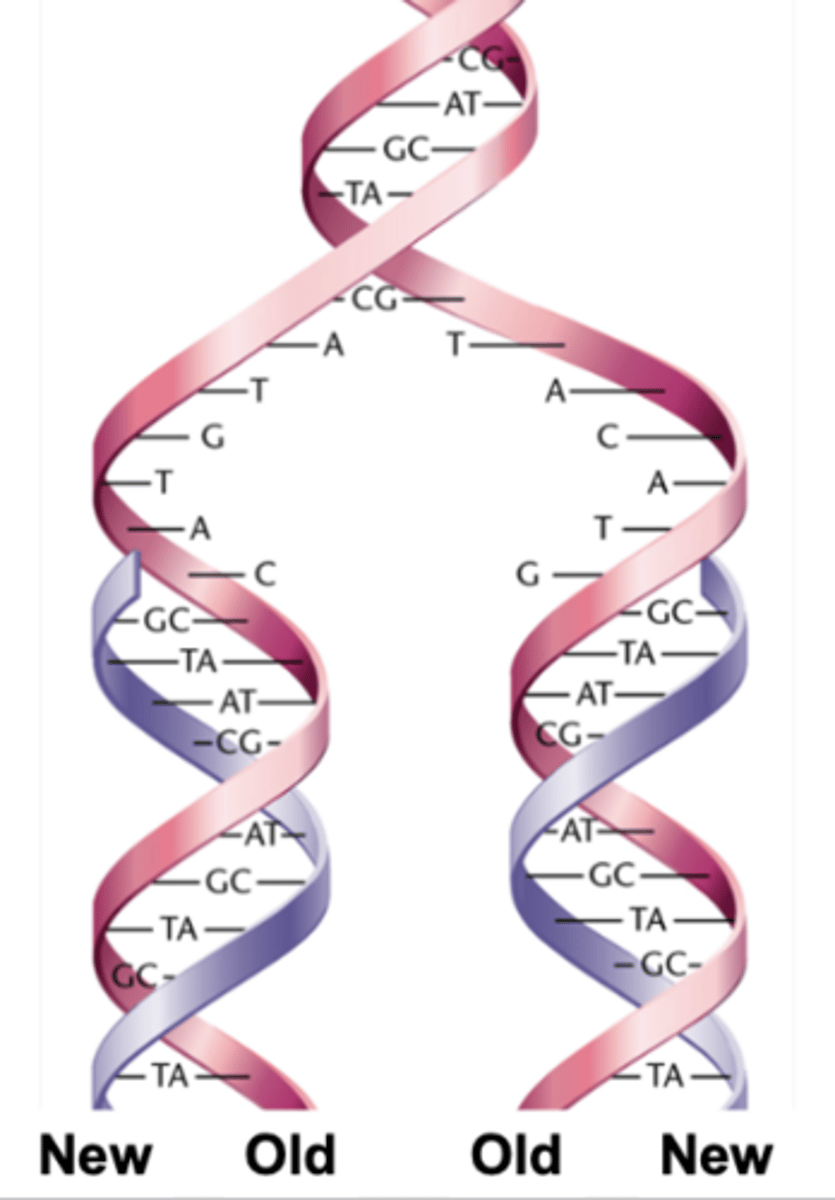
watson-crick model for semiconservative replication of DNA
- the parental(old) strands separate and serve as templates for the synthesis of complementary(new) strands
- the parental strands are retained
- two new DNA molecules each contains an old strand and a new strand
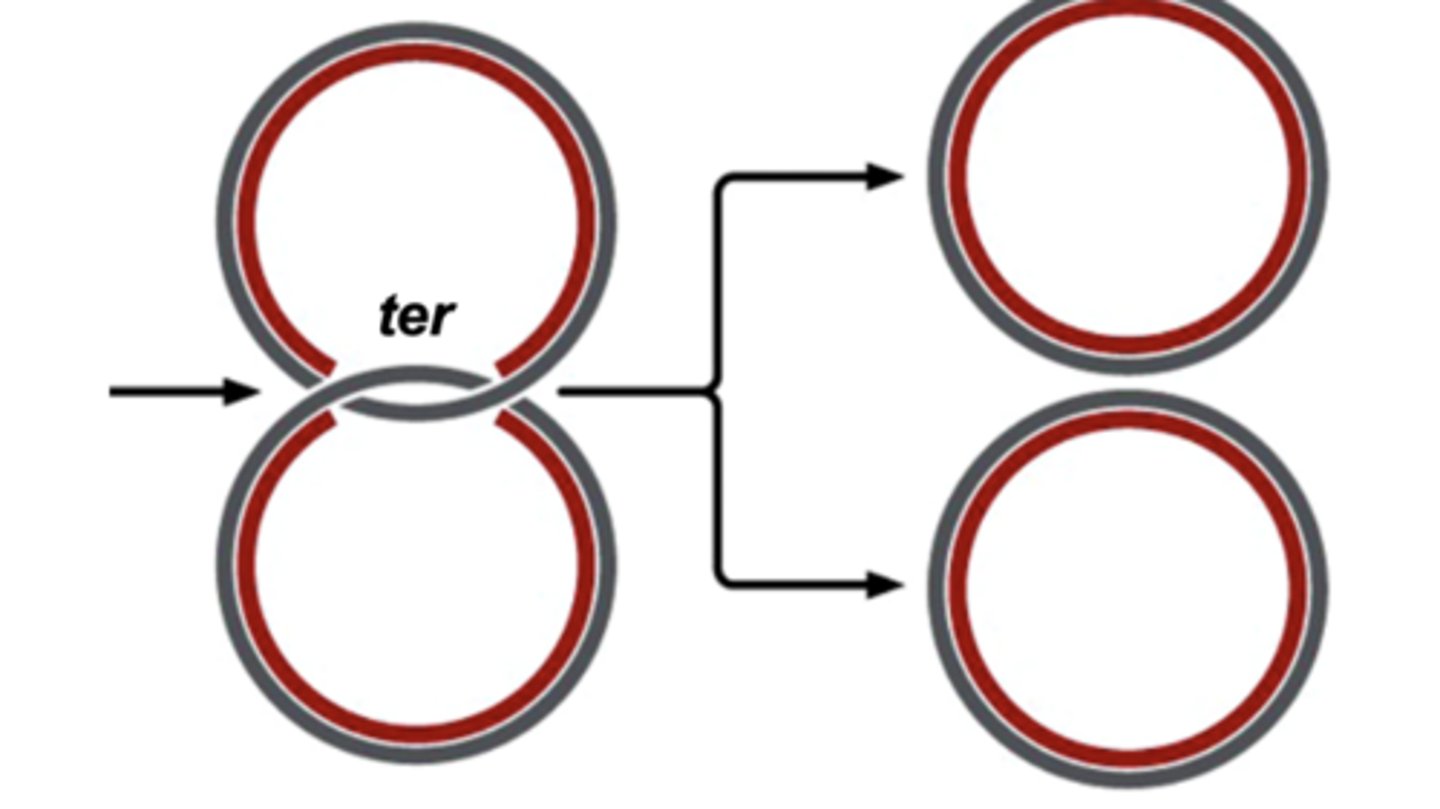
replication of circular chromosomes in bacteria
- origin of replication
- replication fork
- replication bubble
- it is recognized by an initiator protein that opens one replication bubble with two replication forks
- ter: replication termination sit
- self-replicating DNA molecules
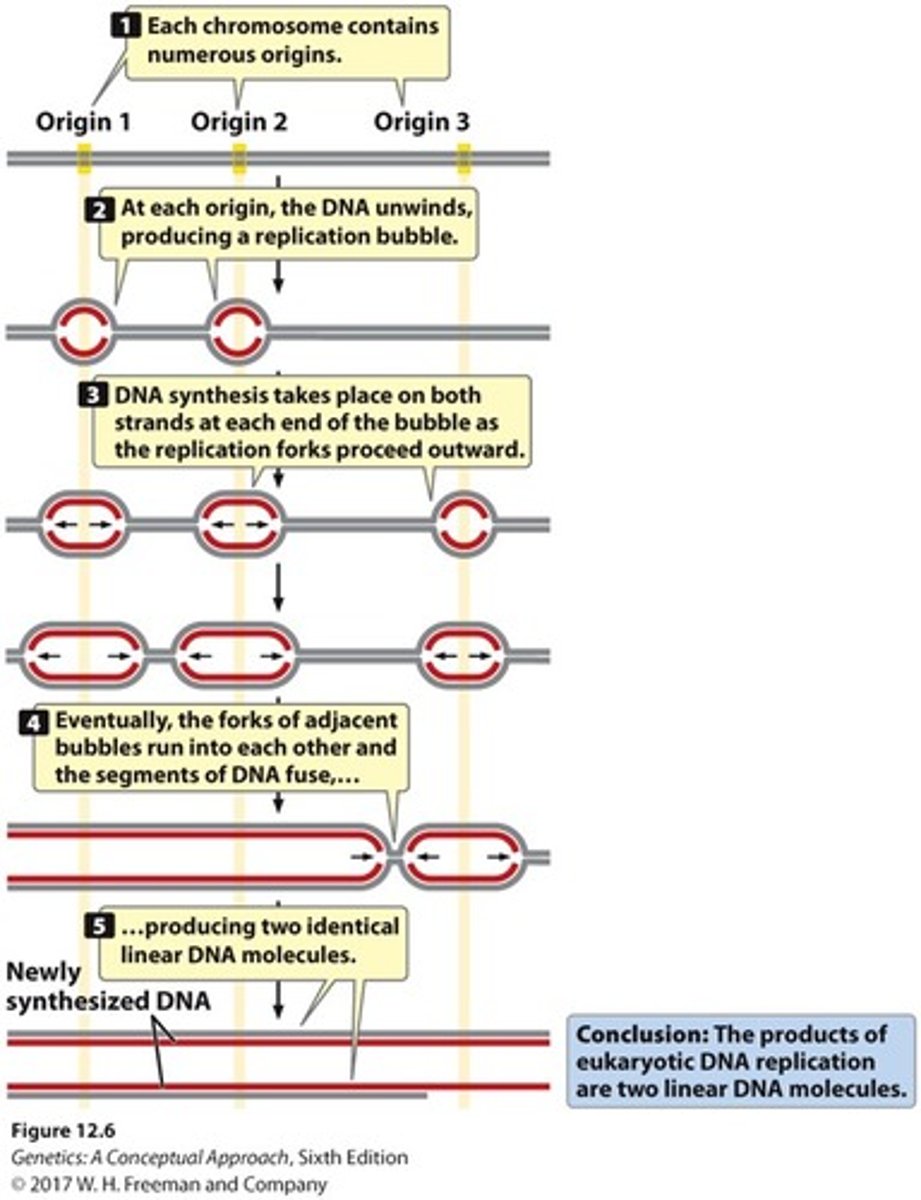
replication of linear DNA in eukaryotes
- contain multiple origins of replication where replication bubbles form
- each replication bubble has two replication forks(DNA unwinds)
- synthesis occurs on both strands until they meet and fuse
- produces two identical linear DNA molecules
DNA synthesis
- must be antiparallel
DNA synthesis is catalyzed by DNA polymerase
- enzyme adds nucleotides to the 3' end of the growing new strand
