genetics save me
1/71
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
72 Terms
define genetics
the study of heredity, and how genes are encoded, replicated, expressed, and evolved.
epistasis
a gene (epistatic) masks/inhibits the phenotype of another gene (hypostatic)
polygenic inheritance
multiple genes affect a single trait
Labrador retriever coat color involves two genes:
• 𝐵 allows black pigment (bb = brown pigment).
• 𝐸 allows pigment deposition in fur (ee prevents deposition → yellow coat).
Consider the cross 𝐵𝑏𝐸𝑒 × 𝐵𝑏𝐸𝑒.
• (a) What phenotypic ratio (black : brown : yellow) is expected in the
𝐹2?
• (b) Which genotype class demonstrates epistasis, and which gene is
epistatic?
a) 9:3:4
b) ee shows epistasis, E locus is epistatic
Holliday model
Double-strand break model
Theta replication
Rolling replication
inversion loops
when a chromosome fragment is flipped 180
paracentric does NOT include the centromere. results in a dicentric chromatid (2 centromeres) and an acentric chromatid (no centromere). usually nonviable.
pericentric includes the centromere, goes around it. results in 2 nonviable recombinant gametes, nonrecombinant with pericentric inversion, and a normal nonrecombinant gamete.
chromosome/DNA counting
DNA replication doubles chromatids, not chromosome number, until they split
What is a Barr body, and how does X‑chromosome inactivation create
mosaicism in heterozygous females?
Inactivated X chromosomes. If females have two X’s, they will delete one (human dosage compensation). If X has different genomes, it will make a mosaic across cells (think calico cats)
complete dominance
one allele completely masks the other
incomplete dominance
neither allele is completely dominant, blend
codominance
both alleles are fully expressed and neither masks the other, patches
lethal alleles
cause death before birth usually
In the human ABO system, the relationship between 𝐼𝐴 and 𝐼𝐵 alleles is:
A. Complete dominance (𝐼𝐴 > 𝐼𝐵) B. Complete dominance (𝐼𝐵 > 𝐼𝐴) C.
Codominance (𝐼𝐴 = 𝐼𝐵) D. Incomplete dominance
C. codominance 𝐼𝐴 = 𝐼𝐵
transition mutation
purine to purine, or pyrimidine to pyrimidine
transversion mutation
purine to pyrimidine, or pyrimidine to purine
insertions/deletions
cause frameshift mutations if less than 3 bases are inserted/deleted
forward mutation
wild type gene to mutant gene
reverse mutation
mutant gene to wild type gene
missense
new codon → new amino acid
nonsense
new codon → STOP codon
silent
new codon → no change in amino acid
tautomeric shifts
hydrogen atoms switch across base
strand slippage
newly synthesized strand loops out and causes addition
template strand loops out and causes deletion
unequal crossing over
homologous chromosomes misalign during crossing over, one product has an insertion and the other has a deletion
depurination
loss of a purine base from a nucleotide leads to base substitution, will often incorporate A into the empty spot
incorporated error
an initial wrong nucleotide placed opposite of the template from mispairing
replicated error
a later, fixed mutation of an incorporated error that survives repair and acts as a template
Which statement correctly compares ultraviolet and ionizing radi-
ation?
A. UV commonly causes pyrimidine dimers, whereas ionizing radiation
can generate free radicals and double-strand breaks.
B. UV mainly causes double-strand breaks, whereas ionizing radiation
mainly causes pyrimidine dimers.
C. Both types of radiation are mutagenic only because they increase tran-
scription.
