intro to tetrad analysis
1/69
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
70 Terms
Why so scientists use recombination frequencies?
They use it between genes to create linkage maps
A linkage map = relative locations of genes along a chromosome
When does recombination occur and what does distance mean?
Recombination occurs during prophase 1 of meiosis and the recombination frequency between 2 genes is proportional to the distance between the 2 genes
Farther apart two genes are on a chromosome = higher the RF for 2 genes
The closer two genes are on a chromosome = lower RF
Does RF reveal actual physical distance in nanometers between genes?
No
Need a physical map to be accurate
How is RF calculated in diploid organisms?

How are haploid RF calculated and whats the model organism used for it?
Calculating RF for haploid organisms - only the recombinant
genotypes.
• In haploid organisms, such as Neurospora:
• Used for tetrad analysis because it produces ordered tetrads
• Fungal asci sometimes include only the 4 haploid products of meiosis to produce a tetrad. Other times, the haploid cells produced by meiosis undergo one round of mitosis to produce an octad
Explain neurospora (fungi)
• 2 mating types (A & a)
• Sexual cycle is initiated by mixing A & a
• Haploid mycelium fuse and form 2n nuclei (only diploid stage A/a)
• Immediately undergo meiosis (4 haploid nuclei 2A and 2a)
• Meiosis followed by mitotic division and form 8 haploid ascospores (produced inside a sac “ascus”)
Whats tetrad analysis used for and hows it measured?
Can be used to calculate the gene-gene distance, and gene centromere distance
In octads ½ recombinant spores, ½ parental spores
In most eukaryotes, recombination analysis cannot be used to map the centromere, why?
Heterochromatic
Little heterozygosity in centromere region (little to no recombination)
‘However, linear tetrads in Neurospora can be used to map the centromere
Take into account crossing over between the centromere and the gene
Explain centromere mapping with linear tetrads
Normally, centromeres are visible microscopically, but cannot be mapped genetically.
However, centromeres can be mapped in species of fungi, that meet 2 conditions:
1. have monocentric centromeres (kinetochore co-localized with the centromere)
2. which retain the 4 cells produced by meiosis (called a tetrad) together in an ordered way within a physical structure called an ascus
What happens when there is crossing over and no crossing over?
Crossing over = 2:2:2:2 A:A:a:a
No crossing over = 4:4 A:a
Understand the following slide
Depending on how homologs line up on metaphase plate, it influences spore patterns (4 second division spore patterns)
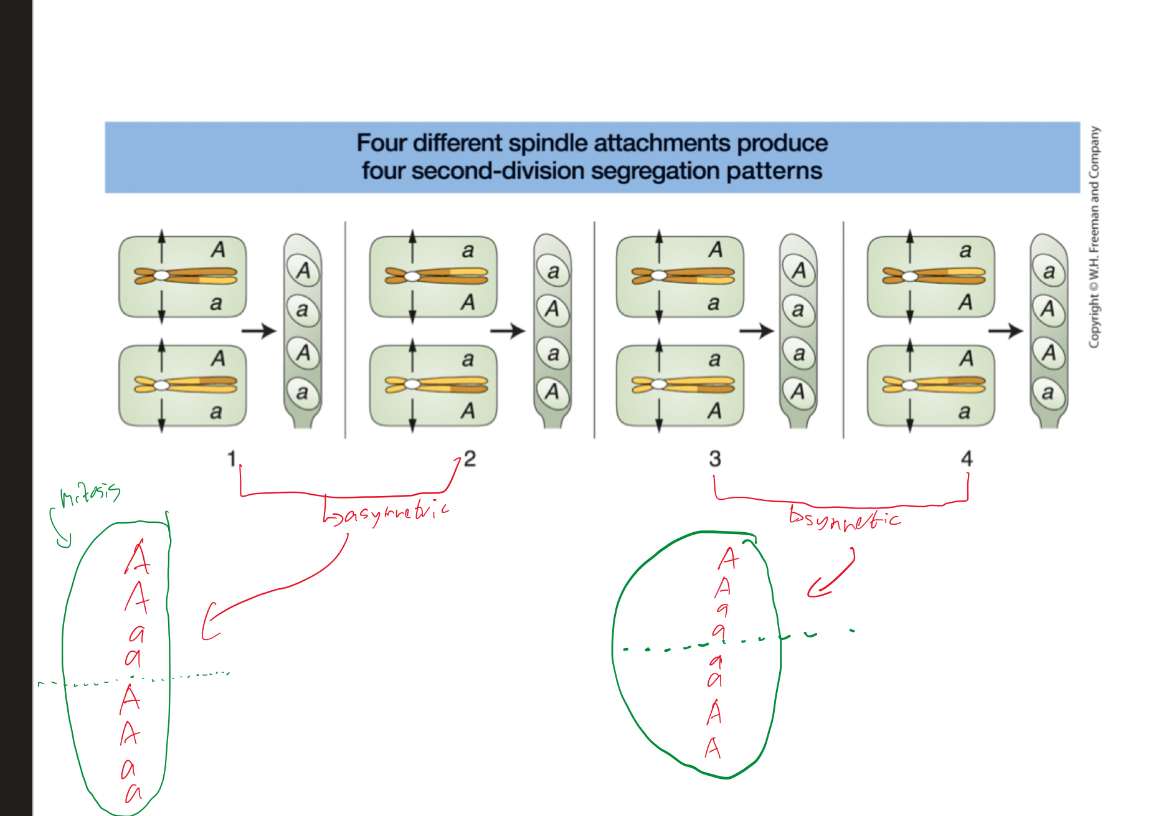
What occurs in divisions when using linear tetras and octads to map centromeres?
