6.5 Regulation of Gene Expression
1/20
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
21 Terms
Gene Expression
the process by which the information encoded in a gene is turned into a functional product (usually a protein, but also RNAs)
not all genes are expressed or turned on in a cell
the combination of genes expressed and the level of expression determine the phenotype of a cell or organism
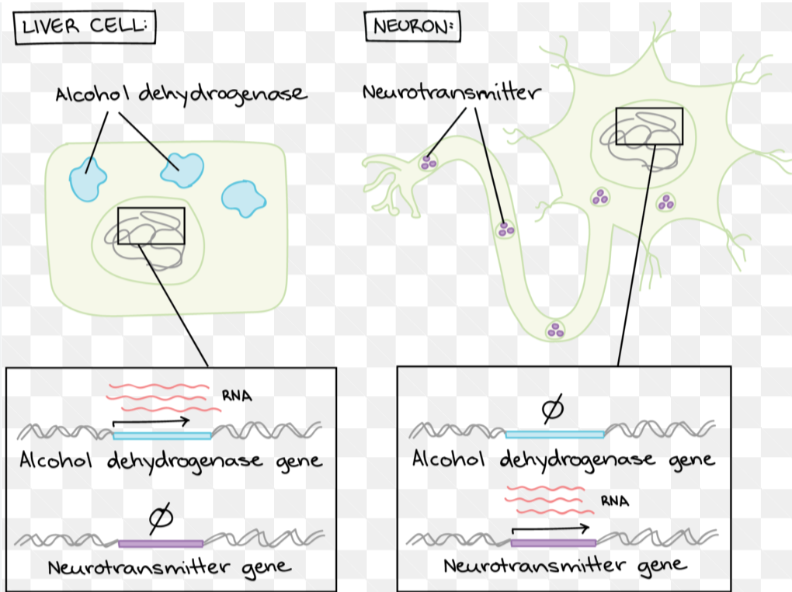
ex. neuron vs muscle cell: same DNA, but different gene expression
ex. butterfly wings: the color pattern is due to cells expressing different genes at different levels across the wings
note: the environment can also impact phenotype
Constitutively Expressed Genes
transcribed at all times
ensures a continuous supply of essential products for basic cellular functions
ex. housekeeping genes in eukaryotes maintain basic cellular functions like glycolysis
Inducible Genes
transcription can be turned “on” or “off” base on environmental and internal cues
ex. rice plants activate genes in response to pathogens for protection
Regulation Benefits
prokaryotes and eukaryotes must be able to regulate gene expression
helps to save energy and resources, respond to changes in their environment, and develop & maintain different cell types (eukaryotes)
prokaryotes: regulation occurs primarily at the level of transcription
eukaryotes: regulation occurs at multiple stages, but many genes are regulated primarily at the level of transcription
Gene Regulation in Prokaryotes
Bacterial Gene Expression: gene regulation in prokaryotes occurs primarily at the level of transcription
