Midterm
0.0(0)
Studied by 3 peopleCard Sorting
1/84
Earn XP
Description and Tags
Last updated 5:10 PM on 10/24/22
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
85 Terms
1
New cards
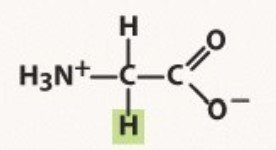
Glycine
Non-polar
2
New cards
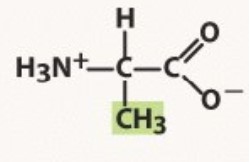
Alanine
Non-polar
3
New cards
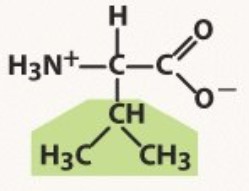
Valine
Non-polar
-shaped like a v
-shaped like a v
4
New cards
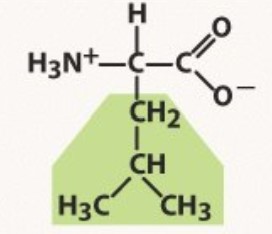
Leucine
Non-polar
-Longer than valine
-Longer than valine
5
New cards
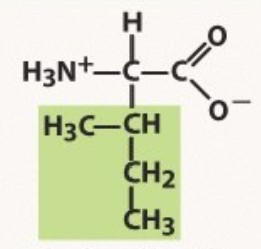
Isoleucine
Non-polar
-ISOmer of LEUCINE
-sorta leucine
-ISOmer of LEUCINE
-sorta leucine
6
New cards
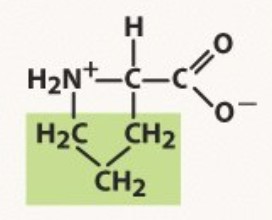
Proline
Non-polar
positive NH2+ because of looping
-PRO amino acid for pro chemists
positive NH2+ because of looping
-PRO amino acid for pro chemists
7
New cards
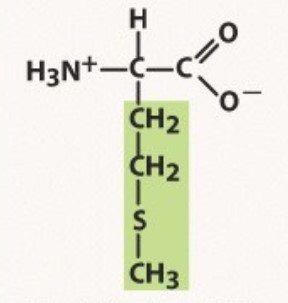
Methionine
Non-polar
start codon
-3 nulceotides in a codon
start codon
-3 nulceotides in a codon
8
New cards
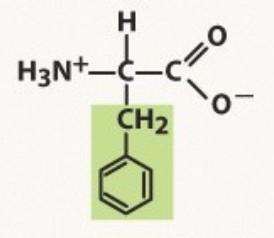
Phenylalanine
Aromatic
-phenyl is a hydrocarbon ring
-phenyl is a hydrocarbon ring
9
New cards
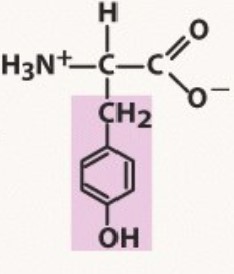
Tyrosine
Aromatic
pKa 10.1
-TYRe with an O following
pKa 10.1
-TYRe with an O following
10
New cards
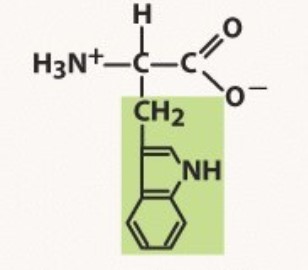
Tryptophan
Aromatic
-you must TRY to remember the largest structure
-you must TRY to remember the largest structure
11
New cards
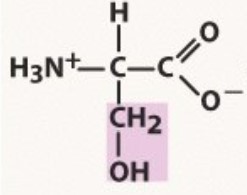
Serine
Polar uncharged
pKa 13.6
-OH no
pKa 13.6
-OH no
12
New cards
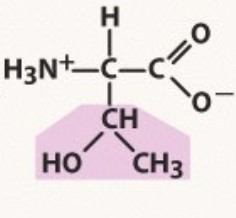
Threonine
Polar uncharged
pKa 13.6
-OH no
pKa 13.6
-OH no
13
New cards
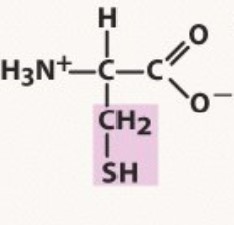
Cysteine
Polar uncharged
pKa 8.3
can create disulphide bonds
-Sistine chapel (two fingers "bonding)
pKa 8.3
can create disulphide bonds
-Sistine chapel (two fingers "bonding)
14
New cards
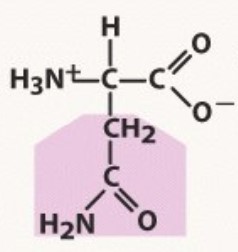
Asparagine
Polar uncharged
-A before G
-A before G
15
New cards
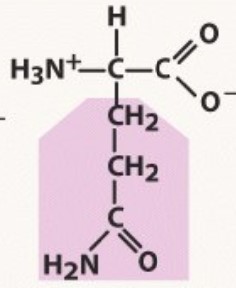
Glutamine
Polar uncharged
-A before G
-A before G
16
New cards
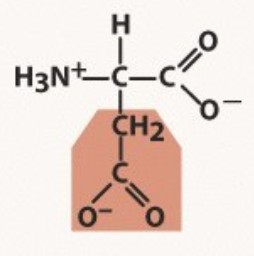
Aspartate
Polar negative
pKa 3.9
-A before G
pKa 3.9
-A before G
17
New cards
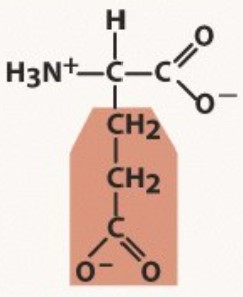
Glutamate
Polar negative
pKa 4.2
-A before G
pKa 4.2
-A before G
18
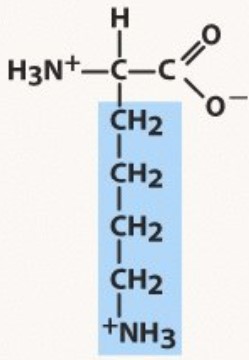
New cards
Lysine
Polar positive
pKa 10.5 (HA)
-LY is a line
pKa 10.5 (HA)
-LY is a line
19
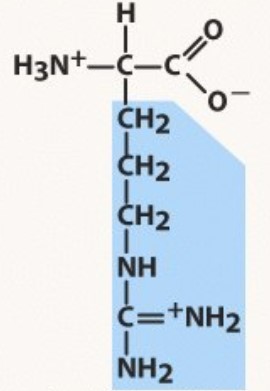
