Genetics M4
1/105
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
106 Terms
What does DNA polymerase need to build a new DNA strand?
1) a template to direct which nucleotide to add via base pairing
2) a 3’ OH to add new nucleotides to (provided by primer)
3) dNTPs
Sanger sequencing also includes low concentrations of chain-terminating ddNTPs, what is needed?
1) a template to direct which nucleotide to add via base pairing
2) a 3’ OH to add new nucleotides to (provided by primer)
3) dNTPs
4) ddNTPs (usually fluorescently labeled ones)
If DNA polymerase adds dNTP, what happens? what about ddNTP being added? How does this relate to Sanger method?
If DNA polymerase adds a dNTP, synthesis can continue… but if a ddNTP is added, synthesis stops
The Sanger method repeats process this many times using the same primer and template.
Each time, the reaction will terminate at a random nucleotide.
If you repeat enough times, what does the Sanger method make?
If you repeat enough times, a “nested set” of fragments of every size will be made
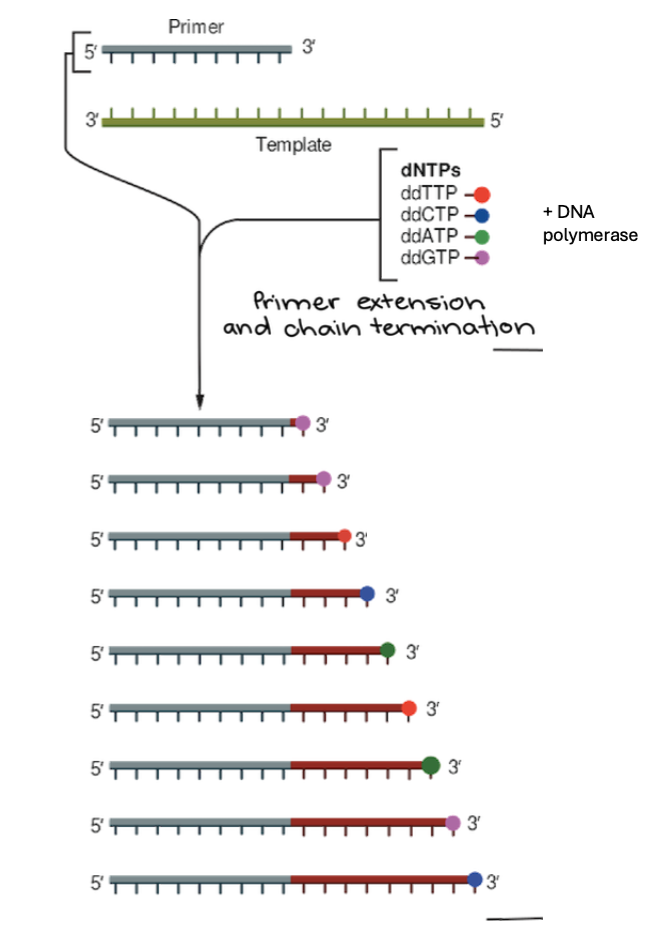
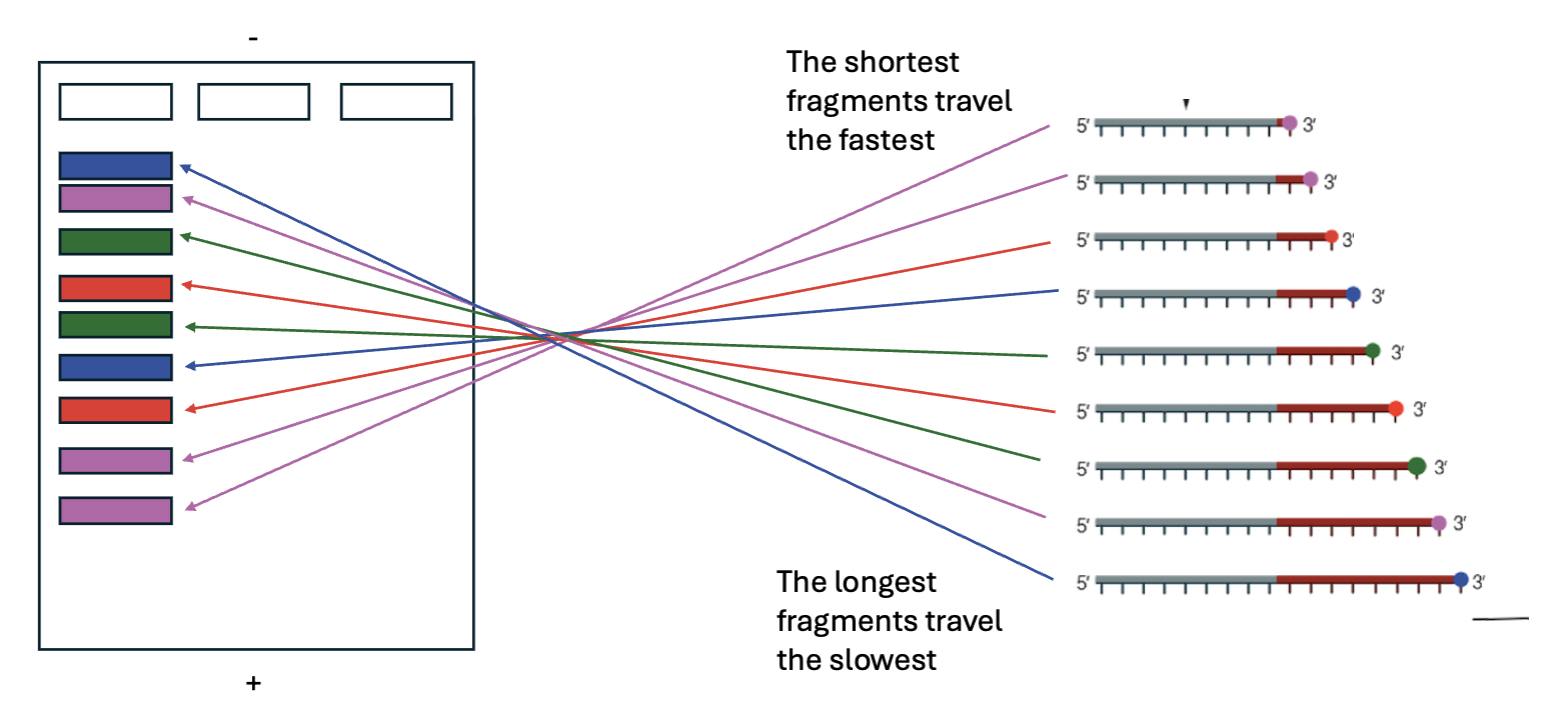
How does electrophoresis separate the fragments?
Electrophoresis separates the fragments by size.
The shortest fragments travel the fastest.
The longest fragments travel the slowest.
Modifications to Sanger sequencing use capillary gel electrophoresis and automated detection, what are the steps?
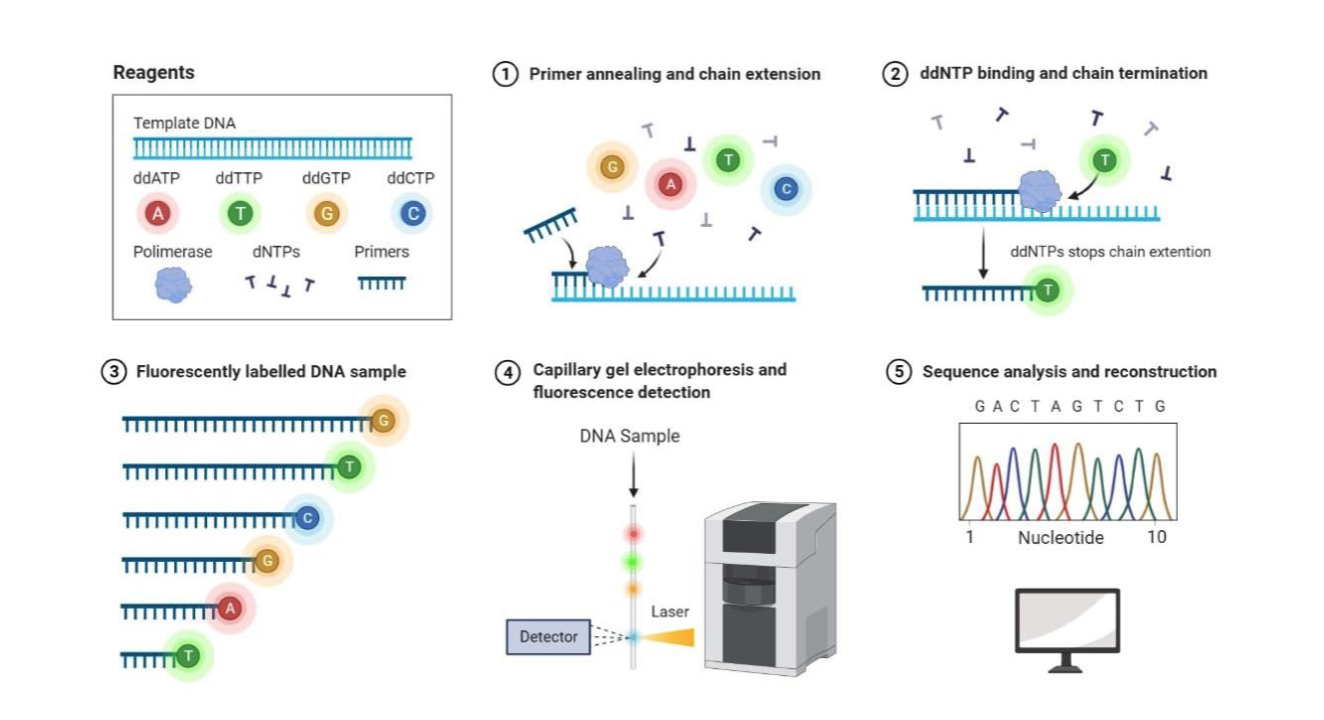
What is the result of sequencing?
