MCB 2210 L14: Protein Targeting: Endoplasmic Reticulum Part 1
1/19
Earn XP
Description and Tags
Slides 1-33
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
Describe where and how protein synthesis primary occurs + main pathway
Protein synthesis @ cytosolic/free ribosomes
Has targeting sequence (address)?
NO → stays in cytosol
YES → guided to specific destination (Mitochondrion, nucleus, chloroplast, endoplasmic reticulum)
Main pathway = ER/Secretion
HAS signal peptide (address)
Co-translational process
Ribosome → ER WHILE protein synthesis occurs
ER = moving bilayer
Protein pushed through pore (translocator) into ER lumen
ER = moving proteins
Protein fold inside ER → vesicles → Golgi apparatus
Golgi sorts protein to final destination (Plasma membrane/lysosome)
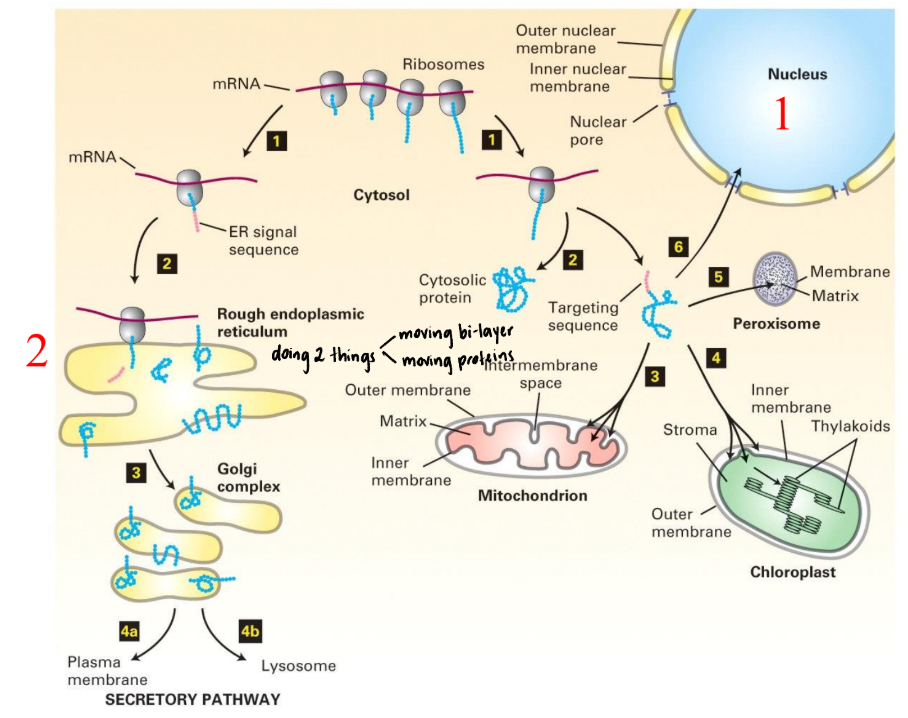
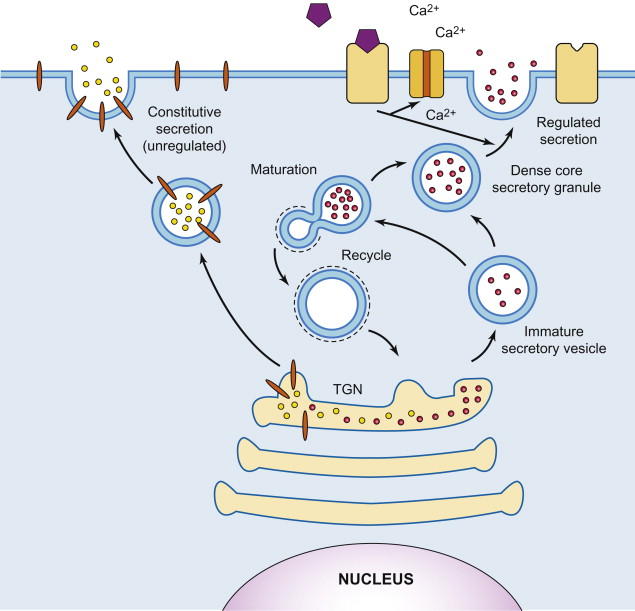
Where are lipids and proteins synthesized in the ER sent? What is this called?
