Module 11: From DNA to Proteins: Transcription
1/108
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
109 Terms
evidence suggests_____was original genetic material
RNA
Ribozymes:
catalytic RNA
The structure of RNA (2)
• Primary structure • Secondary structure
Primary Structure =
RNA molecule folds to form secondary structures, owing to hydrogen bonding between complementary bases on the same strand
Y/N DNA RNA: Composed of nucleotides
Yes, Yes
DNA RNA: Type of sugar
Deoxyribose, Ribose
Y/N DNA RNA: Presence of 2'-OH group
No, Yes
Y/N DNA RNA: Bases
A, G, C, T;
A, G, C, U
Y/N DNA RNA: Nucleotides joined by phosphodiester bonds
Yes, Yes
Y/N DNA RNA: Double or single stranded
Usually double, Usually single
Y/N DNA RNA: Secondary structure
Double helix, Many types
Y/N DNA RNA: Stability
Stable, Easily degraded
(Tp, Location and Function) Ribosomal RNA (rRNA)
Bacterial and eukaryotic, Cytoplasm, Structural and functional components of the ribosome
(Tp, Location and Function) Messenger RNA (mRNA)
Bacterial and eukaryotic, Nucleus and cytoplasm, Carries genetic code for proteins
(Tp, Location and Function) Transfer RNA (tRNA)
Bacterial and eukaryotic, Cytoplasm, Helps incorporate amino acids into polypeptide chain
(Tp, Location and Function) Small nuclear RNA (snRNA)
Eukaryotic, Nucleus, Processing of pre-mRNA
(Tp, Location and Function) Small nucleolar RNA (snoRNA)
Eukaryotic, Nucleus, Processing and assembly of rRNA
(Tp, Location and Function) MicroRNA (miRNA)
Eukaryotic, Cytoplasm, Inhibits translation of mRNA
(Tp, Location and Function) Small interfering RNA (siRNA)
Eukaryotic Cytoplasm Triggers degradation of other RNA molecules
(Tp, Location and Function) Piwi-interacting RNA (piRNA)
Eukaryotic, Nucleus and cytoplasm, Suppresses the transcription of transposable elements in reproductive cells
(Tp, Location and Function) Long noncoding RNA (IncRNA)
Nucleus and cytoplasm Variety of functions
(Tp, Function) CRISPR RNA (crRNA)
Prokaryotic, Assists in destruction of foreign DNA
Which class of RNA is correctly paired with its function?
Transcription Is the…
Synthesis of an RNA Molecule from a DNA Template
Some RNAs are transcribed in both prokaryotic and eukaryotic cells ?
mRNA, rRNA, tRNA
Some RNAs are transcribed only in eukaryotic cells ?
pre-mRNA, snRNA, snoRNA, miRNA, siRNA, IncRNA, piRNA
Some RNAs are transcribed only in prokaryote cells ?
crRNA (crispr RNA)
Some viruses copy RNA directly from…
RNA
All cellular types of RNA are transcribed from…
DNA
The transcribed strand:
template strand
The transcription unit contains:
• A promoter
• RNA-coding sequence
• Terminator
Under the electron microscope, DNA molecules undergoing transcription exhibit…
Christmas-tree-like structures
RNA molecules are synthesized that are complementary and antiparallel to…
one of the two nucleotide strands of DNA, the template strand
RNA is transcribed from one…
DNA strand.
What is the difference between the template strand and the nontemplate strand?
The template strand is the DNA strand that is transcribed into an RNA molecule; the nontemplate strand is not transcribed.
A transcription unit includes:
a promoter, a region that encodes RNA, and a terminator
A transcription unit Drawing
correct
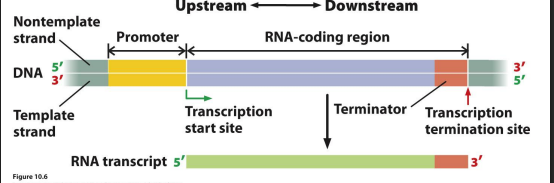
The substrate for transcription
Ribonucleoside triphosphates (rNTPs) added to the 3 end of the RNA molecule
The transcription apparatus
Bacterial RNA polymerase, The sigma factor:
The sigma factor:
binding to the promoter when transcription starts
Ribonucleoside triphosphates are substrates used in…
RNA synthesis
In transcription, nucleosides are always added to the….
