PCR Gel sequencing copy
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
Explain how to find the mutation caused by a gene
1. isolate and copy gene by adding specific DNA prime rthat binds (through complimentary base pairing) to either side of gene of interest through PCR
2. analyze amplified gene on a gel to determine length and expression level
3. sequence the insertion
4. compare sequence to known sequences to determine whats known about that region of the genome
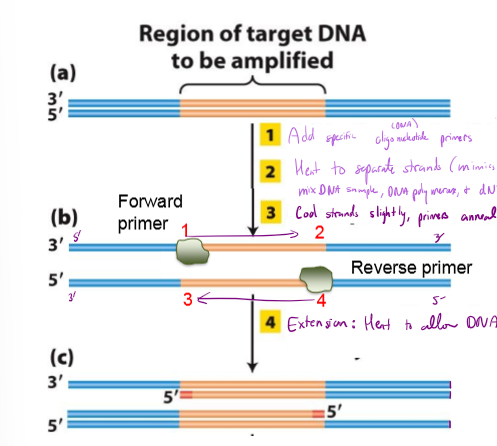
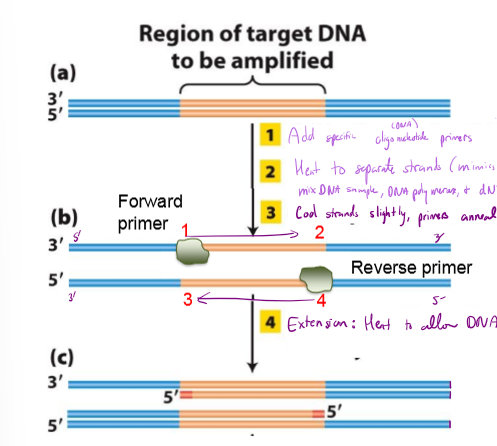
What is PCR? Explain the process
mimics DNA replication to create 10^6 copies of gene of interest
1. Heat DNA to separate strands (Mimics ORC)
2. mix DNA sample, DNA polymerase, synthetic DNA primers, dNTPs (nucleotides), and forward/reverse primers
3. cool strands slightly, allowing forward and reverse primers to anneal (bind to template)
4. heat to allow DNA synthesis
repeat 25 cycles
Why do you amplify DNA?
can analyze on a gel
sequence
clone
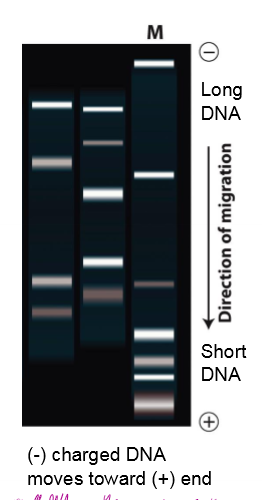
How does gel electrophoresis work? What infromation do you attain?
negatively charged WT and mutant DNA placed on negative end of porous gel
negatively charged DNA moves towards positive end of gel
small DNA moves further through gel (its holey) and long DNA doesn’t move as far
shows length of DNA; shows if theres an insertion/deletion causes a mutation
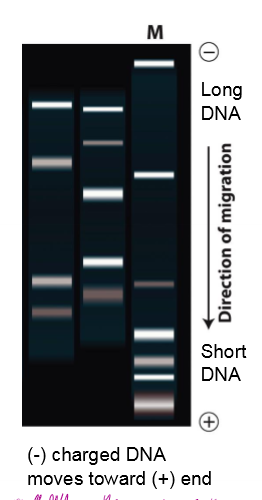
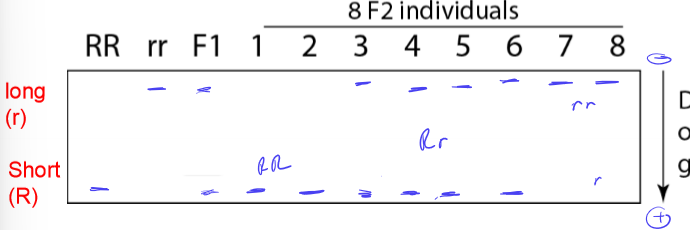
DNA Gel electrophoresis shows that the mutant doesn’t move as far as the wildtype. What does this mean about the mutant?
