BSCI 331: Exam 3 - Regulation of Protein Translation
1/10
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
11 Terms
untranslated regions (UTRs)
information in 5’ and 3’ ends able to regulate translation efficiency as well as mRNA stability
block ribosome from translating RNA
5’ utr RNA structure allow binding of translation repressor protein that blocks ribosome access where rna structure itself may inhibit ribosome scanning
repressors binding to 3’ utr can prevent communication bettween 5’ and 3’ ends of mrna → required for efficient translation initiation
riboswitch
may use binding of an ion or small molecule to switch between translation as “on” and “off” states ONLY at the 5’ end of mRNA
eIF2 (phosphorylation)
initiation factor that binds to met-tRNA to become active once have gtp
inhibits global protein synthesis where unfolded protein is made
uses GTPase motif to mediate binding of initiator met-tRNA to small ribosomal subunit
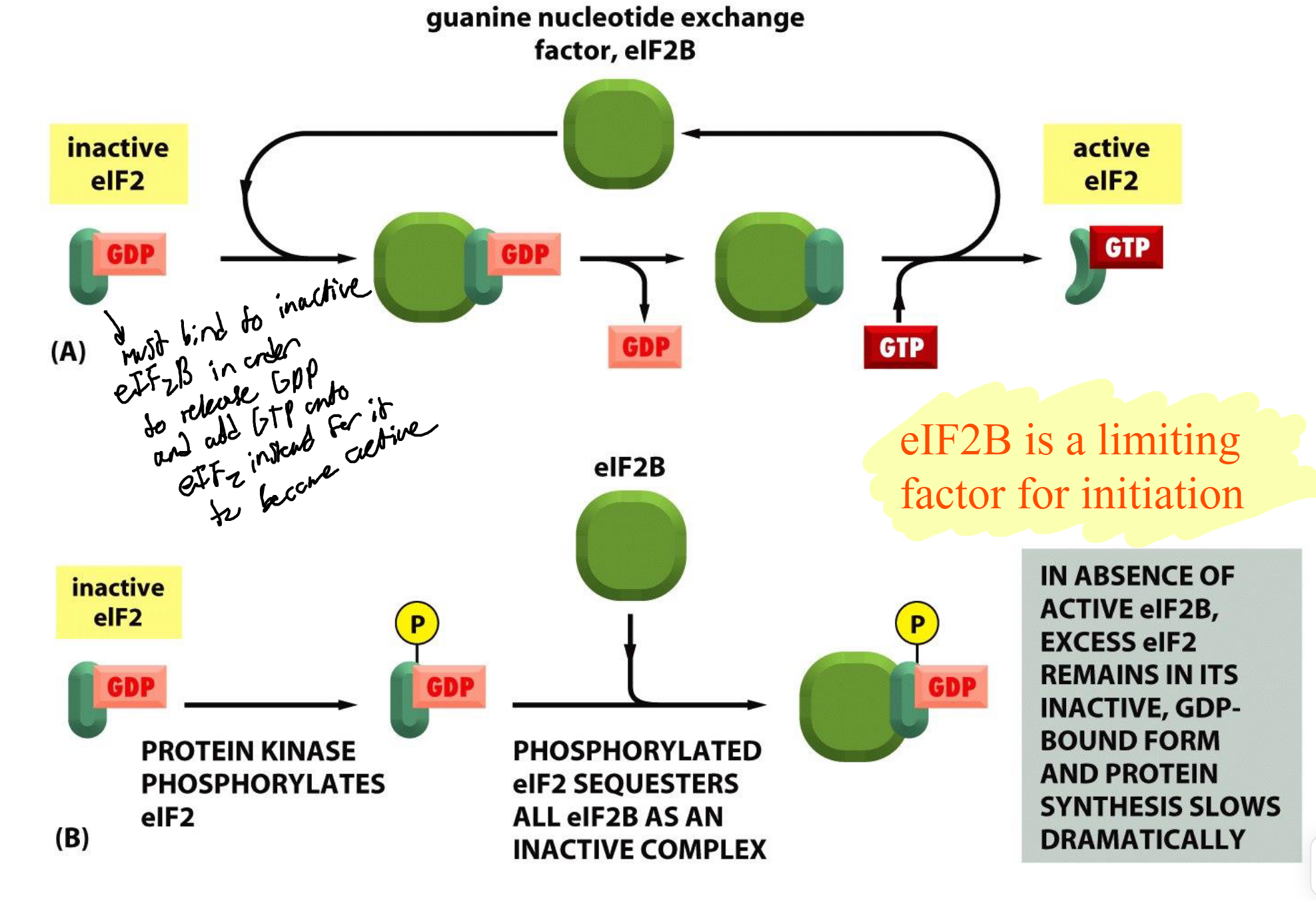
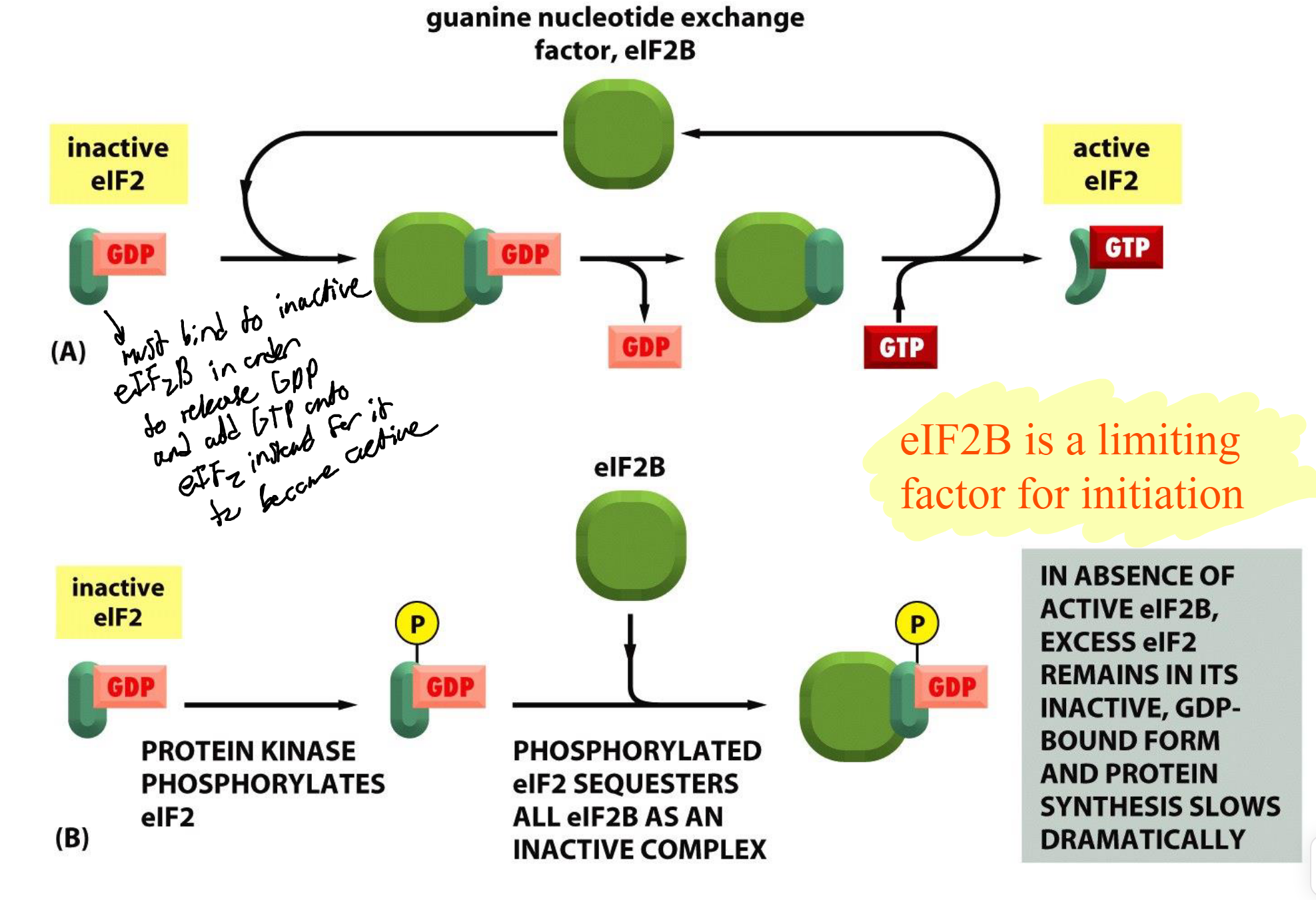
eIF2B
GEF limiting factor that catalyzes exchange of GDP for GTP → activates eIF2
phosphorylation of eIF2 then turns it into an inhibitor of eIF2B → blocks translation initiation and allow cell to recover
bounds to eIF2 complex in order to release GDP and add GtP onto eIF2 to make it be active to inhibit or prevent translation to happen
how much of eIF2 do we need for phosphorylation?
only small amount to be phosphorylated to allow for cell recovery
upstream open reading frames (uORFs)
context surrounding AUG can allow regulation by this
reading frame must be kept the same throughout
