MCB 150 Exam 3 Review
1/77
Earn XP
Description and Tags
Concepts/topics taken from MCB 150 notes and some outside resources
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
78 Terms
Mutations
Heritable changes in the DNA are called mutations
Somatic mutation
Non-inherited DNA changes occurring in body cells after conception, affecting only the individual (not offspring(non sex cell)
Germline mutation
Changes in DNA that is inherited to daughter cell from parent cell
Mutation terminology
Gene- means mutated
Gene+ means normal (or wild type)
Causes of mutations
1.Uncorrected mistakes in DNA replication (accounts for the most mutations)
2.Chemical mutagens
3. High-intensity radiation (x-rays, UV, etc)
Hemimethylated DNA
When one strand does have a methyl group covalently attached to adenine while the other stand does not
Methyl transferase
An enzyme that looks for non-methylated GATC sequences in daughter strands
Ways of fixing (or reducing) mutations
1.Proofreading (in E. Coli) by DNA Polymerase
-An example of 3’ to 5’ exonuclease activity
2.Mismatch Repair system (MMR)
-Scans for mistakes and hemimethylated DNA in recently synthesized DNA
-Mismatch repair enzymes fix problem in unmethylated (new) strand
Impact of proofreading/MMR
Each decrease error frequency by 100x
100 × 100 = 10,000, which is the observed difference
Point Mutation (2 sub-categories)
Base substituion: A point mutation where one nucleotide base in DNA is replaced with another
Frameshift mutation: A point mutation caused by the addition or removal of a small number of bases not divisible by three
The 3 categories of base subtitution mutation
Missense mutation: A change in a nucleotide base that results in the coding of a different protein (than originally)
Nonsense mutation: A change in a nucloetide base that results in the coding of a stop codon
Same-sense mutation: A change in a nucleotide base that reslts in the coding of the same protein
Base subtitution mutation examples
Template DNA: AAC (before mutation)
Missense:
-Template DNA: CAC (after mutation)
-mRNA: GUG (result codes for different protein)
Nonsense:
-Template DNA: ATC (after mutation)