It is a “read”.
The maximum length of a read generated via Sanger sequencing is about 1000 bp. it relies on the random incorporation of chain-terminating ddNTPs, which inevitably causes signal loss and accumulation of errors over longer fragments
What is a genome?
The genome is all the genetic information in an organism
Humans have 24 unique types of chromosomes [22 autosomes + X + Y ]
Each of those chromosomes is a single DNA molecule with a specific sequence of bases.
Within those bases there are genes, regulatory elements, functional and nonfunctional
non-coding bases, and more!
What are reference genomes?
Reference genomes are well-characterized standards used for bioinformatic comparisons
A reference genome is NOT
• the genome of a single individual (usually)
• synonymous with “wild type” or “normal”
• 100% perfectly assembled or complete (it may be improved over time!)
To get the entire human reference genome, how do we do it?
“Shotgun sequencing” – break up chromosomes into fragments small enough to sequence!
Genomic DNA samples can be fragmented using restriction enzymes or mechanical shearing
What do restriction enzymes do?
Restriction enzymes cut (“digest”) DNA at specific palindromic sequences (“restriction sites”).
“palindromic” means that the top strand read 5’ to 3’ is the same as the bottom strand read 5’ to 3’
What do some restriction enzymes produce?
Some restriction enzymes produce blunt ends, others make “sticky ends”
Two sticky ends made by the same restriction enzyme will always be complementary to one another!
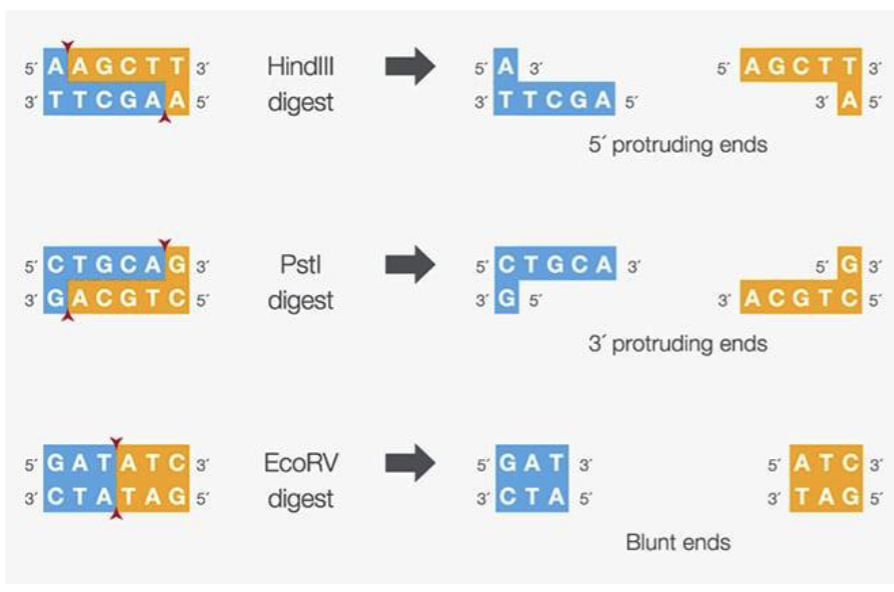
The longer a restriction site is…
the less likely it is to randomly occur in a genome
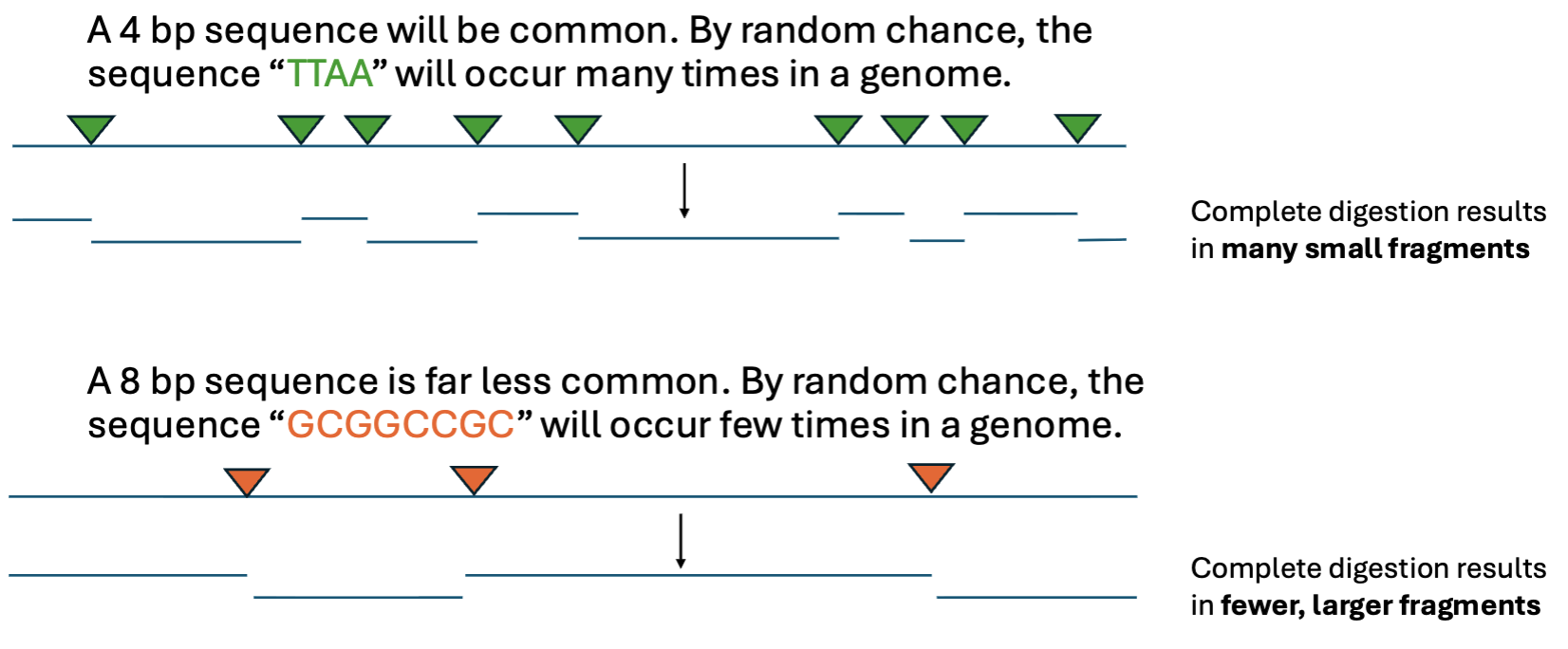
What does incomplete digestion result in?
Incomplete digestion results in fewer, larger fragments
If the digestion is stopped before completion, some restriction sites will be uncut.
What does complete digestion result in?
Complete digestion: If the digestion reaction proceeds for long enough, every restriction site will be cut.
What is produced by incompletely digesting multiple copies of same DNA?
produces overlapping fragments.
Some cut sites are left uncut, but they are all not the same sites that were left uncut on the identical DNA molecule.
What does fragmentation produce?
Millions of fragments with unknown sequences.
This causes:
1) We want to sequence each fragment, but a single fragment is not enough starting DNA for Sanger sequencing.
2) We need to separate the fragments from one another so we can sequence each one separately.
3) We don’t know their sequence, so we don’t know what primer to include to start the Sanger sequencing reaction.
What is the solution to fragmentation?
Clone fragments into plasmids and making a genomic library!
Can the DNA replication machinery in living cells be used to clone DNA?
Yes, Every cell in your body is genetically identical and contains 1 copy of your entire genome → YOU are a cellular clone that has cloned your genome trillions of times!
What does molecular cloning combine?
Fragments of DNA with vectors that can be stored and replicated into many copies
What are cloning vectors? What are the key features?
Are DNA molecules designed for molecular cloning
1) Polylinker (AKA multiple cloning site)
• A section of DNA containing lots of restriction sites for scientists to choose from to insert DNA fragments.
2) Selectable markers (often antibiotic resistance gene)
• Allows for selection of bacteria transformed with the plasmid
3) Origin of replication
• So that the transformed bacteria can replicate (clone) the DNA
What are plasmids?
are small, circular DNA molecules that can be used as cloning vectors.

What are used in plasmids after being used as a cloner vector?
Complementary sticky ends can anneal/hybridize to form a recombinant DNA molecule.