Lipids & proteins synthesized in ER → secreted to plasma membrane/outside world
Constitutive secretion: continuous, unregulated, & default pathway present in all eukaryotic cells that transports proteins & lipids from Golgi apparatus directly to plasma membrane for release
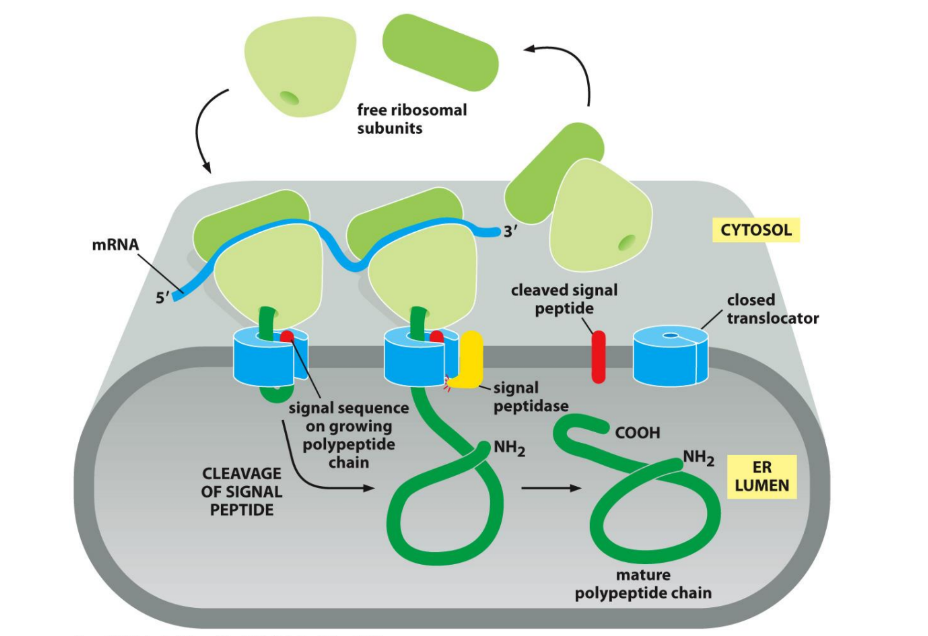
How are proteins targeted to the ER?
Signal sequences @ beginning of protein translation
Signal Recognition Particle (SRP) binds to sequence → PAUSES translation → ribosome directed to ER membrane
SRP binds to SRP receptor on ER membrane → ribosome → translocon
Translocon = protein channel in ER membrane = gateway for newly made proteins
SRP releases ribosome + protein complex → translation resumes → chain pushed through translocon channel into ER lumen or membrane
Signal sequence clipped by signal peptidase → finished protein released into ER by ribosome
What are the 3 ways proteins are moved between compartments within a eukaryotic cell?
Gated Transport = Cytosol → Nucleus (Nuclear Pore Complexes)
Traffic = bidirectional
Transmembrane Transport = Cytosol → non-nuclear organelles (Mitochondria, Peroxisomes, Plastids, ER)
Proteins = unfolded → threaded through protein translocated in organelle’s membrane
Vesicular Transport = Secretory Pathway
Protein enters ER → carried through vesicles from:
ER → Golgi → Late Endosome/Lysosome or Secretory Vesicles → Cell Surface
Protein never re-enters cytosol (protein stays inside vesicle)
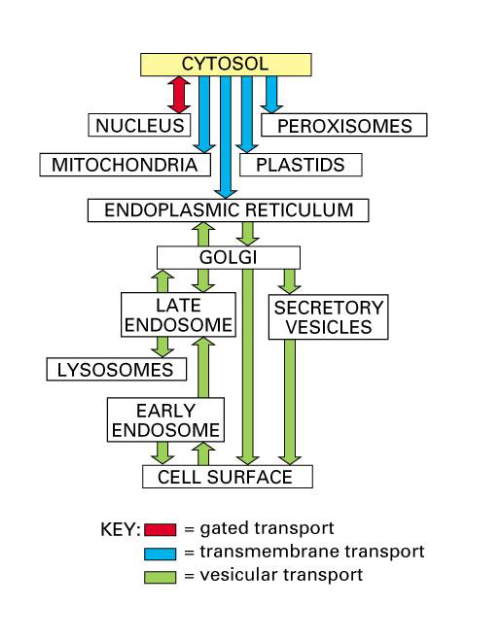
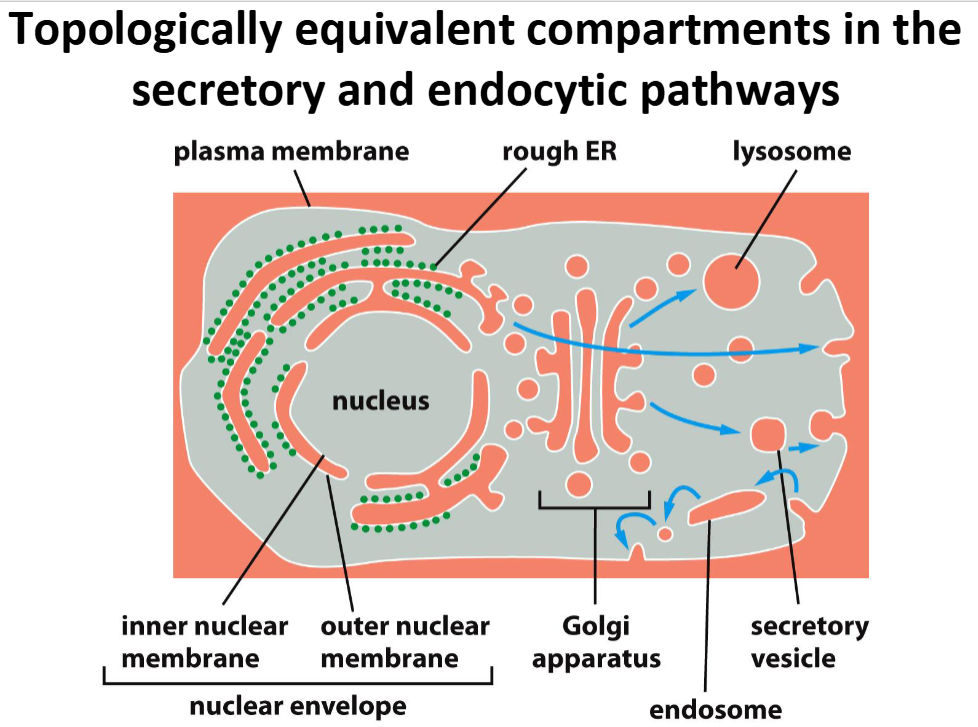
Which cellular compartments are considered “topologically equivalent” to the extracellular space, and why does this matter for vesicular transport?