First division segregants are not recombinant (4:4)
Second division segregants, half are recombinants (2:2:2:2)
Explain and understand the 2 formulas used to calculate distance from centromere in 2nd division segregants
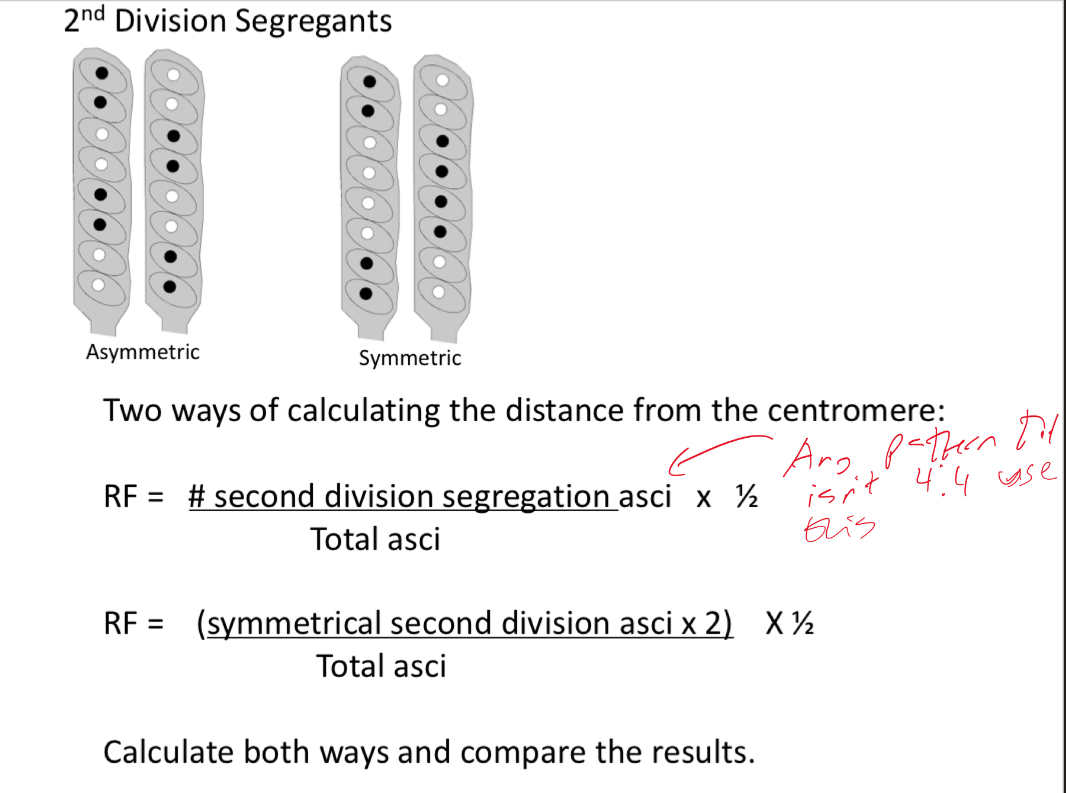
What’re the 3 possible outcomes for a cross in a haploid organism like neurospora and whats the equation for it (linked genes)?