-mRNA: UAG (result codes for stop codon)
Same-sense:
-Template DNA: AAT (after mutation)
-mRNA: UUA (result codes for same protein)
Missense mutation example
If protein changed from Histidine to Arginine, the protein is likely to maintain its function since they are both positively charged amino acids so if one swaps for another, the difference wouldn’t be significant
Replica plating
A techinque used by scientists, Joshua and Esther Lederberg, in which a replica of bacterial colonies was transfered from one petri plate to a new plate
-Proved that mutations are random, spontaneous events
Frameshift mutation
The addition or removal of a small number of bases not divisible by three
-Disrupts the normal reading frame in the mRNA
-Results in all codons downstream of the change being altered
-Usually results in premature appearence of a stop codon (called truncation)
-Diseases like Cystic Fibrosis caused by frameshift mutation
Chromosomal-level mutations
Large-scale changes that affect chromosome structure or number, rather than individual genes
Types of chromosomal-level mutations
Deletion: Loss of a segment of a chromosome
Insertion: Addition of extra base pairs
Inversion: A segment of a chromosome that is reversed end to end
Duplication: A repeat of a segment of a chromosome
Translocation: Segment of chromosomes are swapped
The Ames Test (for mutagenicity)
A test that uses a special strain of salmonella typhimurium (bacteria)
-Proved mutations happen spontaneously & randomly
Cytoskeleton
The dynamic system of protein fibers of various sizes and physical support & movement in a cell
Actin microfilaments
The building blocks of the cytoskeleton, composed of a form of action called G-actin
G-actin
-It is made of 43 kDa protein & 375 amino acids
-Has a nucloetide binding stire that can accommadate ATP or ADP
-Has one side that is a minus/pointed and another side that is plus/barbed (Polarity)
-Highly conserved
F-actin
F-actin (filamentous actin) is formed from G-actin (globular actin) through a process called polymerization, G-actin is its monomer
-Has helical structure
Nucleation