Topology equivalent compartments = compartments where molecules can move b/w them w/o having to cross a physical membrane
“Equivalent” Compartments = nuclear envelope, lumens of ER, Golgi apparatus, endosomes, lysosomes, and peroxisomes
Form endomembrane system = collection of interrelated membranes & organelles that work together to synthesize, modify, package, and transport proteins & lipids
Vesicular transport = allows proteins to move between “topologically equivalent” compartments
Secretory vesicle fuses w/ plasma membrane → vesicle’s interior = cell’s exterior
What are the 2 functions of the Endoplasmic Reticulum (ER)?
Synthesis for endomembrane system
Proteins in plasma membrane, transmembrane, GPI-linked (lipid-anchored), soluble proteins in lumen of endomembrane system
Storage
Calcium (Ca2+)
Enzymes (some cell types)
Ex. detoxifying enzymes in liver cells
ER as a dynamic structure
ER = moving network of tubules & sheets
Stretches from nucleus to outer periphery of cell (plasma membrane)
During cell division:
ER breaks into → vesicles → split into daughter cells
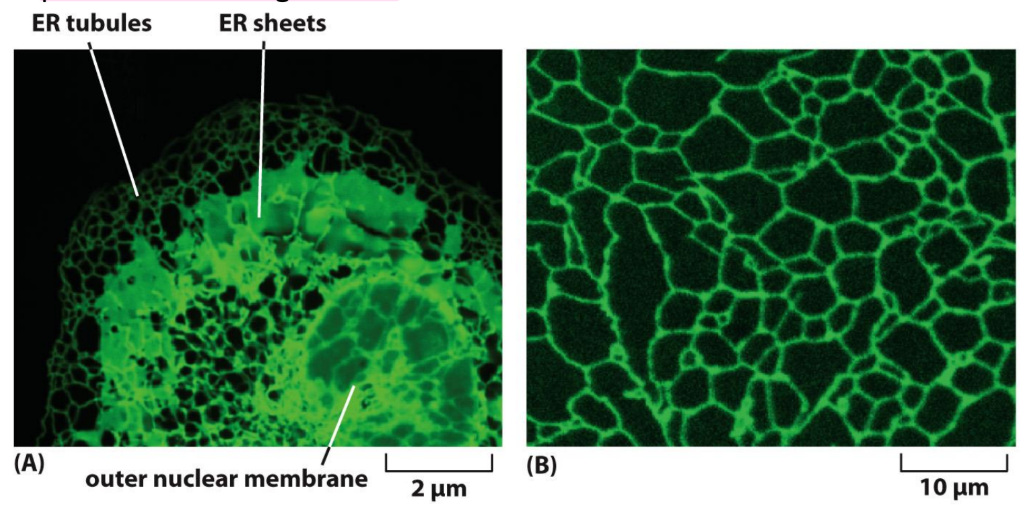
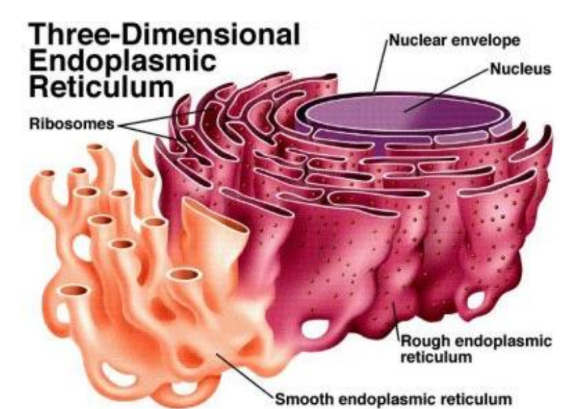
Endoplasmic reticulum structure + location within cell
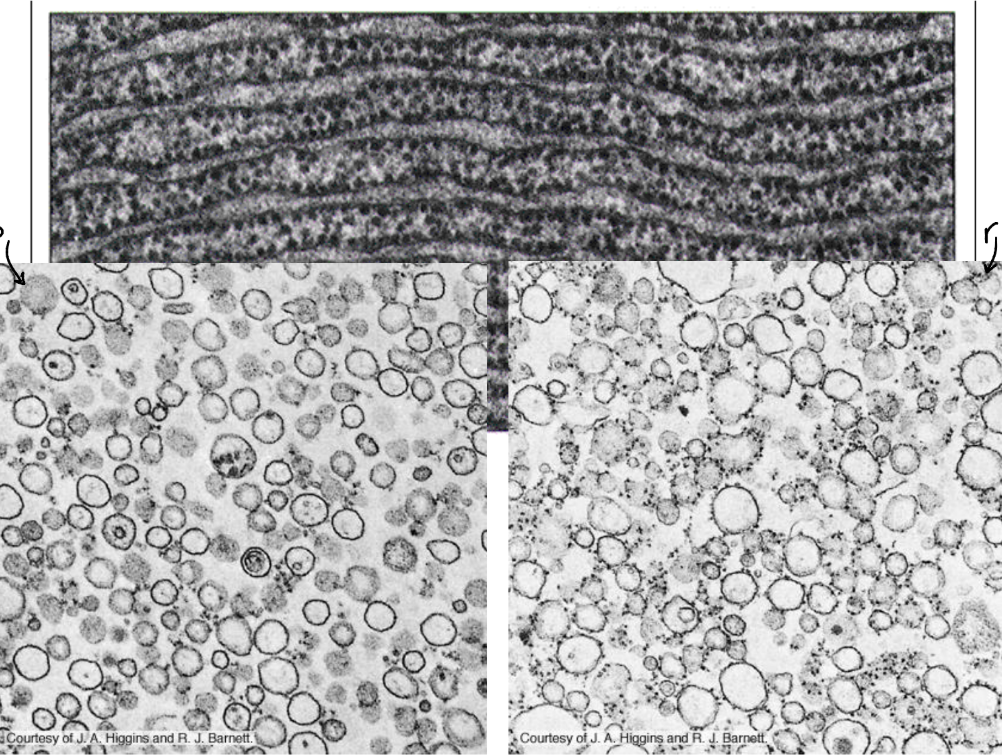
Two types of ER membranes (viewed through electron microscope)
Rough ER = studded with ribosomes → rough → produces & process proteins
Closer to nucleus
Smooth ER = NO ribosomes → smooth → produces lipids, steroids, & detoxifies drugs
Peripheral (outer) regions of cell
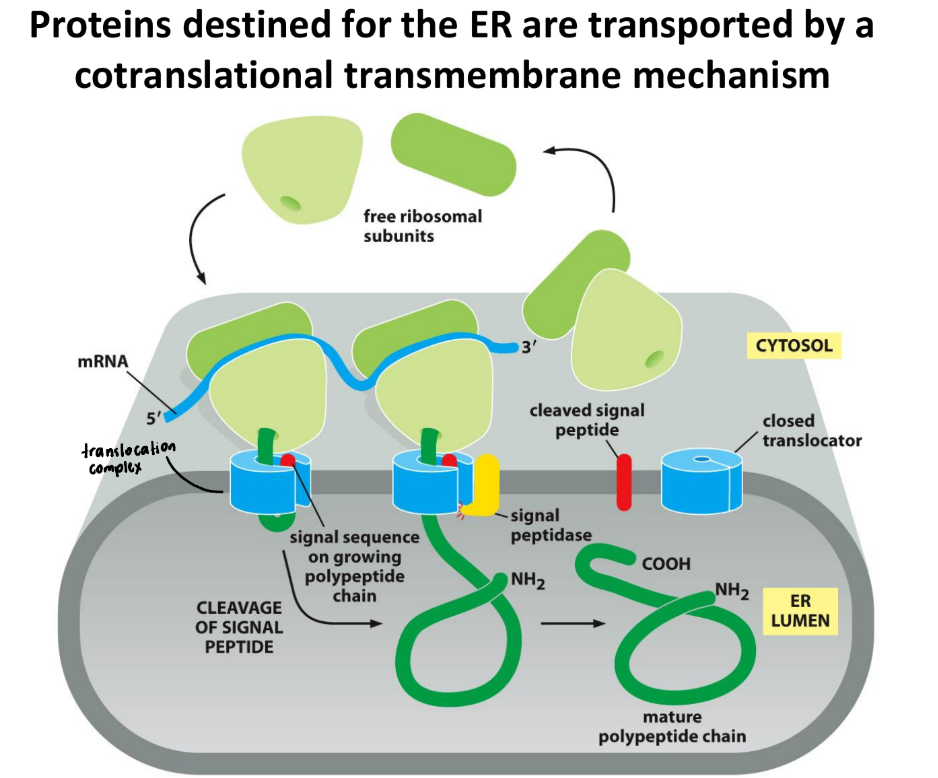
How are ribosomes bound to the Rough Endoplasmic Reticulum membrane?