- this is the 5'->3' polymerase activity
formation of phosphodiester bond
- new DNA is synthesized from deoxyribonucleoside triphosphates(dNTPs)
- incoming nucleotides must be in the triphosphate form before they can be incorporated into DNA
- the hydrolysis of a phosphoanhydride bond between phosphates releases the energy necessary for the making of a new bond
proofreading activity
- error in DNA replication can lead to a permanent mutation
- the 3' to 5' exonuclease activity of DNA polymerases
- enzyme stops polymerizing, goes back, removes the incorrectly placed nucleotide, then resumes DNA polymerization
- even with this ability, DNA polymerases still leave occasional mistakes
DNA synthesis in a replication fork
- two DNA polymerases are needed: one for each strand
- new DNA strands are always synthesized antiparallel to template strands
lagging strand
- discontinuous DNA synthesis
- runs out of template
okazaki fragments
- in the lagging strand
- short fragments of DNA that are discontinuous
replication bubble in DNA replication
- the leading and lagging strands are defined for each fork, not for the entire bubble
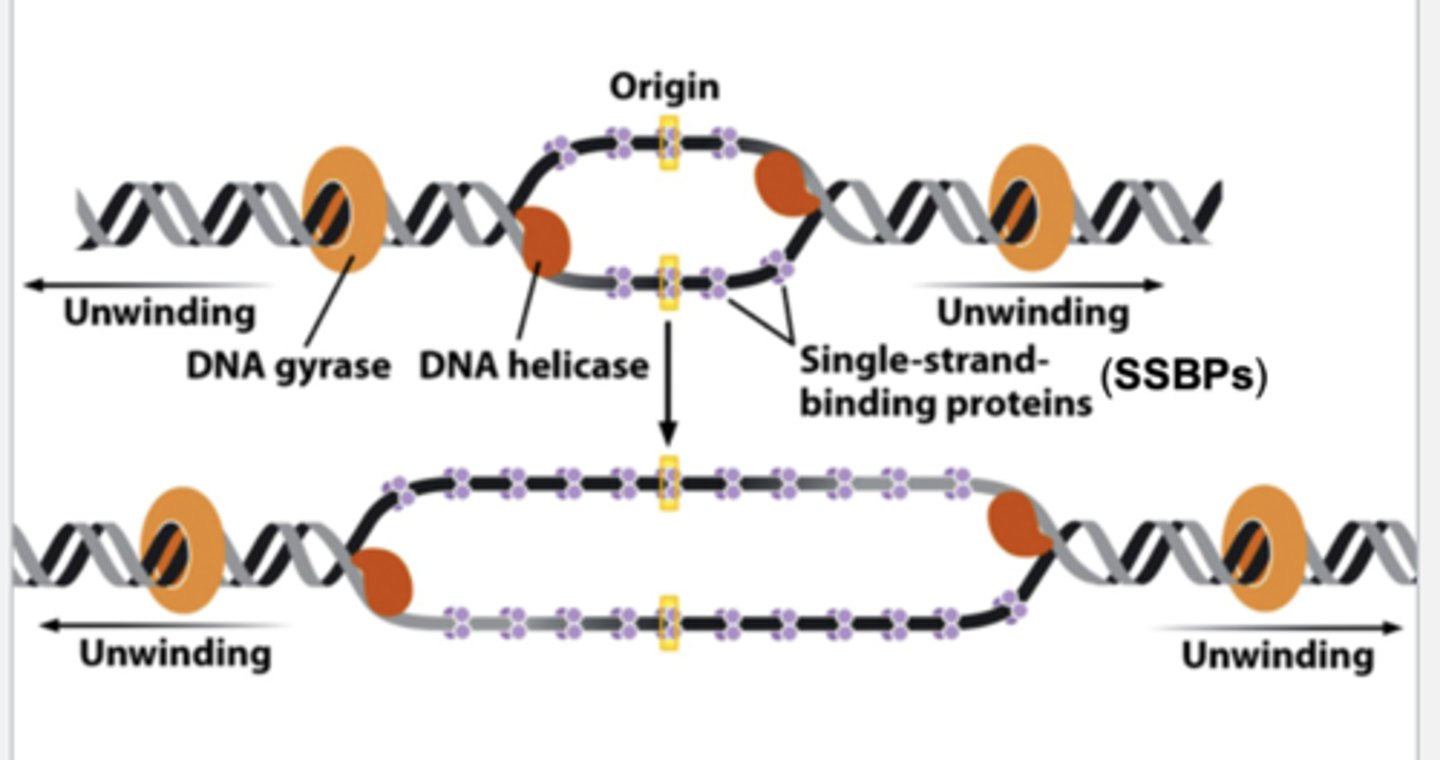
other participants in DNA replication
- DNA helicase: unwind the strands after they are separated
- single-stranded DNA binding proteins(SSBPs): bind to the separated DNA strands to stabilize the open conformation
- gyrase: unwinds supercoiled DNA ahead of the replication fork. in bacteria, makes single strand cut, allows supercoiled helix to unwinds, and reseals the cut
- sliding clamp: prevents the polymerase from "falling off"
- primer extension: the synthesis of the new strand of DNA
DNA polymerase III
- the main polymerizing enzyme in bacteria
- extends DNA using RNA primers
- can extend an existing strand of nucleic acid
- cannot initiate a brand-new strand "from scratch"
the initiation of DNA synthesis requires a primer
- DNA polymerase III
- primase: makes a small RNA primer to which DNA polymerase adds deoxyribonucleotides
- after RNA primer is used, it is removed with the 5' to 3' exonuclease activity of DNA polymerase I and replaced with DNA by the same enzyme
