DNA Technology
1/43
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
44 Terms
5 key concepts
Synthesis of recombinant DNA
PCR to amplify specific sequences of DNA
Recombinant DNA can be used to form protein fusions
DNA sequences can be specifically manipulated
DNA sequencing technology has increased in speed and fidelity
Recombinant DNA
Restriction enzymes
Plasmid Vectors
PCR based technologies
Oligonucleotide primers
Recombinant fusion proteins
Epitopes
Affinity tags
GFP
Manipulating DNA sequence
Quick Change
CRISPR/CAS9
Measuring gene expression
Microarray
RT-PCR
DNA sequencing technology
Sanger
Sequencing by synthesis
identifying and purifying dna
Chargoff determines that the ratio of A:T and G:C
(Purines:Pyrimidines) is always 1
Characterizing Properties of DNA Sequences
Remember: It takes more energy to separate C – G pair
Melting Temperature (tm) is____________
increases as __________________ in DNA sequences
the specific temperature at which HALF of the DNA strands separate
INCREASES as the number of G–C pairs increases in DNA sequences.
Recombinant DNA
Study the genome in fragments: how?
DNA formed artificially by combining constituents from different organisms.
Digest genome with restriction enzymes into random pieces and ligate them into prokaryotic plasmids.
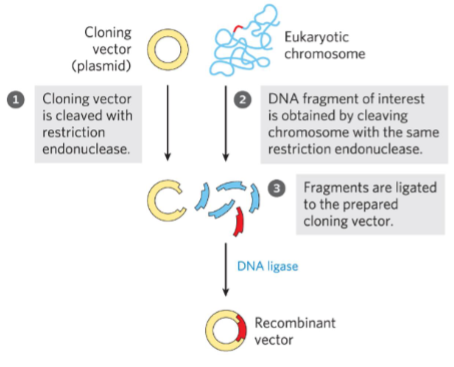
DNA is cleaved at specific sequences by
restriction endonucleases
Restriction endonucleases (restriction enzymes) often bind and cut the phosphodiester backbone at ___________________
palindromic sequences.
Some enzymes leave overhangs (____________) example?
sticky ends, HindIII digest
Other enzymes cut at the same site on both strands (________) example?
blunt ends e.g. EcoRV digest
Plasmids are
circular pieces of DNA that only contain a few genes.
Selecting for recombinant plasmids
Plasmids include a selection marker that commonly confers ___________________________
Plasmids are transformed into __________.
Only cells that take up the plasmid_____________________.
antibiotic resistance
bacteria
express resistance to ampicillin
PCR is? steps? MAE
1. Melt: Heat up to separate strands
2. Anneal: Short oligonucleotide primers bind
3. Elongation: DNA polymerase synthesizes new DNA strand.
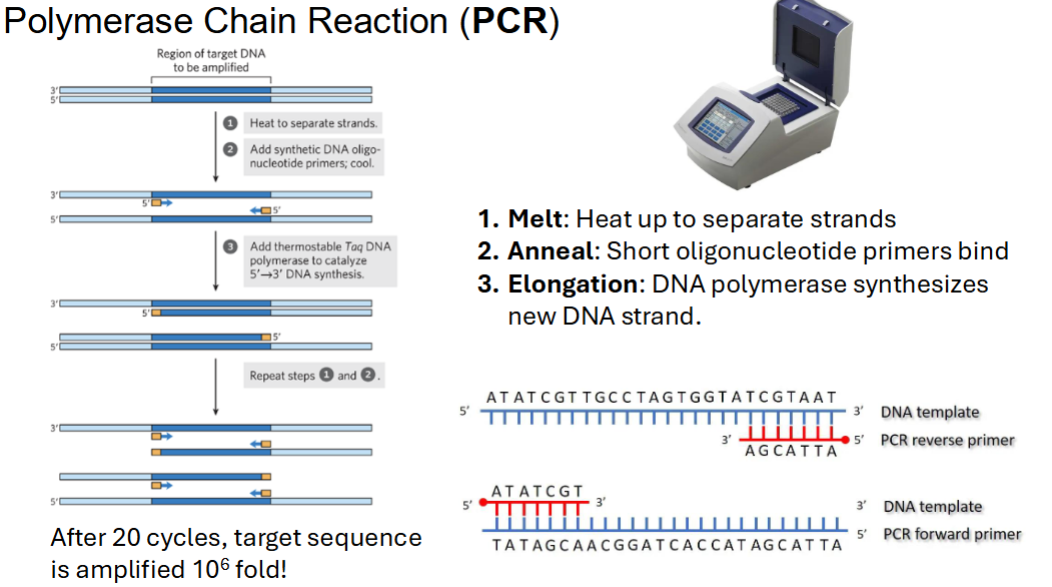
Design primers engineered with a restriction enzyme cut site so THAT
your PCR fragment can be cut and ligated into a plasmid.