Outcomes
Parental ditypes (PD)
Non-parental ditypes (NPD)
Tetra types (T)
When calculating RF for haploid organisms use only the recombinant genotypes
In haploids, the recombinant gametes, the nonparental ditypes (NPD), and half of the tetra types (T) are products of crossover during meiosis
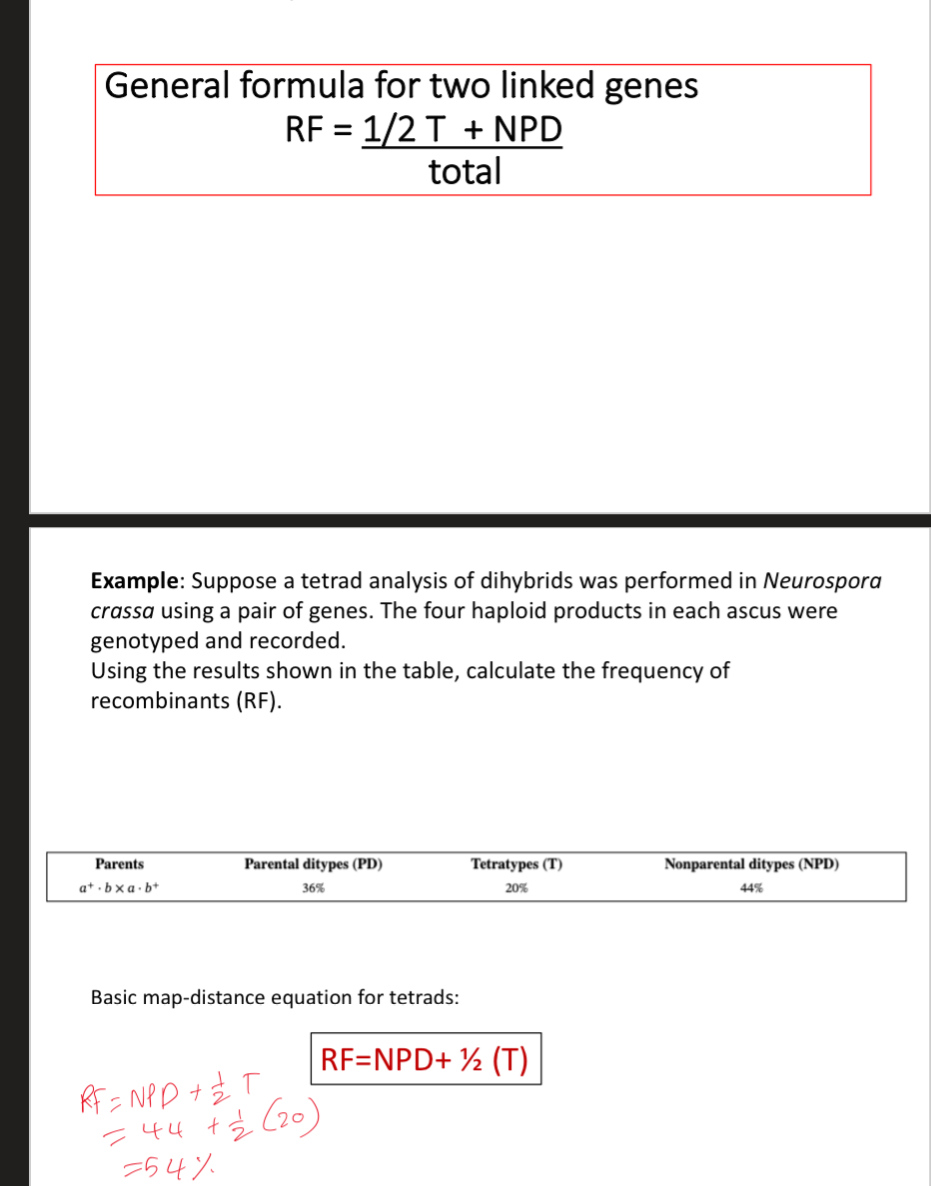

What gets produced in the different crosses in linked genes and whats most common?
