BIOC 302 - Decoding the genetic code
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
The genetic code is a blueprint for ___ which is a blueprint for ____
RNA
Protein
what amino acid are all proteins born with?
methionine, but not all proteins have it at the start after bc proteases remove some aas from the sequence
Describe the experiments that Nirenberg performed to crack the code
He fed poly(U) and 20 radiolabeled amino acids to E. coli. They found that the radioactive polypeptide only formed in the tube containing *Phe, thus, UUU codes for Phe. They did the same approach to reveal that CCC codes for proline, AAA codes for lysine, etc.
why does GGG not generate any polypeptides?
bc it forms tetraplexes that cannot be bound by ribosomes. However GGG do have aa that they code for but having huge stretches of G’s forms these tetraplexes.
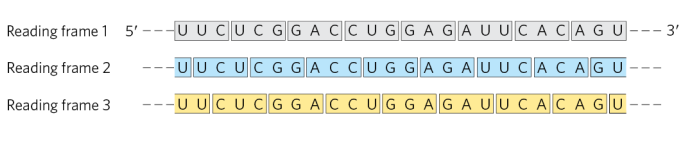
What is the implication of the genome having multiple reading frames?
Because codons are read in groups of three with no punctuation, a single mRNA sequence can be read in three different reading frames, each producing a completely different amino acid sequence. The correct reading frame is determined by the start codon, and choosing the wrong frame results in a nonfunctional protein.
what is the initiation codon? what does it do?
AUG. it signals the beginning of a polypeptide in all cells. It also codes for Met in internal positions of polypeptides
what are termination codons?
codons that normally signal the end of polypeptide synthesis. Also called stop codons or nonsense codons. Do not code for any known amino acids.
what are missense mutations?
mutations in which a single new base pair replaces another. This is the most common type of mutation. In the third position of the codon, single base substitutions change the amino acid about 25% of the time
what are silent mutations?
mutations in which the nucleotide is different but the encoded amino acid stays the same
what is a transition mutation?
a missense mutation in which a purine is replaced by a purine, or a pyrimidine by a pyrimidine. All 3 codon positions have some resistance to these mutations
what is the effect of a mutation in the first base of a codon?
a mutation in the first base of a codon usually produces a conservative substitution. For example, changing GUU (Val) to AUU (Ile) substitutes one hydrophobic amino acid for another
what is a nonsense mutation?
nucleotide change that results in a premature stop codon
Explain Francis Crick’s adaptor hypothesis
he proposed that there are small RNA adaptors (tRNA) that binds a specific amino acid and recognizes a complementary codon on the mRNA, bringing the correct amino acid to the ribosome for protein synthesis
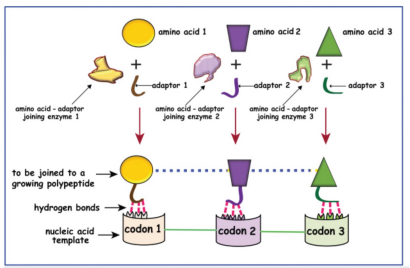
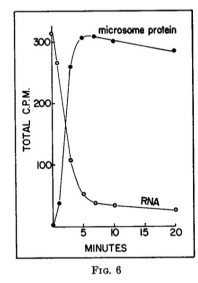
interpret the experiment that demonstrated that tRNAs are the adaptor molecules needed to decode the genetic code
Used hot leucine (radioactive). At the beginning of the experiment, the leucine was associated with a small molecule of RNA (can tell it’s small bc they used centrifugation which separates based on size). They quickly observed that there was transfer of this leucine to the growing polypeptide chain. Transfer of leucine from a small thing to a big thing. (RNA c.m.p. goes down as leucine is tranferred off of it, while polypeptide chain gets bigger as leucine is added to it)
what are the common structural elements of tRNA?
8+ residues have modified bases and sugars (many are methylated derivatives)
have a guanylate (pG) residue at the 5′ end
have the trinucleotide sequence CCA(3′) at the 3′ end
have a cloverleaf shape in 2-D and a twisted L shape in 3-D
tRNA has an amino acid arm, what does it do?
carries a specific amino acid esterified by its carboxyl group to the A residue at the 3’ end of tRNA
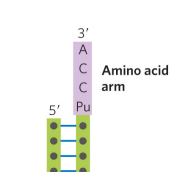
what is the anticodon arm in tRNA?
arm that contains the anticodon
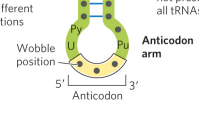
what is an anticodon?
a 3-base sequence on the tRNA that base pairs with mRNA codons through hydrogen bonding. The alignment of the 2 RNA segments is antiparallel.