New cards
Arginine
Polar positive
pKa 12.5 (HA)
pKa 12.5 (HA)
20
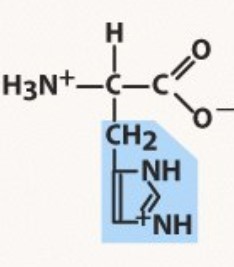
New cards
Histidine
Polar positive
(A-)
(A-)
21
New cards
Main Biomolecules
Proteins
Lipids
Carbohydrates
Nucleic Acids
Lipids
Carbohydrates
Nucleic Acids
22
New cards
Biochemistry
principally concerned with the chemistry and molecular interactions of biological molecules associated with living systems
23
New cards
Enzymes
Proteins
catalyze biochemical reactions
-composed of amino acids
catalyze biochemical reactions
-composed of amino acids
24
New cards
Metabolism
Catabolism -yield energy, degrade
Anabolism -take energy, assemble
25
New cards
Electronegativity
ability to attract electrons
H, P < C, S < N < O
-bond strength
26
New cards
Biochemical transfromation
1. Group transfer, add or remove
2. Oxidation-Reduction
3. Rearrangement, function changes
4. Cleavage, break, add water
5. Condensation, form, remove water
27
New cards
Saturated hydrocarbons
C-H, C-C
non-polar
non-reactive
non-polar
non-reactive
28
New cards
Structure of liquid water
H-bonds are continually broken and reformed
29
New cards
Hydrogen bonding
electrostatic interaction between
O-H, N-H, F-H
explains why polar molecules can dissolve in water
strongest -linear
breaking -requires enthalpy
O-H, N-H, F-H
explains why polar molecules can dissolve in water
strongest -linear
breaking -requires enthalpy
30
New cards
Electrostatic interactions
between oppositely charged ions
TdS >> dH
TdS >> dH
31
New cards
van der Waals interactions
short range, very weak interaction
induced dipole attraction
induced dipole attraction
32
New cards
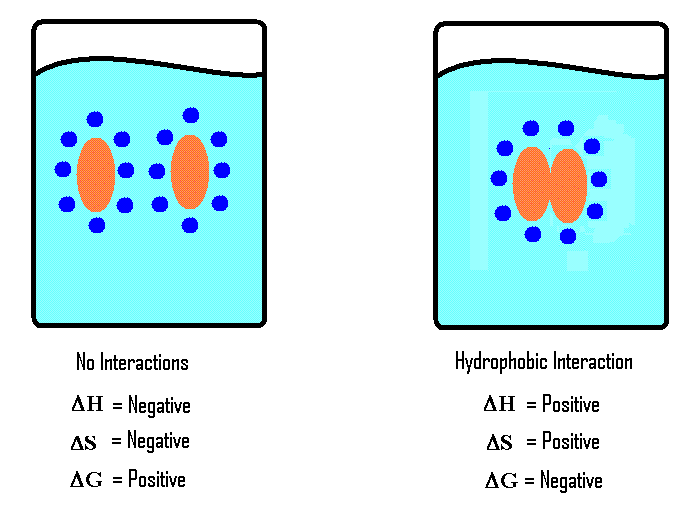
Hydrophobic Effect
non-polar hydrocarbon dissolves in water:
broken -van der Waals and H bonds
formed -H bonds in a cage
entropy is reduced
broken -van der Waals and H bonds
formed -H bonds in a cage
entropy is reduced
33
New cards
Amphipathic
polar and non-polar groups
-detergents, lipids, proteins
lowest free energy state is in hydrophobic clusters
-detergents, lipids, proteins
lowest free energy state is in hydrophobic clusters
34
New cards
Detergents
-sodium dodecyl sulfate
micelle -hydrophobic core
micelle -hydrophobic core
35
New cards
Ionization of water equations
pH = -log[H+]
pOH = -log[OH-]
pOH = -log[OH-]
36
New cards
Strong acid
complete dissociation
HA -> H+ + A-
HA -> H+ + A-
37
New cards
Strong base
complete dissociation
BOH -> B+ + OH-
BOH -> B+ + OH-
38
New cards
Weak acid
incomplete dissociation
proton donors
proton donors
39
New cards