The backbones can be ligated together with ligase.
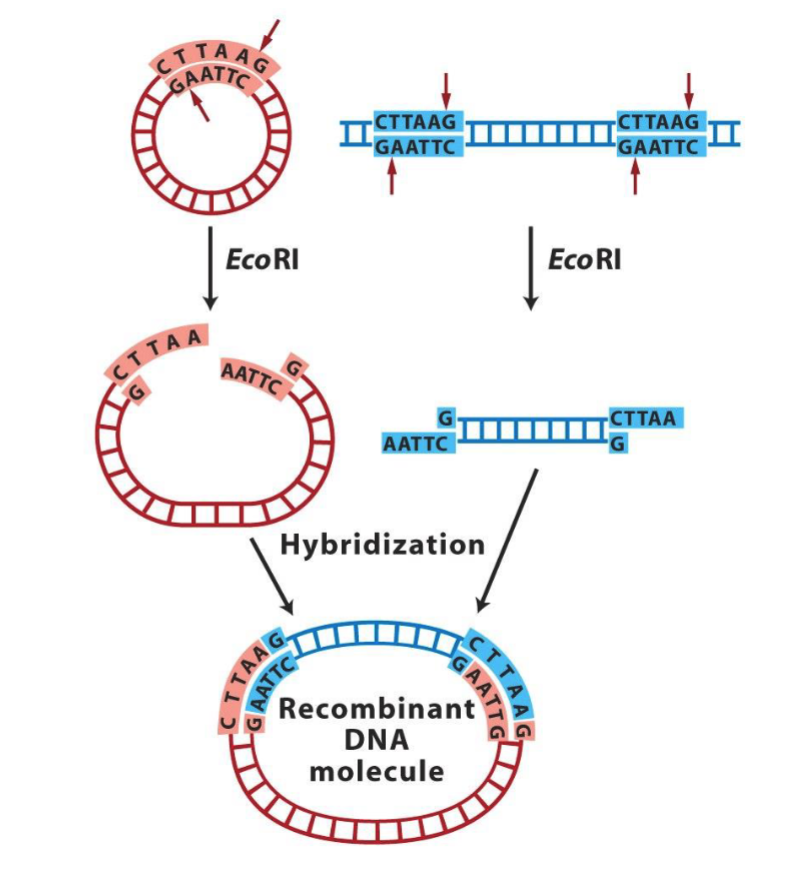
What can we do with vectors and fragments?
Combining many digested vectors with all the fragments from a genome generates a diversity of recombinant plasmids
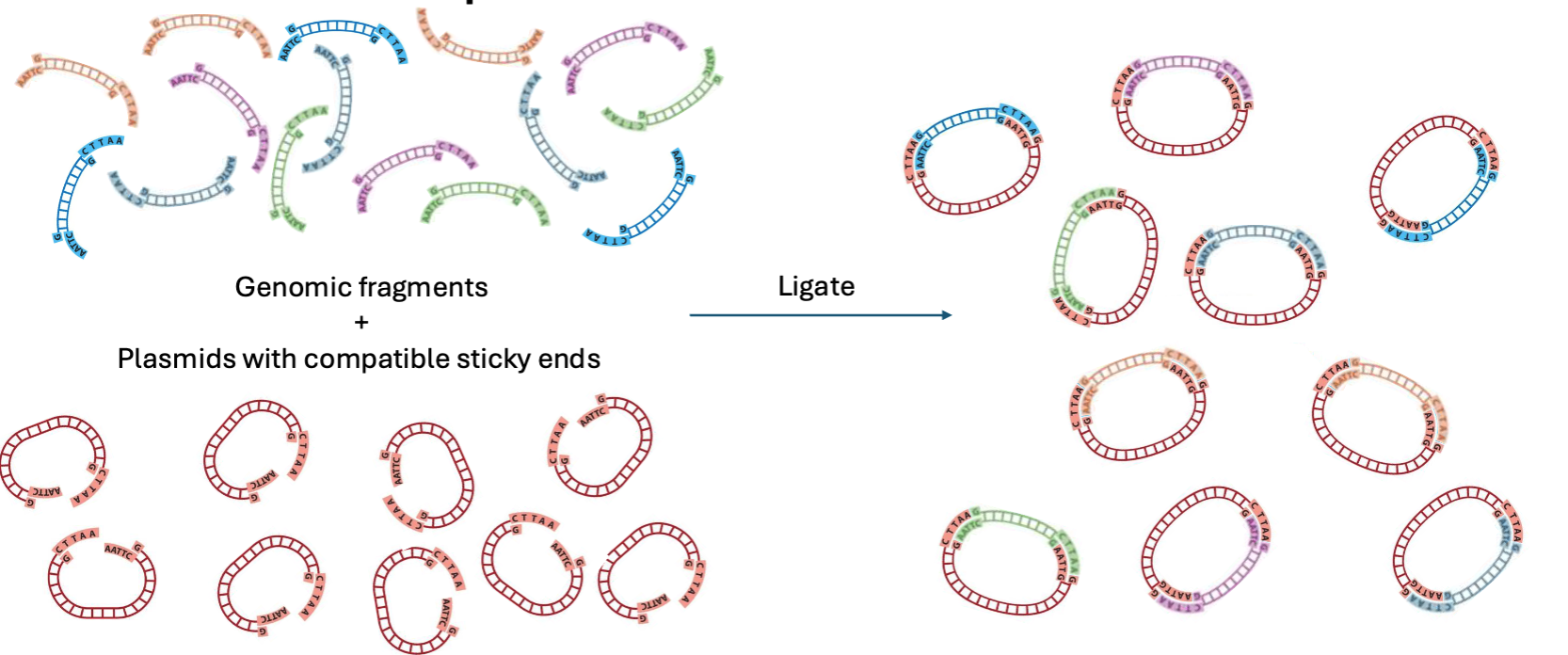
Can recombinant plasmids be transformed into bacteria?
Yes, Transformation is rare enough that most bacteria do not uptake a plasmid. The few bacteria that do get transformed will have 1 plasmid containing a random genomic fragment, which they can copy by DNA replication.
Antibiotic selection eliminates what?
Eliminates untransformed bacteria, so only the rare bacteria we want survive
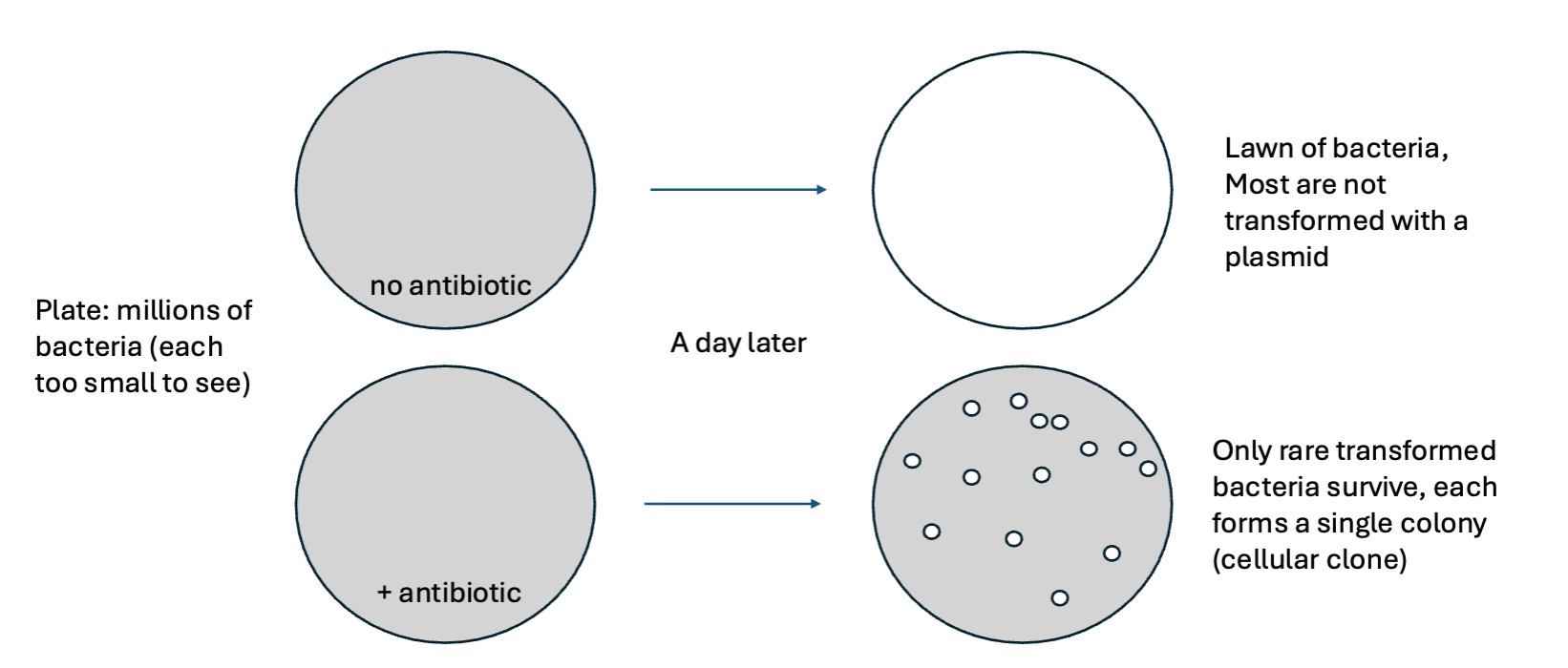
What is the collection of cloned genomic fragments in the transformed into cells is called what?