Ribosomes bound through nascent chains
Nascent chains = protein molecule in process of being synthesized by ribosome
Proteins = co-translationally inserted into membrane
Signal Recognition Particle (SRP) recognizes specific signal sequence on protein → pauses translation
Ribosome + nascent protein chain directed to SRP receptor on ER
Ribosome = anchored to ER membrane by translocon
Translocon = protein channel in ER membrane
Translation resumes when ribosome = bound to translocon
Pushes protein directly into lumen → embedded in ER membrane
Describe how a cell sample is prepared for gradient centrifugation and the two types of ER membranes that can be separated
Mechanically break cell → microsome formation
Microsomes = small, artificial vesicles created when cells are broken in lab (formed from ER fragments)
Rough Microsomes = DENSE due to ribosomes
Smooth Microsomes = less dense due to lack of ribosomes
Centrifuged in sucrose density gradient
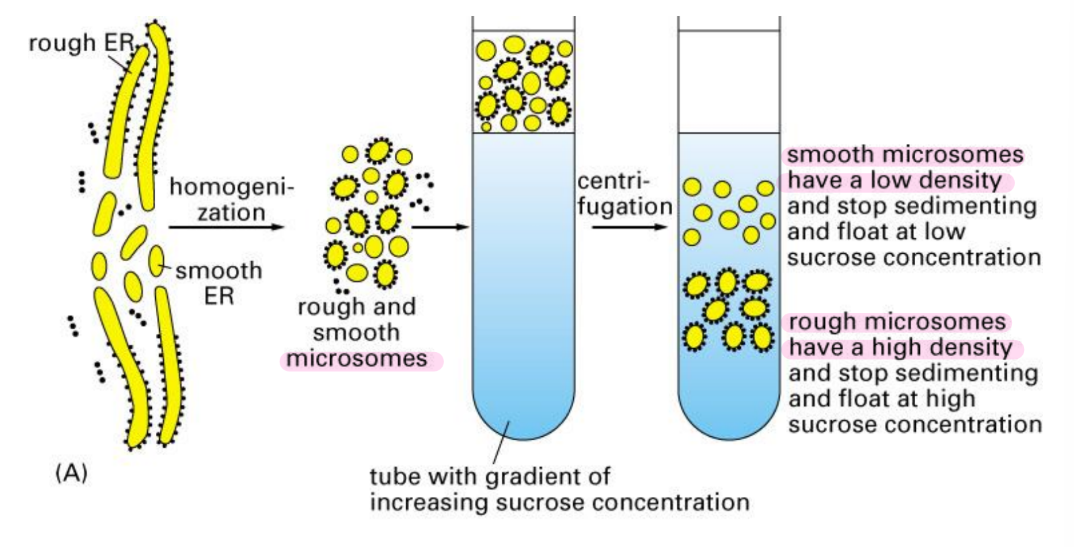
Microsome
Fragments of rough or smooth ER that seal themselves into tiny, artificial vesicles when cells are mechanically broken in a lab.
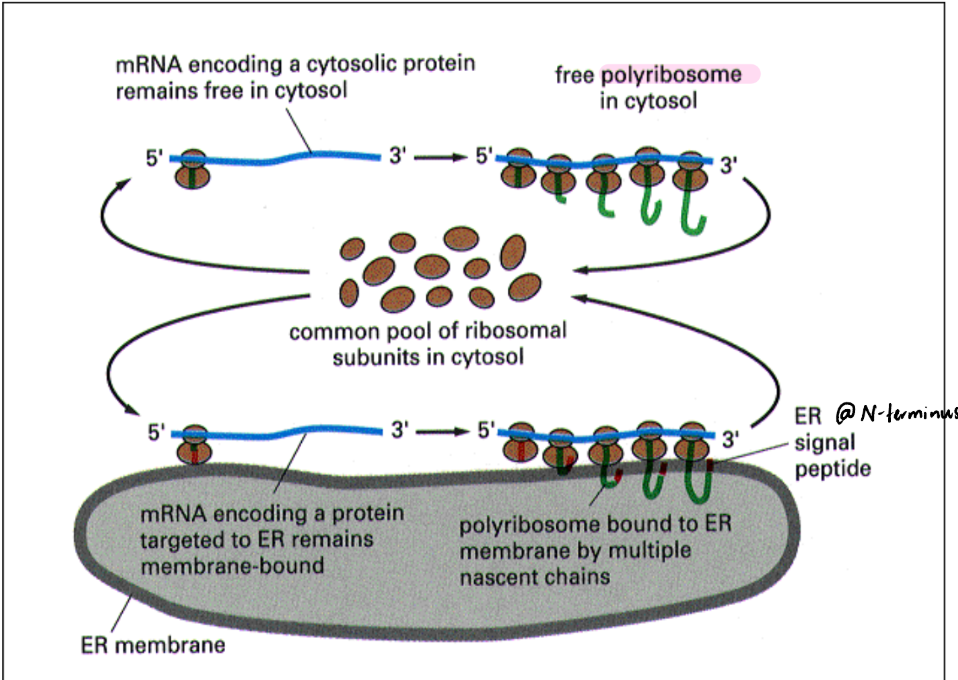
Free vs. Bound Polyribosomes
Protein translation = slow process
mRNA = long
Multiple ribosomes can attach to a single mRNA = polyribosome/polysome
Polysome = single mRNA molecule being read by multiple ribosomes simultaneously. Appears like “beads on a string”
Protein HAS ER targeting signal ribosome & growing peptide chain (nascent chain) go to ER membrane → bound polyribosome/polysome
Difference b/w free vs. bound polyribosomes = localization & peptide synthesized
Rough Endoplasmic Reticulum (RER) = efficient
