journal 2 post quiz
1/88
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
89 Terms

question
what is the open reading frame (ORF) seq for human SOX7 (=part of dna seq that does not have a stop codon so that rna code is continuously read) + compare to mouse sox7 to see if conserved

method
pcr

result
orf seq has hmg box. 87.4% similarity btn human and mouse sox7. 100% overlap for hmg box

control
mouse sox7 establish seq to compare human seq to

conclude
high fx similarity btn human and mouse. fx of mouse sox7 protein=fx of human sox7 protein. conserved

classify
establishing, expression****
question
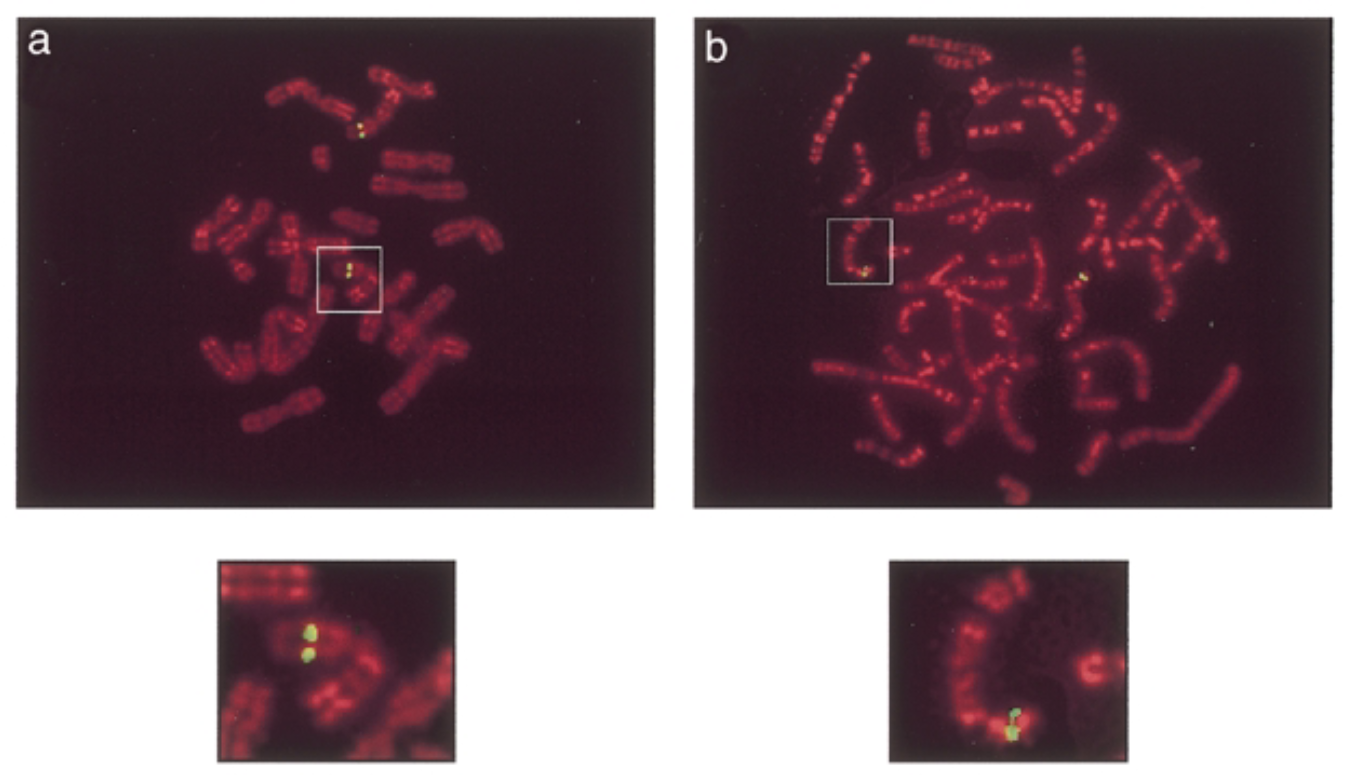
where are sox7 genes in mouse and human chromosomes
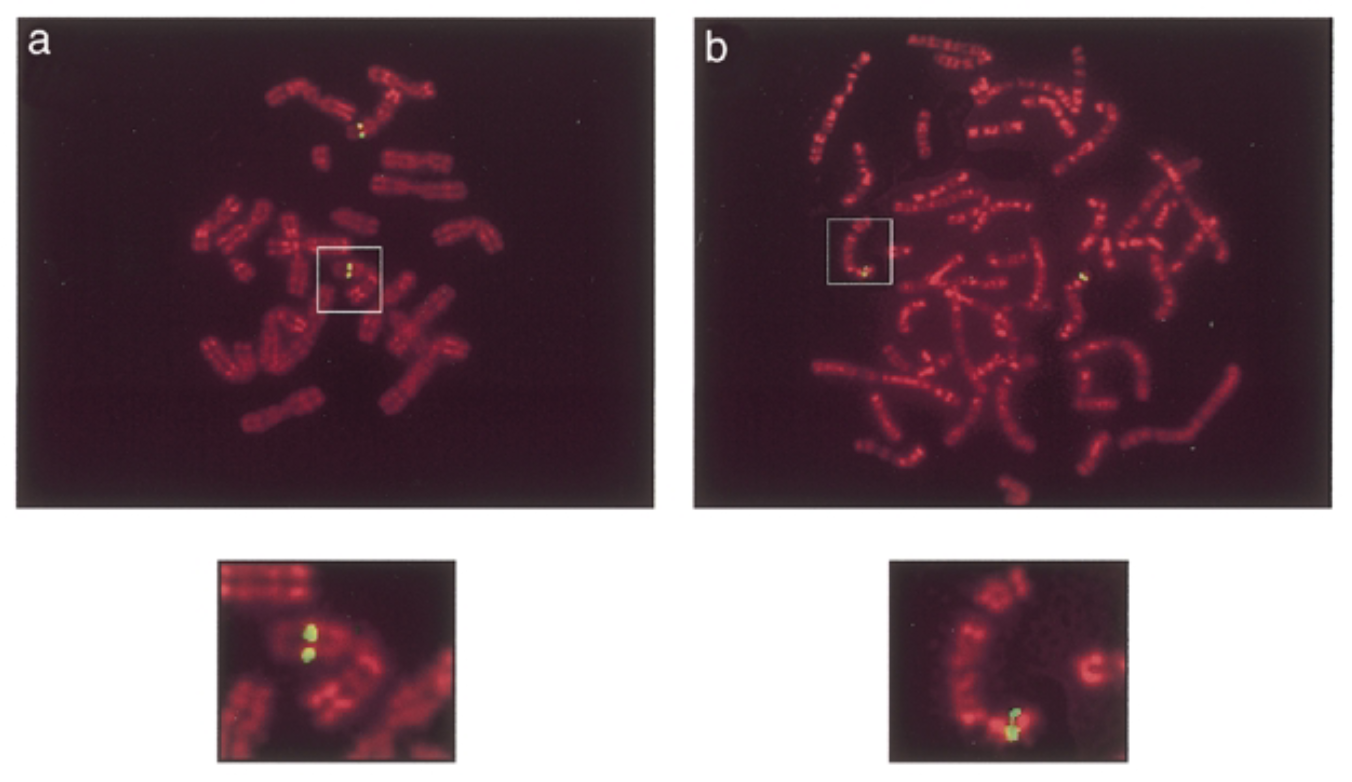
method
fish mapping
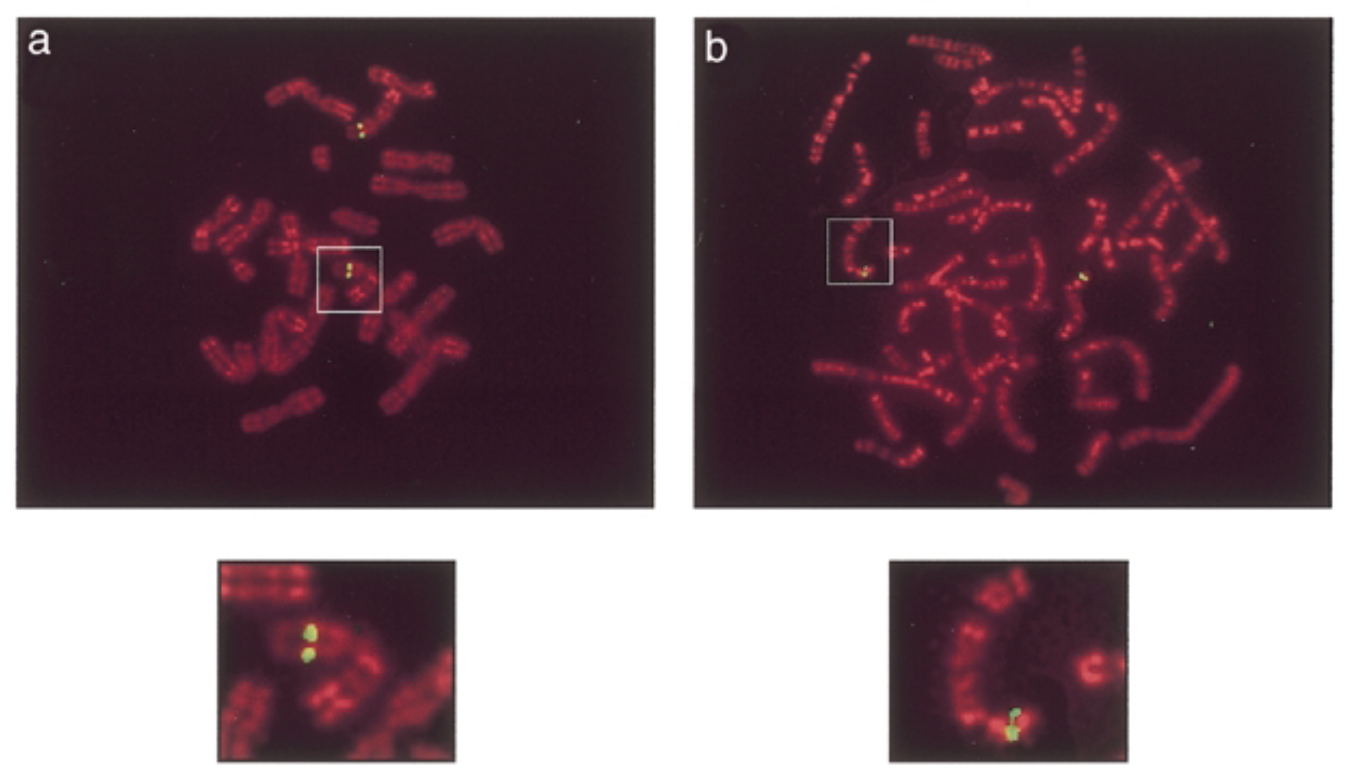
conditions
human bac and mouse pac