A genomic library.
Each cellular clone contains only one random fragment of the total information within the genome…but collectively, a complete library will have every fragment of the genome present in at least one colony.
How do we find and annotate genes in a genome?
Bioinformatics tools and databases allow genomes to be explored and compared
What can long open reading frames identify?
Can identify candidate genes in a genome
If you locate a reading frame that is open for hundreds of bp, what are you likely looking at?
At a part of a protein coding sequence
How can we use common ancestry as a way of gene prediction?
The more recently two species shared a common ancestor, the more similar their genomes will be (less time for mutations to accumulate that are the source of genetic diversity!)
Although new mutations can arise randomly anywhere…changes that affect genes are more likely to affect fitness and be eliminated by natural selection.
What does sequence conservation methods identify?
Identify areas of the genome subject to selection pressure.
What happens to functional regions as a result of selection pressure?
They are typically under evolutionary constraint.
What does mutations do to a genes function
Mutations disrupting a gene’s function will likely impact fitness and be subsequently eliminated.
What does conservation reflect in selection pressure?
Conservation reflects selection against new variation when it arises, not a resistance to mutation occurring in the first place!
What can transcriptomic data confirm?
Can confirm whether a predicted gene is functional.
What does mRNAs help us identify in genes?
Knowing what mRNAs are present in a cell can help identify genes that are actively transcribed.
mRNAs are prone to degradation, so one way to study the transcriptome of a cell is to make a DNA copy of all the mRNAs.
The sequence information can be preserved, but in a much more stable molecule.
What do cDNA libraries contain?
Only contain sequences found in mature mRNAs made by that cell
A cDNA library will only contain the genomic information that ends up in the mature mRNAs made by that cell
Will not have enhancers, promoters, but will contain protein-coding sequences.
What does alternative splicing mean?
Alternative splicing means that not all exons will be present in mature mRNAs
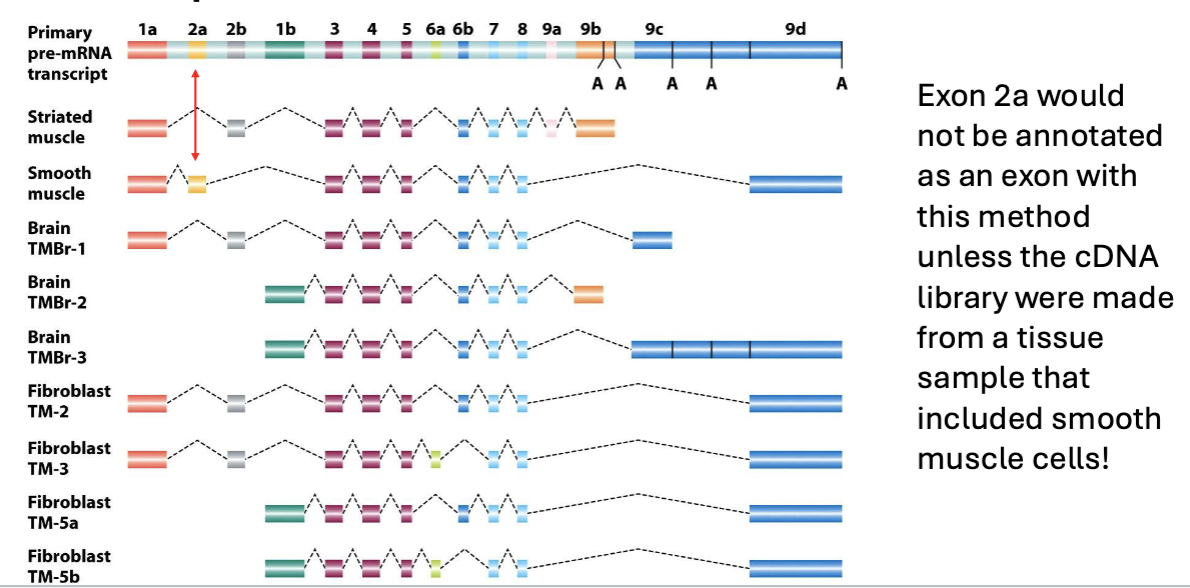
How much of the human genome are genes?
Genes (introns + exons) make up ~50% of our genome
Exons are only 1-2% of the total genome!
What does human chromosomes contain? What is the function of telomeres, and centromeres?
Contain functional DNA interspaced between a LOT of repetitive.
Telomers protect the ends of chromosomes
Centromeres hold sister chromatids together after DNA replication and are attachment points for spindles during mitosis/meiosis.
Other repeats do not have known functions.
What can structural rearrangements generate?
Can generate new genes and pseudogenes through duplication and divergence.
What is exon shuffling?
Exon shuffling is when a new gene is created from the components of multiple existing genes.
What does aligning do to a reference genome?
It simplifies future genome sequencing. Genome sequencing of individuals takes advantage of the fact that there will be many similarities to a reference human genome
How can we use aligning?
Aligning reads to a reference genome can be used to sequence new genomes quickly, with lower depth of coverage required.
We can figure out what part of the genome a read maps to even if there are gaps between reads and/or lack of overlapping reads
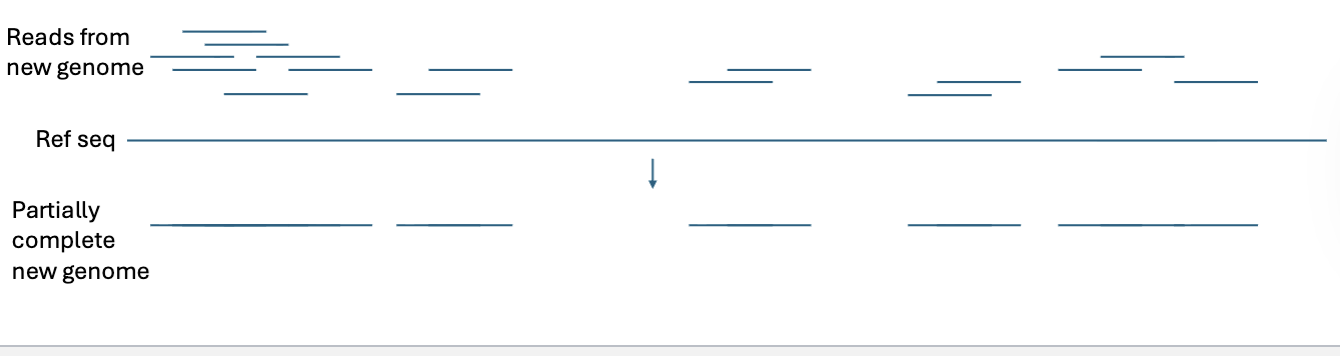
Where does genetic variation occur?
Short answer → anywhere in the genome
What are the 4 major types of genetic variants?
1. Single nucleotide polymorphisms (SNPs)
2. Simple sequence repeats (SSRs)
Short tandem repeats (STRs)
3. Insertions/Deletions (InDels)
4. Copy number variants (CNVs)
Why study genetic variation?
- Drives disease and health outcomes
- Essential for modern science and medicine
- Critical in Forensic Science
- Ancestry and evolution studies
What is an SNP?
A single base change
Most common genetic variant
3-4 million SNP differences between individuals
Used in ancestry inference
Population-specific SNP patterns
Can work with fragmented DNA
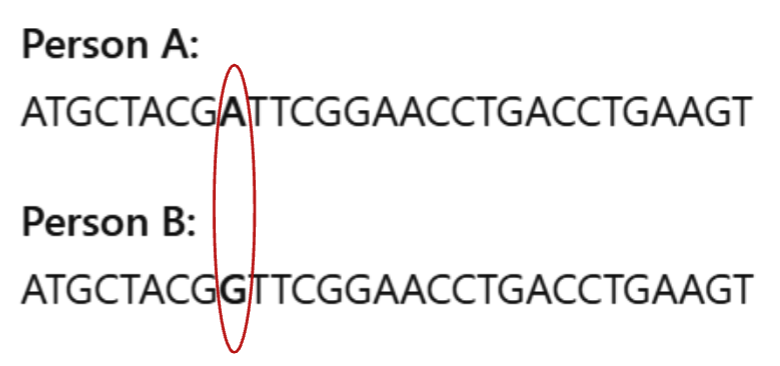
Calling a SNP typically requires a what?
Requires using a reference genome
There is no such thing as a wild type DNA sequence
What is a SSR (aka STR)?
Short DNA sequences (2-6 bases) repeated in tandem
Number of repeats varies between individuals

What is InDels?