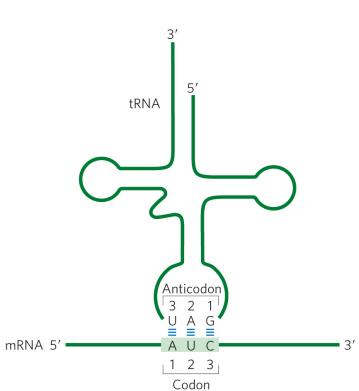
the genetic code is redundant. what does this mean?
64 codons means some amino acids are encoded by more than one codon (most actually are)
why can some tRNA molecules recognize more than one codon?
bc of the “wobble” at the first position on the tRNA (last nucleotide in the codon). The wobble position allows for non-canonical base pairing that will satisfy the system (e.g., U with G).
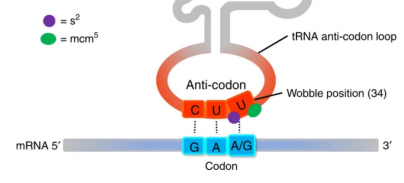
the anticodons in some tRNAs include the nucleotide inosinate (designated I), why?
bc inosinate can form weak hydrogen bonds with A, U, and C (not G bc too similar in structure).
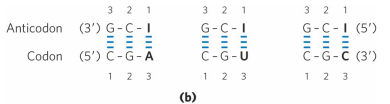

2 bc:
can use inosinate at the wobble position would be able to bind with A, U, and C
need a second one bc can’t use inosinate to bind to G, so need smth else at the wobble position
According to the wobble hypothesis, how many tRNAs are needed to translate all codons, and why?
At least 32 tRNAs are needed to translate all 61 codons. Codons that differ in the first two bases need different tRNAs, but the wobble at the third codon position allows a single tRNA to recognize multiple codons that vary only in that position.
what do aminoacyl-tRNA synthetases do?
they catalyze the reaction between the tRNA and a specific amino acid in the cytosol. It activates the carboxyl group of each amino acid and establishes a link between each new amino acid and the information encoding it in the mRNA
what does the “second genetic code” refer to?
refers to the interaction between aminoacyl-tRNA synthetases and tRNAs. Ind. aminoacyl-tRNA synthetases must be specific for a single amino acid and for certain tRNAs. Nucleotides in the tRNA that confer binding specificity are concentrated in the amino acid arm, the anticodon arm, and the nucleotides of the anticodon itself.
Draw + explain step 1 of the aminoacyl-tRNA synthetase reaction
step 1 forms the enzyme-bound intermediate, aminoacyl adenylate (aminoacyl-AMP)
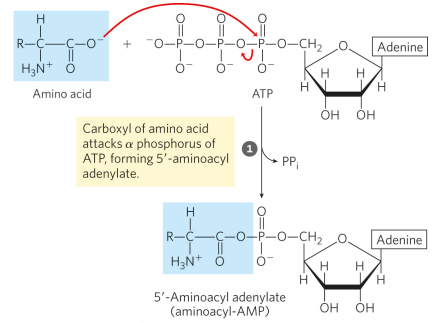
draw + explain step 2 of the aminoacyl-tRNA synthetase reaction
step 2 transfers the aminoacyl group from enzyme -bound aminoacyl -AMP to its corresponding specific tRNA – mechanism depends on the enzyme class.