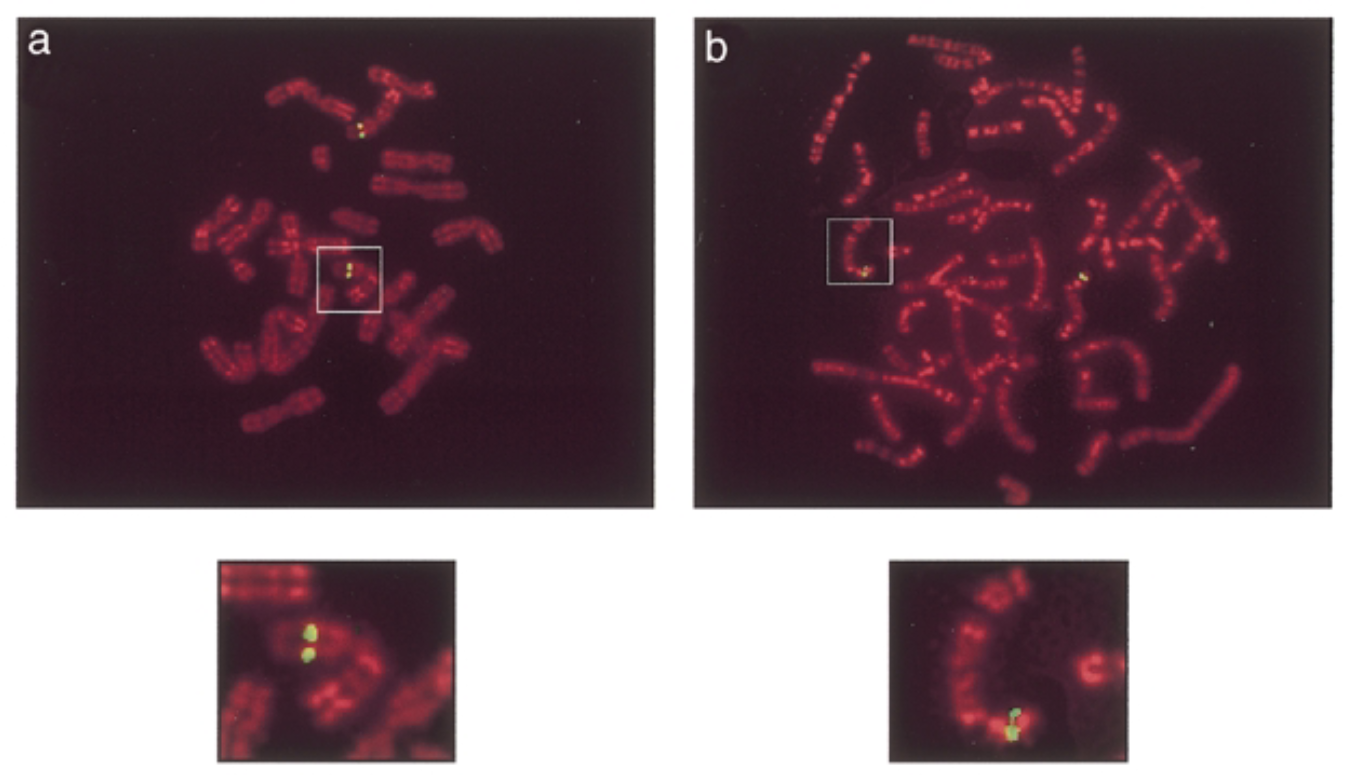
result
green signals: mouse sox7 at chromosome 14 band D. human sox7 at chromosome 8 p22 which is similar seq
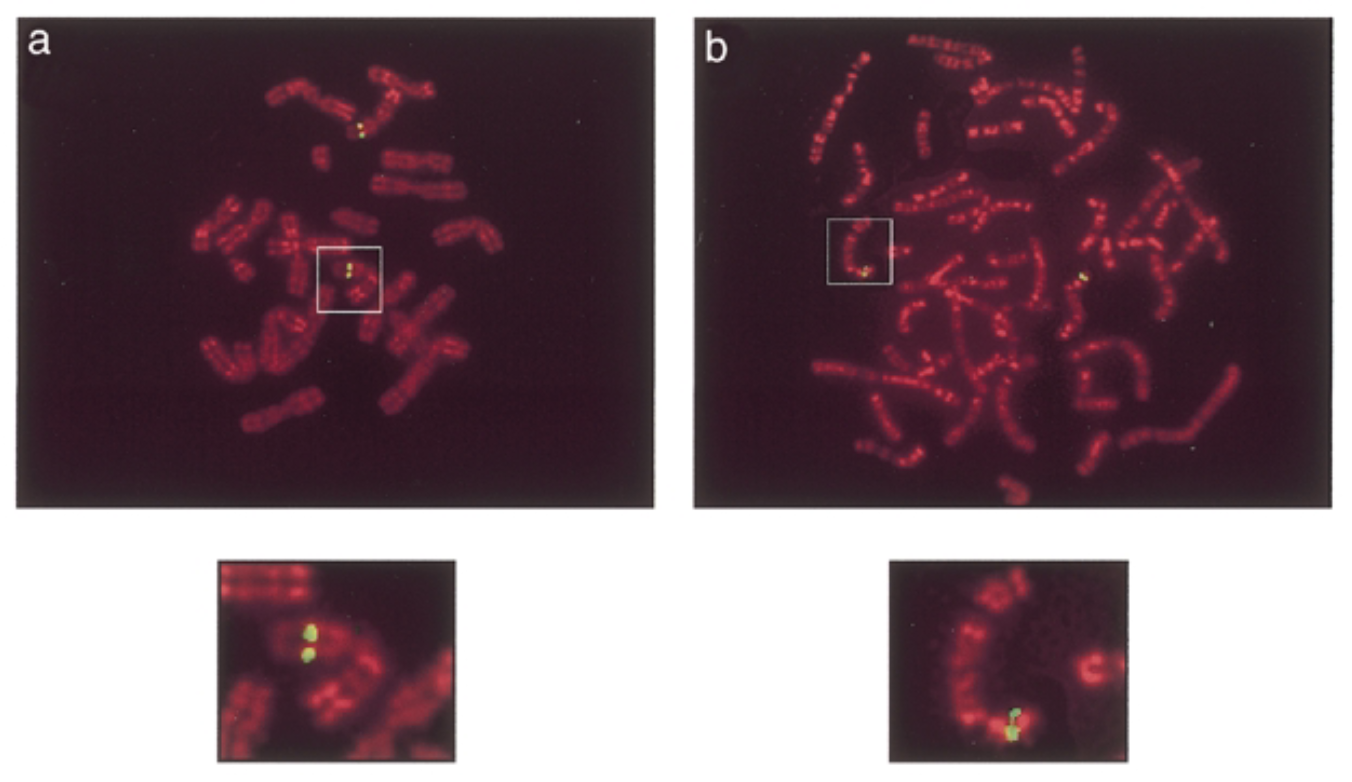
control
mouse pac establish baseline localization to compare human loc to