Weak base (Conjugate base)
incomplete dissociation
proton acceptor
proton acceptor
40
New cards
Weak acid equations
Ka = [H+][A-]/[HA]
pKa = -log[Ka]
pKa = -log[Ka]
41
New cards
Buffer questions
1. M1/C1 = M2/C2
2. [A-] = x, [HA] = M2 - x
3. pH = pKa + log[A-]/[HA]
4. concentrations
5. A- & HA +/- change
6. pH
2. [A-] = x, [HA] = M2 - x
3. pH = pKa + log[A-]/[HA]
4. concentrations
5. A- & HA +/- change
6. pH
42
New cards
Henderson-Hassalbalch equation
pH = pKa + log[A-]/[HA
43
New cards
Non-polar AA
Gly, G
Ala, A
Val, V
Leu, L
Ile, I
Met, M
Pro, P
Phe, F
Trp, W
Ala, A
Val, V
Leu, L
Ile, I
Met, M
Pro, P
Phe, F
Trp, W
44
New cards
Polar Uncharged AA
Ser, S
Thr, T
Asn, A
Gln, Q
Tyr, Y
Cyc, C
His, H *
Thr, T
Asn, A
Gln, Q
Tyr, Y
Cyc, C
His, H *
45
New cards
Polar + AA
anionic
Arg, R
Lys, K
His, H *
Arg, R
Lys, K
His, H *
46
New cards
Polar - AA
Glu, E
Asp, D
Asp, D
47
New cards
pKa of AA
a - COOH 2.2
Aspartate 3.9
Glutamate 4.2
Histidine 6.0
Cysteine 8.3
a - NH3+ 9.5
Tyrosine 10.1
Lysine 10.5
Arginine 12.5
Serine 13.6
Threonine 13.6
Aspartate 3.9
Glutamate 4.2
Histidine 6.0
Cysteine 8.3
a - NH3+ 9.5
Tyrosine 10.1
Lysine 10.5
Arginine 12.5
Serine 13.6
Threonine 13.6
48
New cards
Sterochemistry
conformation -rotation
configuration -breaking/forming
configuration -breaking/forming
49
New cards
Isomers
same formula different arrangement
stereoisomer -same formula, different faces (up vs down)
stereoisomer -same formula, different faces (up vs down)
50
New cards
Chirality
non-superimposable mirror images
-enantiomers D or L
AA are mostly L
-enantiomers D or L
AA are mostly L
51
New cards
Zwitterion
hybrid ion
-no net charge
-no net charge
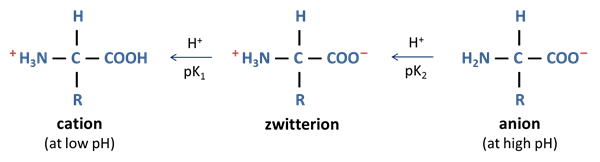
52
New cards
Titration of glycine
pKa 1 = 2.4 (COOH)
pI = 6
pKa 2 = 9.6 (NH3+)
pI = 6
pKa 2 = 9.6 (NH3+)
53
New cards
Isoelectric point
pI = pKa 1 + pKa/2
54
New cards
Calculating pH
1. determine form (A >6, B
55
New cards
Titration of AA
1. protein mols
2. NaOH or HCL mols (x)
3. determine form and direction (pH or pI)
4. pH = pKa + log (A-/HA)
2. NaOH or HCL mols (x)
3. determine form and direction (pH or pI)
4. pH = pKa + log (A-/HA)
56
New cards
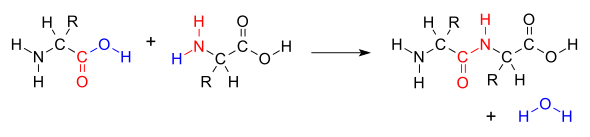
Peptide bond
amide bond that joins two amino acids
57
New cards
residue
amino acid units in a protein
58
New cards
Amino acid sequence
N-C (NH3+ to COO-)
order of AA
order of AA
59
New cards
Sanger protein sequencing
1. hydrolysis of polypeptide into small bits
2. separating bits
3. arranging bits to find the sequence
2. separating bits
3. arranging bits to find the sequence