includes operons and regulatory genes
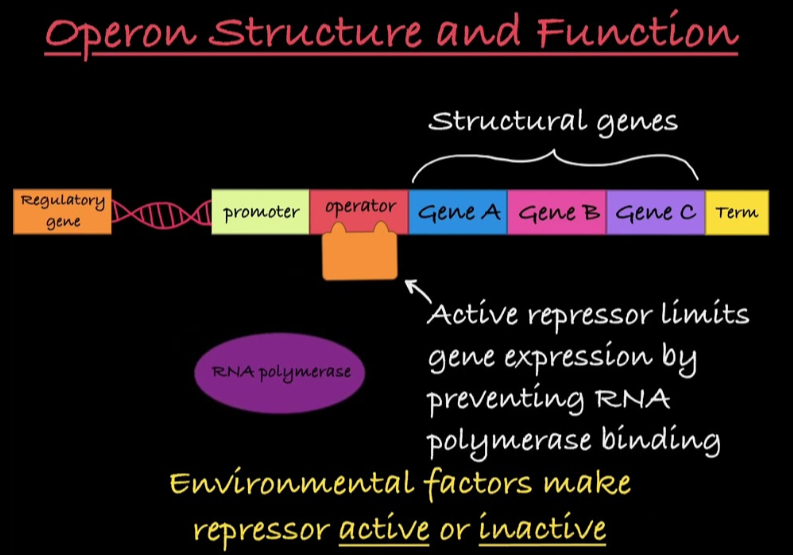
Operons
cluster of related genes that can be regulated and transcribed together under a single promoter
can be repressible or inducible
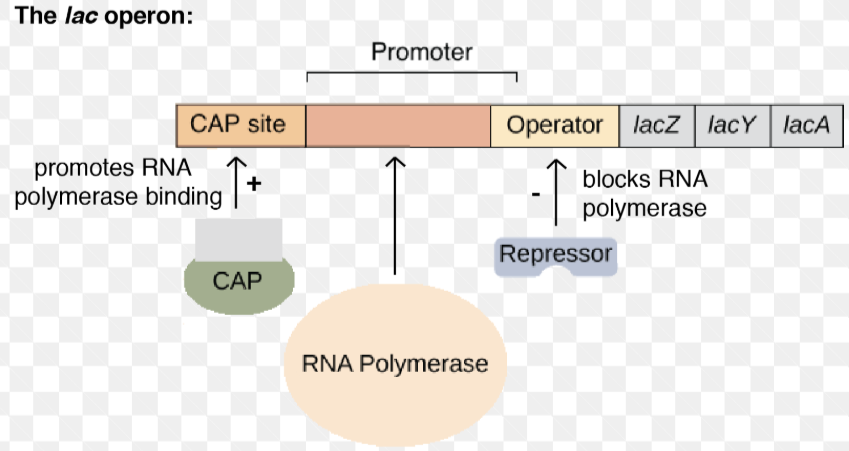
3 parts: promoter, operator, and structural genes
Promoter
RNA polymerase binding site
Operator
the on/off switch
Structural Genes
code for related enzymes in pathway
Regulatory genes
produce regulatory proteins (like repressors and activators) that control the expression of other genes
usually separate from the operon
Repressors
regulatory proteins that reduce transcription when bound to operator
negative regulation
Activators
regulatory proteins that increase transcription when bound to DNA
positive regulation
Repressible Operon
transcription is usually on, but can be turned off (repressed)
on→off
ex. trp operon in E.coli
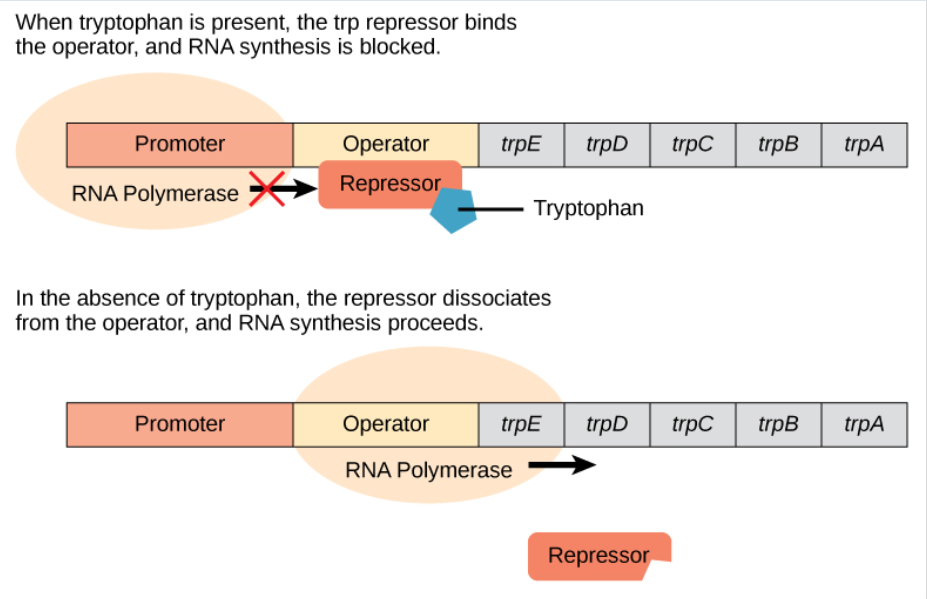
the trp operon in bacteria controls the synthesis of tryptophan (contains genes needed to make tryptophan)
the trp operon is normally “on” (synthesizing tryptophan
when tryptophan levels build up, tryptophan(a corepressor) binds to the repressor, changing its shape
the repressor can now bind to DNA to temporarily shut off transcription for tryptophan, so the cell does not waste energy
Inducible Operon
transcription is usually off, but can be turned on (induced)
off→on
ex. the lac operon in E.coli
has both positive and negative regulation
glucose is the preferred energy source, but if levels are low/absent, E.coli can switch to lactose
the lac operon includes 3 genes that encode enzymes for the use/breakdown of lactose