NCO → PD
SCO → T (1/2 parental ½ recom)
DCO → NPD (all recombinant)
Understand how to figure these out
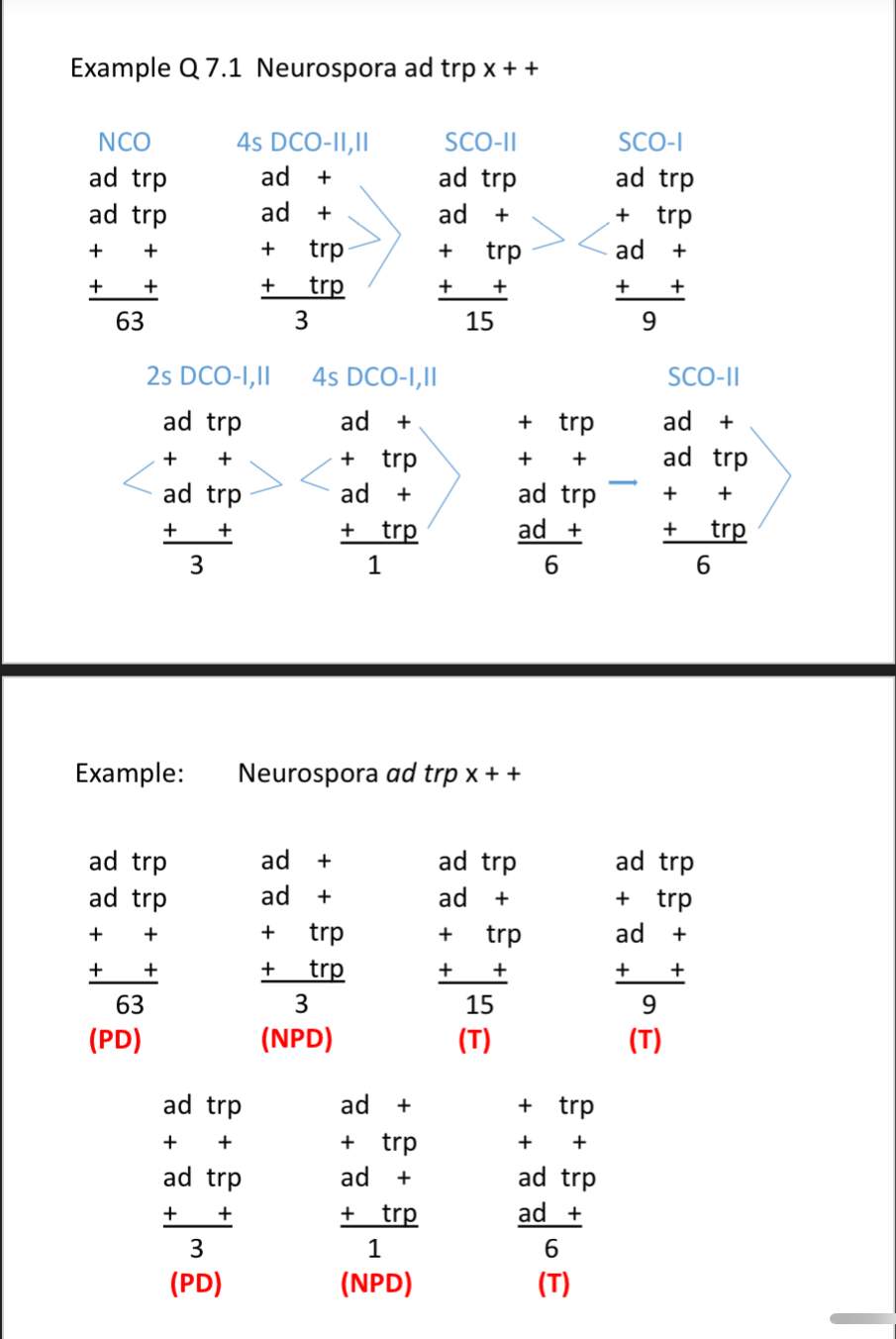

Look at all the equations and examples and know how to use them (the ones at end of slide pack 2 in ch 5)
Whats homologous recombination and whats the accepted model?
The exchange of genetic material between homologous DNA molecules
The accepted model is the double stranded break model (“holiday model”)
Where does homologous recombination happen in eukaryotes?
In prophase 1 of meiosis
Explain the double stranded break model
homologous recombination is initiated by controlled double stranded DNA breaks
Double stranded breaks that initiate recombination are not spontaneous, but are generated in a programmed manner by the activity of special enzymes
Current evidence indicates that meiotic recombination is initiated by double stranded breaks on the same/ one duplex
The heteroduplex DNA and a crossover are both produced by a double-stranded breaks in the DNA of one of the chromatids participating in the crossover
A crossover is initiated by a double stranded break in the DNA of a chromatid at meiosis. A series of molecular events ensures that eventually produces crossover DNA molecules
What’re the major steps of the double stranded break model and what happens in them?