Agarose gel electrophoresis
________ _______ is a common stain for DNA
what does it do?
DNA is normally __________ charged
Ethidium bromide
It intercalates between bases, causing it to fluoresce under UV excitation.
Remember: DNA is normally negatively charged
1. BamH1 cleaves the following sequence. Does this result in a blunt end or sticky end?
2. Bcl1 cleaves the following sequence. Write the sequence of the overhang after digestion.
3. How many fragments will result from the cleavage of the following sequence with BamHI and Bcl1?

Cloning the coding sequence of genes
DNA contains all introns and exons:
How can you create recombinant DNA with just exons?
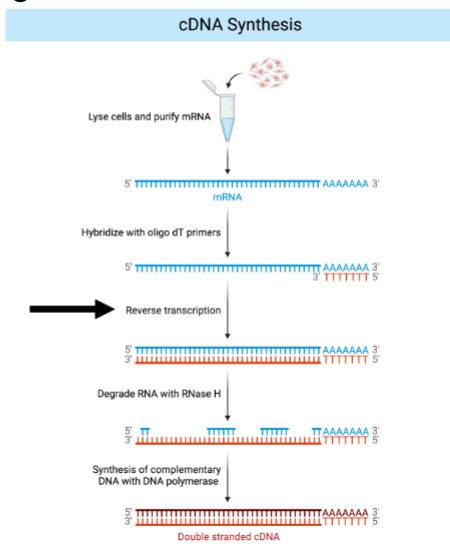
Complementary DNA (cDNA) is generated from mature mRNA.
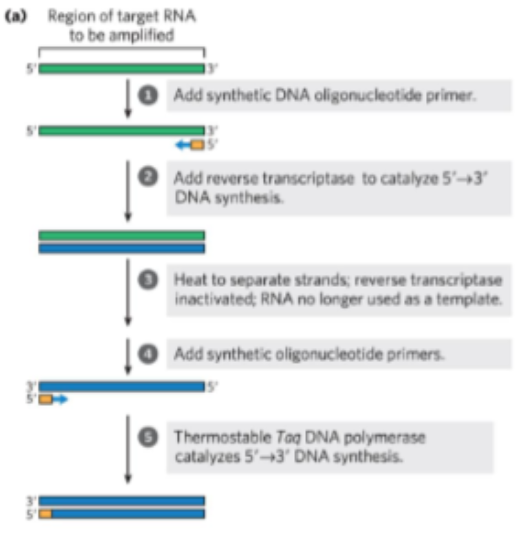
steps for cDNA synthesis
Remember: Reverse transcriptase is an RNA-dependent DNA polymerase.
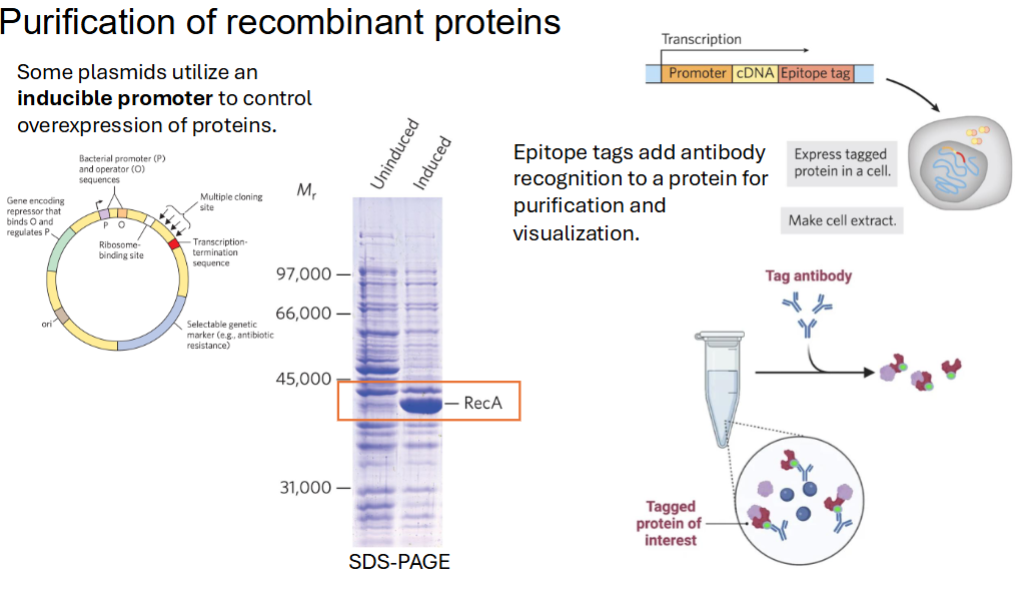
Purification of recombinant proteins
Some plasmids utilize an __________________ to control overexpression of proteins.
