BIOS 301 - ETC 1
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
Locations of major metabolic processes
Glycolysis → cytoplasm
PDC, CAC → mitochondrial matrix
Electron transport, proton gradient, O2 reactions, ATP synthesis → inner mitochondrial membrane
Structure of a mitochondrion
Outer membrane
Relatively porous, allows passage of small molecules, ions, and metabolites
Intermembrane space
Similar environment to cytosol
Higher proton concentration (lower pH)
Inner membrane
Relatively impermeable, with proton gradient across it
Location of electron transport chain complexes
Convolutions (called cristae) increase surface area
Matrix
Location of CAC and parts of lipid and amino acid metabolism
Lower proton concentration (higher pH)
Mitochondrial quality control
Two pathways for unhealthy mitochondria
Sustained depolarization → mitophagy
Transient depolarization → fusion with healthy mitochondria → allows for salvation of partly defective mitochondria through dilution
Depolarization
Involves loss of electric potential across inner mitochondrial membrane
Number of protons pumped according to electron carrier
10 H+ per NADH + H+
6 H+ per FADH2
3 H+ pumped yields 1 ATP
What determines flow of electrons in ETC?
Electrons flow from a negative to a more positive reduction potential
Delta E must be positive (for negative delta G)
Delta E is calculated by: Eacceptor - Edonor
Energy required for ATP synthesis
30.5 kJ/mol or 0.2-0.3 volts
Terminal reaction of ETC
Requires 4e- transfer to O2 to give 2 H2O
Complex IV (cytochrome oxidase) stores 4 e- just prior to binding and reducing O2
Actual vs. Theoretical yield of ATP from ETC
More than enough energy is made available by passing 2e- through ETC to make 3 ATP
However, this theoretical yield of > 3 ATP is diminished by:
Proton gradient used for transport of other molecules
Controlled leakage of protons to the cytosol
Excess energy dissipated as heat
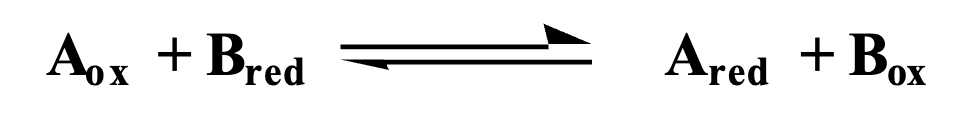
Relationship of reduction potentials and equilibrium constant
Oxidant = e- acceptor, Reductant = e- donor
If reduction potential of A>B, then Keq > 1, and current flows from left to right (electrons move B→A)
If reduction potential of A<B, then Keq<1, and current flows from right to left (electrons move A→B)
Coenzyme Q/Ubiquinone
Lipid soluble compound that readily accepts electrons
Upon accepting 2e-, ubiquinone picks ups two protons to generate an alcohol, ubiquinol
Can freely diffuse in the membrane, carrying electrons with protons from one side of the membrane to the other