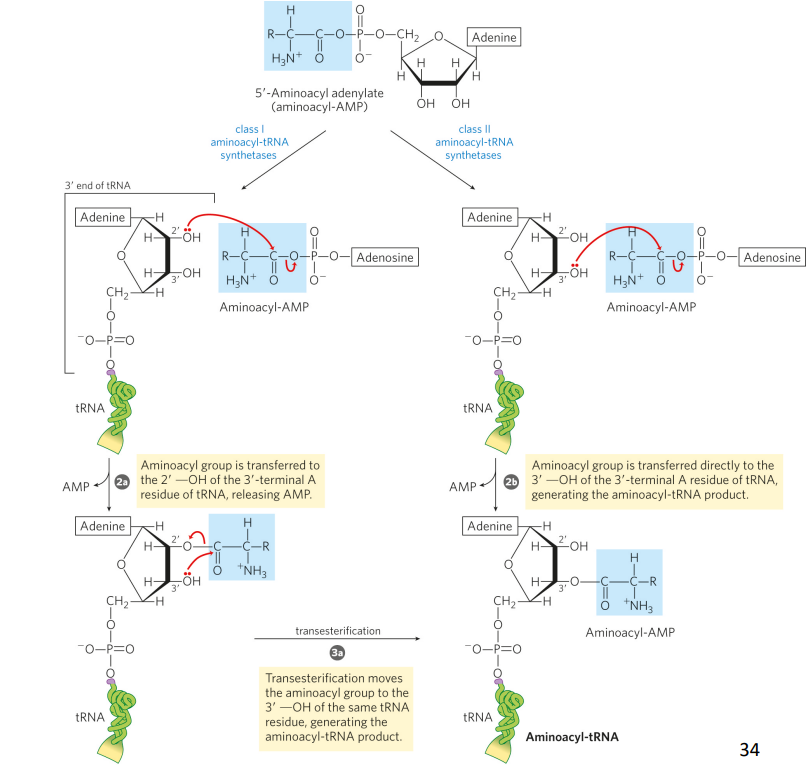
what part of the tRNA could be used in the amino acyl-tRNA synthetase recognition?
amino acid arm - (specific base pairs like G=U in the amino acid arm of tRNA for alanine)
3’ end (CCA tail)
anticodon loop
(full tRNA structure not always needed for amino acyl-tRNA synthetase recognition if key identity elements are present)
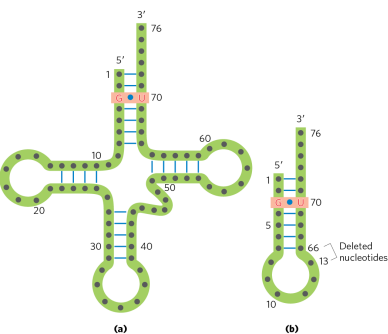
True or False: Since mulFple tRNA molecules are charged with the same amino acid. Aminoacyl-tRNA synthetases can add an amino acid to mulFple tRNAs
true
some aminoacyl-tRNA synthetases have proof-reading activity. Describe this proof-reading activity of Aminoacyl-tRNA synthetases with an example
valine and isoleucine are very similar in structure, therefore they may be able to fit into active site of tRNA synthetase even if its not meant for it (e.g., Val-AMP fits into the active site Ile-tRNA synthetase). However, Ile-tRNA synthetase has a proofreading function to distinguish between Val and Ile. Val-AMP fits into the hydrolytic (proofreading) site of Ile-tRNA synthetase and so it is hydrolyzed. Thus, tRNA does not become aminoacylated to the wrong amino acid. Basically, wrong amino acid → editing site → gets removed
What are the components that make up the ribosome?
rRNA
Proteins
organized into 2 subunits: 30S (small) and 50S (large) - bacteria. 60S and 40S in eukaryotes
describe the structure of bacterial ribosomes
the 50s and 30s subunits come together to form a cleft that allows for the passage of mRNA (where peptide bond forms). Where the peptide bond forms is actually an RNA compartment, so there’s no protein around that region (the active site for peptide bond formation). Therefore, we consider a ribosome a ribozyme bc the business part of it is made of RNA.