The process of forming nuclei (happens outside of the nucleus, in the cytoplasm)
Polymerization/Depolymerization
The addition of monomers is called polymerization, favored at the barbed (+) end since its easier to add at this end
The “falling off” of monomers is called depolymerization, favored at the pointed (-) end since its harder to add at this end
Monomer binding proteins
Proteins that have the ability to hide actin protein not needed at the moment in order to keep actin protein from polymerizaing all the time at both ends
Filament severing proteins
Proteins that have the ability to cut actin protein
-If protein is becoming too long and you don’t want to wait for depolymerization to happen, filament severing can cut down
Filament bonding proteins
Link actin protein into tight, stiff, rod-like structures
Filament crosslinking protein
Link actin protein into loose, flexible structures
Filament capping proteins
Proteins with the ability to stop the bind of the ends of actin filaments, prevents the filament from growing or shrinking at a given time
Filament anchoring proteins
Proteins with the ability to physically link the actin to integral membrane proteins (can act as signals for the cell)
Myosin
A molecular motot that converts chemical energy (ATP) into mechanical energy, generating force and movement
-Actin and myosin together are responsible for cell division, cell crawling, some vesicle movement, and muscle contraction
Note: Myosin only works on actin & actins only work on myosin motor proteins
Cell Movement (CASE 1)
-Myosin moves closer to barbed end of the actin filament, causing actin filament to slide toward pointed end
Cell Movement (CASE 2)
-Myosin is fixed so actin filaments moves towards barbed end
Steps of Cell Crawling
1. Lamellipodium (leading edge of cell) extends in the direction that cell wants to move (via actin polymerization at the barbed ends)
2. Focal adhesions (the “glue”) form at the lamelliupodium which glue to the subtraction (whatever surface the cell is on) to keep lamellipodium extended
3. Myosin pulls cell body forward (cell movement case 1)
4. Rear Adhension points of the trailing edge releases its grip, rest of cell body moves forward