Epitope tags add ________________ to a protein for ___________ and _____________
Some plasmids utilize an inducible promoter to control overexpression of proteins.
Epitope tags add antibody recognition to a protein for purification and visualization.
Recombinant DNA can create ________________ for what?
fusion proteins for affinity chromatography
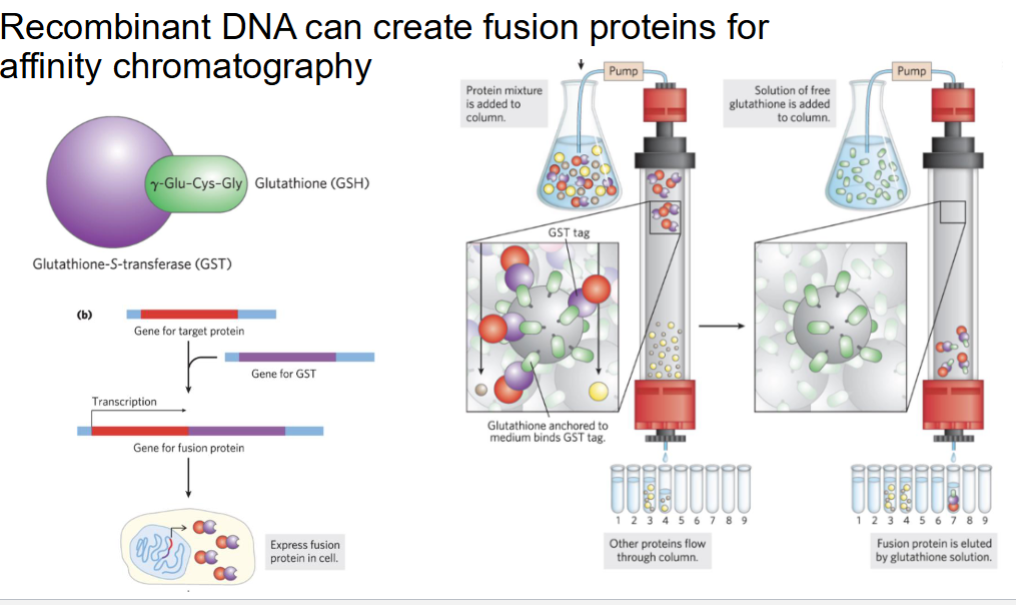
Recombinant DNA can create _______________ that aid in what?