insertion in mutant relative to WT
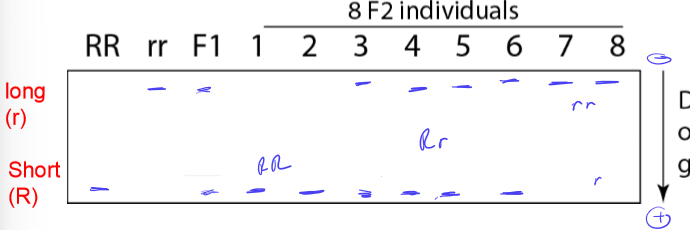
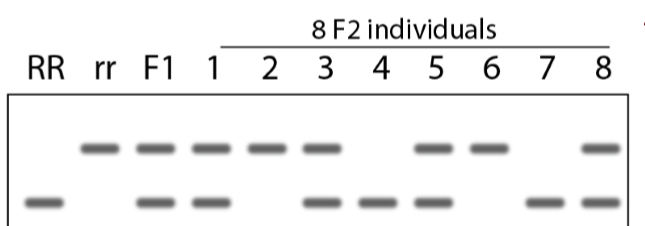
How do homozygous and heterozygous alleles appear on DNA gel electrophoresis?
homozygous: 1 band
heterozygous: 2 bands
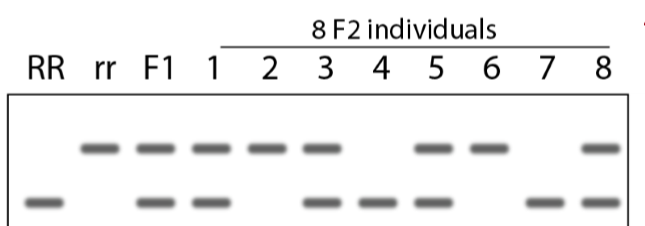
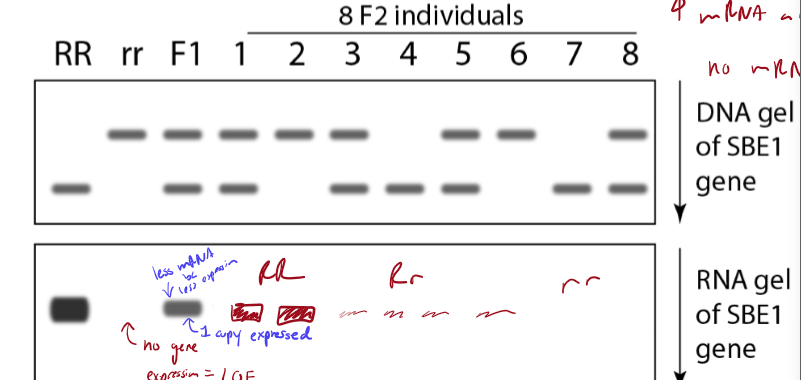
What do DNA gel and RNA gel detect and determine/compare?
DNA gel - detects DNA fragment size; helps determine genotype
RNA gel - detects mRNA abundance (expression level); tool fo rcomparing gene expressoin
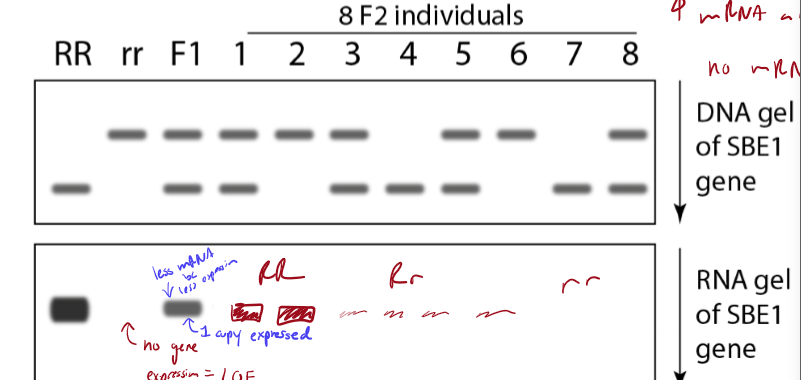
What do a dark adn light band represent after RNA gel electrophoresis?