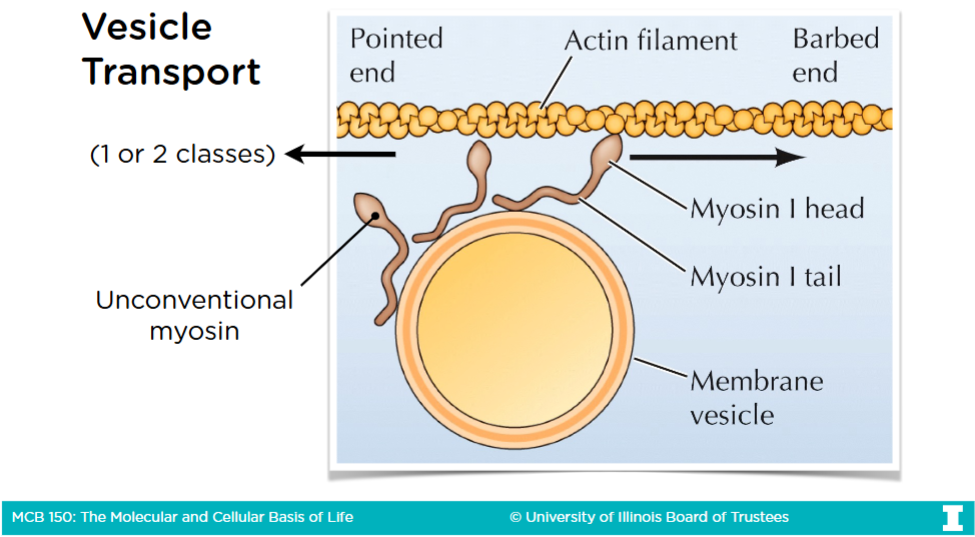
Vesicles
A type of phospholipid bilyater used to move things from one organelle to another driven by motor proteins (Myosin, Kinesin, Dynein) walking along the cytoskeletal tracks
Vesicle Transport
Muscle fibers
-Muscle cells are called muscle fibers
-Muscle fibers are multi-nucleate cells
-Long & thin cells (as opposed to the typical ballon-shaped cells)
-Organelles pushded towards periphery (outer edge) of cell
Muscle fibers continued (and contraction)
-The cytoplasm of muscle fibers are called sarcoplasm
A sacromere is th structural unit of contraction
-The repeating units of proteins within a myofibril are called sacromere
-Myofibrils are rod-like bundles of proteins are fill the sarcoplasm
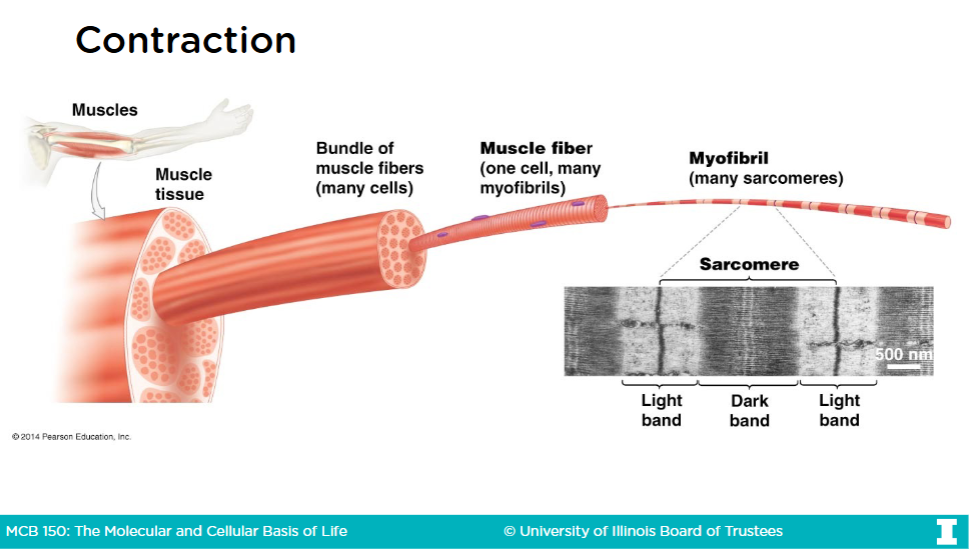
Myofibril structure
[View Image]
![<p>[View Image]</p>](https://assets.knowt.com/user-attachments/5c54f8a4-65e4-44b1-987c-6cf153505728.png)
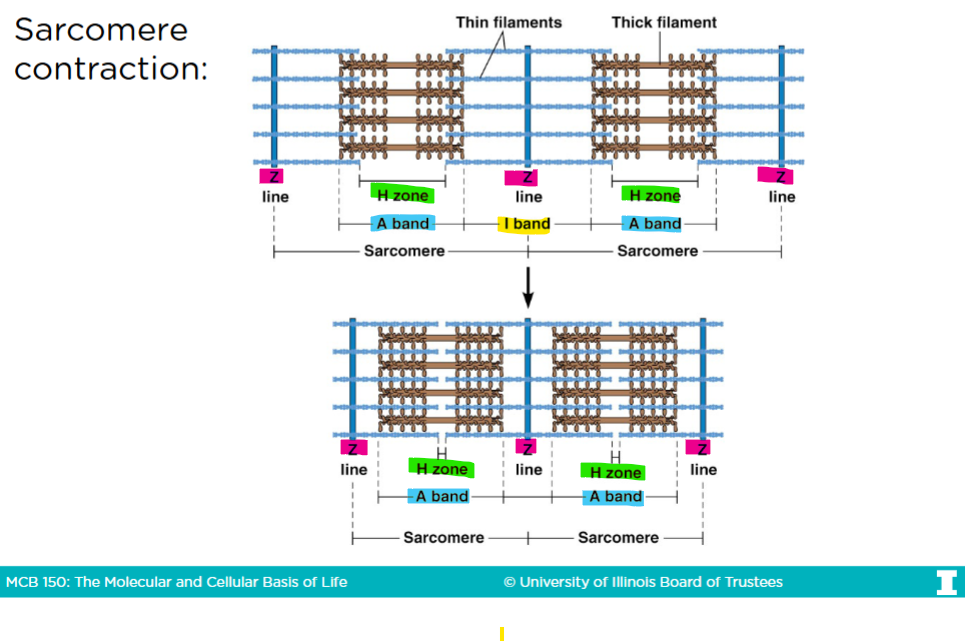
The sacromere
-Only actin is in the I-band
-Only myosin is in the H-band
-The A-band is the region that represents th entire length of myosin (thick filaments
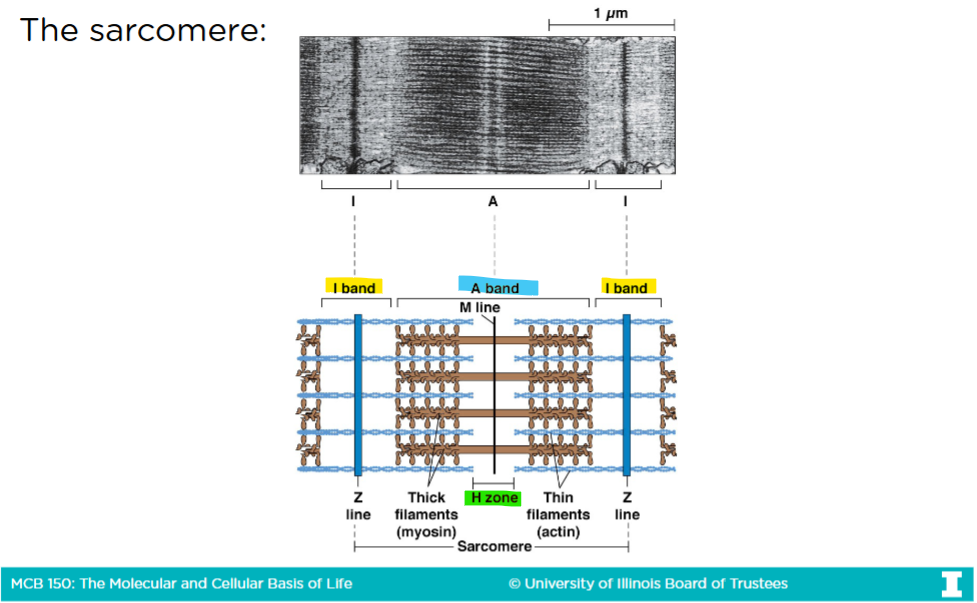
Sacromere contraction
-In an uncontracted sacromere, there is a very big H-zone
-When myosins are given ATPs, they get closer to the barbed (+) end of the actin filaments
The barbed end in the actin filaments are anchored into the Z-lines (or disk)
-After sacromere contraction, the H-zone virtually disappears, the I-band and the sacromere itself shrinks, but the A-band remains the same
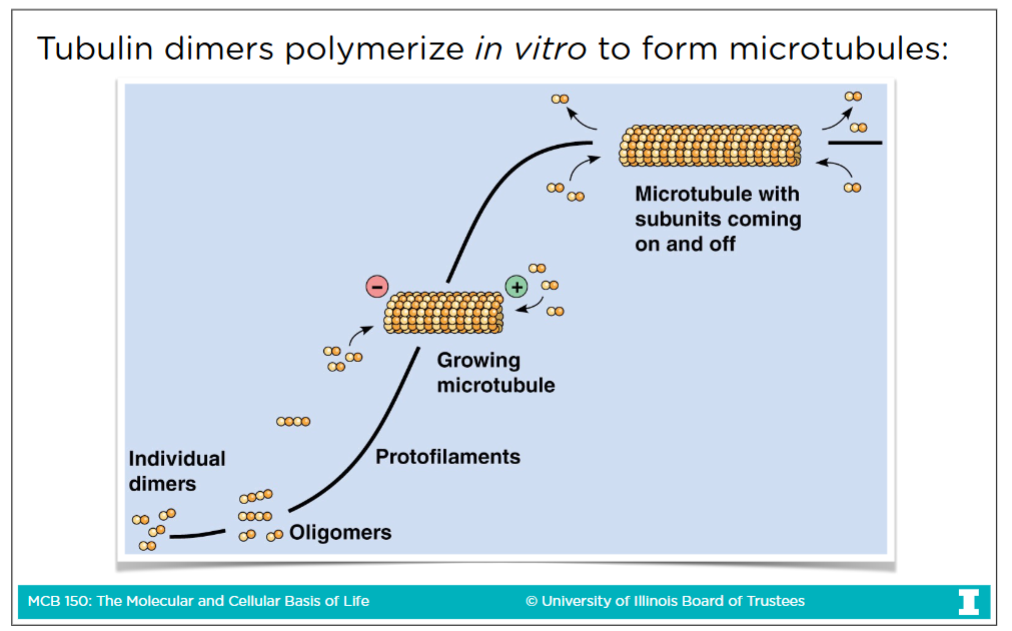
Microtubules (MTs)
-MTs are rigid, hollow tubes made of tubulin
-Movemment of cells or movement of things across a cell done by microtubules
-A verison of microtubules aclled spingle fibers segrate chromosome during mitosis
MTs characteristics
-Made of 13 linear protofilaments (which are a-tubulin and b-tublic) arranged in a hollow cylinder
-MTs have polarity, a plus end and a minus end
-Polymerization and depolymerization possible at both ends at the same rate