3’ end of the RNA molecule
In transcription, nucleosides are always added to the 3’ end of the RNA molecule Steps:
Initiation of RNA synthesis does not require a primer
New Nucleotides are added to the 3’ end of the RNA molecule
DNA unwinds at the front of the transcription bubble and then rewinds
What is the function of the sigma factor?
The sigma factor controls the binding of RNA polymerase to the promoter.
Bacterial Transcription Consists of…
Initiation, Elongation, and Termination
Initiation • Bacterial promoters
Consensus sequences:
Consensus sequences:
sequences that possess considerable similarity
Consensus sequences: examples
• –10 consensus: 10 bp upstream of the start site
• Pribnow box: • 5 TATAAT 3 3 ATATTA 5
• –35 consensus sequence: TTGACA
The consensus sequence comprises the most …
commonly encountered nucleotides at each site
In consensus sequences: Pyrimidines indicated by..
Y
In consensus sequences: Purines indicated by..
R
In consensus sequences: N means….
that no particular bases is more common
In consensus sequences: C/G means….
Cytosine and Guanine are equally common
A consensus sequence consists of the most….
commonly encountered bases at each position in a group of related sequences
In bacterial promoters, consensus sequences are found…
upstream of the start site, approximately at positions -10 and -35
Initial RNA synthesis: primer?
no primer is required
The location of the consensus sequence determines the…
position of the start site
Transcription in bacteria is catalyzed by….
RNA polymerase, which must bind to the sigma factor to initiate transcription
RNA transcription is initiated when …
core RNA polymerase binds to the promoter with the help of sigma
RNA transcription initiated in Bacteria 6 steps
The sigma factor associates with the core enzyme to form a holoenzyme
which binds to the -35 and -10 consensus sequences in the promoter
The holoenzyme binds the promoter tightly and unwinds the double-stranded DNA
A nucleoside triphosphate complementary to the DNA at the start site serves as the first nucleotide in the RNA molecule
Two phosphate groups are cleaved from each subsequent nucleoside triphosphate, creating an RNA nucleotide that is added to the 3’ end of the growing RNA molecule
The sigma factor is released as the RNA polymerase moves beyond the promoter
RNA elongation is carried out by….
the action of RNA polymerase
Termination: Rho-dependent termination:
uses rho factor
Rho-independent termination:
hairpin structure formed by inverted repeats, followed by a string of uracils
The termination of transcription in some bacterial genes requires the…
presence of the rho factor
Rho-independent termination in bacteria is a multistep process (steps)
inverted repeats are transcribed into RNA
The string of U’s causes the RNA polymerase to pause
the inverted repeats in RNA fold into a hairpin loop
this destabilizes the DNA-RNA pairing
The RNA transcript separates from the template, terminating transcription
Rho-independent terminator contains…
an inverted repeat followed by a strong of approximately 6 adenine nucleotides
Conclusion of Rho Independent termination
Transcription terminates when inverted repeats form a hairpin followed by a string of uracils
Number of nucleotides in a gene should be proportional to…
the number of amino acids in the encoded protein
DNA _____ than mRNA;
much longer
DNA much longer than mRNA; demonstrated through…
hybridization
The concept of colinearity suggests that a continuous sequence of nucleotides in DNA encodes a…
continuous sequence of amino acids in protein.
A continuous sequence of nucleotides in the DNA codes a ….
continuous sequence of amino acids in the protein
The noncolinearity of eukaryotic genes was discovered by…
hybridizing DNA and mRNA
Is the coding sequence in a gene always continuous?
Coding sequences in a gene may be interrupted by noncoding sequences
Experiment: coding sequence in a gene always continuous?
