Protein Targeting
1/32
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
33 Terms
What synthesizes proteins and where does it take place
Ribosomes and in the cytosol
Where can proteins be targeted to
They can exit the cell
Integrate into membranes
Enter subcellular compartments
What are signal sequences and what do they do
Short amino acid motifs that serve as addresses guiding proteins to specific locations
Why does protein targeting occur exclusively in eukaryotic cells
It allows for subcellular organization which aids in compartmentalizing metabolic pathways
What signal sequence targets ribosomes synthesizing proteins and directs them into the ER
N-terminal signal sequences
Features of N-terminal signal sequences
13-20 amino acids in length
Contains about 10-15 hydrophobic amino acids
Includes one or more positively charged amino acids near the N-term
Has a polar short sequence near the C-term
Contains a cleavage site
Signal Recognition Particle (SRP)
A conserved ribonucleoprotein complex that directs specific proteins into the ER by binding to signal sequences of nascent polypeptides
Nascent polypeptide
A newly synthesized amino acid chain that emerged from the ribosome
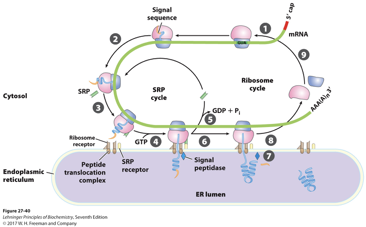
How does the SRP pathway work
The SRP recognizes and binds to a hydrophobic N-terminal signal sequence from a nascent polypeptide
Upon binding, the SRP’s Alu domain interacts with the ribosomal GTPase center, pausing translation, preventing protein misfolding or aggregation in the cytosol
The SRP- Ribosome-Nascent Chain (RNC) complex moves to the ER membrane, where the SRP binds to the SRP receptor SR (GTPases)
The RNC is transferred from the SRP/SR complex to the Sec61 translocon
The SRP and SR dissociate following GTP hydrolysis. Translation resumes and the growing polypeptide is threaded directly into the ER lumen or integrated into the ER membrane
What is the overall goal of the SRP pathway
To translocate proteins to the ER co-translationally
Where do proteins undergo post-translational modifications and list an example
In the ER and Glycosylation
What happens to proteins after undergoing post-translational modifications
They get packaged into transport vesicles and are taken to the Golgi apparatus to get sorted
How does the Biosynthetic / Secretory pathway work
In the ER: Ribosomes on the rough ER synthesize proteins targeted for secretion or membrane insertion
ER Exit Sites (ERES) & ERGIC: Proteins destined for transport are captured into COPII-coated vesicles. These vesicles leave the ER at specialized sites (ERES) and travel to the ER-to-Golgi intermediate compartment (ERGIC)
Golgi Complex: Vesicles from the ER fuse to form the cis-Golgi , where proteins are further processed, sorted, and packages, and move through the Golgi cisternae toward the Golgi network (TGN)
Post-Golgi Transport (TGN): The TGN acts as a sorting hub, packaing proteins into specific vesicles directed to their final destinations: lysosomes, the plasma membrane, or extracellular space
What is the main goal of the biosynthetic secretory pathway
To synthesize, modify, and transport proteins / lipids to the plasma membrane, lysosomes, or extracellular space by moving cargo through the ER and Golgi in an anterograde manner
Glycoprotein
A linkage between an oligosaccharide to a side group on a peptide
N-linked glycosylation
Occurs on the nitrogen atom of Asparagine (Asn) or Arginine (Arg) side chains
O-linked glycosylation
Occurs on the oxygen atom of Serine (Ser), Threonine (Thr), and Tyrosine (Tyr)
Core oligosaccharides are built on dolichol phosphate by the successive addition of what sugar units
monosaccharides
Where do oligosaccharides get transferred to and within what
Asn residues in the ER lumen
Tunicamycin function
To block step 1, the initial step of N-linked glycosylation
Where can O-linked glycosylation take place
In the Golgi or cytosol
Golgi Apparatus function
Responsible for modifying, sorting, and packaging proteins and lipids from the Endoplasmic Reticulum (ER) into vesicles for delivery to specific destinations
Why will a protein get directed to the plasma membrane
For incorporation as membrane proteins or secretion
Lysosome function
They break down waste materials, foreign invaders (bacteria/viruses), old cell parts (autophagy), and facilitate programmed cell death (apoptosis) and repair the plasma membrane.
Where do proteins destined for the mitochondria and chloroplasts bind chaperone proteins
In the cytosol, they are then delivered to receptors on the exterior of the organelle
Chaperone protein function
To maintain proteins in an unfolded state suitable for import
What do proteins targeted to the nucleus carry
A nuclear localization sequence (NLS)
Characteristics of a nuclear localization sequence (NLS)
4-8 amino acids long
Contains several consecutive basic residues (Arg, Lys)
Not removed after import
Can be located anywhere within the protein sequence
Targeting to the nucleus mechanism step 1
A protein with an appropriate NLS is bound by a complex of importin α and β
Targeting to the nucleus mechanism step 2
The importin (α and β)-protein complex binds to a nuclear pore and translocates into the nucleus
Targeting to the nucleus mechanism step 3
Inside the nucleus, dissociation of importin β is promoted by the binding of Ran-GTP
Targeting to the nucleus mechanism step 4
(nucleus)- Importin α binds to Ran-GTP and CAS (cellular apoptosis susceptibility protein), releasing the nuclear protein
Targeting to the nucleus mechanism step 5
(nucleus)- Importin α and β and CAS are transported out of the nucleus and recycled. Ran is transported back into the nucleus