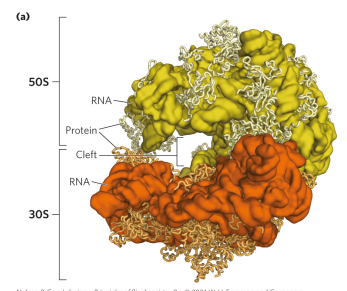
describe the structure of eukaryotic ribosomes
60S and 40S subunits come together to form a cleft that allows for the passage of mRNA (where peptide bond forms). Where the peptide bond forms is actually an RNA compartment, so there’s no protein around that region (the active site for peptide bond formation). While eukaryotic ribosomes have similar structure to bacterial ribosomes, they have somewhat increased complexity
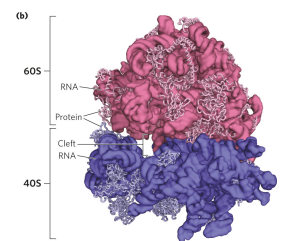
Compare bacterial and eukaryotic ribosome
they have an overall similar structure. However, eukaryotic ribosomes are larger (around 80S) and more complex than bacterial ribosomes, and have 2 subunits (60S and 40S on average), while bacterial are smaller (around 70S), and have 2 subunits (50S and 30S). Chloroplasts and mitochondria have simpler ribosomes than bacteria.
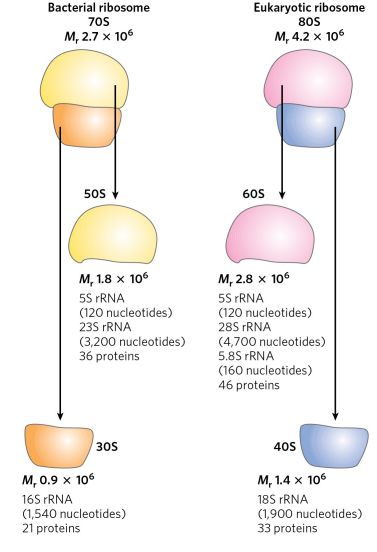
What is conserved in the small subunit rRNAs across all organisms?
The folding patterns (secondary structure) of rRNAs are highly conserved in all organisms, allowing for ribosomes to function properly
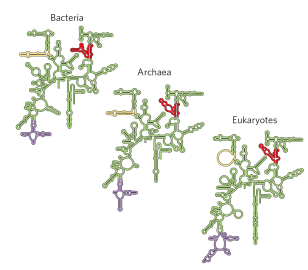
Explain how ribosomes are assembled in eukaryotes
in the nucleolus, Pol I and Pol III are going to make different rRNA molecules
processing of the pre-rRNAs occurs within large ribonucleoprotein complexes
mRNA thats going to make the ribosomal protein is made from Pol II in the nucleoplasm
The mRNA made by Pol II is spliced and processed and sent into the cytoplasm for translation
ribosome translates ribosomal proteins and the ribosomal protein is going to fold in the cytoplasm and then go back into the nucleus through nuclear pore and into then the nucleolus
In nucleolus, ribosomal proteins are packaged with rRNAs into large and small ribosomal subunits.
assembly occurs hierarchically, as r-proteins are incorporated while rRNAs are synthesized
dozens of proteins and snoRNAs are involved in final maturation of the ribosome
large and small subunits leave the nucleus separately and than are joined together in the cytoplasm
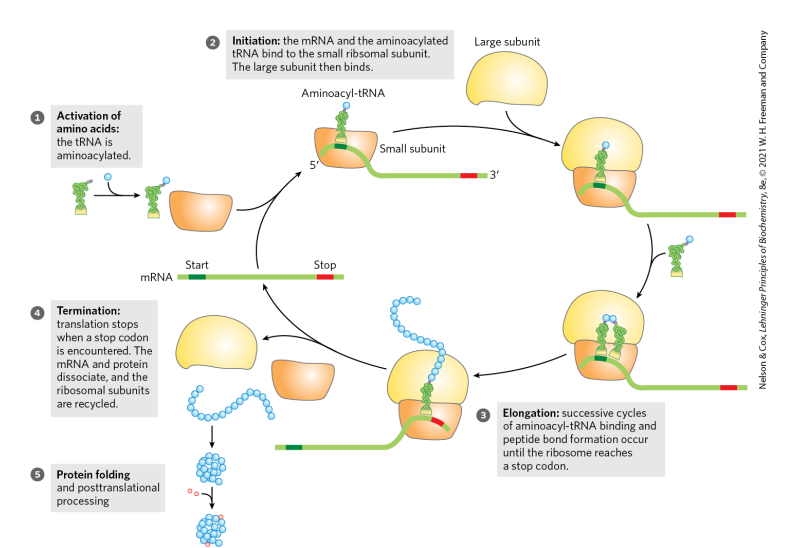
what are the 5 stages of protein synthesis?