60
New cards
Fragmentation
Trypsin - Lys, Arg (+) KR
Chymotrypsin -Phe, Tyr, Trp (aromatic) FYW
CNBr - Met (start) M
Chymotrypsin -Phe, Tyr, Trp (aromatic) FYW
CNBr - Met (start) M
61
New cards
Secondary structure
H-bonds between C=O & H=N
a-helix
-right handed spiral (1 turn is 3.6 residues)
b-sheet
a-helix
-right handed spiral (1 turn is 3.6 residues)
b-sheet
62
New cards
a-Helix
peptide is planner (cis or tans)
H bond between every 4th aa
disruptions:
-electrostatic repulsion
-bulky R groups
-small R groups (favour b)
-proline
H bond between every 4th aa
disruptions:
-electrostatic repulsion
-bulky R groups
-small R groups (favour b)
-proline
63
New cards
b-Conformation
peptide is planner (cis or trans)
H bond between adjacent strands
-antiparallel 7
-parallel 6.5
small R groups
H bond between adjacent strands
-antiparallel 7
-parallel 6.5
small R groups
64
New cards
Tertiary structure
globular form
-hydrophobic core (hydrophobic effect)
stablizing:
-disulfide
-electrostatic/ionic
-h bonds
-metal chelation
-hydrophobic core (hydrophobic effect)
stablizing:
-disulfide
-electrostatic/ionic
-h bonds
-metal chelation
65
New cards
Disulfide bonds
crosslink protein with covalent bonds
Cys
Cys
66
New cards
Metal chelation
salt bridge to stabilize charges
N(+) |||| Me++ |||| O-
N(+) |||| Me++ |||| O-
67
New cards
Quaternary structure
interaction of globular polypeptides
-same factors as 3
-same factors as 3
68
New cards
Protein folding
dependent on aa sequence
69
New cards
Protein denaturation
native protein converted to unfolded
1. heat
2, change in solvent
-few proteins can reverse spontaneously (need chaperons)
1. heat
2, change in solvent
-few proteins can reverse spontaneously (need chaperons)
70
New cards
Prostheic groups
attached non-amino acids
cofactor
-organic (coenzymes)
-inorganic
cofactor
-organic (coenzymes)
-inorganic
71
New cards
Ion exchange chromatography
protein purification
1. beads with sulfonic acid (anionic)
2. AA in buffer at 2
3. AA washed with higher pH
4. pH is pI of AA it will wash away
-salt can remove AA (compete for binding sites)
1. beads with sulfonic acid (anionic)
2. AA in buffer at 2
3. AA washed with higher pH
4. pH is pI of AA it will wash away
-salt can remove AA (compete for binding sites)
72
New cards
Size-Exclusion (Gel Sieving) chromatography
1. bead with pores
2. mixture of proteins
Big proteins -quick, can't go through pores
Small proteins -slow, go through beads
2. mixture of proteins
Big proteins -quick, can't go through pores
Small proteins -slow, go through beads