Double stranded break
Both strands of the chromatid break in the same location
Erosion (degradation of 5’ end)
5’ ends of both strands (blue) are degraded and create 3’ single-stranded DNA regions (3’ overhang)
Invasion and displacement
One 3’ ends invades the DNA of the other duplex (enters the centre of the helix and base pairs with the homologous region) which displaces the other strand and forms the displacement (D) loop
Polymerization
Invading strand uses the adjacent strand as a template for new polymerization and DNA synthesis extends the invading 3’ end until 5’ end is reach and then ligation occur which results in one Holliday junction
Replicated ends are sealed so theres 2 single stranded junctions called Holliday junctions → complete double stranded crossover
Resolution
Will be discussed next slide but can be either opposite sense resolution or same sense resolution
Explain the differences between the two ways step 5 can occur
Opposite sense regulation
Involves cutting and rejoining of DNA strands in one of the Holliday junctions and cutting and rejoining of the two strands outside the second Holliday junctions
1 north-south cut and 1 east-west cut
The result with opposite sense resolution is heteroduplex DNA in both recombinant chromosomes
Opposite sense regulation is more common than same sense regulation
Same sense regulation
involves cutting and rejoining of the DNA strands in BOTH Holliday junctions (both cuts are east-west)
Genetic recombination doesnt take place, though heteroduplex regions remain flanking genes DO NOT RECOMBINE and the resulting chromosomes are NOT recombinant
Same sense resolution is rare
Explain the steps of the double stranded break model of meiotic recombination with the proteins used In yeast
According to this model, recombination is initiated by the protein Spo11 which is a nuclease that makes a double stranded break
It generates an asymmetric double stranded cut in one chromatid, and then the proteins Mrx and Exo1 associate with Spo11 and help trim the cut strands (Exo and Mrx binds to 5’ ends and starts degrading them)
The proteins Rad51 and Dmc1 which help facilitate strand invasion and D loop formation
The 2 strands that appear to cross over one another form a Holliday junctions; theres also a heteroduplex region
D loop formation and formation of the first heteroduplex. The 2 strands that appear to cross over one another form a Holliday junction, assisted by proteins Rad52 and Rad59
Strand extension further displaces D loop, pairs with top strand → form second heteroduplex
Top strand extended using D loop as template → ligated to free 5’ end
Ligation and formation of 2 Holliday junction
For Holliday junction resolution when connections between homologs get resolved before metaphase 1 what do bacteria and eukaryotes use?
Bacteria use RuvAB and RuvC
Eukaryotes use Rad51c-XRCC3
Whats is a crossover event initiated by?
Double stranded DNA break
LOOK AT ALL EXAMPLE QUESTIONS
Explain how gene conversion is directed Mismatch repair in heteroduplex DNA
Gene conversion is a process of DNA sequence change in which one allele is altered to another allele
Double stranded DNA is not completely complementary in heteroduplex region → can include nucleotide mismatches Therefore non complementary DNA is repaired by mismatch repair during meiosis 1
One strand will randomly act as template to correct the other strand
Gene conversion is a rare meiotic mismatch repair mechanism in heteroduplex region of heterozygous loci and is most readily observed in fungi that form as us and produce aberrant ratio of spores
Understand all 3 options of mismatch repair and what they produce
The pattern of mismatch repair determines the ratios in the eight cell ascus
Repair option 1 = if both strands are repaired using A1 as template it’ll produce a 6:2 ratio
Repair option 2 = if only one strand is repaired using A1 as template will produce a 5:3 ratio
Repair option 3 = no repair of either strand will produce an abberant 4:4 ratio in the form of 3:1:1:3
How are other patterns possible?
Other patterns are also possible since spindle orientation is random (spindle overlap)
Explain restriction enzymes
Exist in bacteria for their own protection (bacterial diffuse mech)
First identified in bacteria cells, where they function as protection against viral infection
They have DNA modification (like methylation) protects host DNA
Recognizes a specific DNA sequence at which it cuts both strands of the sugar phosphate backbone of DNA
The sequences recognized by restriction enzymes vary from 4 to 8 bp
Explain the cuts from restriction enzymes
The single stranded segments produced by some restriction enzyme cuts - sticky ends and can base pair with complementary sequences (EcoR1 recognizes palindromic sequence)
2 DNA molecules with complementary sticky ends can thus be combined by complementary base pairing
Sticky ends cause overhangs and are single stranded segments at the ends of each fragment
Some restriction enzymes leave blunt ends with no single stranded overhangs
Whats the probability of cut occuring in random sequence?