The addition (insertion) or removal (deletion) of one of more nucleotides in a DNA
sequence
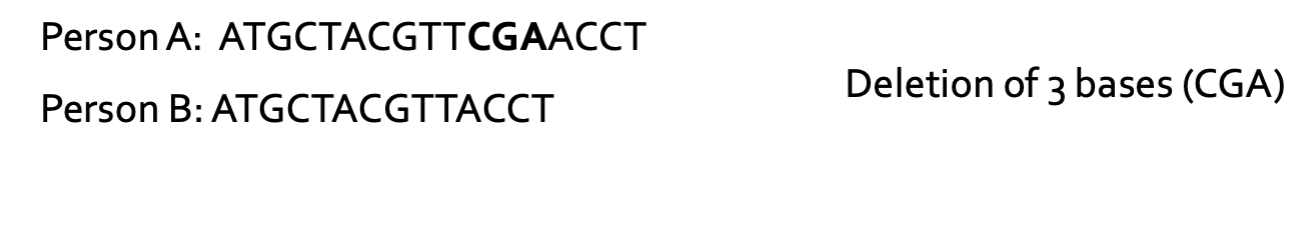
What is a CNV?
Large regions of DNA that are duplicated or deleted
Can involve thousands to millions of base pairs
Can include entire genes
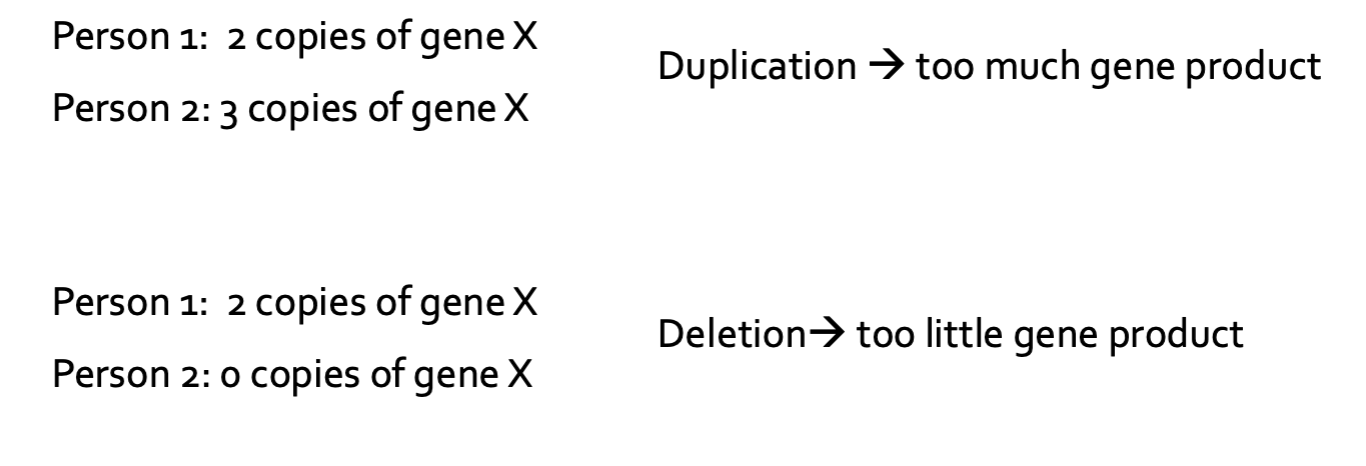
How can CNVs be caused?
CNVs can be caused from misalignment during recombination
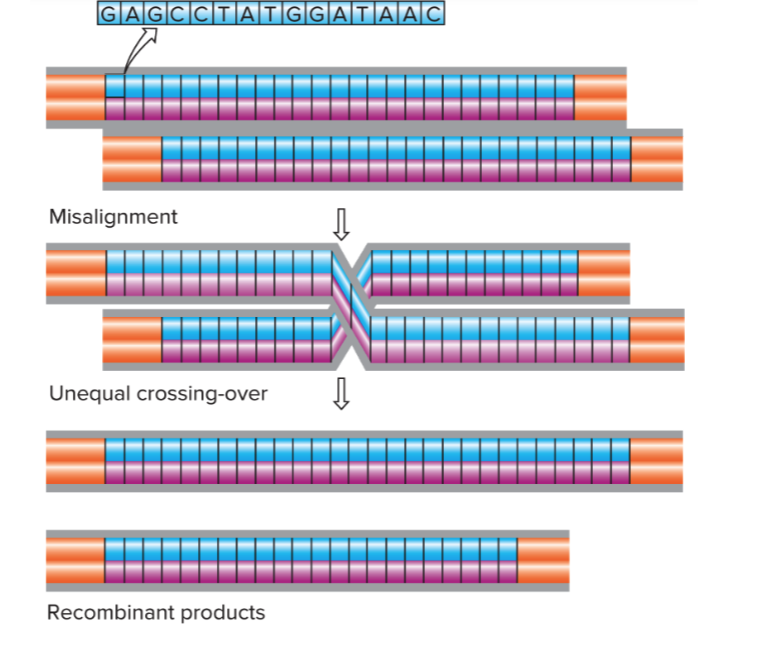
Difference Nuclear DNA vs mitochondrial DNA
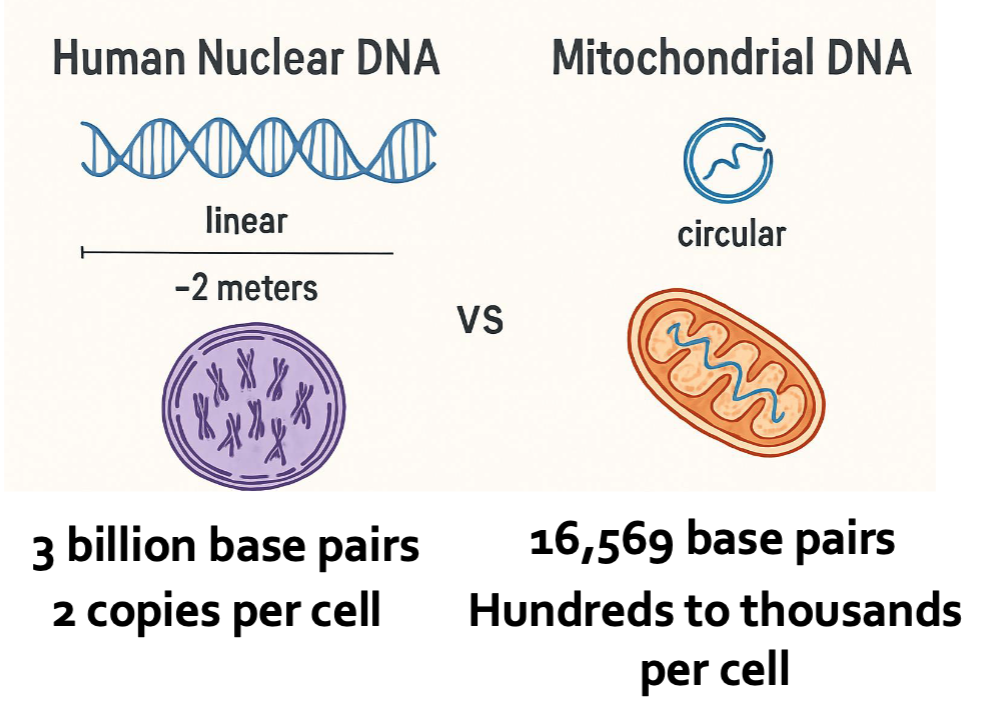
How do we use mitochondrial DNA in forensic science?
Only analyzing from noncoding regions
Resistant to degradation
More copies per cell
Small and circular
Why is mitochondrial DNA less effective at distinguishing between individuals?
Passed down from mother
Individuals in same maternal line have the same mitochondrial DNA
Lower variation between individuals
Groups of people have the same mtDNA sequence
Cannot uniquely identify one person
What is reference-based sequencing?
Compare DNA reads to a reference genome
Identify variants
What is Whole genome sequencing (WGS)
Sequence the entire genome (with or without a reference)
Identify all variants, including novel ones
What is the growth hormone produced by?
Growth hormone is produced by the pituitary and regulates body size
What is a major symptom of GH deficiency in children?
Short stature.
What is CJD and how is it caused?
CJD (the human version of “mad cow”) is a devastating and incurable neurodegenerative disease.
It is caused by Cadaver-derived hGH contaminated with prions gave patients Creutzfeldt-Jakob Disease
What is an expression vector designed for? What features must be present?
An expression vector is designed to express a transgene.
The vector/plasmid must have all the features of a cloning vector:
• Origin of replication → so plasmid can be replicated inside bacteria
• Selectable marker → so we can select for transformed bacteria
• Multiple cloning site → so we can insert DNA (e.g. a transgene)
• Sometimes located in lacZ gene, which enables blue-white screening
Many bacterial expression vectors have the what located inside the lacZ gene? How does it effect the lacZ gene?
Have the polylinker/MCS
The location will be such that…
• An inserted sequence will be transcribed under the control of the lac regulatory elements
• The transcript produced will retain key features of the lacZ 5’UTR (e.g. a Shine-Dalgarno sequence)
• Insertion of a transgene prevents expression of functional lacZ from the plasmid (enables blue-white screening)
How do we isolate our gene of interest?
Do PCR to amplify the specific sequence we want.
Because we want the hGH ORF without any introns, we should use cDNA rather than genomic DNA.
cDNA only contains sequences found in the mature mRNAs expressed in that cell.