these genes are only expressed when lactose is present
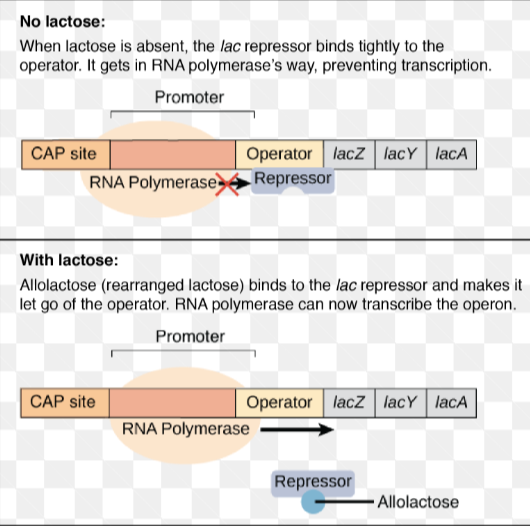
Lac Operon: negative regulation
when there is no lactose present, the lac repressor is bound to the operator, which prevents transcription
when lactose is present, the cell converts lactose to allolactose, which acts as an inducer for the lac repressor(because it changes the shape and allows RNA polymerase to initiate transcription)
allolactose binds to the lac repressor→ repressor changes shape→detaches from operator
RNA polymerase can now bind to the operator, but only loosely until it has help from a protein
transcription is weak until there is positive regulation
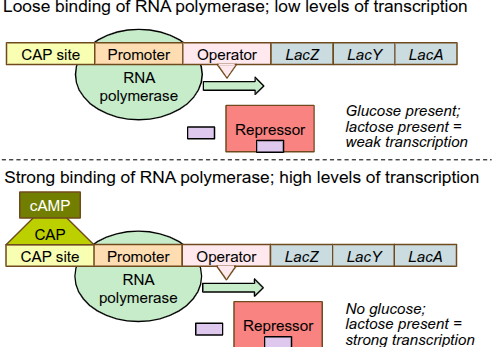
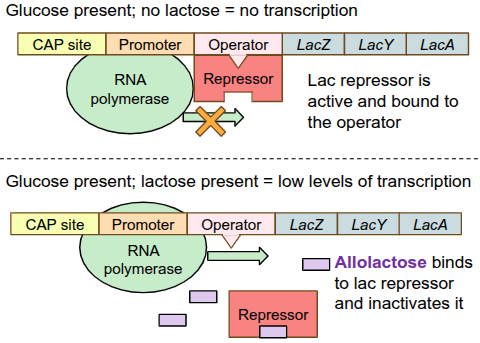
Lac Operon: positive regulation
for strong transcription of the lac operon, RNA polymerase needs help binding to the DNA from a protein called CAP, BUT this protein needs to be activated first
when glucose is very low/absent, E.coli gets “hungry” and produces a small molecule called cAMP
cAMP binds to CAP, activating it
cAMP-CAP complex can now bind to the DNA, which enhances RNA polymerase binding, triggering high levels of transcription
Gene Expression in Eukaryotes
during early embryonic development, cells receive internal and external cues that lead to differentiation
think of these cues as the first instructions about which genes need to be turned on or off in the cells
cues establish early gene expression patterns
Internal cues: cytoplasmic determinants
External cues: induction
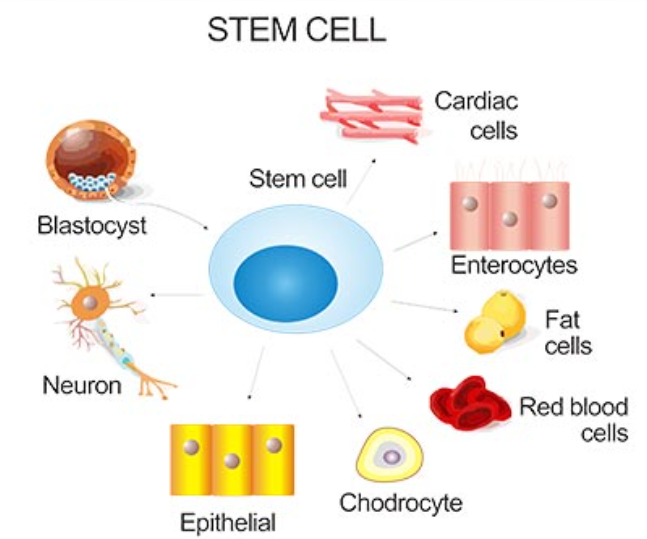
Differentiation