What is the number and size of fragments produced by digesting with a given restriction enzyme dependent on?
The size of the genome and the relative abundance of each nucleotide
Restriction digests with more than one enzyme, including double digests, help generate accurate maps
What does the southern blot quantify for and what’s the probe?
Quantifies DNA
Probe = DNA or RNA fragments
In vitro
What does the northern blot quantify for and what’s the probe?
Quantifies RNA
Probe = DNA or RNA fragments
In vitro
What does the PCR quantify for and what’s the probe?
Quantifies DNA
Probe = DNA primer
In vitro
What does the reverse transcription quantify for and what’s the probe?
Quantifies RNA
Probe = DNA primer
In vitro
What is PCR and what does it require?
PCR is an automated version of DNA replication in a test tube
PCR requires:
A double-stranded DNA template containing the target sequence to be amplified
A supply of the 4 DNA nucleotides
A heat-stable DNA polymerase → needs to be heat stable cuz PCR involves a heating process = the most common DNA polymerase Taq, is isolated from Thermus aquaticus which naturally occurs in hot springs
2 different single-stranded DNA primers which are short oligonucleotides and primer should be the complementary sequences to the DNA sequence flanking the area of interest to be amplified
A buffer solution
What’re the 3 general steps of PCR?
Denature (approx) 95C
Anneal 55C
Extend 72C
Whats the order of events of PCR?
The reaction mixture is heated to 95C to unwind DNA and break hydrogen bonds
The double stranded DNAs denatured to from single stranded DNA → need to separate strands otherwise primers wont anneal
The temperature of the reaction mixture is lowered to allow primers to anneal to the DNA template
The reaction mixture is heated to 72C (72C is ideal temp for taq polymerase)
Taq polymerase extends the primer
Why do we use Reverse transcription PCR and what is it?
Used cuz regular PCR doesnt work with RNA
Its used to see expression of gene cuz can be quantified by RNA