D. Neither type of radiation changes DNA directly.
A
repair pathway for pyrimidine dimer
nucleotide excision repair, removes bulk and fills with polymerase + ligase
nucleotide excision repair
strands are separated and section of DNA with the bulky lesion is removed, polymerase fills in gap and ligase seals it
repair pathway for O^6 methyl guanine
direct repair restores it to original guanine structure
direct repair
does not replace mistake, but fixes it to its original form
repair pathway for uracil from cytosine deamination
base excision repair removed the damaged base using glycosylase
base excision repair
a modified base is removed, and the whole nucleotide is replaced
repair pathway for mismatch after replication
mismatch repair corrects the mismatched base that escaped proofreading
mismatch repair
corrects incorrectly inserted nucleotides that escape proofreading by DNA polymerase during replication
transposable element
DNA sequences that move in the genome and often cause mutations
flanking direct repeat
staggered cut is made into DNA, transposable element inserts itself but there are gaps. polymerase fills in the gaps to make the repeats
replicative transposition
copy/paste
nonreplicative transposition
cut/paste
retrotransposons
transpose through an mRNA intermediate and copied back into DNA (reverse transcription)
restriction enzymes
recognize specific nucleotide strands in DNA and make double-stranded cuts at the sequences. makes sticky/blunt ends that can combine fragments
CRISPR-Cas9
modifies genomes
Cas 9 and sgRNA combine to make an effector complex
this associates with PAM sequence (target) and unwinds the DNA
pairs with complementary sequence
Cas protein cleaves DNA
nonhomologous end joining (NHEJ) or homology directed repair (HDR)
nonhomologous end joining (NHEJ)
CRISPR repair, joins ends and introduces duplications or deletions that cause frameshifts
homology directed repair (HDR)
CRISPR repair, repairs by inserting donor DNA
western blot
separates proteins by mass using anitbodies
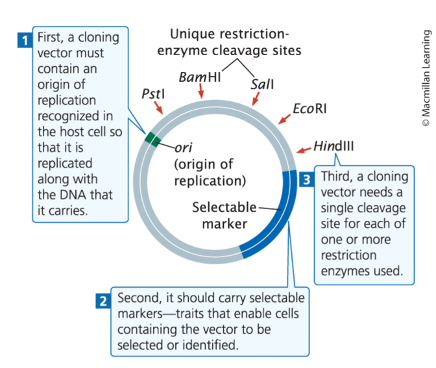
cloning vectors
bacteria, have an origin of replication, 1+ selectable markers, and recognition sites for 1+ restriction enzymes
plasmid vectors
ecoRI cuts plasmid to insert a foreign DNA segment, sticky ends join
expression vector
has an operon sequence that allows the DNA to be transcribed and translated, can turn on/off the sequenced gene
LacZ gene
screens for bacteria with recombinant plasmids. Foreign DNA is inserted into the partial lacZ gene.
nonrecombinant OG plasmids: blue, intact lacZ
recombinant: white, disrupted lacZ
no plasmid: not grown
Sanger sequencing
reads DNA sequences
sequences have dNTPs added on, but there’s no 3’OH
fragments terminated by ddNTPs
reads fragments from shortest to longest
PCR
DNA amplification
requires: template DNA, primers (3’OH end), thermostable polymerase, dNTPs, buffer
denaturation separates strands
annealing lets primers bind to complementary sites
extension copies from primers and grows
grows 2^n from the starting molecule
repeat
Illumina sequencing
accurate short reads
fluorescently tagged dNTPs are added onto the primer
tag is excised with a labor and fluoresces.
read by a computer
tag/reversible terminator removed, and repeated
PacBio HiFi
long reads with high accuracy
emits light
Nanopore
very long reads, portable field use
electrical current
GWAS
tests markers (SNP) to find associations with a trait, can detect diseases caused by complex interactions
SNP can be causal or tag a nearby causal variant
linkage disequilibrium
nearby variants are likely inherited together often by chance
RNA sequencing
can detect sequences without prior probe designs, way more useful, reads expressed RNAs
makes cDNA from mRNA, broken into fragments, amplification adaptors are added and undergo PCR. assembled into RNA transcripts
DNA microarray
hybridization to predesigned probes
orthologs
genes in different species descended from a common ancestral gene.
paralogs
related genes produced by duplication within a lineage
synteny
neighboring genes retained in the same order across genomes.
C-value paradox
genome size does not scale with organism complexity
map based sequencing
detailed genetic and physical maps to align sequenced fragments (not really used)
shotgun sequencing
uses sequencing overlap to align sequenced fragments, overlap creates clones
X-linked recessive inheritance
sons receive X from mother and Y from father
daughters recieve X from both
affected father passes mutant X to all daughters and Y to all sons
linked genes
largest classes are parentals
coupling: dominant alleles together
repulsion: dominant alleles on opposite homologs
<50%
meiotic nondisjunction
failure of sister chromatids or homologous chromosomes separating before fertilization, zygote starts with abnormal chromosome number
mitotic nondisjunction
failure of sister chromatids or homologous chromosomes separating after fertilization, only descendent cells carry change
affect larger fraction of the body