activation of amino acids: tRNA is aminoacylated
initiation
elongation
termination
protein folding
what are the 3 sites on ribosomes that bind tRNAs?
aminoacyl (A) site
peptidyl (P) site
exit (E) site
what is the A site for in ribosomes?
site where incoming aminoacyl-tRNAs (other than fMet-tRNA) bind. IF1 binds and prevents tRNA binding at the A site during initiation
what is the P site for in ribosomes?
site where amino acids are added to the growing chain (only site where fMet-tRNA can bind)
what is the E site for in ribosomes?
binds uncharged tRNAs
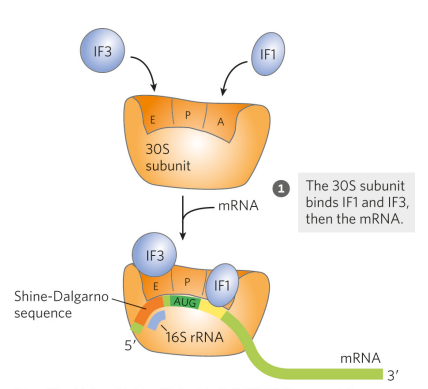
What is the first step of initiation for protein synthesis in bacteria?
The 30S subunit binds IF1 (at the A site) and IF3 (at the E site), then the mRNA. IF3 prevents the 30S and 50S subunits from combining prematurely. On the mRNA, the Shine-Dalgarno sequence forms W-C interactions with 16S rRNA (part of the ribosome), allowing for the initiating (5’)AUG to its correct position (the P site) (the mRNA is complementary to a sequence in the rRNA)
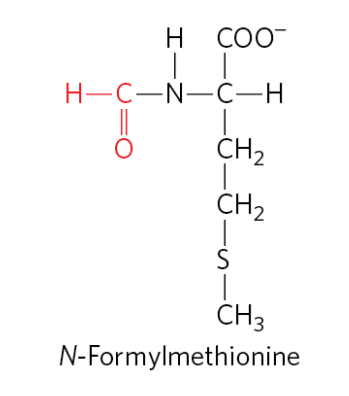
what are the two tRNAs specific for methionine in bacteria, and what are their roles?
fMet-tRNA^fMet inserts N-formylmethionine (fMet) at the amino-terminal end (start of the protein) in response to the (5’)AUG initiation codon.
Met-tRNA^Met inserts methionine in interior positions of a protein in response to interal (5’) AUG codons
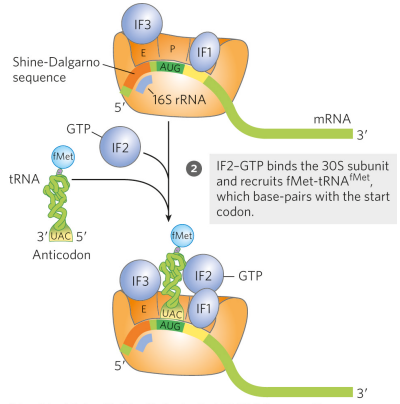
What is the second step of initiation for protein synthesis in bacteria?
the anticodon of fMet-tRNA^fMet is correctly paired with the mRNA’s initiation codon at the P site. IF2-GTP binds the 30S subunit and recruits fMet-tRNA^fMet, which base-pairs with the start codon (IF2 will only bind if methionine is formylated)
What is the third step of initiation for protein synthesis in bacteria?
The 50S subunit associates, IF2 hydrolyzes GTP, and IF1, IF2, and IF3 dissociate, leaving the 70S initiation complex (initiation complex = functional 70S ribosome containing mRNA and the initiating fMet-tRNA^fMet)
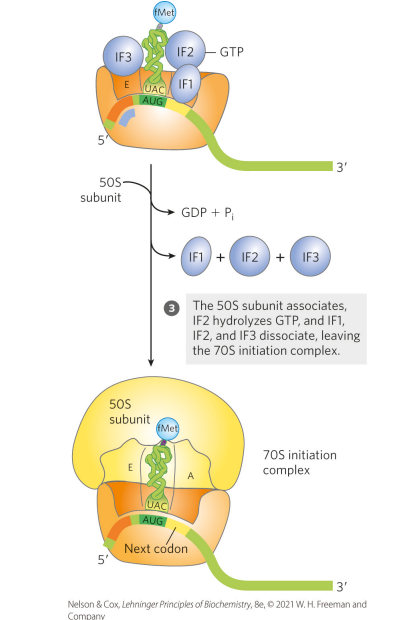
what is the first step of elongation for protein synthesis in bacteria?