dark: high expression
light: less expression
RNA gel electrophoresis is performed. What do a dark, light, and no band in a mutant relative to the WT indicate?
dark band: high expression - GOF
light band : less expression - LOF (recessive)
no band: low expression - LOF
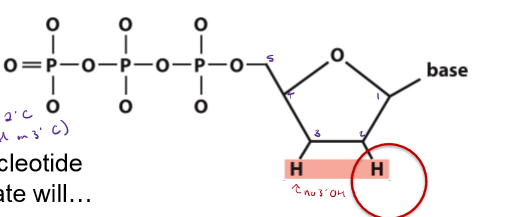
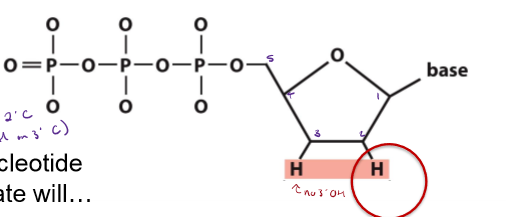
What is different about a dideoxynucleotide triphosphate (ddNTP) used in Sanger sequencing from a normal dNTP? Why is ddNTP used in Sanger sequencing?
does not have an OH on the 3’ C
will not allow strand elongation once incorporated into a growing DNA stand
not base-pair with the complementary strand
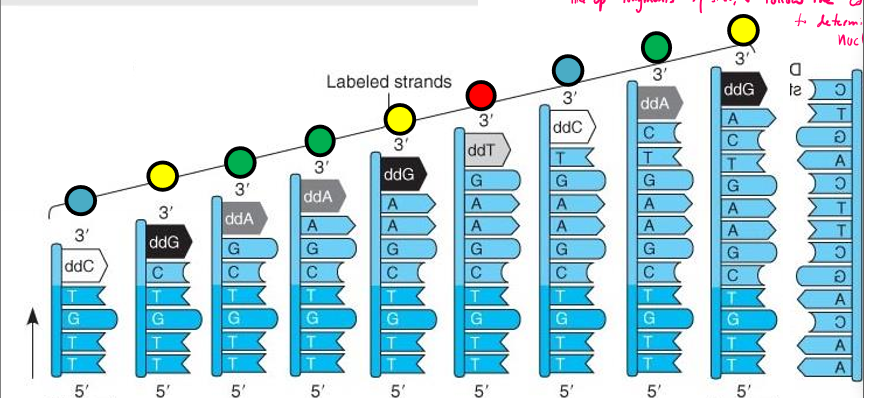
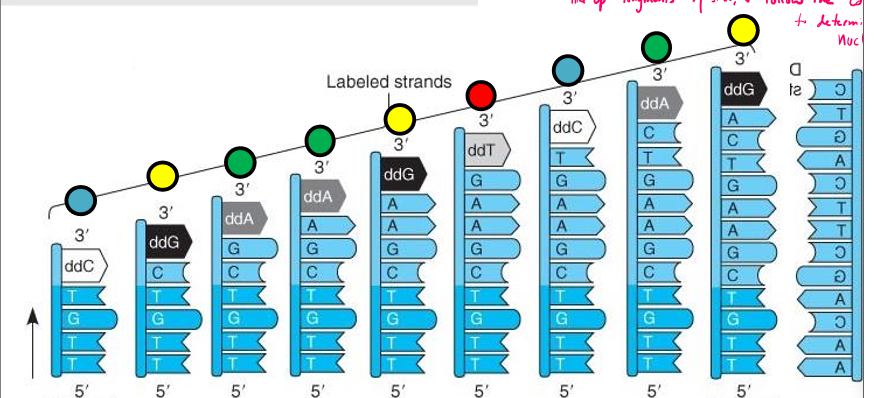
Describe Sanger sequencing
Start with lots of PCR as template strand
mix in lots of DNA template strands, synthetic primer, DNA polymerase, flourescently labeled ddNTPS, and more dNTPs
thermocycle to denature, anneal, extend and repeat
dNTPs/ddNTPS randomly incorporated by DNA poly
results in strands of varying length with flourescently labeled ddNTPs corresponding to a specific nucleotide at the end
line up fragments by size using gel electrophoresis, and follow the color to pattern to determine of nucleotides
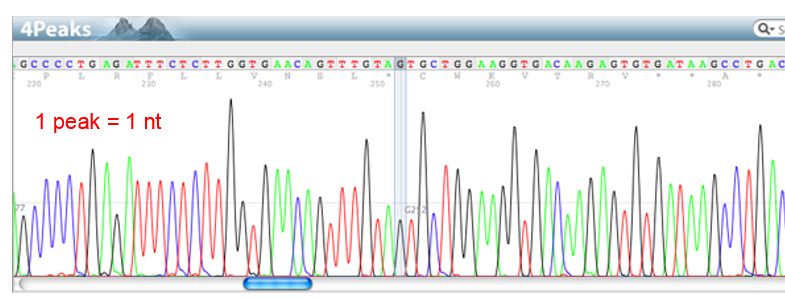
What is a chromatogram?