fusion proteins that aid in visualizing cellular location
Green Fluorescent Protein (GFP) ____________________-
emits light upon excitation.
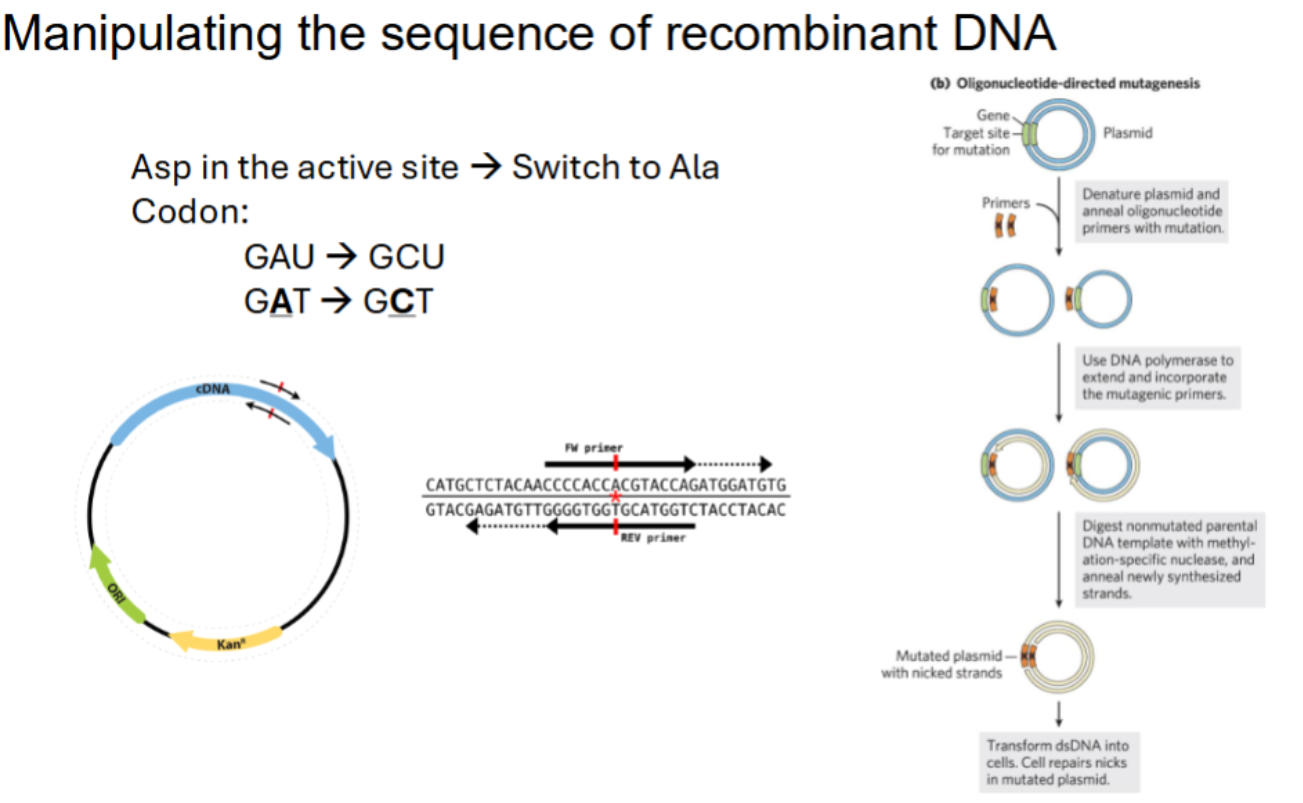
Manipulating the sequence of recombinant DNA
what is oligonucleotide directed mutagenesis? end goal?
Asp in the active site → Switch to Ala
Codon:
GAU → GCU
GAT → GCT
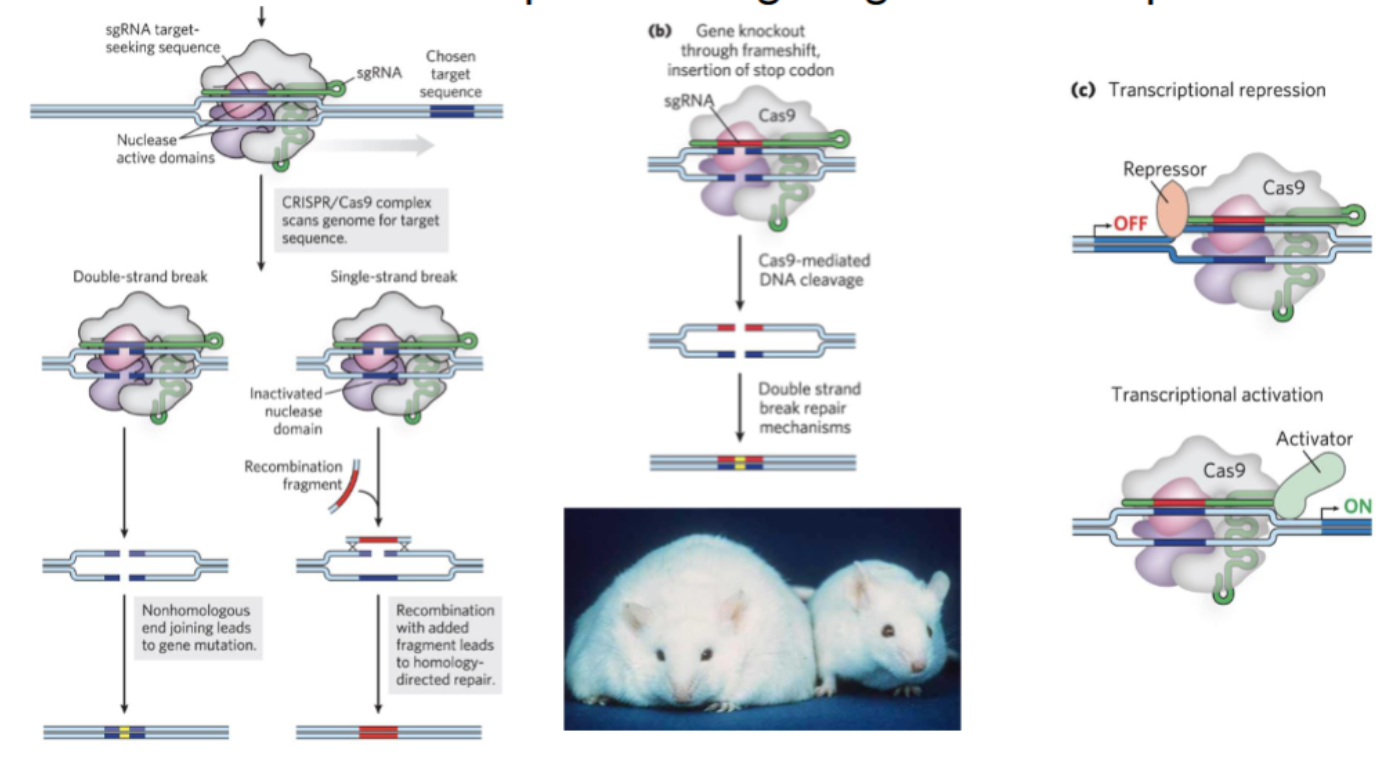
CRISPR/Cas9 enables what
name three purposes of Cas9 complex
precise targeting of DNA sequences