Once we isolate our desired insert by PCR, what do we do?
We can insert it into the polylinker/MCS region
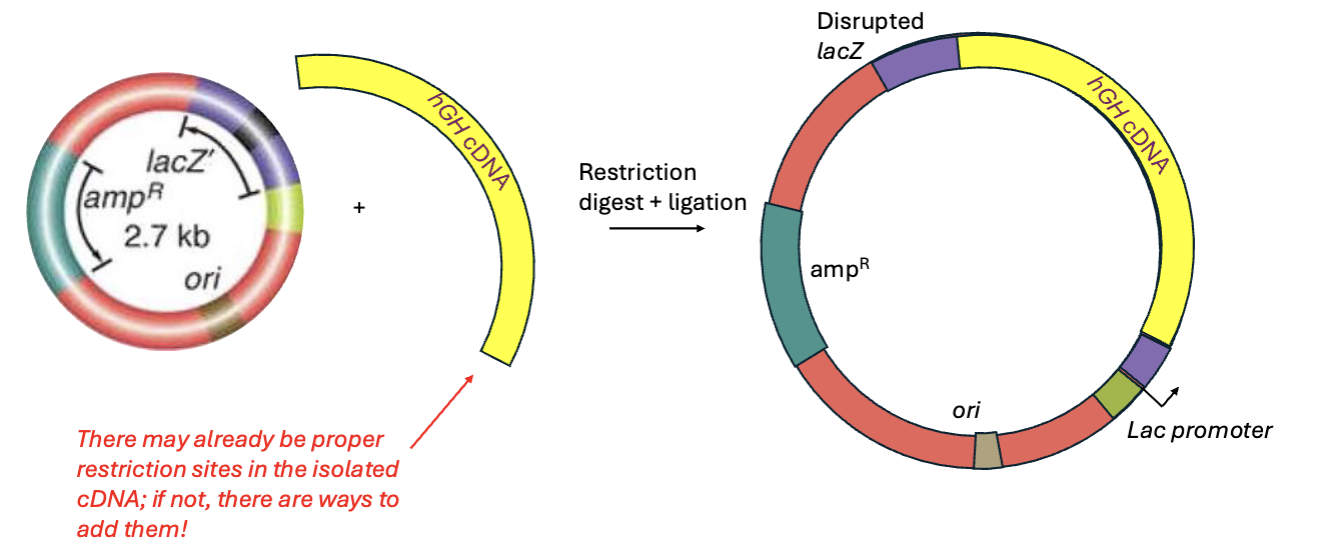
Selection and screening are strategies geneticists use to identify the bacteria of interest. What is selection and screening?
Selection:
Only those that satisfy a certain criterion survive
Screening:
Evaluate to see which that satisfy a certain criterion
Agrobacterium tumefaciens can do what?
Can insert pieces of its Ti plasmid into plant genomes

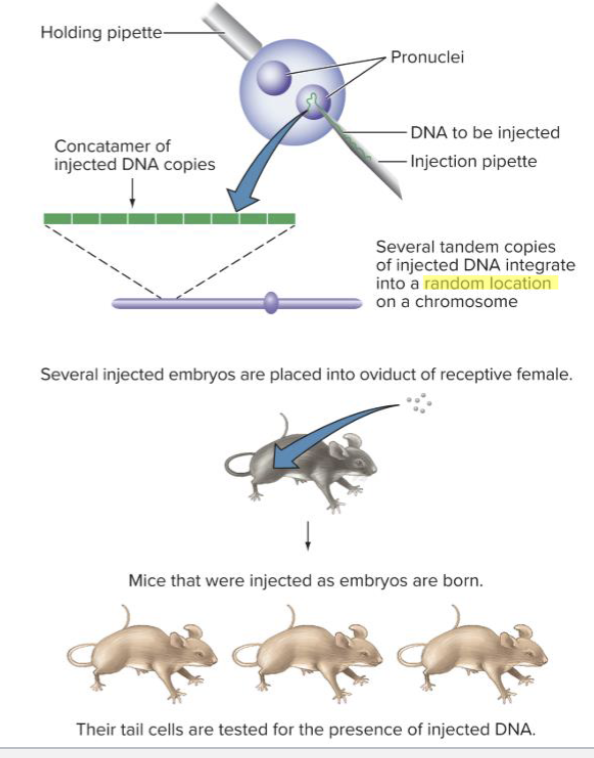
How can transgenic mice be created? and what can they be used for?
through pronuclear injection. The insertion occurs in a random location
Can be used to model disease
What are some of the uses of DNA technology?
Any organism that contains DNA that has been recombined from multiple sources is considered a “recombinant” organism.
Or, in more common terms: Genetically modified organisms (GMOs)
GMO tech is generally safe, but case-by-case product safety concerns exist
New GMOs must be evaluated for any safety concerns specific to the particular genetic modification made
The overwhelming scientific consensus is that GM food is no more or less safe overall than non-GM food
What can gene targeting be used to generate?
Can be used to generate transgenic animals
Unlike the pronuclear injection method, another method allows researchers what?
Insert/delete DNA in a targeted region of the genome.
Mouse “knockout” or “knock-in” models are used regularly in biomedical research
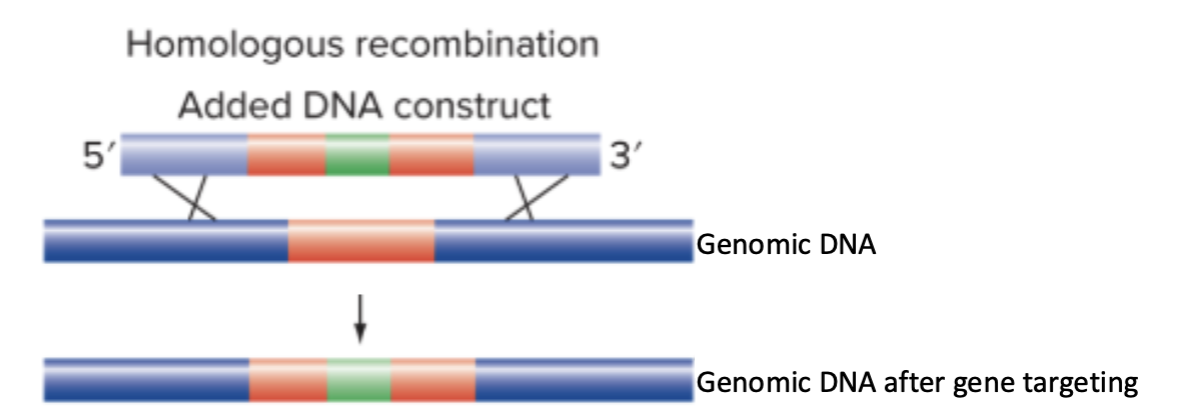
Gene targeting can what with a genomic sequence?
Gene targeting can replace a genomic sequence in with the sequence from a donor DNA construct
If the goal is to knock out the endogenous gene, we can introduce a version of the allele has an insertion in the middle that renders the allele nonfunctional
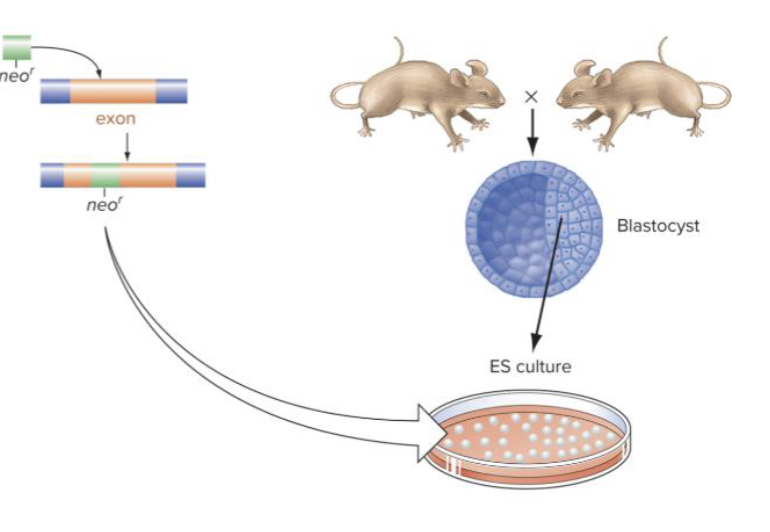
Is gene targeting by homologous recombination a rare event?
Yes it is, we use Mouse embryonic stem cells!
What does the homology template contain? What does this accomplish?