mRNA near membrane (many ribosomes = recycled)
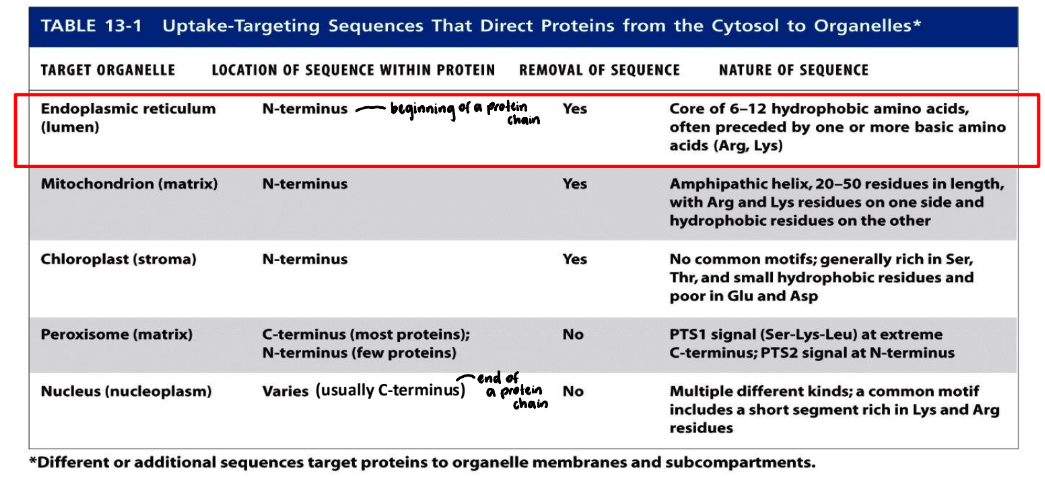
ER signal sequence
ER signal sequence = hydrophobic amino acids
N-terminal signal sequence
Sometimes label = internal for membrane proteins (hydrophobic)
Necessary & sufficient for ER membrane targeting
Only need sequence → ER membrane
Co-translational translocation across ER membrane = protein transported to ER as protein = translated
Signal peptidase = cleaves ER signal sequence after targeting
Signal ≠ found in mature protein
Protein = folds in lumen w/ chaperones + other molecules
Protein = free → soluble in ER lumen OR bound to ER membrane
What is ER targeting signal recognized by?
Signal Recognition Particle (SRP): factor that allows attachment of pure ribosomes to ER membranes
Ribonucleoprotein particle = RNA + 6 proteins
Binds to signal sequence & ribosome
Binding of SRP to nascent peptide chain → stops translation (peptide = 70-100 AA long) = long enough to protrude from ribosomes
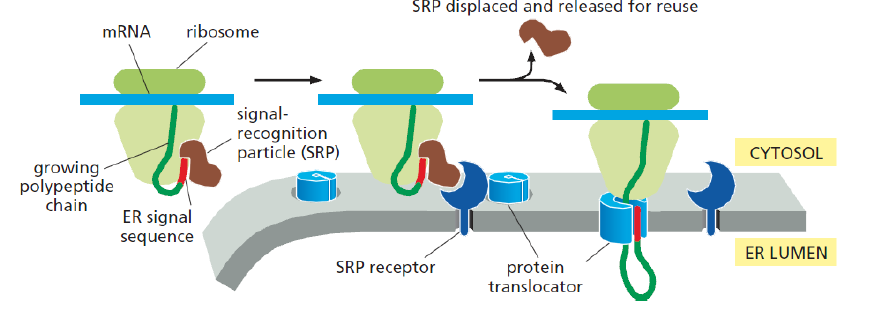
Co-translational translocation of proteins → ER
Signal sequence = out of ribosome
Associates w/ Signal Recognition Particle (SRP)
Binds to signal sequence + ribosome
SRP associates with SRP receptor in membrane
Ribosome docks onto translocon channel
Protects proteins from protease in assays
Translocon pore though membrane opens (stays closed to prevent leakage of small molecules)
Signal sequence + nascent chain → pore
SRP + SRP receptor RELEASED from ribosome & translocon
Translation proceeds through membrane pore → protein enters lumen
GTP binding/hydrolysis of BOTH SRP + SRP receptor regulate:
SRP/SRP-receptor binding
Ribosome docking
SRP/SRP-receptor release