73
New cards
Affinity Chromatography
1. beads with ligand
2. mixture of proteins
3. hexokinase binds, others washed
4. ATP added (takes binging site) to unbind hexokinase
2. mixture of proteins
3. hexokinase binds, others washed
4. ATP added (takes binging site) to unbind hexokinase
74
New cards
SDS-PAGE
1. crosslinked polyacrylamide gel
2. detergent binds to proteins
3. electrical potential moves proteins
4. staine
2. detergent binds to proteins
3. electrical potential moves proteins
4. staine
75
New cards
Catalyst
speeds up the rate without being consumed
S + E -> E.S -> E.S# -> E.P -> E + P
S + E -> E.S -> E.S# -> E.P -> E + P
76
New cards
Apoenzyme/Apoprotein
protein
77
New cards
Holoenzyme
protein + coenzyme
78
New cards
Enzyme substrate complex
E.S or E.P
non-covalent complex
non-covalent complex
79
New cards
Transition state theory
reaction rate = (constant)Te^-dG/RT
dG is out activation energy S#
dG is out activation energy S#
80
New cards
Enzyme lowering the S#
formation of E.S is spontaneous
-binding energy used to fund
-weak but favorable interactions
-binding energy used to fund
-weak but favorable interactions
81
New cards
E.S formation
decrease of entropy
desolvation -removing water cage
induced fit
alignment of reaction groups
desolvation -removing water cage
induced fit
alignment of reaction groups
82
New cards
Recognizing substrates
shape, consistency, or fit -lock and key
electrostatic consistency -correct bonds within site
thermodynamic consistency -flexing to fit
electrostatic consistency -correct bonds within site
thermodynamic consistency -flexing to fit
83
New cards
Binding energy is used for
entropy reduction -hold substrates
desolvation -cage must be removed to react
strain reduction -cushion that uncomfortably
desolvation -cage must be removed to react
strain reduction -cushion that uncomfortably
84
New cards
Specificity
Chiral specificity -D vs L
Geometric specificity -trans vs cis
Geometric specificity -trans vs cis
85
New cards
Chymotrypsin
Trp, Tyr, Phe
1. substrate enters active site
2. His N: -> H of S OH -> C of substrate C=O -> O of C=O
3. H to N+ -> C-HN of substrate to H -> - to bond
-tetrahedral intermediate
4. N of His -> H of water -> C=O -> O
5. O- -> bond O-C -> H -> N+
-tetrahedral intermediate
1. substrate enters active site
2. His N: -> H of S OH -> C of substrate C=O -> O of C=O
3. H to N+ -> C-HN of substrate to H -> - to bond
-tetrahedral intermediate
4. N of His -> H of water -> C=O -> O
5. O- -> bond O-C -> H -> N+
-tetrahedral intermediate