binding of an incoming aminoacyl-tRNA. An incoming aminocyl-tRNA binds a GTP-bound EF-Tu complex. The aminoacyl-tRNA–EF-Tu– GTP complex binds to the A site of the 70S initiation complex. After GTP hydrolysis (which makes sure correct AA in A site), EF-Tu– GDP complex leaves the ribosome.
what is the second step of elongation for protein synthesis in bacteria? (describe + sketch)
The formation of a peptide bond. The N-formylmethionyl group is transferred to the amino group of the aminoacyl-tRNA in the A site to form a dipeptidyl-tRNA (alpha-amino group of the amino acid in the A site acts as the nucleophile). N-terminus faces the carbonxyl end of the last amino acid. tRNA shift to a hybrid binding state where the 3’ and 5’ ends of tRNA^fMet are in the E site and the 3’ and 5’ ends of the peptidyl-tRNA are in the P site
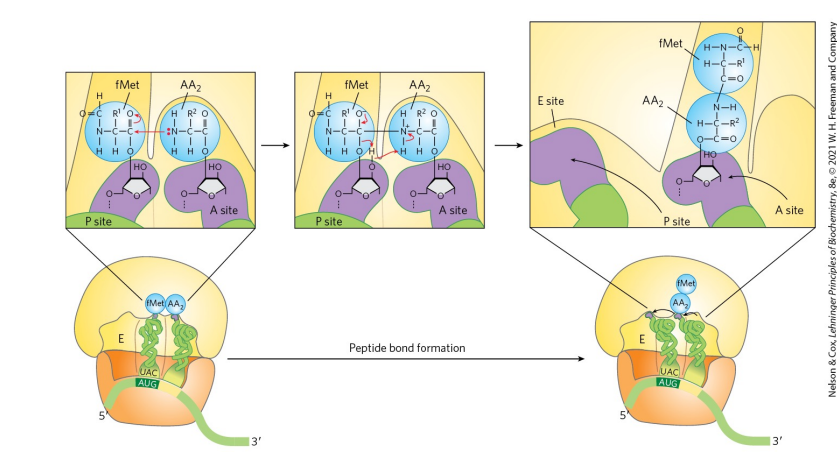
how does tRNA processing of the 3’ end contribute to the formation of the peptide bond?
It ensures the tRNA has the correct length and shape so the amino acid aligns properly in the ribosome for peptide bond formation
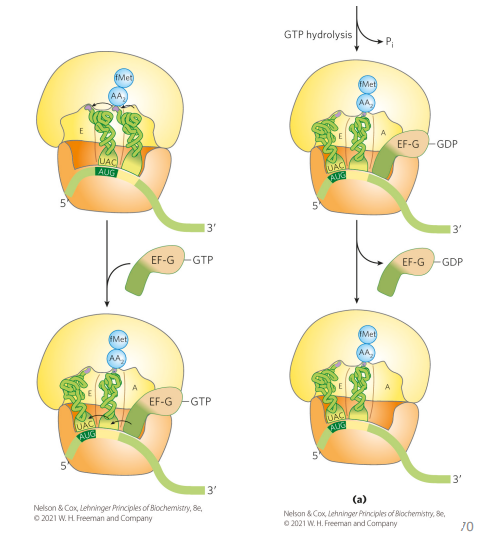
how does translocation during translation occur in bacteria?