RNA needs to converted to its DNA copy = RNA gets converted to DNA via reverse transcriptase
Resulting DNA - complementary DNA (cDNA)
The rest of the amplification steps are the same
In real-time PCR/RT PCR, a probe is added during amplification which can give off fluorescence when a new cDNA molecule is formed which helps quantify gene expression/ RNA
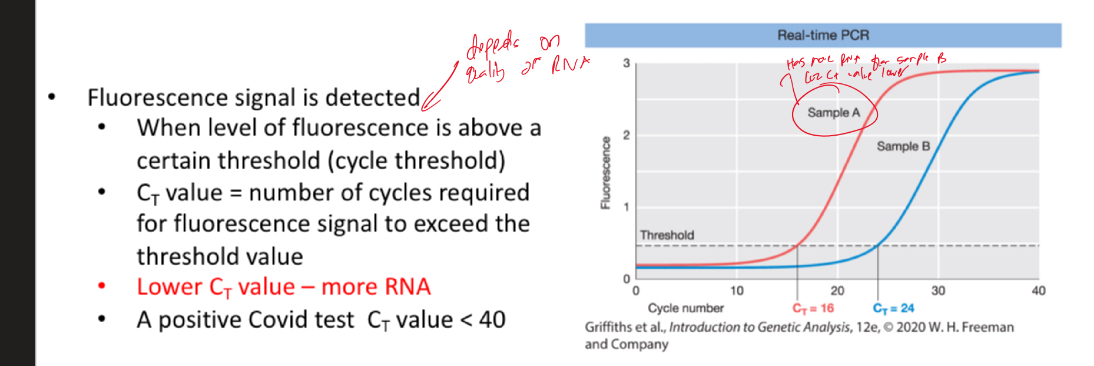
Explain and understand this
Lower Ct value - more RNA
Explain northern blotting
Detects a specific RNA
RNA is extracted from cells and purified and loaded onto a gel that separates the RNA transcripts according to their size
RNA from the gel are then blotted onto a nylon membrane and probed with a labeled fragment of DNA
RNAs that are complementary to the DNA probe are detected as labeled bands
Explain southern blotting
it detects a specific DNA
Has an extra step where gel us soaked in NaOH to denature the double-stranded DNA into single-stranded DNA prior to the membrane transfer step
Probes are radioactive (32P-labeled) single stranded nucleic acids
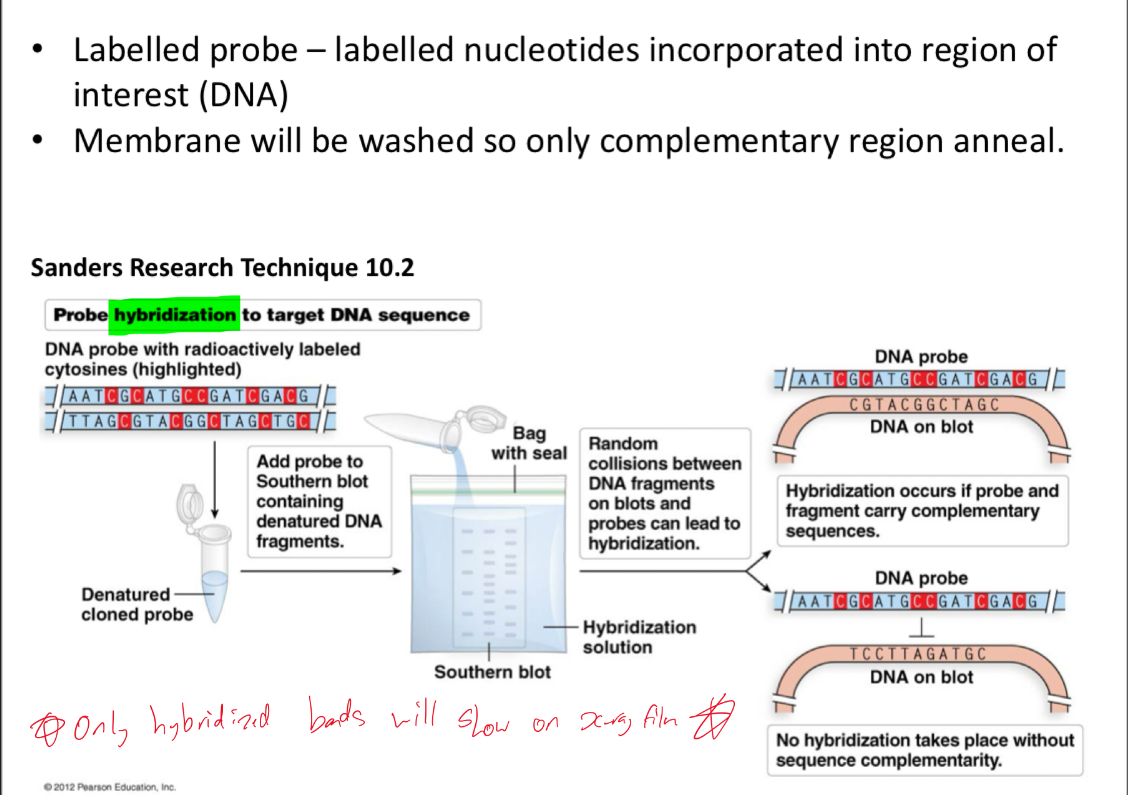
Explain and understand this
Explain mapping with molecular markers
Molecular markers allow direct detection of differences between individuals
Take into account the variations in DNA polymorphisms - genetic DNA markers
Genetic markers are typically in noncoding regions of the genome
Advantage: inherited in Mendelian fashion
Not subject to environmental influences, pleiotropy, epistasis, penetrance and expressivity
What’re the 2 types of markers and what are their subgroups?
SNPs = RFLP, ASO
Repetitive sequences = STRs, VNTRs
Explain SNPs
~3.3 million SNPs in human genome
They involve variants where one base pair is substituted by another base pair
They typically occur in a noncoding region (no detectable effect on genotype)
However, sometime they occur in expressed regions of genes (variation can affect phenotype)
Ex: APoE is in an exon
Whatre some applications of SNPs?
– gene mapping
– crime scene DNA analysis
– paternity testing
How are SNP variations identified by?