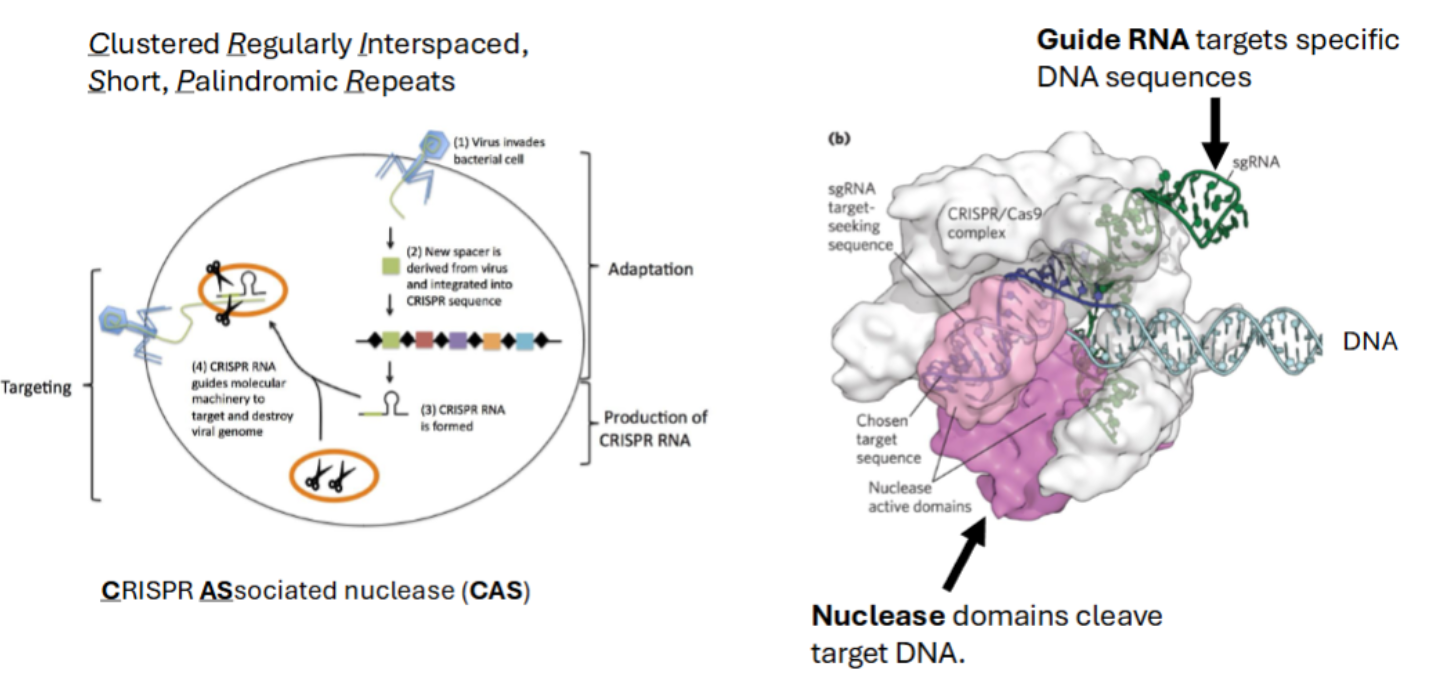
Clustered Regularly Interspaced, Short, Palindromic Repeats
scans genome for target sequence
nonhomologous end joining = gene mutation
recombination with fragment = homology-directed repair
gene knockout through frameshift → insert stop codon
double strand break repair mechanisms
transcriptional repression/activation
CRISPR ASsociated nuclease (CAS) do two things
Guide RNA targets specific DNA sequences
Nuclease domains cleave target DNA.