cells become specialized in their structure and function through the expression of different genes for tissue-specific proteins
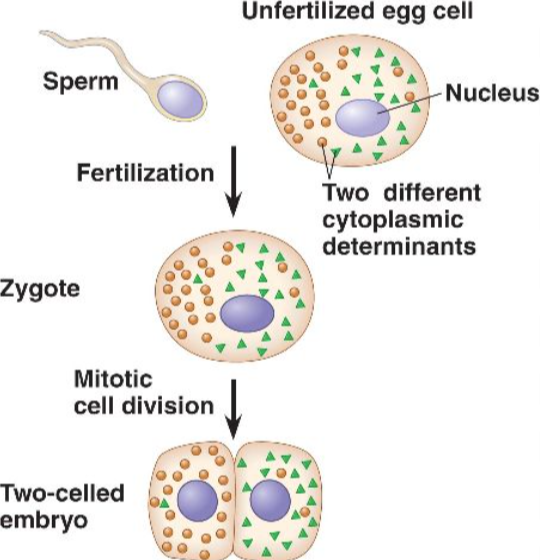
Cytoplasmic Determinants
mRNA and proteins (including transcription factors) from the maternal egg cytoplasm that direct early animal development
unequally distributed in the cytoplasm of an egg cell
when a sperm fertilizes an egg. it forms zygote
as the zygote divides, daughter cells receive different cytoplasmic contents, which means each nucleus is exposed to different cytoplasmic determinants, and that leads to different patterns of gene expression
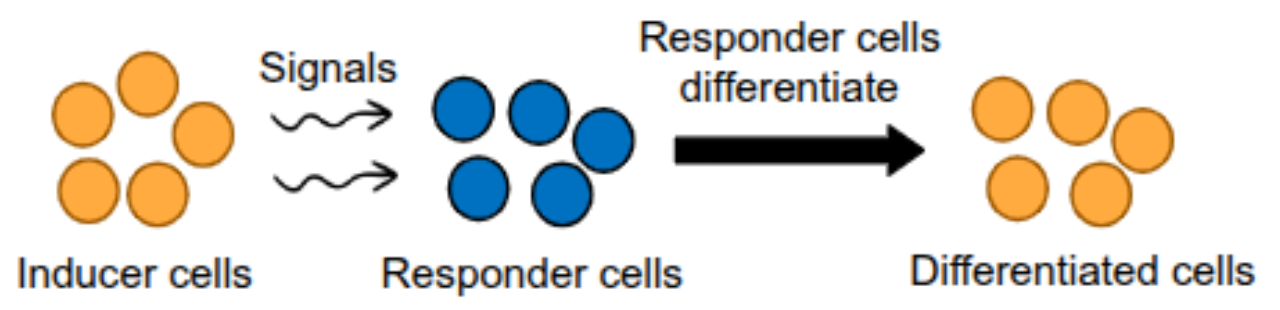
Induction
cell to cell signals in early development that cause a change in gene expression in nearby cells
one group of cells (inducers) send inductive signals to another group of cells (responder cells) via cell-cell contact or paracrine signaling(unit 4)
triggers signal transduction pathway in the responder cells that activate transcription factors, which change gene expression
some inductive signals act as morphogen
the induction of transcription factors during early development (from both internal/external cues) will turn genes on/off
some genes may code for other transcription factors, which leads to sequential gene expression, ensuring the correct order and timing of development
Morphogen
specific type of signaling molecules that diffuse through developing tissues, forming a concentration gradient
a cell’s response is dependent on the concentration of the morphogen it was exposed to
leads to differential gene expression between cells