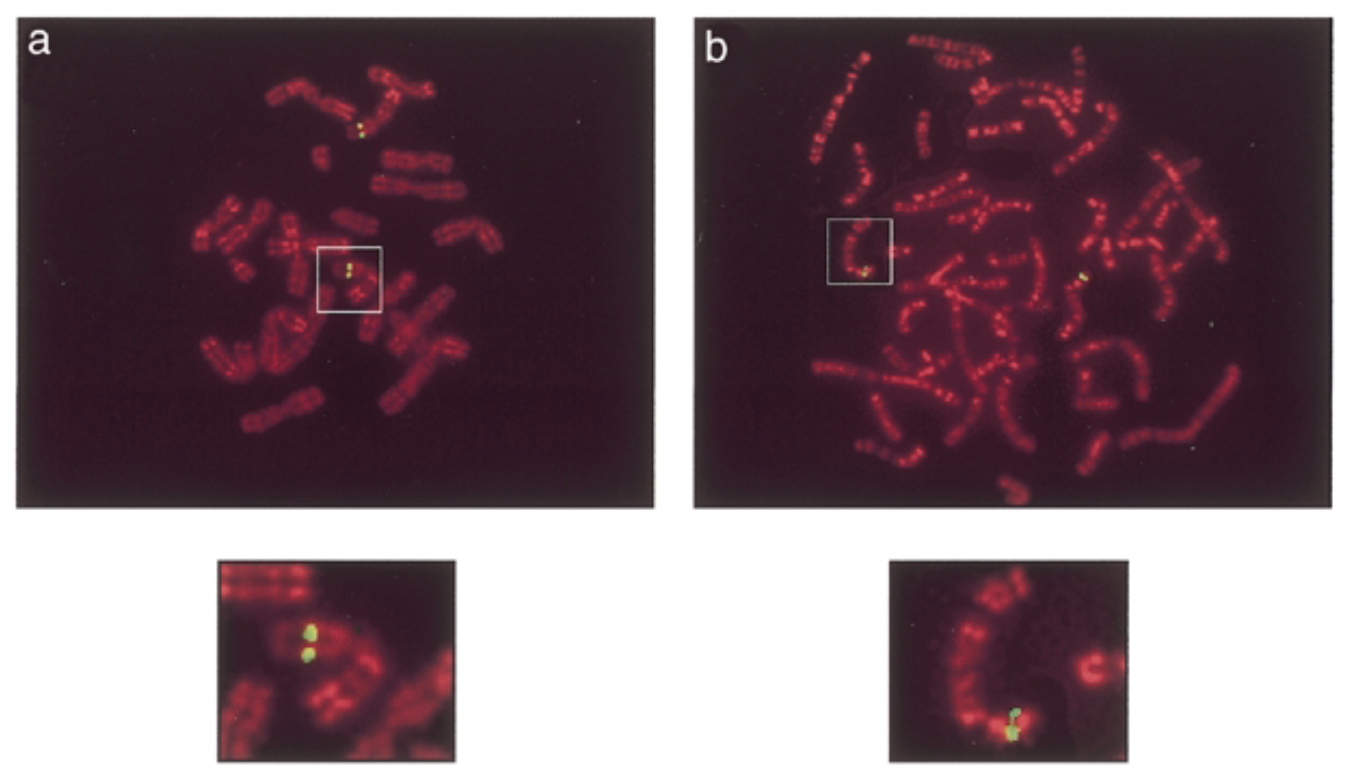
conclude
locs are similar. fx of mouse sox7 protein=fx of human. loc of sox7 gene on human is where syndromes→ sox7 associated with syndromes
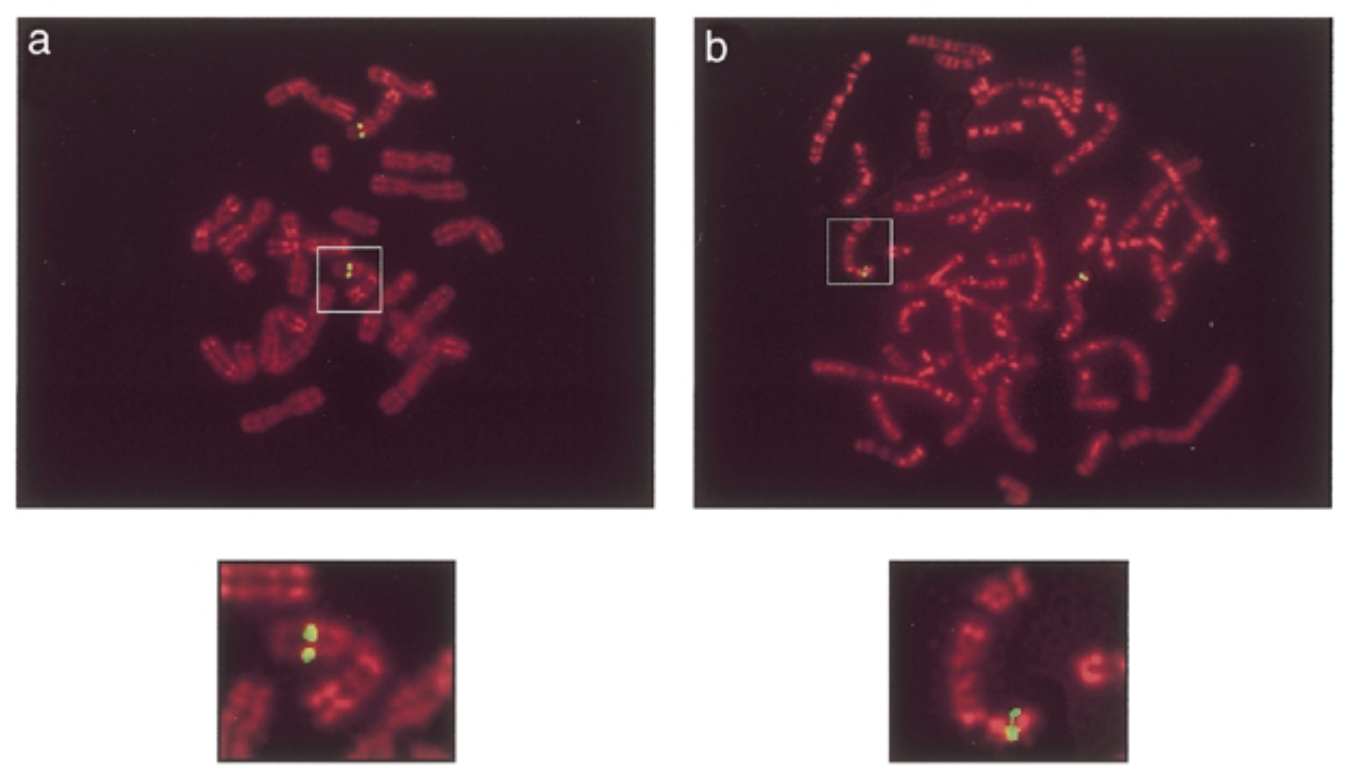
classify
establishing, other=flourescene ver of southern blot
question