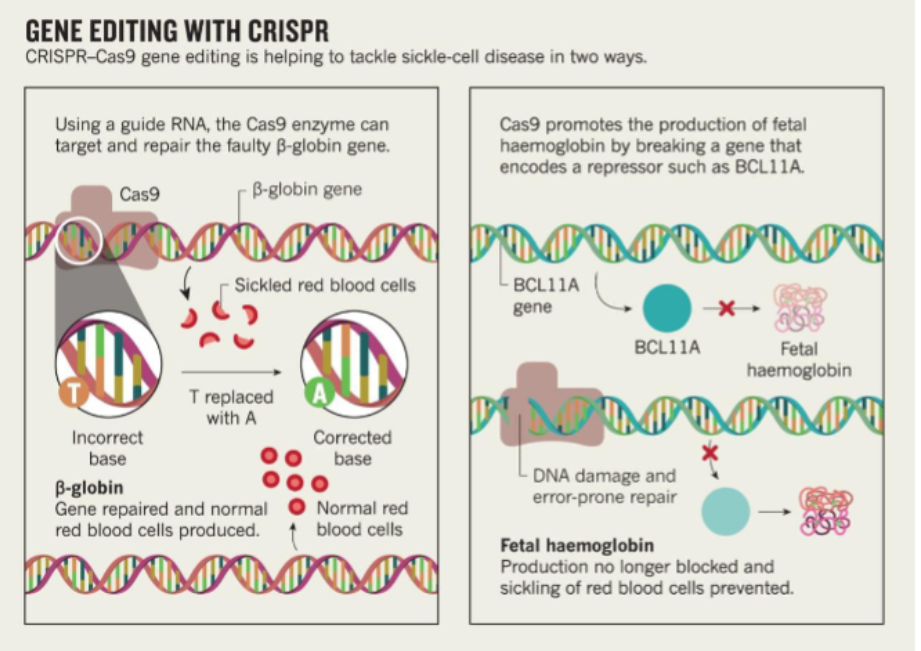
Gene editing with CRISPR (3 steps)
Isolate patient’s Hematopoietic Stem and Progenitor Cells (HSPC)
Edit cells in culture with CRISPR
Re-establish stem cell population in bone marrow
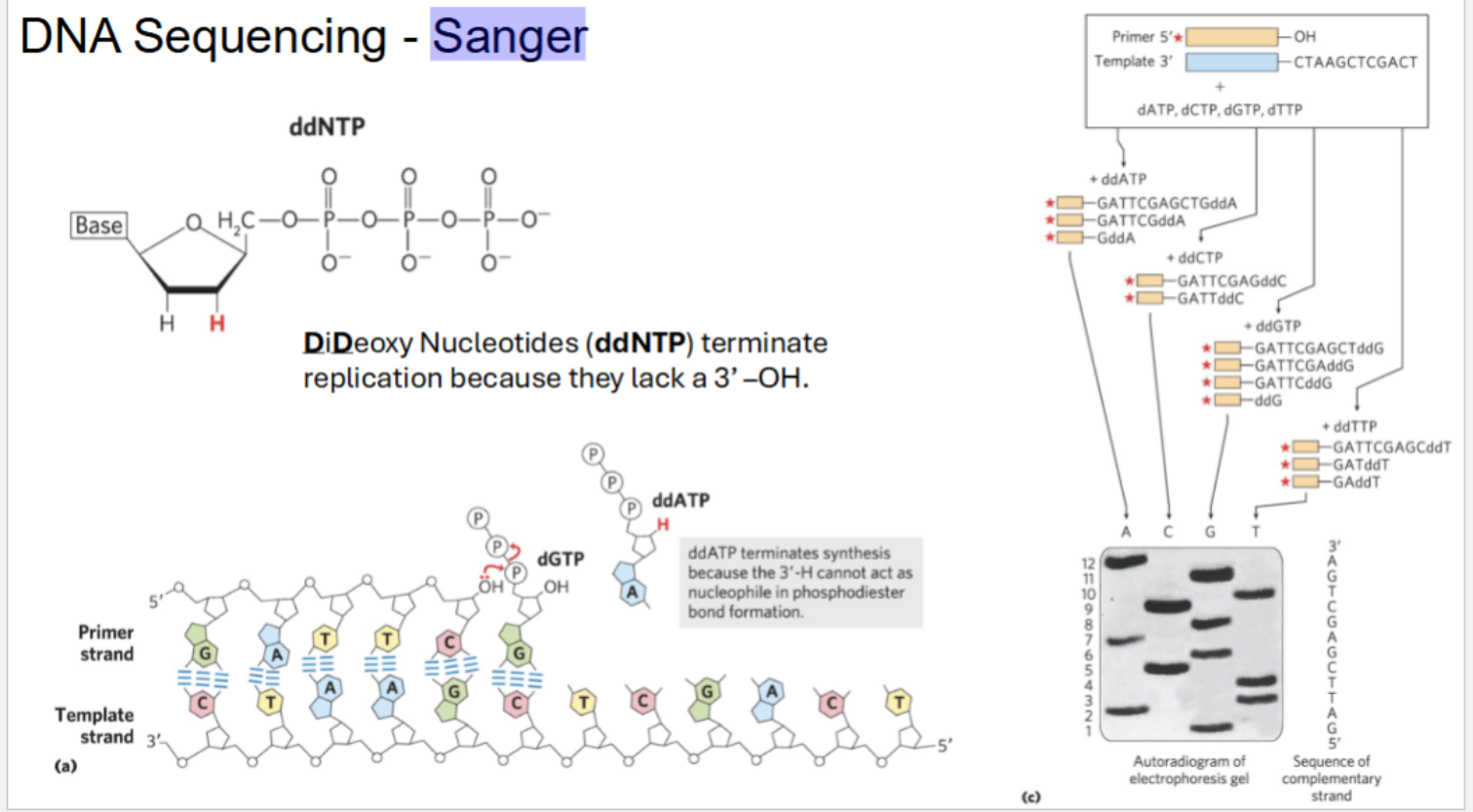
Sanger Sequencing
what is ddNTP? why does it terminate replication?