Transports electrons from Complexes I and II to Complex III
Chloroplast structural similarities to mitochondrion
Calvin Cycle (CO2 fixation) occurs in matrix
Proton gradient, ATP synthesis takes place across inner membrane
Electron transport and photoreactions occur in thylakoid membranes
Chemiosmotic Theory
Reduced substrate (e.g. NADH+, FADH2) donates e- fuel to ETC
Electron carriers pump H+ from matrix to intermembrane space as electrons flow to O2
Energy of e- flow is stored as electrochemical potential
ATP synthase uses electrochemical potential (i.e. flow of protons down concentration gradient) to phosphorylate ADP
Characteristics of membranes that perform chemiosmotic energy coupling
Must be impermeable to ions to stably establish a proton gradient
E.g. plasma membrane in bacteria, inner membrane in mitochondria, thylakoid membrane in chloroplasts
Must contain proteins that couple the downhill flow of electrons in ETC with uphill flow of protons across membrane
Must contain a protein that couples the downhill flow of protons to the phosphorylation of ADP
Redox centers
Specific chemical groups within the protein complexes of ETC that physically capture and release electrons
E.g. flavin mononucleotide (FMN), flavin adenine dinucleotide (FAD), cytochromes a-c, iron-sulfur cluster
Electrons move through centers with low reduction potential to high reduction potential
Each jump releases a small amount of energy that is sued by ETC protein complexes to pump H+ against concentration gradient
FMN exists in Complex I because it can simultaneously accept 2e- from NADH and pass them along one-by-one to subsequent series of Fe-S clusters
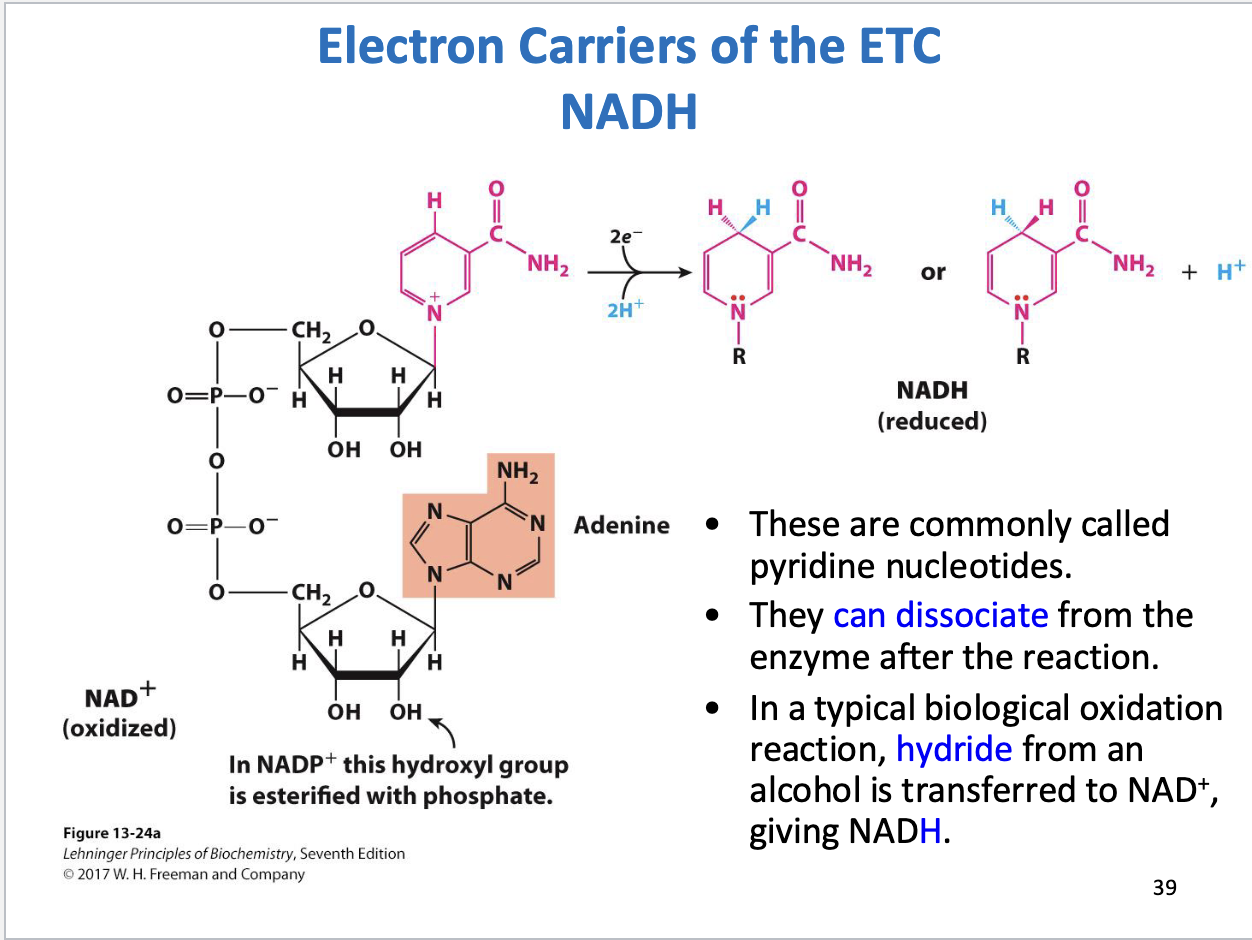
NAD+/NADH structure, function
Dissociate from enzyme after reaction
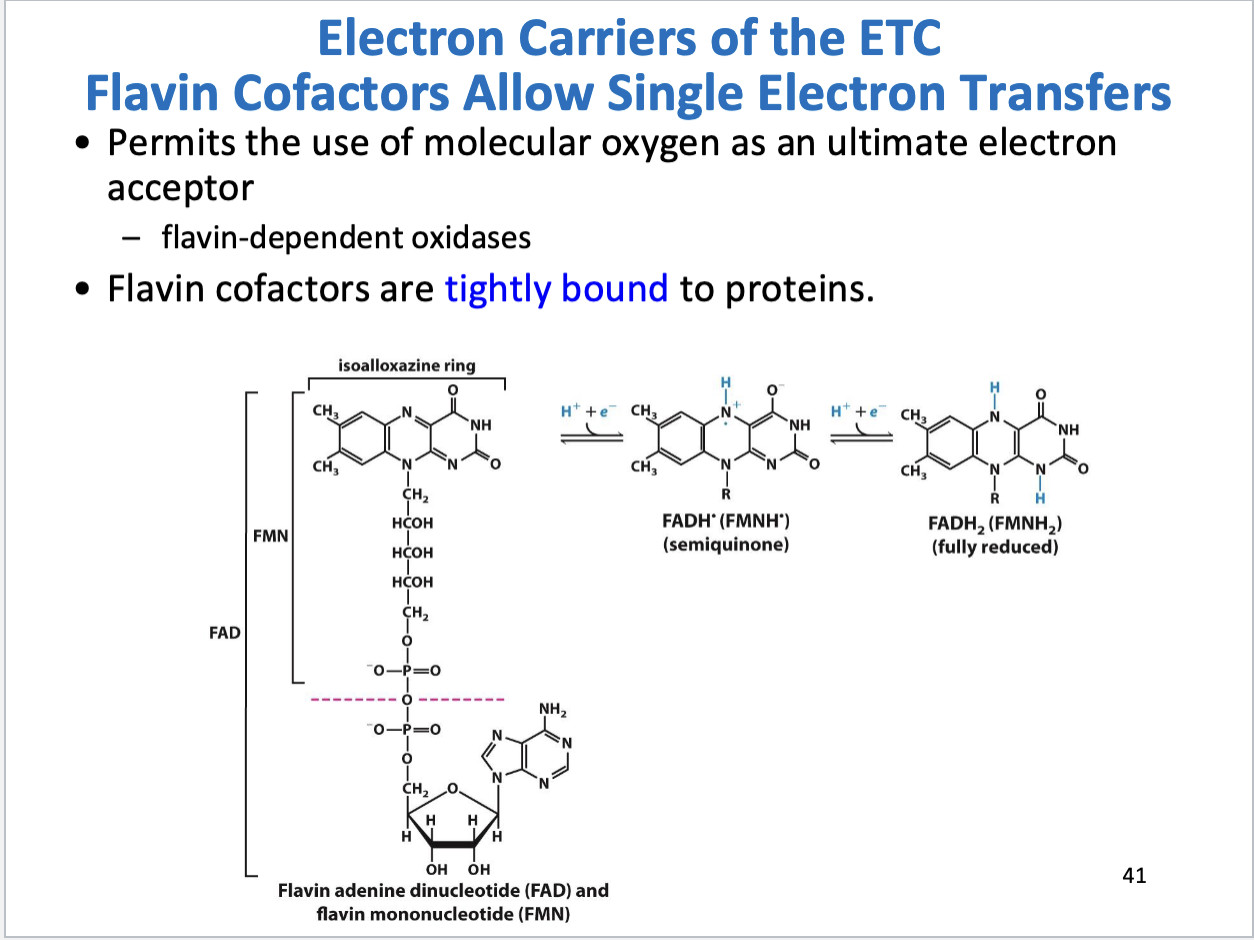
FAD/FMN structure, function
Allow for single electron transfers
Tightly bound to proteins (prosthetic group)
Iron-Sulfur clusters structure, function
One electron carriers
Coordinated by cysteines in the protein
Have a brownish color when oxidized
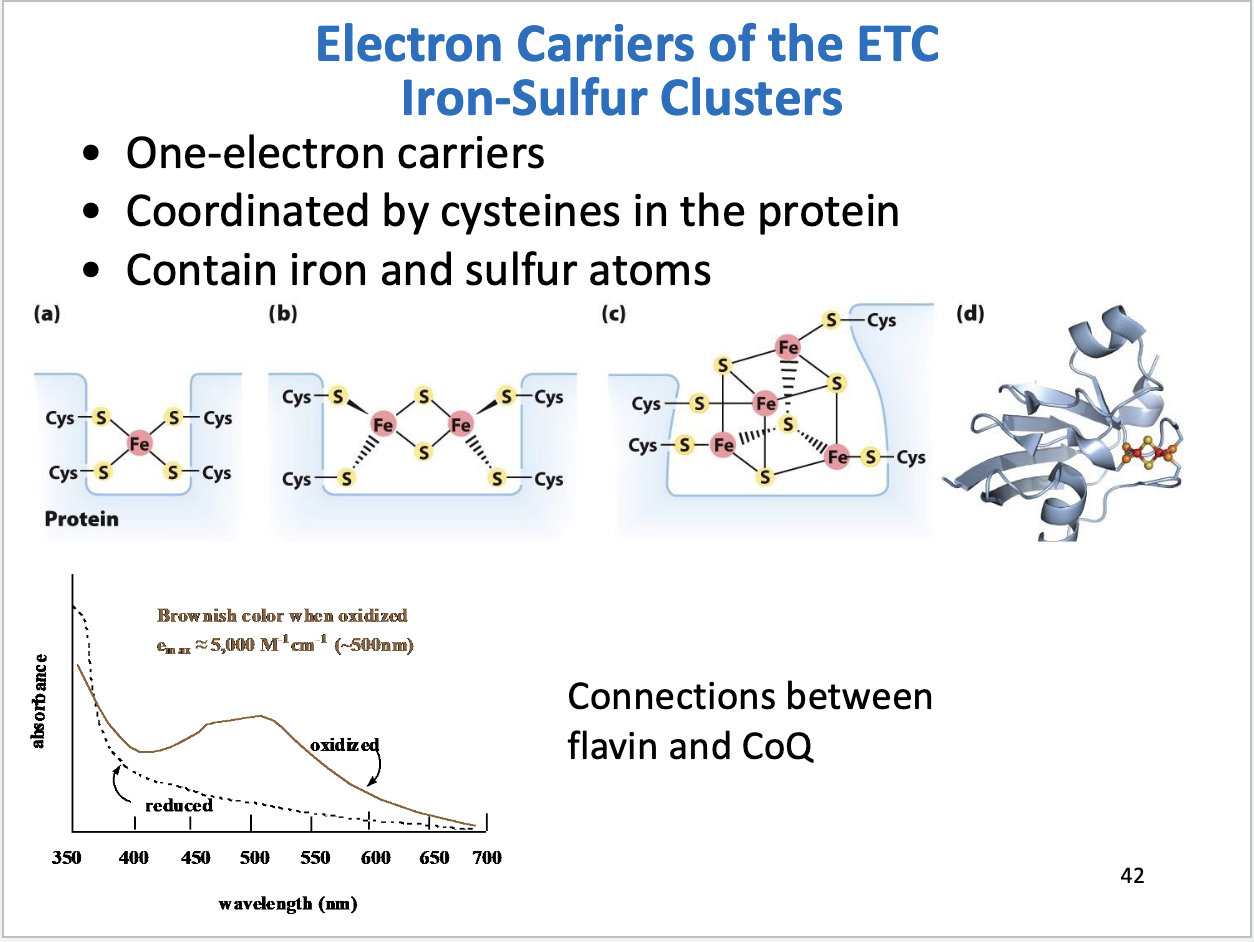
Cytochromes structure, function
One electron carriers
Contain porphyrin derivatives coordinated with iron
Cytochromes a,b,c differ by addition of chemical groups to the porphyrin ring
Oxidized and reduced forms differ in relative light absorption
The wavelength at which alpha absorption peak occurs characterizes the identity of the reduced cytochrome