- DNA ligase: seals the resulting nick
- the fragments of the new DNA strands can be extended to replace the RNA primers
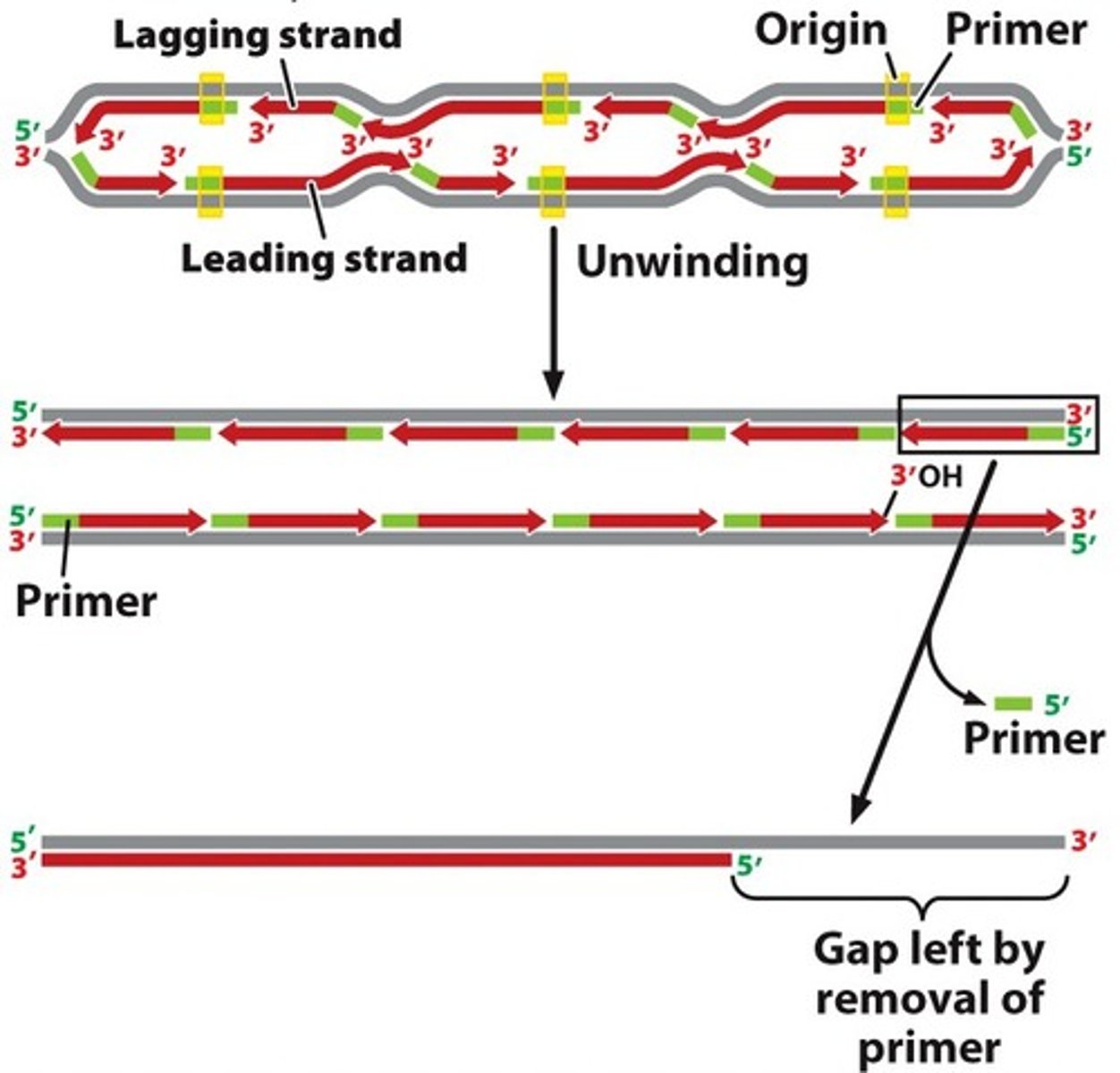
replication of linear DNA molecules of eukaryotes
- a gap left by removal of primer
- gap cannot be filled because there is no 3' template to accommodate a primer
- results in the shortening of the chromosome with every round of DNA replication and cell division
telomerase activity
- the enzyme telomerase is a ribonucleoprotein: made of protein and a short piece of RNA that serves as both guide and template for synthesis of complementary DNA(cDNA)
- telomere has a protruding end with a G-rich repeated sequence where RNA template attaches to telomerase that add nucleotides then is removed
telomerase is present in:
- single cell eukaryotes
- germ cells
-stem cells in both animals and plants
- activated lymphocytes
- cancer cells
- absent in most normal somatic cells of multicellular animals
- in the absence of telomerase activity, chromosomes are shortened with each round of DNA replication and cell division
the polymerase chain reaction(PCR)
- method for rapid cloning of DNA
- results in DNA amplification: the synthesis of many copies of a desired sequence
- the reaction mixture: template DNA, dNTPs mixture, specific pair of primers, Taq DNA polymerase
- typical cycle: denatures DNA, anneals primers, extends primers
PCR
- thermal cyclers automatically change the temperatures for the set times
- primers are designed to recognize a desired target sequence
DNA polymerase from Taq is very heat-resistant because the bacterium lives in hot springs
RNA transcription in all cells
- messenger RNA(mRNA)=>proteins
- ribosomal RNA(rRNA)
- transfer RNA(tRNA)
RNA transcription in eukaryotes
- pre-messenger RNA(pre-mRNA)
- small nuclear RNA(snRNA)
- small nucleolar RNA(snoRNA)
- microRNA(miRNA)
- small interfering RNA(siRNA)
- piwi-interacting RNA(piRNA)
RNA transcription in prokaryotes
- CRISPR RNA(crRNA)
- precursor
transcription unit
- a segment of DNA that encodes an RNA molecule
- contains a promoter, the RNA-coding region(the gene), and a terminator as part of the gene