sequence starting with AUG starting codon and ending with a stop codon, theoretically able to encode a polypeptide (small and short)
does not mean it will go all the way through translation pathway and reach main coding sequence → eventually will release or cleave off towards middle
if reach to main coding sequence then able to bind to eIF2
what happens if the ribosome begins to translate a uORF?
it will terminate with the stop codon and fall off the mRNA before reaching the main coding sequence → only will “scan through” and not necessarily reading it to selectively increase a few proteins during stress conditions such as amino acid starvation
eIF2 turns decreases global translation initiation that allows some ribosomes to “read through” uORFs to reach the main coding sequence
ATF4
transcription factor involved in responses to various stresses
what happens to atf4 under non stress conditions?
translation of atf4 is inhibited by uORFs
what happens to atf4 under stress conditions?
eIF2 gets phosphorylated which reduces initiation at uORFs (“scan through”)
internal ribosome entry site (IRES)
allow ribosomes to skip the first AUG codon by binding to an internal site → internally initiates translation
allows two different protein sequences to be derived from a single mRNA
may sit between two separate ORFs, allows independent simultaneous translation of two completely different proteins from one mRNA
different initiation sites may lead to skipping of a signal sequence (required for secreted/transmembrane proteins), and so switching between cytosolic and secreted forms of a protein