In vivo (in all living organisms), all polymerization and deploymerization occurs at the plus end
Tubulin
-Tubulin is globular protein made of two different subints called a(alpha)-tubulin and b(beta)-tubulin
-Both a-tubulin and b-tubulin have GTP binding sites
-Only GTP in the beta subunit is hydrolzed and GTP hydrolysis weakes the affinity of other tubulin
Tubulin expanded
Affinity: the strength of attraction or “desire” of a protein
-If you hydrolyze the alpha subunit, you would lose your affinity
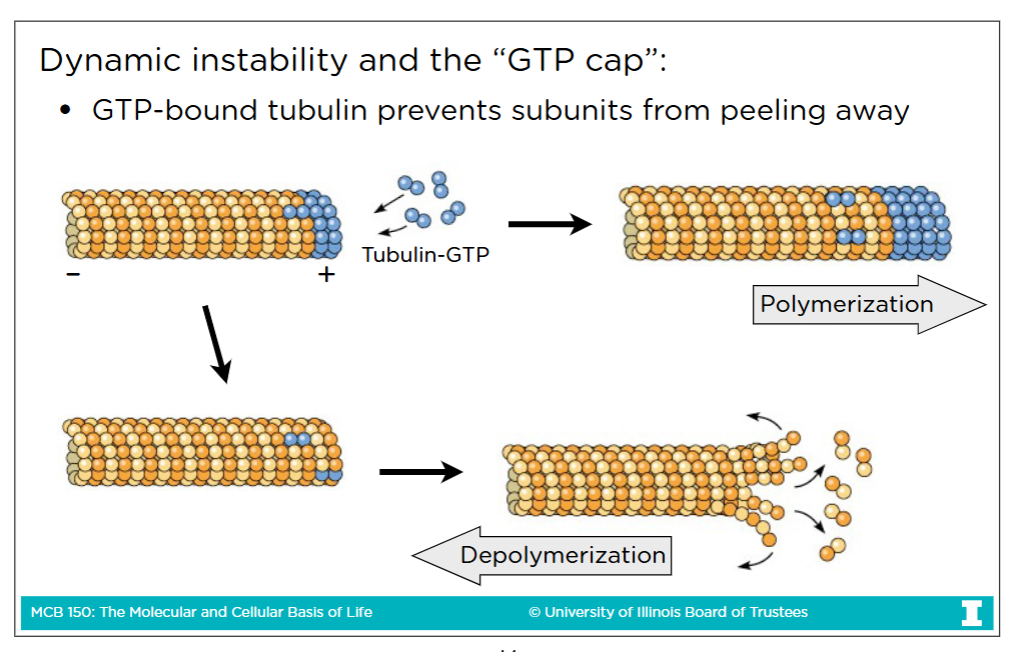
The “GTP” Cap
GTP-bound tubulin prevents subunits from peeling away
Binding tubulin
-Binding tubulin is the process in which a drug binds to tubulin, causing tubulin levels to drop
-Binding tubulin induces depolymerization
Binding tubulin drug types
Colchicine and Colcemid are drugs that affect all cell types and all MTs
Vincristine and Vinblastine are drugs that affect only rapidly-dividing cells
Binding microtubules
-Binding microtubules is the process in which a drug sits at the end of a microtubule, preventing it from growing or shrinking
-This stabilizies microtubules
Binding microtubules drug (Taxol)
Taxol is a drug that binds to microtubules and keeps from being depolymerized
Fluorescence
In Fluorescence microscopy, brighter means more
In (most) fluorescence images, tubulin are strained in green and DNA is strained in blue
Microtubules origin
Microtubules originate from Microtubule-Organizing Centers (MTOCs)
NOTE: The most precise answer to where microtubules come from is the y(gamma)-tubulin
Microtubule-Organizing Centers (MTOCs)
-In animal cells, the primary MTOC is the centrosome
In microtubule formation, the minus end of the microtubules stays anchored in the centrosome
This is why in vivo, you tend to see polymeriazation and depolymerization at ONLY the plus end
The centrosome: the primary MTOC
-Centrioles are found within a centrosome and they run perpendicular to each other
-The material surrounding the centrioles is called pericentriolar material (PCM)
-The definition of a centrosome is a pair of centrioles and PCM
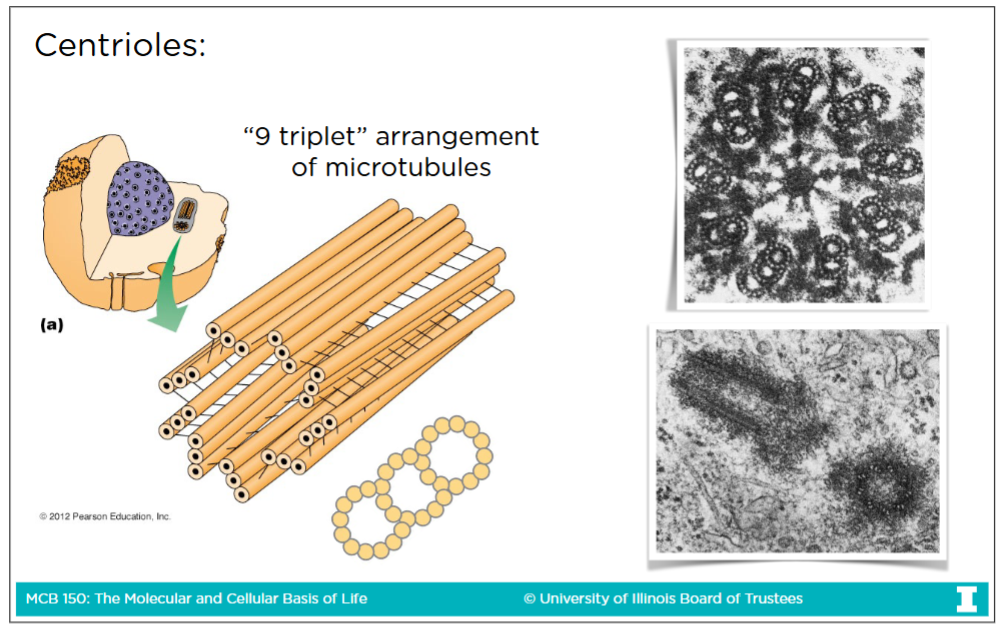
Centrioles structure