Process is not well known
Signal peptidase cleaves N-terminal ER signal sequence
Protein synthesis = completed (ribosome docked)
Translocation channel closes → ribosome undocks
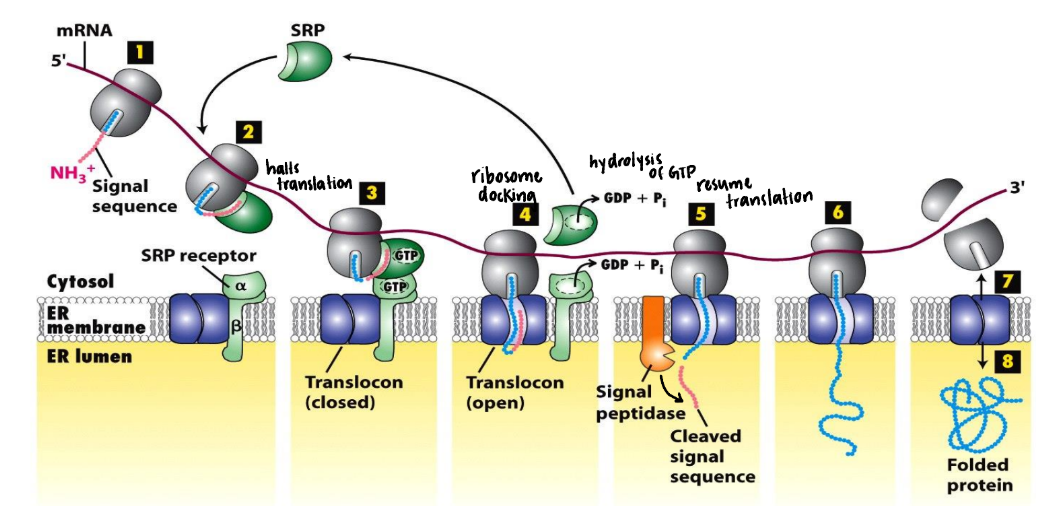
How do you demonstrate co-translational transport?
Use microsomes & protease protection experiments → determine if proteins = exposed
Mix mRNA encoding ER-targeted protein + ribosomes + ATP + tRNAs + Amino Acids = synthesize protein (in vitro or in lab)
During/after protein synthesis + protease → protein = degraded by protease (through gel)
During/after protein synthesis + protease + Microsomes → protein ≠ degraded
During/after protein synthesis + detergents → protein = degraded
Conclusion:
Proteins = protected during synthesis
Proteins = transported to lumen of membranes DURING synthesis → co-translational translocation
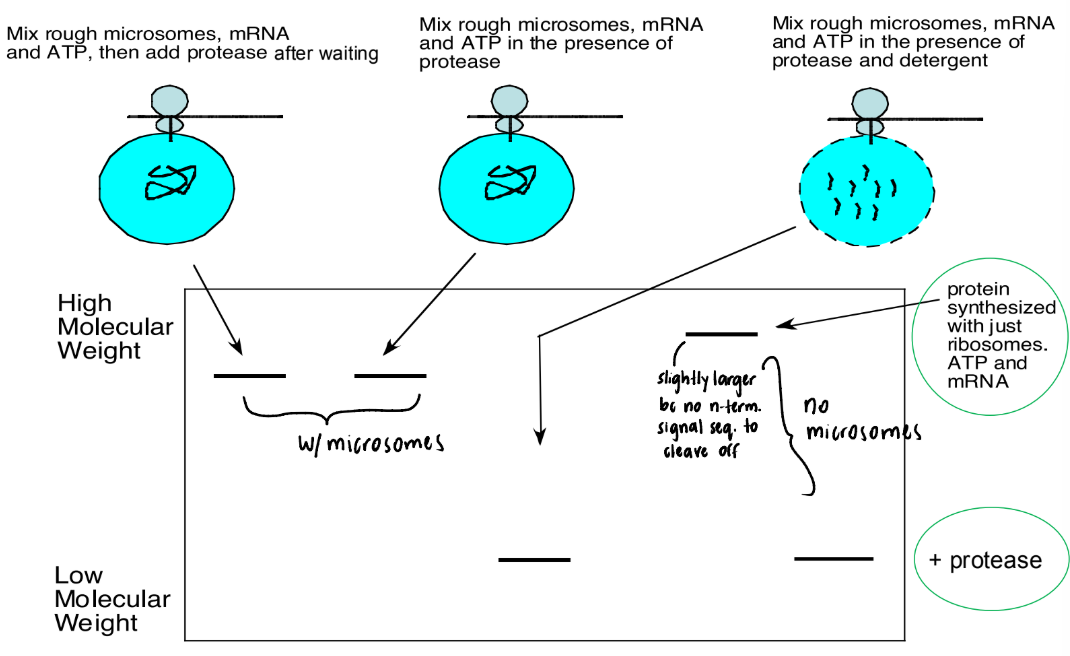
Explain what is happening in this picture
Lane 1 = Protein synthesized → microsome
Protein = safe from protease inside microsome
Lane 2 = protein synthesized while protease = present
Ribosome plugged into microsome → protein threads into microsome
Protein = safe from protease inside microsome
Lane 3 = detergent added → microsome membrane = dissolved
Protein = exposed ≠ safe from protease
Lanes 1-3 = smaller than Lane 4 (farther along lane) → N-terminus signal sequence = CLEAVED OFF by signal peptidase
Lane 4 = protein synthesized w/o microsomes
No microsomes = no signal peptidase → N-terminal signal sequence = STILL ATTACHED
Only appears if no protease is added
Lane 5 = + protease
No microsomes = protein = unprotected ≠ protected from protease
How is the full sequence of events of protein targeting to ER demonstrated?