elongation factor G (bound to GTP) goes on A site and bumps everything over (displacing the peptidyl-tRNA). Hydrolysis of GTP allows for elongation factor to leave, so we end up with an empty A site
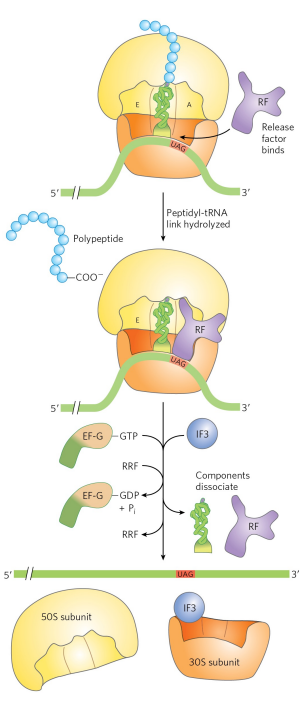
Explain how termination of protein synthesis occurs in bacteria
There are 3 distinct proteins that recognize the 3 distinct stop codons (humans have 1 protein that recognizes the stop codons). Stop codon is recognized by release factor. Release factor fits into A site and recognizes stop codon, and it brings water to hydrolyze the last peptide. Ribosome recycling factor (RRF) and energy from GTP hydrolysis aid in ribosome dissociation. IF3 promotes dissociation of the tRNA. The complex of IF3 and the 30S subunit is then ready to initiate another round
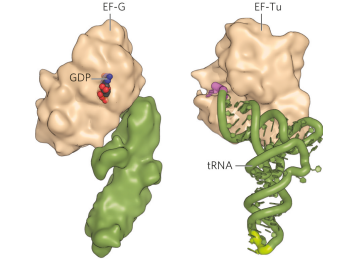
what is the similarly between the EF-G and EF-Tu-tRNA complex?
the structures of both are very similar.
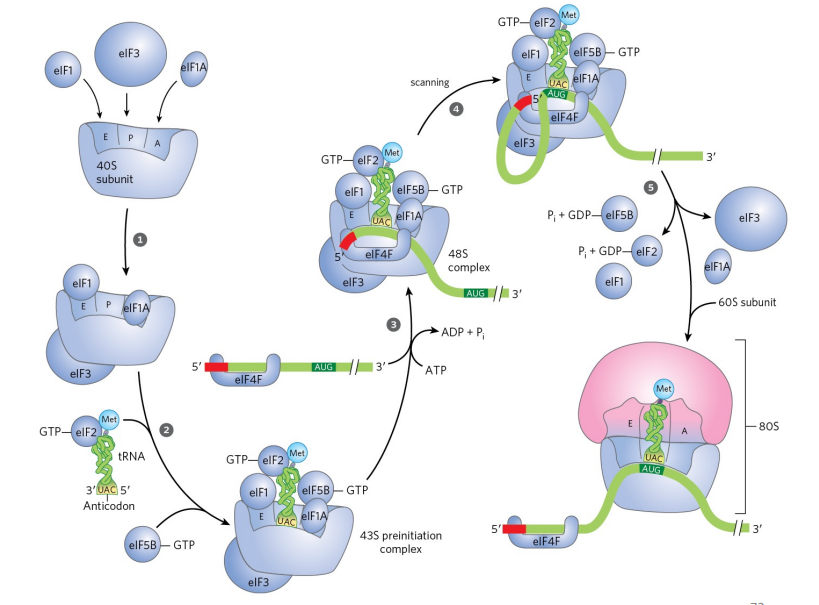
Explain how eukaryotic protein synthesis is initiated
40S subunit is protected by initiation factors
first, tRNA with methionine is loaded into 40S subunit (small subunit) at P site (no formulation of methionine required, and no Shine-Dalgarno)
5’ cap and 3’ poly(A) tail are recognized. Recognition is required so that the ribosome doesn’t waste its time on something that’s fragmented (want a full mRNA)'
the mRNA & protein gets loaded up onto the small subunit (ATP hydrolysis required)
RNA gets pulled out (scanning) when first AUG is hit, we get the W-C interactions (no wobble interactions with methionine).
after GTP hydrolysis, elongation factors leave and the big subunit gets added
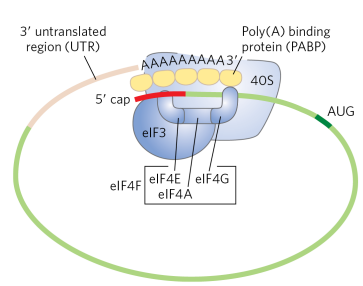
How does circularization of mRNA help in eukaryotic translation?
The small ribosomal subunit, along with initiation factors, binds both the 5’ cap and the polyA tail to align the mRNA at the correct AUG start codon. This ensures only intact mRNAs are translated (quality control).
True or False: Formation of a circularized mRNA molecule is conserved between Prokaryotes and Eukaryotes
false