-Centrioles are 27 microtubules, found in nine triplets of microtubules
-On the image the most upper-right triplet represents the intact microtubule where all 13 profilaments are present
This is a-tubulin
-The remaining 2 triplets represent incomplete microtubule where 11-12 profilaments are present
This b-tubulin
-There are always a pair of centrioles and they are perpendicular to each other
Pericentriolar material (PCM)
PCMs are a amorphous (without clear shape) collection of protein which MTs spread out from
Microtubules and their associated motor proteins
-Microtubules have plus end and minus end (no BARBED end or POINTED end like in actin filaments)
-Dynein is a motor protein of micrtubules that goes toward the minus end
-Kinesin is a motor protein of microtubules that goes toward the plus end
Note: Myosin is ONLY a motor protein within actin filaments
BUT for the purposes of MCB 150, dyneins & kinesins essentially work the same as myosin
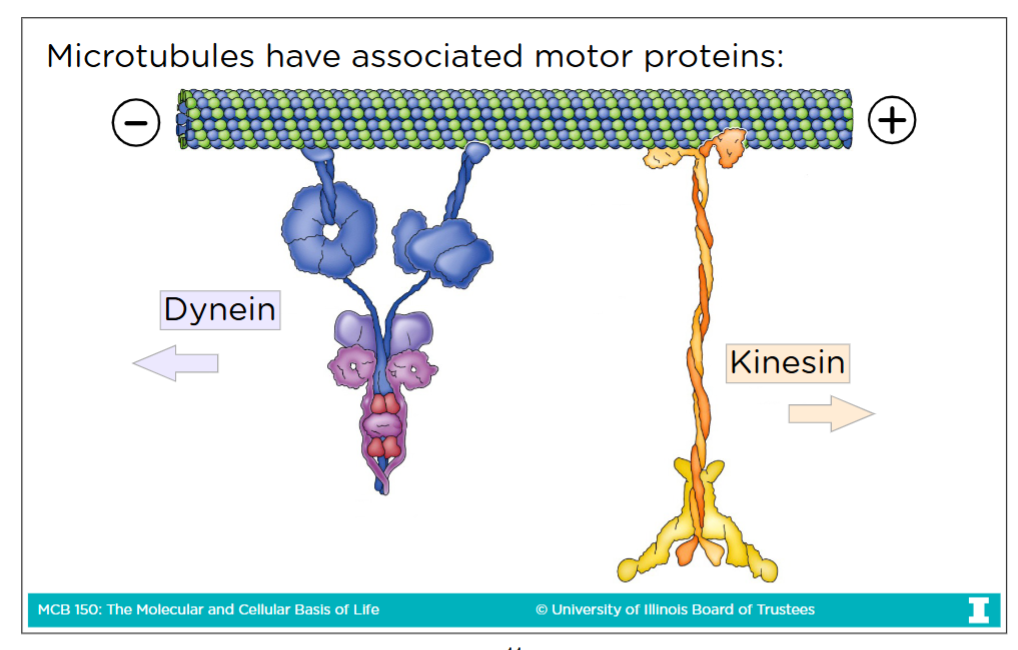
Microtubule motor proteins functions
-Intrecelluar vescile transport
-Organelle movement and positioning
-Bonding of cilla and fragella
-Separation of sister chromatids and centrosomes (during M-Phase)
Stable arrangements of microtubules
-Stable arrangments of microtubules can be used mutility via cilia and flagella
Ex: Sperm cells use (one) flagella
The “9+2” arrangment of axoneme
-The top of the flagella/cilia is the axoneme
Consists of 9 peripheral paired microtubules surrounded two central microtubules
-The bottom of the flagella/cilia is called the basal body
The minus ends of axoneme MTs are anchored to basal body
-Dynein are located in between the paired microtubules, and move towards the minus end
Cilia or flagella movement
-The dynein head groups have their tailed anchored into the a-tubule of one doublet and reach out and grab the b-tubulin of the neighboring doublet
For more details: Search “How Do Cilia and Flagella Move?” on YT
The eukaryotic cell cycle
-Our cultured human cells divide once every 24 hours
-Cell life is 95% of the time in Interphase and 5% of the time in M-Phase
11 hours in G1, 8 hours in S, 4 hours in G2, 1 hour in M (M-phase)