By using purified components
A = (-) SRP, (-) SRP receptor, (-) microsomes
= Complete polypeptide w/ signal sequence
B = (+) SRP, (-) SRP receptor, (-) microsomes
= Polypeptide elongation paused @ 70-100 AA
C = (+) SRP, (+) SRP receptor, (-) microsomes
= Complete polypeptide + signal sequence + SRP & SRP receptor released from polypeptide (GTP hydrolysis)
D = (+) SRP, (+) SRP receptor, (+) microsomes
= Polypeptide synthesized in microsome through transmembrane channel + signal sequence = cleaved + found in microsomes
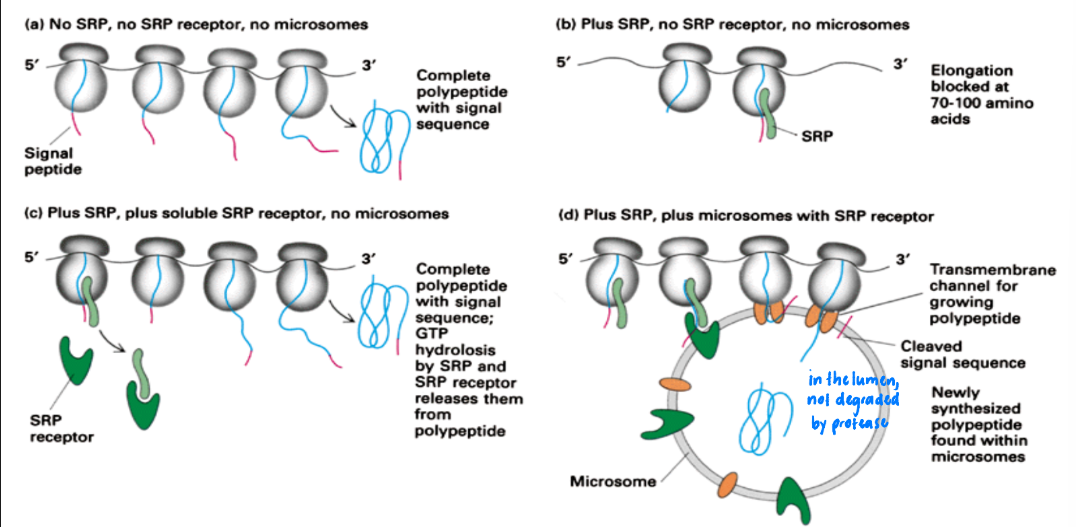
Purified components + protease protection
Protein Outcomes:
Ribosome → (+) full length protein, (-) protease protection
Protein = floats in liquid
Ribosome + SRP → (-) full length protein, (+) protease protection
SRP = pauses translation @ 70-100 amino acids
Ribosome + SRP + Receptor → (+) full length protein, (-) protease protection
SRP = released from protein
No membrane of microsome present to go inside
Ribosome + SRP + Microsome → (+) full length protein, (+) protein protection
Protein threaded into → microsome
Shorter because signal sequence = cut off (signal peptidase)
Stripped Microsomes (sMicro) = microsomes (-) ribosomes (-) SRP (+) SRP receptor (+) translocon (+) signal peptidase
SRP = chemically removed → Protein = outside (sensitive to protease)
Rough Microsomes = contain “hitchhiking” SRP
Protein can go inside (protected from protease)
SRP is always required for translocation
Signal peptidase
Protease that recognizes N-terminus signal sequence
Embedded in ER membrane & associated w/ translocon complex
Releases soluble proteins into lumen of ER
Can be secreted or reside in ER, Golgi, lysosomes (w/ appropriate signals)
Signal ≠ removed → protein stuck in membrane held by signal sequence