Genome sequencing
Short restriction sequences
Explain restriction fragment length polymorphisms (RFLP)
DNA molecules with multiple restriction sequences are cut with a restriction enzyme - many DNA fragments are produced- these fragments are characteristic of a given DNA molecule
Inherited variability in the number or length of restriction fragments is called restriction fragment length polymorphism (RFLP)
When a SNP variation affects a restriction site, the length of restriction fragments (measured in kilobases/kb) may also change (1kb = 1000 nucleotide bases)
Separated fragments are transferred to membrane and radiolabelled probes are added to identify the fragments of our interest
Look at markers in notes
Explain an example of an application of RFLP in detecting a disease
Mutation in hemoglobin gene – Sickle Cell Disease (SCD)
– The single-nucleotide DNA difference is the result of a SNP that alters the sixth DNA triplet of the coding sequence
– Result: Glutamic acid in the normal protein is replaced with valine
LOOK AT EXAMPLE IN NOTES
Explain southern blot analysis of beta-globin gene variation
• SCD mutation destroys a restriction sequence, leading to an RFLP that can be detected by Southern blot analysis
• The b-globin gene
– WT allele - three restriction sites for the restriction enzyme DdeI
– SCD allele – two restriction sites for DdeI
Look at examples on slide
Understand the slide of RFLP in paternity testing
Whatre some advantages of RFLP?
• Inherited as codominant markers – can distinguish heterozygote from homozygote
• Probes are sequence specific – can target specific DNA locus
• Restriction sites are distributed throughout the genome in abundance
• Provide reproducible results
Whatre some limitations of RFLP markers?
• High quality DNA and large amounts of DNA required
• Use radiolabelled probes
• Molecular markers are often functionally neutral (do not result in phenotype)
• Also functionally important mutations rarely alter restriction sites
Where would functionally neutral SNPs be and provide examples
Functionally neutral SNPs closely linked to gene/ phenotype of interest
• Eg1 : Dominant Breast cancer allele (BRCA2) in unknown gene
• Eg 2: RFLP marker in flanking gene used to test for PKU mutants
– Presence of marker indicates the inheritance of disease allele
– Marker must be closely linked to reduce the chances of recombination
Look at drawn example slide on notes
Whatre some characteristics of markers based on repetitive sequences?
• Higher level of change in non-coding regions
• Very abundant, highly polymorphic
• Inherited in a codominant manner
Whatre the 2 types of markers based on repetitive sequences?
1. Microsatellites or STR (Short Tandem Repeats)/SSR (Simple Sequence Repeats)
2. Minisatellites or VNTRs (Variable Number Tandem Repeats)
Explain STR markers
• STR markers are polymorphic DNA loci that contain a repeated nucleotide sequence.
– STR repeat unit - 2-6 nucleotides in length
• The number of repeat units at an STR locus may differ, so alleles of many different lengths are possible.
• Polymorphic STR loci are therefore very useful for human identification purposes.
• Eg: Trinucleotide repeats in fragile X, Huntington's disease
• STR loci can be amplified using PCR process and the PCR products are then analyzed by electrophoresis to separate the alleles according to size.
• PCR amplified STR alleles can be detected using various methods (fluorescent dye labelling)
Explain VNTRs
• Fewer than STRs but longer than STRs
• VNTRs consist of short DNA sequences; usually 7-30 nucleotide repeats
• Different chromosomes can carry different repeat numbers of the sequence
• Can be detected via PCR or Southern blot
– PCR analysis based on the number of repeats within amplified fragment
– Southern blot analysis based on cuts at flanking regions resulting in fragments of different lengths depending on the number of repeats
Explain the variable nimber of repeat sequence blocks that VNTRs contains
(a) A chromosome pair in which one homolog has 6 repeats and the other has 10 repeats.
(b) VNTRs are inherited in a codominant manner.
(c) Single nucleotide polymorphisms (SNPs) are single base-pair sequence variants, also inherited in a codominant manner
Whatre some advantages of STRs and VNTRs?
• Highly polymorphic
• Highly abundant and randomly dispersed across the genome
• High throughput and can be automated
Explain absence and presence of DNA markers
Absence of marker excludes suspect
However presence of marker doesnt prove guilt, it just simply fails to exclude
Probability of marker being contributed by someone else depends on allele frequency in population
Understand example of CODIS
Explain RAPD
They don’t require any specific knowledge of the DNA sequence of the target → we use this when we don’t know target sequence
Advantage is that can be used when sequence of locu of interest not available → need only low amount of DNA, inexpensive as well
Use short (about 10 nucleotides) random primers that can bind to many sites in genome
Look at example of RAPDs
How to find RAPD markers?
Screen RAPD primers-monitor-co segregation of RAPD bands with a particular trait
Allows genotyping by molecular rather than phenotypic analysis eg: plant breeding
Works best when a single gene trait and true breeding parents available
What’re the limitations of RAPD?