DiDeoxy Nucleotides (ddNTP) terminate replication because they lack a 3’ –OH (3’ H CAN’T act as a nucleophile for phosphodiester bond formation)
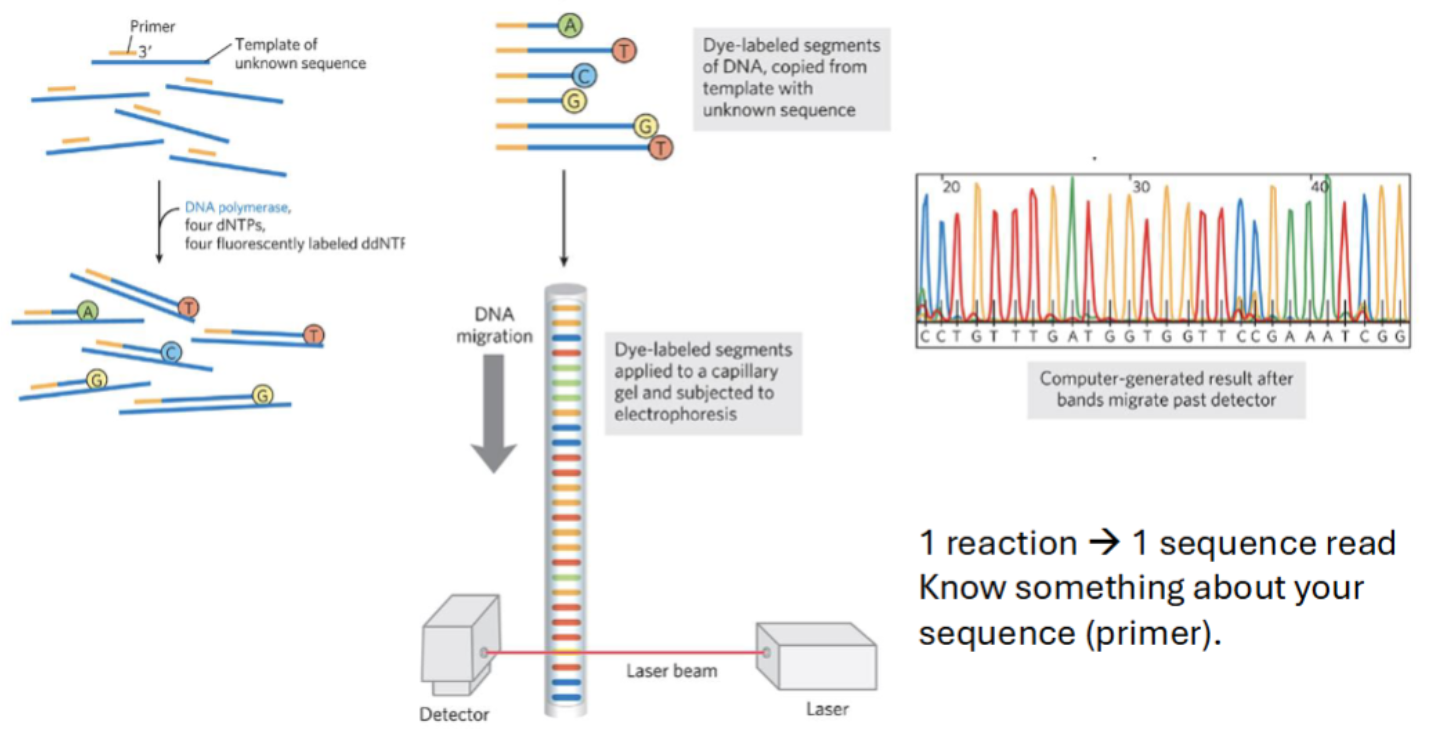
1 reaction → 1 sequence read
know what about your sequence?
Know something about your sequence (primer).