The homology template contains a copy of the targeted gene disrupted by a neomycin resistance gene
This setup accomplishes two things at once:
• Knock out the target gene
• Enable drug selection so only successfully-edited cells survive
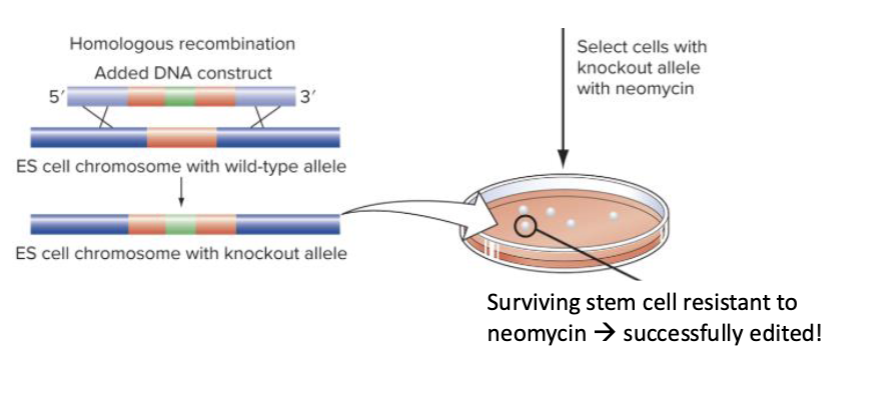
What does Neomycin selection do?
Kills cells that did not incorporate the transgene via HDR
What are edited stem cells injected into and form what?
Edited stem cells are injected into a blastocyst which will develop in the womb of a surrogate.
The stem cells are pluripotent, which means they can develop into any kind of cell type given the proper cues.
As a result of embryonic stem cells being pluripotent, what can it do when returning to a blastocyst?
They can contribute to any part of the chimeric organism.
What is a chimeric organism?
Some parts of the body will have cells that contain the genome of the host blastocyst cells
Some parts of the body will have cells that contain the genome of the gene-targeted ES cells
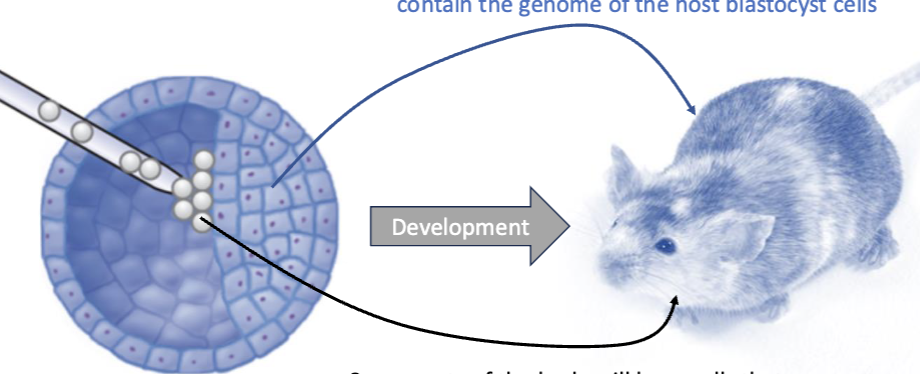
What are Mosaic pups?
Mosaic pups" (or mosaic animals) refer to individuals that possess two or more genetically distinct cell lines within a single body.
Some cells descend from the original blastocyst. Other cells descended from the edited stem cells.
Mosaic pups are crossed with WT mice to produce offspring
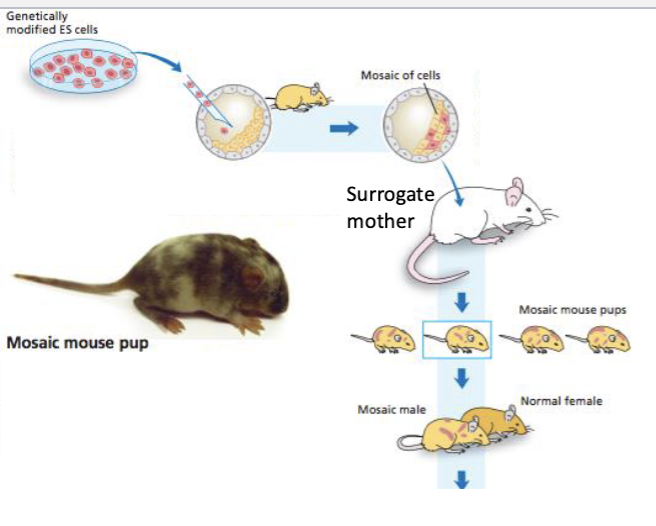
What are Knock-ins?
Replacing an allele (goal is not to simply disrupt the endogenous allele, but to replace with a new one)
What are some mechanisms to repair DNA if a double-stranded break occurs?
1. Non-homologous end-joining (NHEJ)
often introduces mutations
2. Homology-directed repair (HDR)
If DSB is induced, what can it lead to if NHEJ is used?
Random indels can be introduced at the targeted site.
This technique is often used to generate LOF mutations. This relies on cutting the gene in a place where a random indel is likely to disrupt function
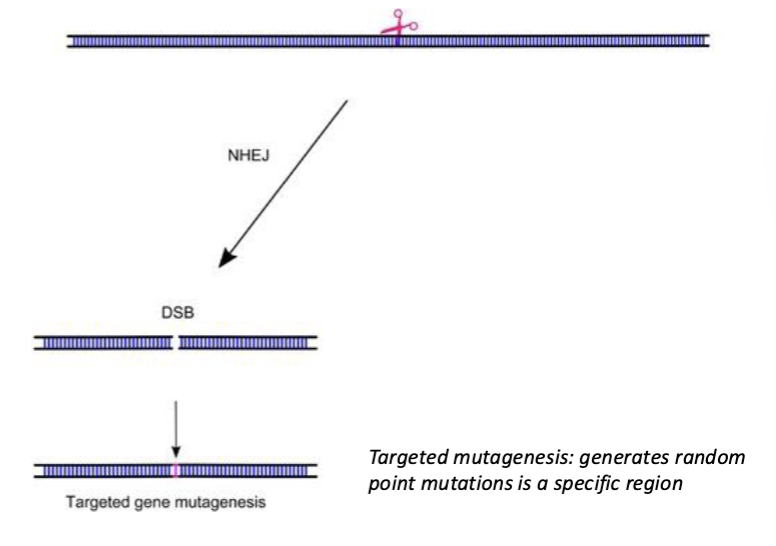
If DSB is at a precise location, what does it stimulate?
Homology-directed repair (HDR)
HDR normally relies on a sister chromatids. BUT…if a repair template with homology arms is provided, the cell may use that instead.
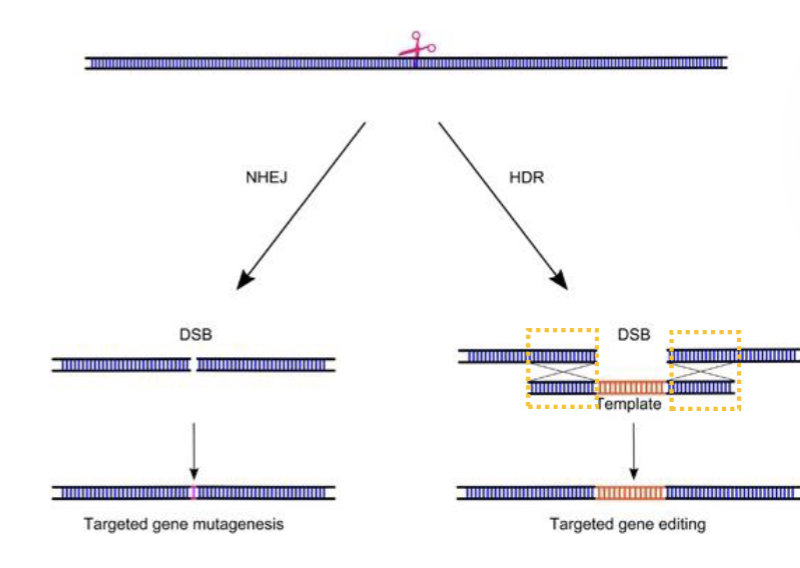
What is CRISPR - Cas9?
Is a bacterial defense system against viruses
Bacteria integrate bits of viral genetic material into their genomes. These encode for RNAs that can bind to viral DNA by complementary base pairing, allowing the bacteria to detect and destroy viruses.
What does CRISPR - Cas9 do?
cut DNA at precise locations
What does the guide RNA do for Cas9?
The guide RNA directs Cas9 to cut at specific locations based on base pairing between the spacer region and the target sequence
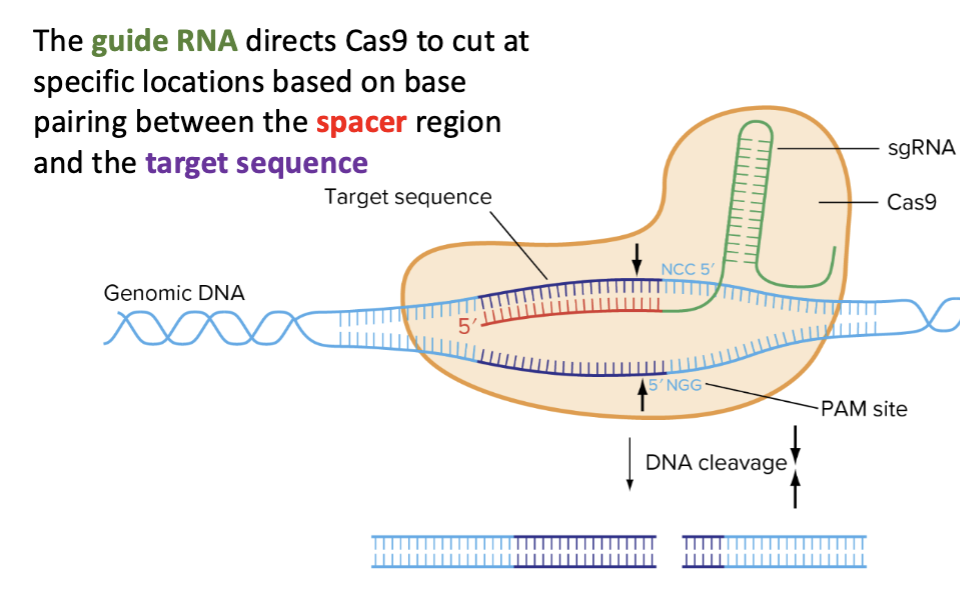
What is Cas9?