visual representation of the flourescence detected by the DNA sequencer
each peak corresponds to 1 nucleotide
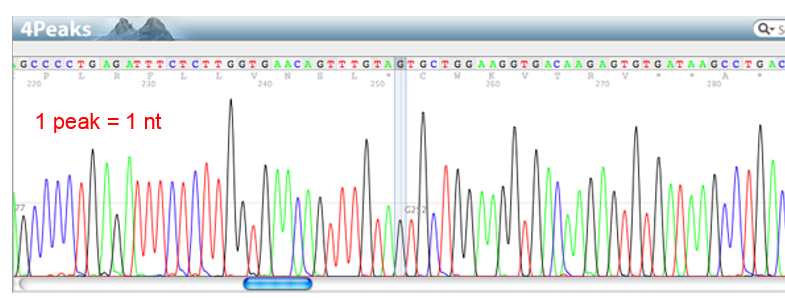
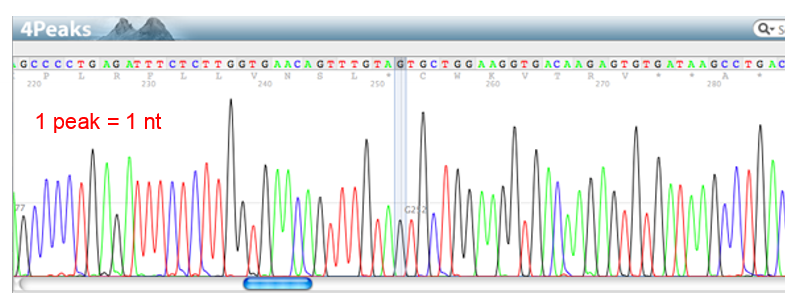
What is read length? What limits it?
number of base pairs in a row that be sequenced in a single reaction
DNA polymerase processivity (how long DNA poly can stay on before falling off)
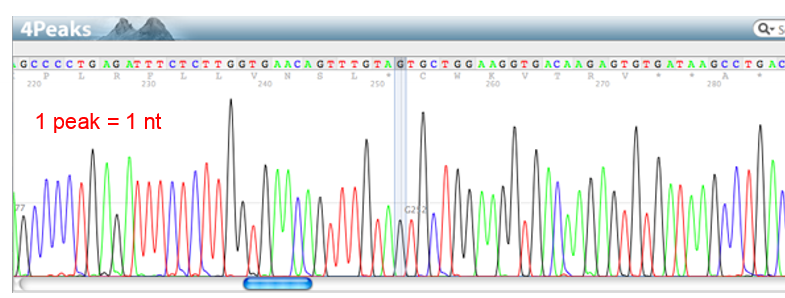
What is a transposable element?