Next Generation Sequencing – Sequencing by Synthesis
A picture of the entire flow cell is taken after each fluorescent base is added.
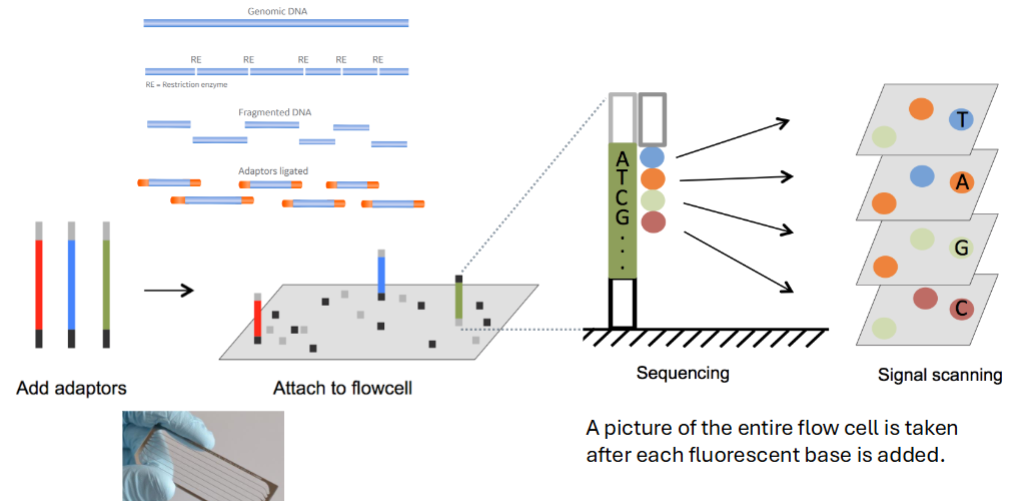
Each spot is a _____________ and the color changes each image as new bases are added to the __________________
Overlapping reads are built into a ____________________________ and mapped onto a reference genome.
DNA fragment
complementing strand.
Overlapping reads are built into a contiguous sequence (contigs)
Measuring changes in gene expression
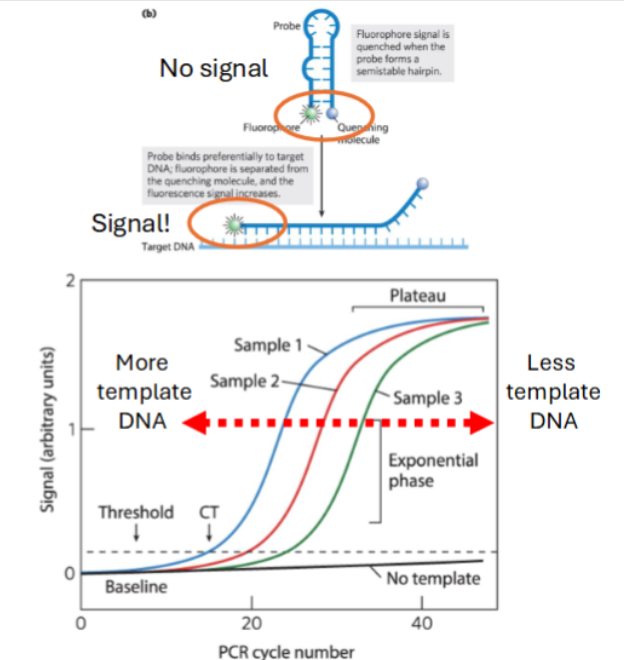
Quantitative Real-Time – Polymerase Chain Reaction (qRT-PCR) does what?
utilizes fluorescent probes to monitor amplification.
steps of qRT-PCR (5)
Isolate mRNA and convert to DNA
Amount of DNA produced should correlate with mRNA levels.
Lower Cycle Threshold (CT) value
indicates higher mRNA levels (higher expression)
Measuring changes in gene expression
Microarray analysis allows you to ________________________________
compare expression across an entire transcriptome.
Note: Comparing two conditions of a cell. (well fed vs starved, control vs stressor, etc.)
Current approaches utilize NGS to get ____________________ analysis
highly sensitive transcriptome analysis – even the transcriptome of a single cell!
Polymorphisms:
Small variations in gene sequence that lead to changes in phenotype and variation in populations.
Genome analysis identifies ____________________?
Haplotypes are ______________________
Single Nucleotide Polymorphisms (SNPs)
groupings of multiple SNPs
Tag SNPs are _______ and __________
Sequencing technology helps _____________ and more accurately _______________
unique and define the haplotype.
identify haplotypes and more accurately draw phylogenetic trees.
DNA sequencing will allow the ________________________ and ____________________
identification of unique populations within cancerous cells and prescribe more effective treatments.