how is sox7 mrna expressed in mice
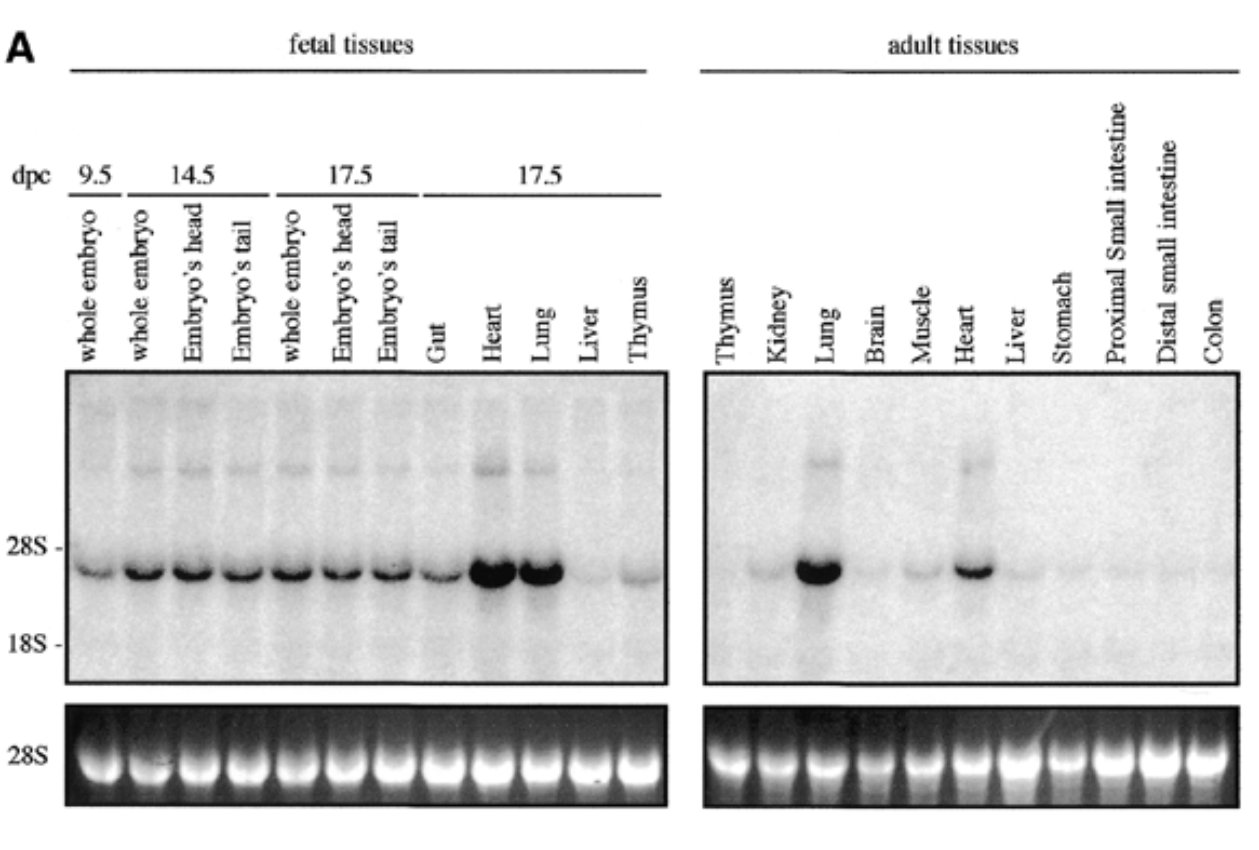
method
northern blot
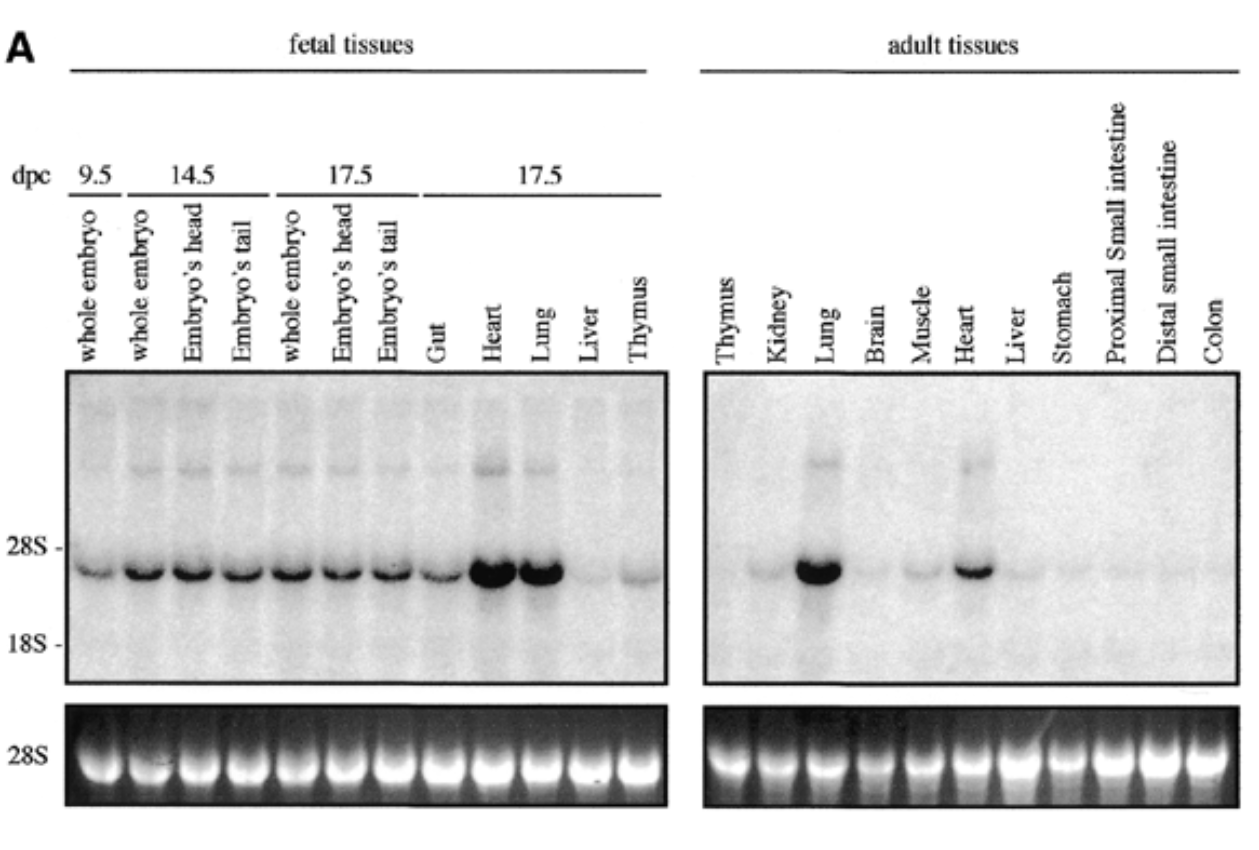
conditions
fetal and adult tissues of mice + probe
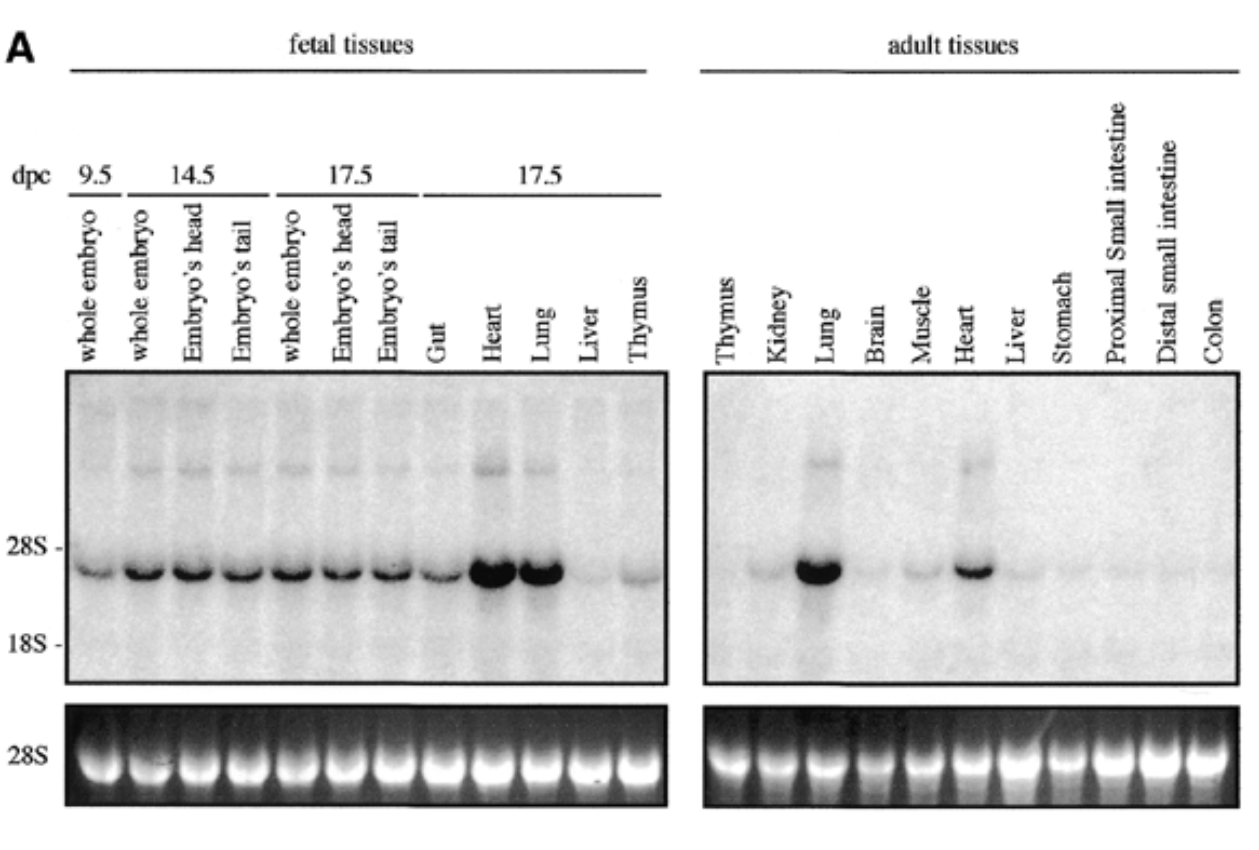
result
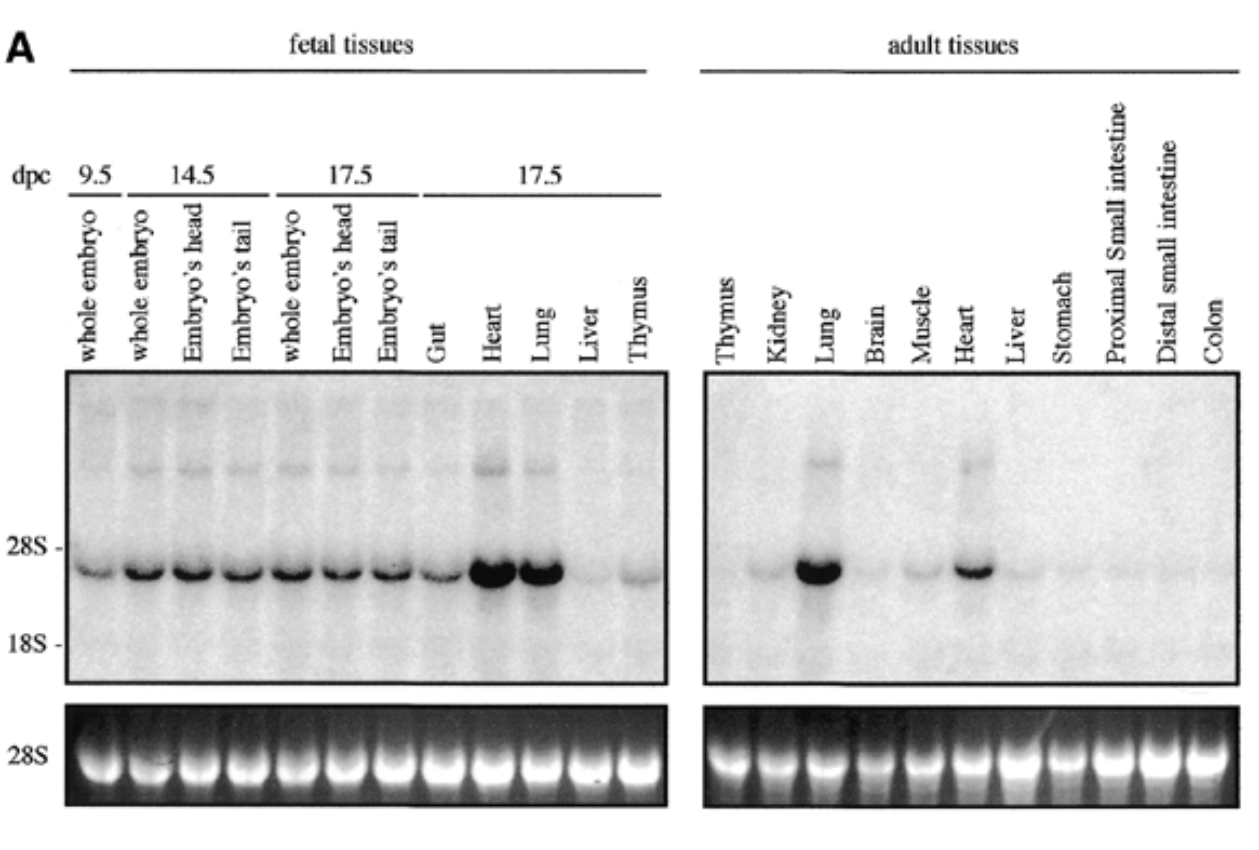