DNA region that moves around genome to change expression of a gene
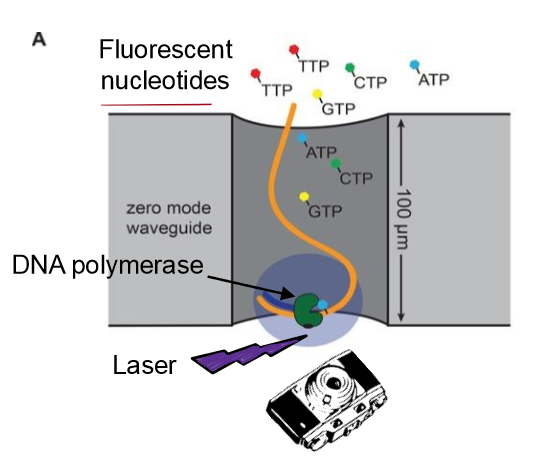
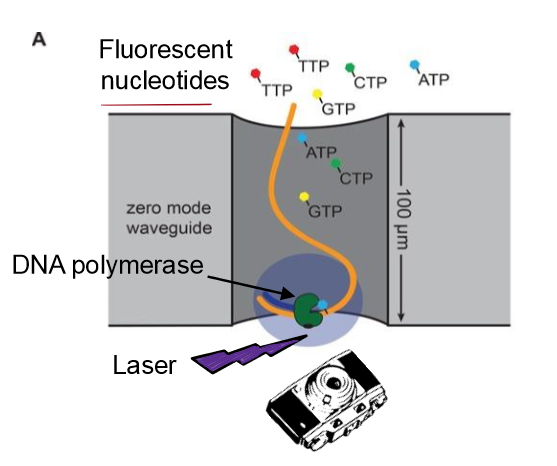
Describe next-gen DNA sequencing, specifcially SMRT sequecning
for sequencing an entire genome
SMRT sequencing (single-molecule real time sequencing)
long read length (high processsivity)
accurate
large outputs achieved by simultaneous analyis
uses fluorescently labeled dNTPS (regular 3’ OH) for continuous syntehsis of strands and records color as DNA is lengthened
results in a chromatograph with the order of nucleotides
can do for RNA as well to see what genes are being expressed at that moment
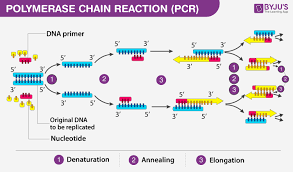
Distinguish between replicating DNA and amplifying DNA, including:
What performs the reaction and what’s being copied
Which occurs in a cell and which occurs in vitro?
What is the outcome of replication and amplification?
DNA replication is performed by a complex of enzymes (DNA poly 3, primase, SSB, helicase, etc.), proceeds from origin of replication
DNA amplification usually targets a specific gene/region of DNA and uses PCR, using heat stabilized taq-polymerases . uses heatcycling to denature, and help anneal and extend
replication occurs within a cell, amplifying occurs in a lab (in vitro)
replication results in two identical daughter DNA molecules, each containing one original strand and one new strand
amplification results in millions of copies of a specific DNA fragment
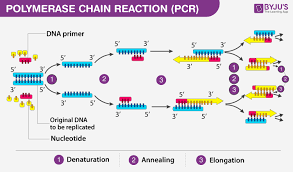
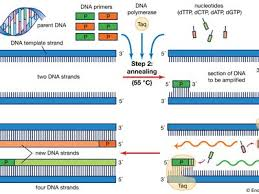
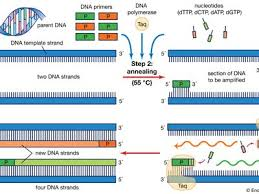
What events occur at each different temperature during the polymerase chain reaction (PCR)?
heating - denaturing/separating strands
cooling - allows primers to anneal
heating - allows DNA polymerase to extend
Sanger sequencing involves DNA polymerase, a primer, and nucleotides. What is special about the nucleotides used and how does this allow the DNA sequence to be determined (dideoxy, fluorescent)
uses fluourescently labeled ddNTPs that lack a 3’ OH
color corresponds to nucleotide
can’t extend chain once ddNTPS added
How does SMRT technology sequence billions of base pairs in a single run of the machine? (output, read length, parallel)
directly photographs/record DNA synthesis in real ime
simultaneously records polymerase additions on multiple nucleotide fragments
continuously synthesizing polymerase has long read length