G0 Phase (Quiscence)
G0: When cells don’t need to divide and replicate DNA
-Ex: Skin cells are temporaily in G0 but enter G1 when they need to produce a new layer of skin tissue
-Ex: Muscle cells are permenantly in G0 as they don’t need to continue dividing after inital development
Kinetochrores
The type of proteins attached to the centrosome of a chromatid
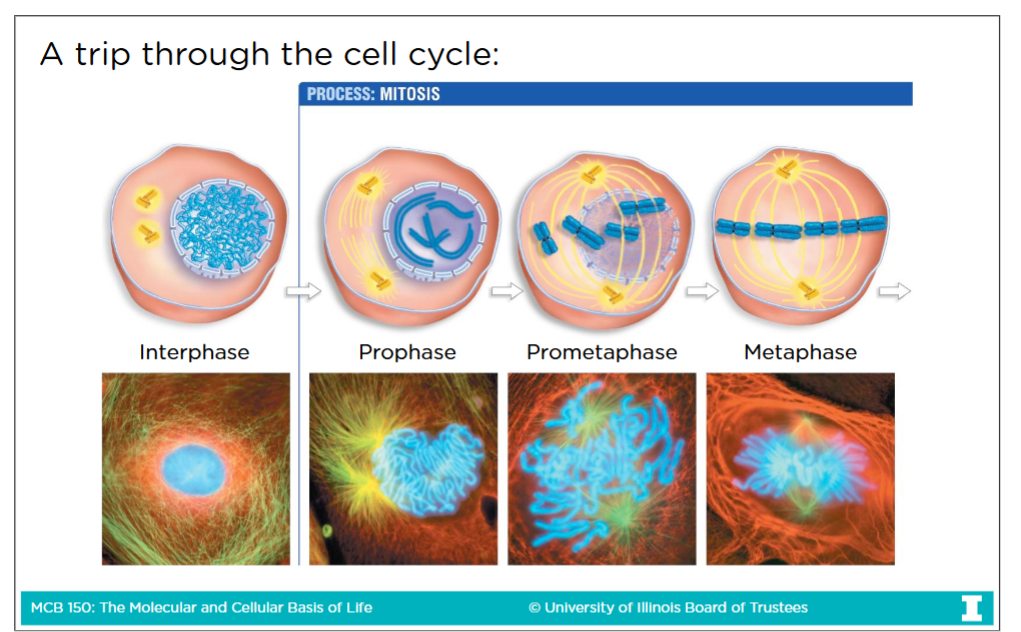
M-Phase: (Mitosis and cytokinesis)
There are 5 stages of Mitosis:
-Prophase:
Nuclear envelope disociates
Chromatins condense
Centrosomes move to opposite sites of cell
-Prometaphase:
Kinetochore MTs move pairs of sister chromatids back and forth until they reach the metaphase plate (in the middle)
-Metaphase:
All pairs of sister chromatids are broken on the metaphase plate, connections between chromatids are broken
-Anaphase:
Anaphase A: kinetochrore MTs separate sister chromatids
Anaphase B: Polar MTs “push” the poles apart, while astral MT “pull” poles
-Telophase:
Phase where everything in prophase is undo
-Cytokinesis:
Cytoplasmic division using contractile ring of actin and myosin, the cell physically divides into two cells
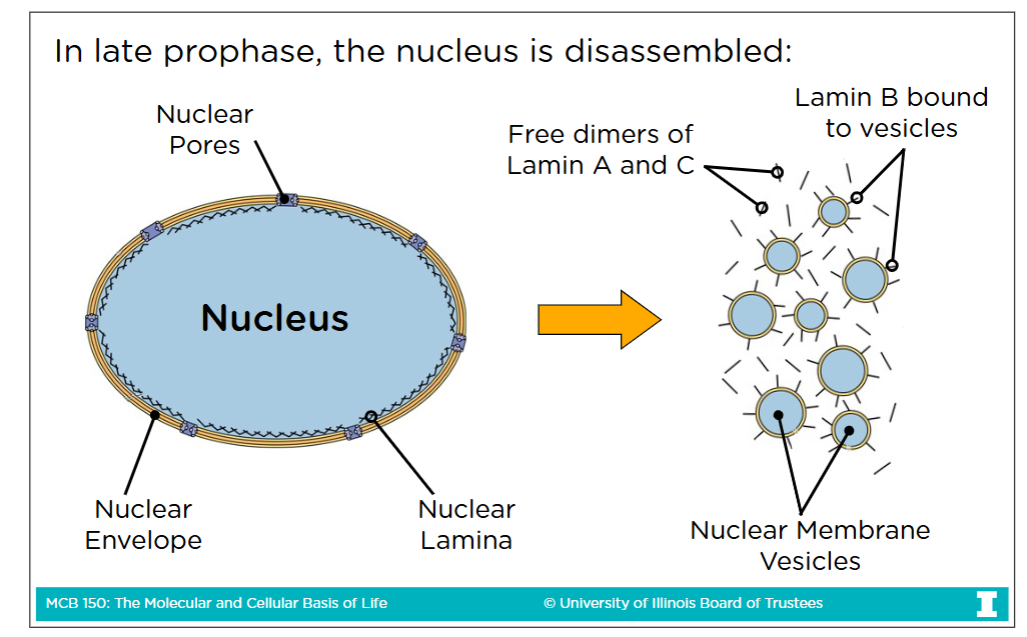
Nuclear lamina
A system of rope-like proteins fibers
During late prophase, the nuclear lamina is broken via phosphoralyation in order to disassemble the nucleus
Phosphoralyation breaks the lamina because adding the negatively charged phosphate group causes the laminas to lose their affinity for each other
(What you do in late porphase, you undo in telephase so you dephosphorylate them to form the nucleus once again)
Anaphase A
Movement of sister chromatids to opposite poles via kinetochrore MTs
-Dyneins are used to make Anaphase A happen (dynein get closer to minus end) cause the sister chromatids to be pulled toward the poles
Anaphase B
Spindle fibers distance themselves from each other, further separating sister chomarids
-Duing Anaphase B, one kinesis grabs on to one polar MT and another kinesis grabs on to another polar MT, getting closer to the plus end
-The kinesis can’t move with means the rods have to move toward plus end (with are the poles)