Mix DNA with Complimentary RNA and heat to a seperate DNA strands
cool the mixture complementary, sequences pair
Experiment: coding sequence in a gene always continuous? Results
DNA may reanneal with its complimentary strand
or with RNA (Noncoding regions of DNA are seen as loops)
What evidence indicated that eukaryotic genes are not collinear with their proteins?
When DNA was hybridized to the mRNA transcribed from it, regions of DNA that did not correspond to RNA looped out.
exons
coding sequences
introns
noncoding sequences
The coding sequences (exons) of many eukaryotic genes are disrupted by…
noncoding sequences (introns)
The gene includes:
DNA sequences that code for all exons and introns
• Those sequences at the beginning and end of the RNA that are not translated into a protein, including the entire transcription unit
Those sequences at the beginning and end of the RNA that are not translated into a protein, including the entire transcription unit
• The promoter
• The RNA coding sequence
• The terminator
Many RNA Molecules Are Modified After…
Transcription in Eukaryotes
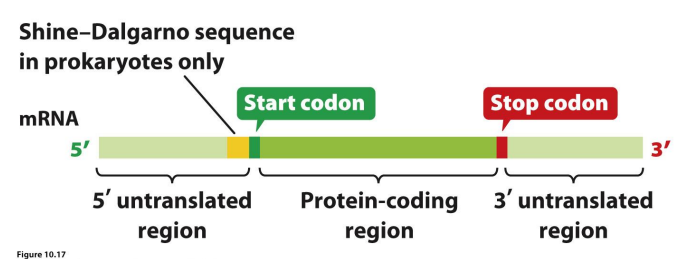
A mature mRNA contains…
5′ untranslated region (5′ UTR, or leader sequence), protein coding region, 3’ untranslated region
Shine-Dalgarno sequence in….
Prokaryotes only
5′ untranslated region aka.
Shine–Dalgarno sequence
Draw a mature mRNA
correct
The addition of the 5′ cap
A nucleotide with 7-methylguanine; 5′-5′ bond is attached to the 5′- end of the RNA
The addition of the poly(A) tail
~ 50 to 250 adenine nucleotides are added to the 3′-end of the mRNA
Most eukaryotic mRNAs have a…
5’ cap
Most eukaryotic mRNAs have a…
3’ poly (A) tail
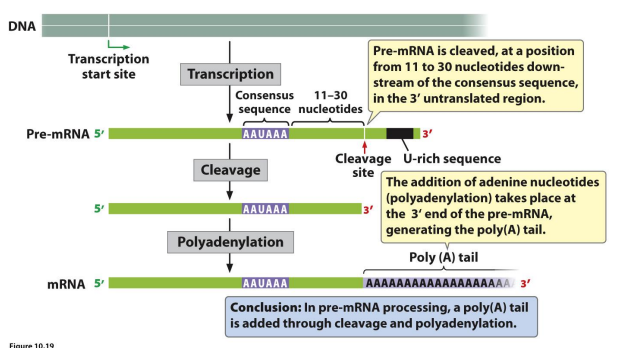
In pre-mRNA processing, a poly(A) tail is added through…
cleavage and polyadenylation.
pre-mRNA processing drawn
correct
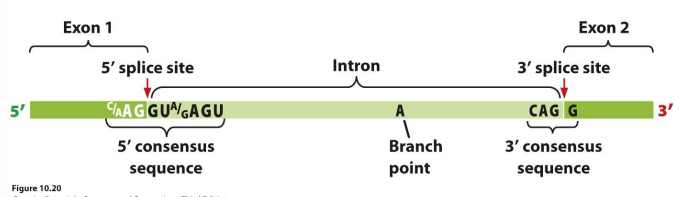
RNA splicing
Consensus sequences, Spliceosome:
Consensus sequences
• 5′ consensus sequence
• 3′ consensus sequence:
• Branch point: the adenine “A”: ~18- 40 nucleotides upstream of 3′- splicing site
Spliceosome:
five RNA molecules + 300 proteins
The splicing of pre-mRNA requires…
consensus sequences.
Critical consensus sequences are present at the….
5’ splice site and the 3’ splice site.
Draw the splicing of pre-mRNA
correct
The splicing of pre-mRNA introns requires a ____ process
two-step