Cas9 is an enzyme that cuts DNA to form double-stranded breaks
In bacteria, how is Cas9 used?
Cas9 uses two separate RNAs. Scientists adapted it for biotechnology by creating a single guide RNA which functions the same as the two separate RNAs
CRISPR-Cas9 is what in genome editing?
CRISPR-Cas9 is specific AND programmable, which enables targeted genome editing
Specific: it will cut only at particular a DNA sequence
Programmable: The DNA sequence it cuts is not set in stone, but rather can be determined (by the sequence of the guide)
Two conditions must be met for Cas9 to recognize and cut DNA, what are they?
1) The single guide RNA base pairs to the DNA
2) The DNA in the non-complementary strand contains a PAM sequence in the correct location (directly adjacent to where the guide binds, on the side closest to the 3’ end of the guide)
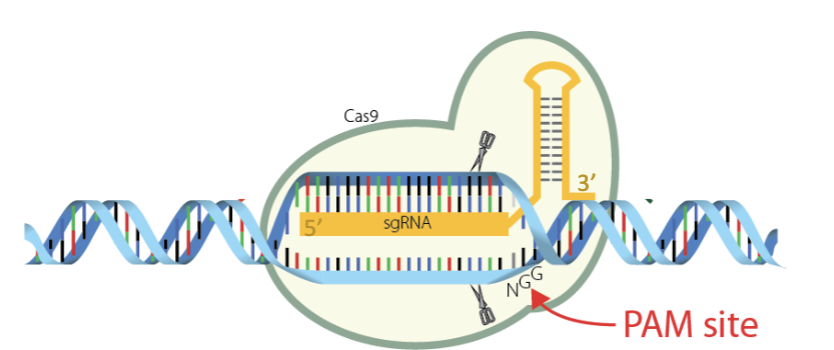
What does sgRNA contain?
Contains the same sequence (except Us instead of Ts) as the bottom strand in this diagram and it does not contain the PAM sequence
Complementary to the top strand, meaning it has the same sequence as the bottom strand.
Is the guide RNA complementary to the CRISPR locus of the genome?
Yes it is, the PAM is not complementary to the sequence transcribed from the bacterial genome to make the crRNA.
Cas9’s dependence on a PAM prevents the bacteria from targeting the complementary region in its own genome!
Hemoglobin is a what?
Is a protein in red blood cells that carries oxygen
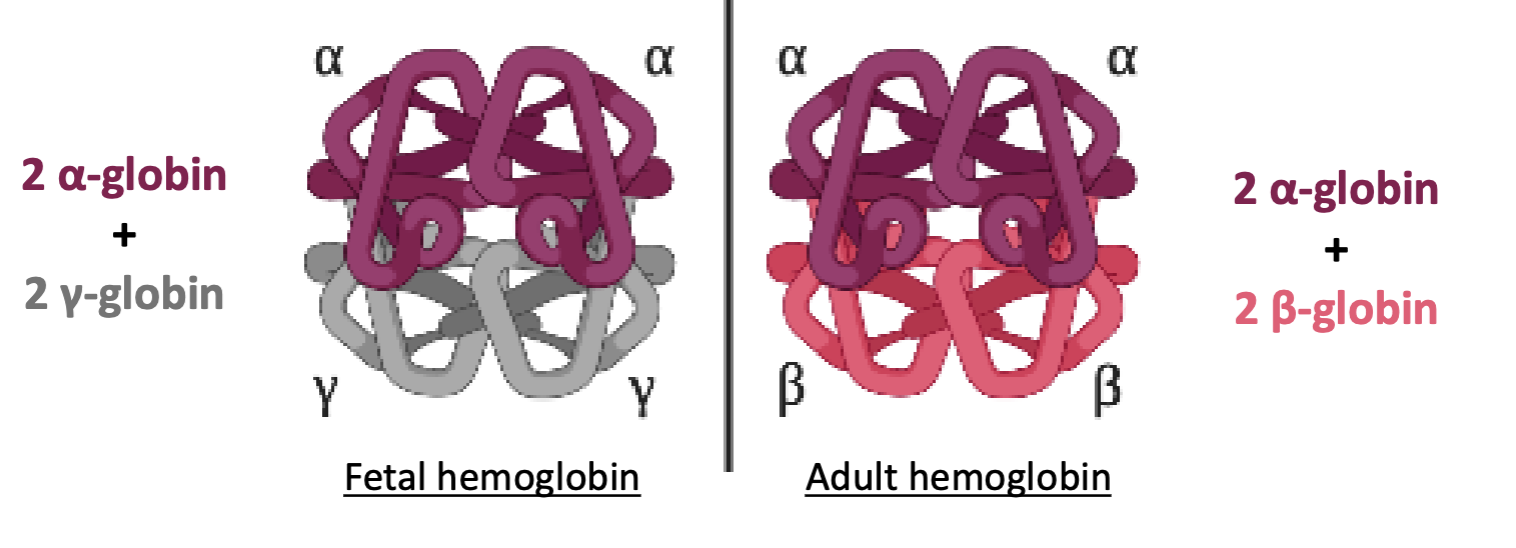
What are the four subunits of Hemoglobin?
Four protein subunits: 2 a-globin + 2 B-globin
Each subunit is encoded by its own gene
Which globin results in two serious blood disorders?
Mutation in the B-globin
1. A specific missense mutation changes the 6th amino acid from glutamate to valine, which causes protein aggregation and sickle cell disease.
2. Any null or amorphic mutation that reduces the amount of β-globin produced causes to β-thalassemia.
Fetuses have what type of hemoglobin?
Fetal hemoglobin, which incorporates y-globin instead of B-globin
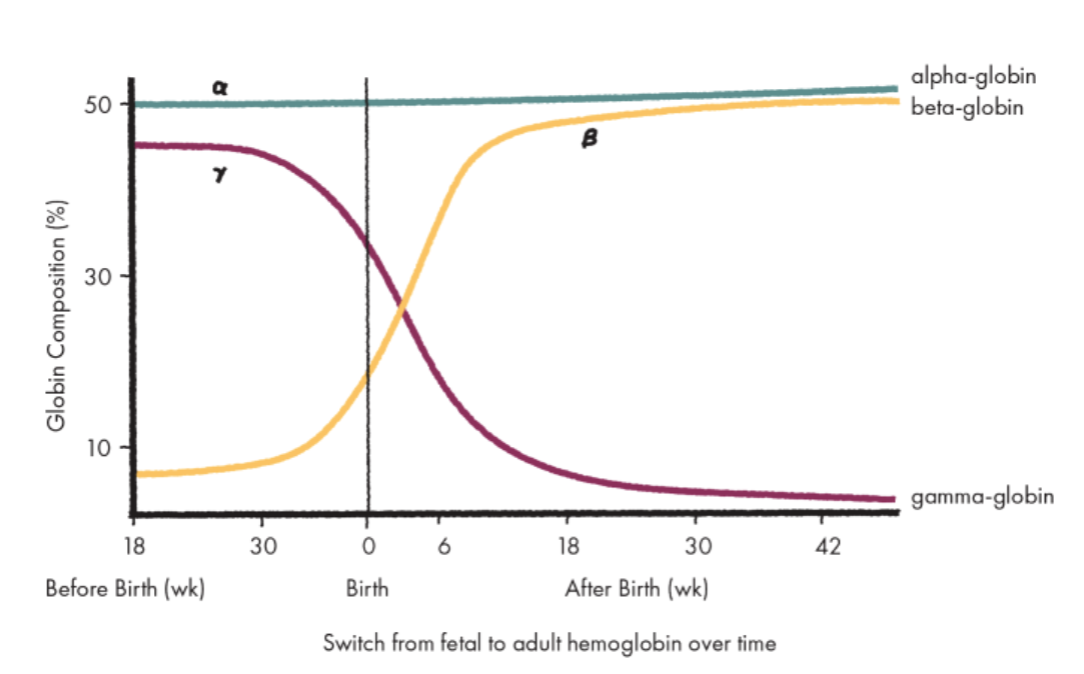
Around birth, what globin is turned off and what begins to be expressed?
Around birth, γ-globin expression mostly turns off and β-globin begins being expressed
Where are all blood cells derived from? And what does bone marrow transplants do?
All blood cells are derived from blood stem cells in the bone marrow.
Bone marrow transplants can replace a person’s blood stem cells
A recipient’s stem cells are purposefully killed by chemotherapy, then replaced by donor cells.