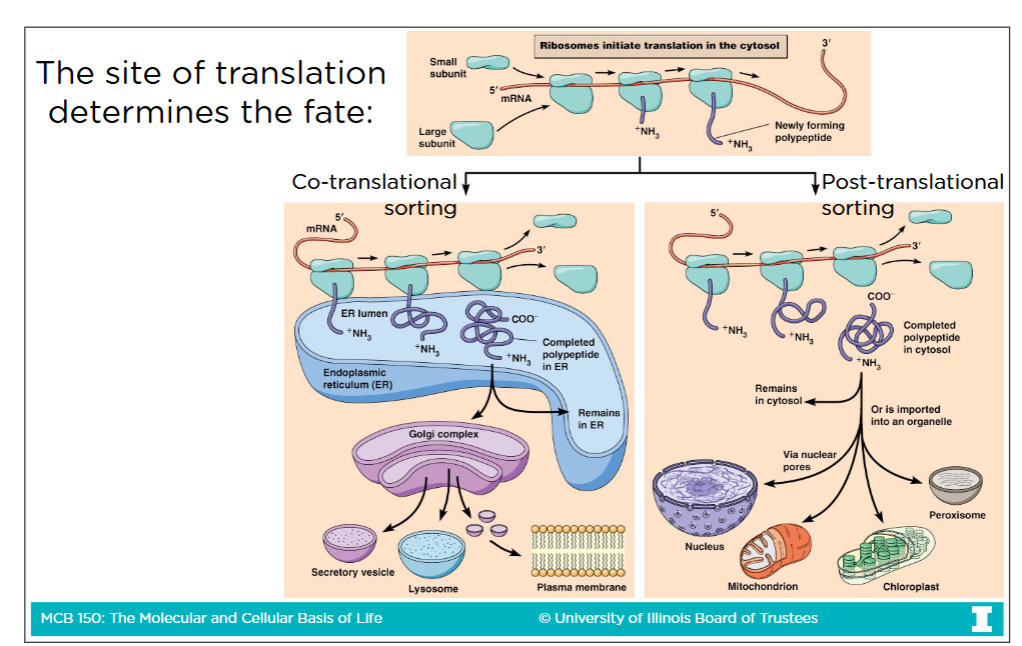
Free cytoslic ribosome
A protein that is initated in the cytoplasm and translated by a ribosome and doesn’t signal to the cell to be moved anywhere else (so remains in cytoplasm
Free cytoslic ribosome location restrictions
Free cytoslic ribosome can end up in the following locations (ONLY):
-Nucleus
-Mitochondria
-Chloroplasts
-Peroxisomes
Membrane-bound ribosome
If a protein presents a signal early that it does not want to stay and continue in the cytoplasm, then the rest of its translation continues on a membrane-bound ribosomes
Membrane-bound ribosome location restrictions
Membrane-bound ribosome can end up in the following locations (ONLY):
-The Endoplasmic Reticulum
-The golgi appartus
-Lysomoses or endosomes
-The plasma membrane
-Exported outsite cell (secretion)
Post-translational sorting
Free cytosolic ribosome (the completed protein) that go to one of their associated destinations
Co-translational sorting
Membrane-bound ribosome (the completed protein) that go to one of their associated destinations
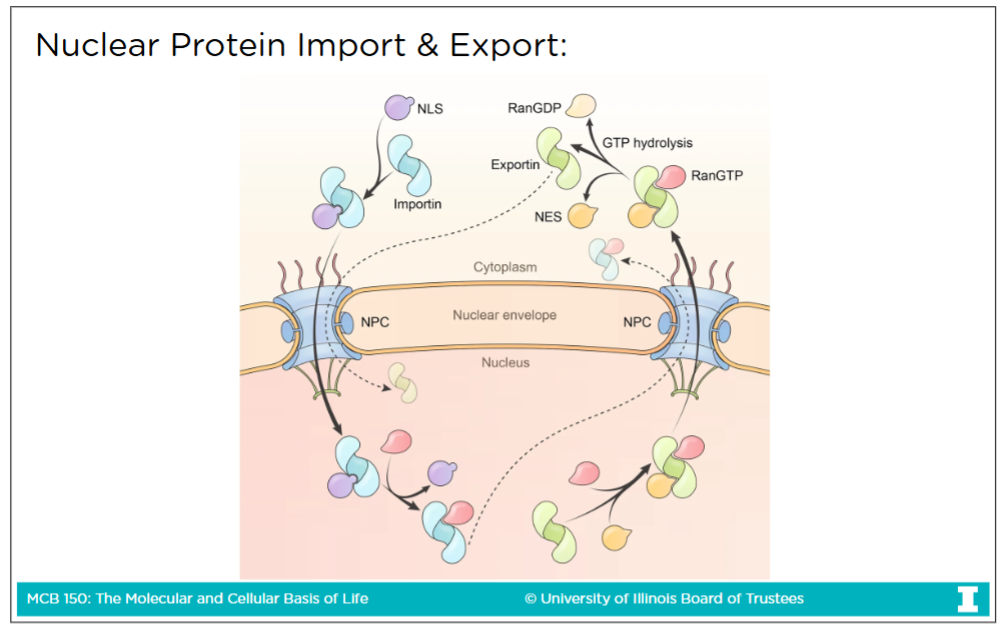
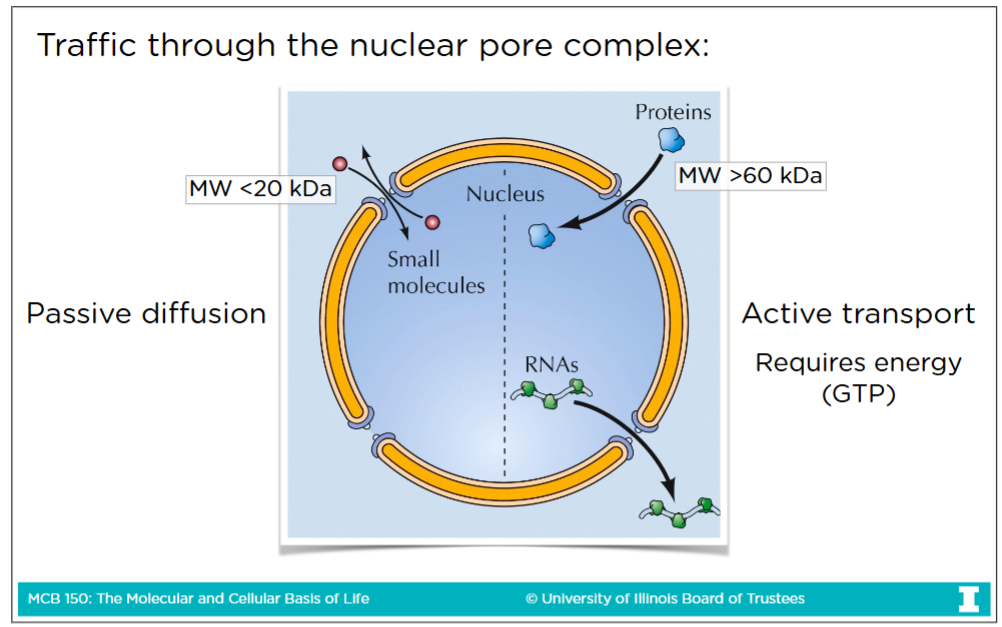
Traffic through the nuclear pore complex
-Transport through nuclear pore complex is the way to get to and from cytoplasm & nucleoplasm
Passive diffusion: The spontaneous movement of small molecules (< 60 kDA sized) and proteins through the pore without need of energy
Active transport: Transport of large molecules (> 60 kDA sized) like large protein and RNA. This process requires energy (GTP) to go through nuclear core complex
Nuclear localization signal (NLS)
The signal that tells a protein to be imported from the cytoplasm to the nucleus
-NLS is both necessary and sufficient
Necessary: removing it prevents import (protein stays in cytoplasm)
Sufficient because the signal sequence itself carries all “instructions” needed for cell’s transport
Nuclear export signal (NES)
The signal that tells a protein to be exported out of the nuclues into the cytoplasm
-NES is both necessary and sufficient
If your don’t have an NES, then you don’t get from the nucleus to the cytoplasm