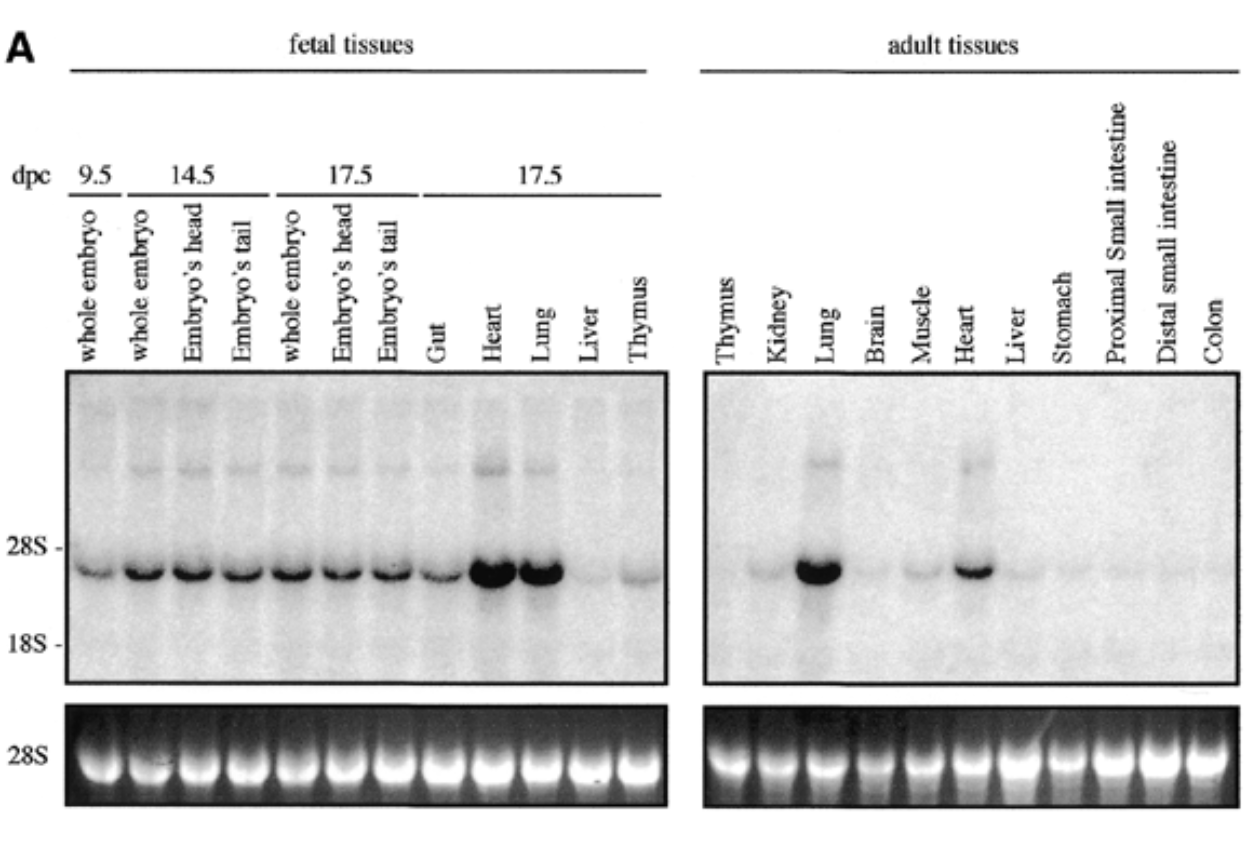
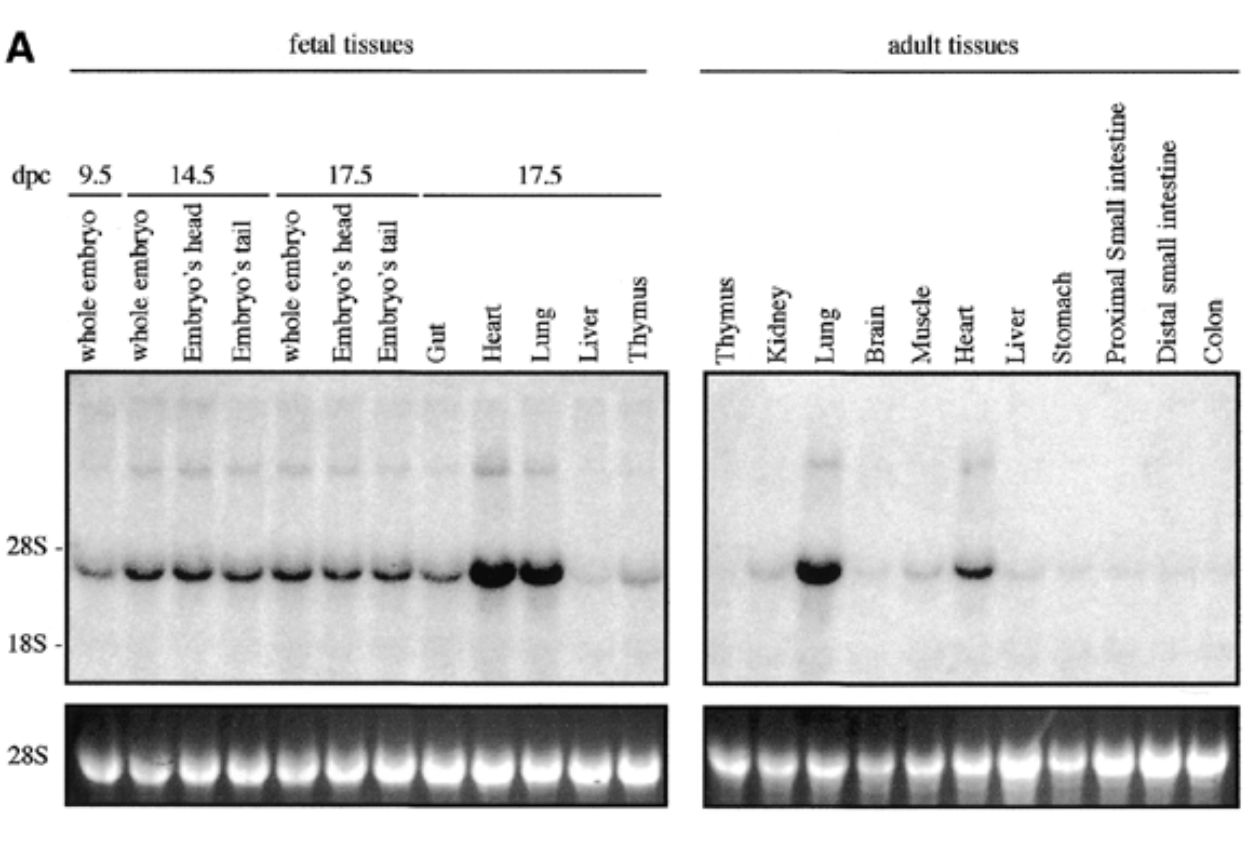
sox7 expressed in all fetal and adult mouse tissue at 4kb with higher exp in heart and lungs for both embryonic and adult
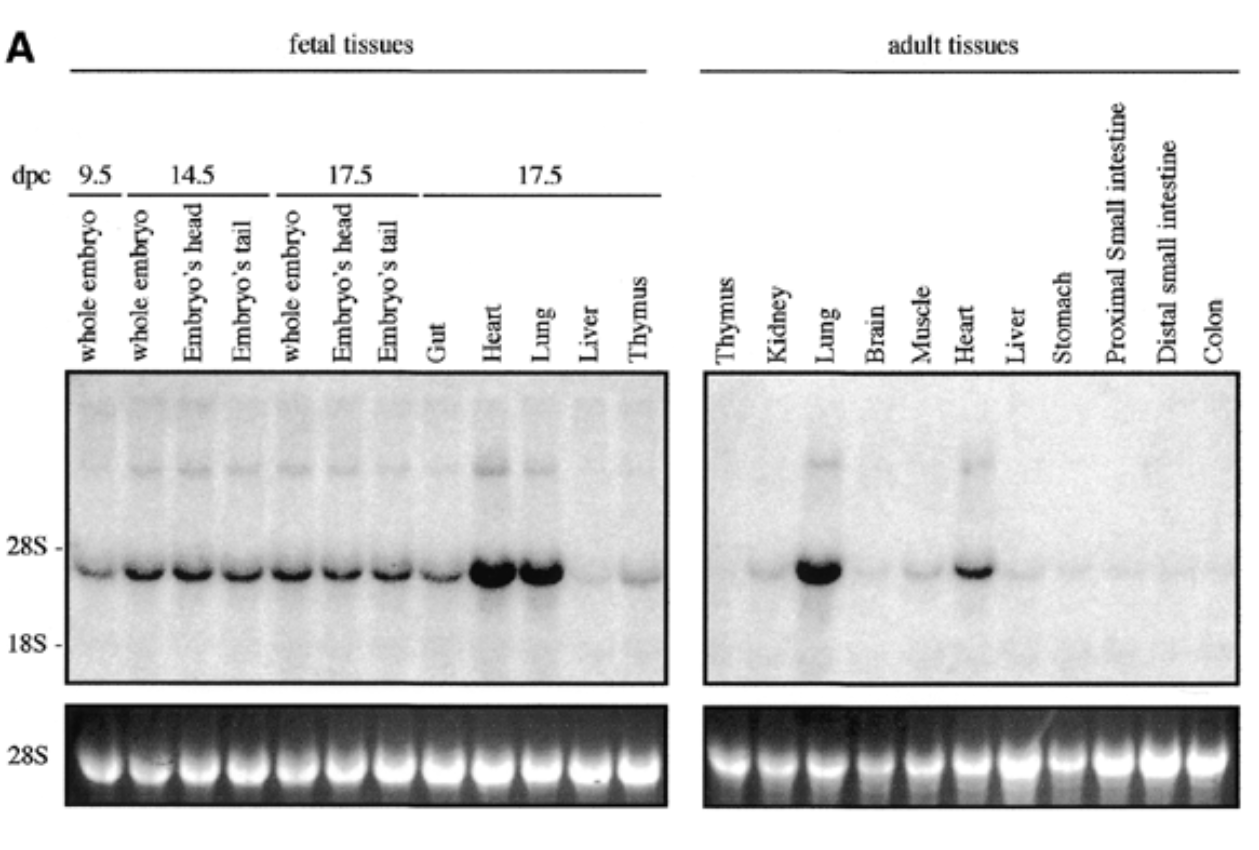
control
28s is rrna that is always exp at same levels → shows equal loading + ethidium bromide staining to show if equal amt of rna in each lane
conclude
sox7 plays a role in all tissues (contrary to prev study) but is particularly significnat in dev of heart and lungs
classify
hypothesis/establishing, expression
question
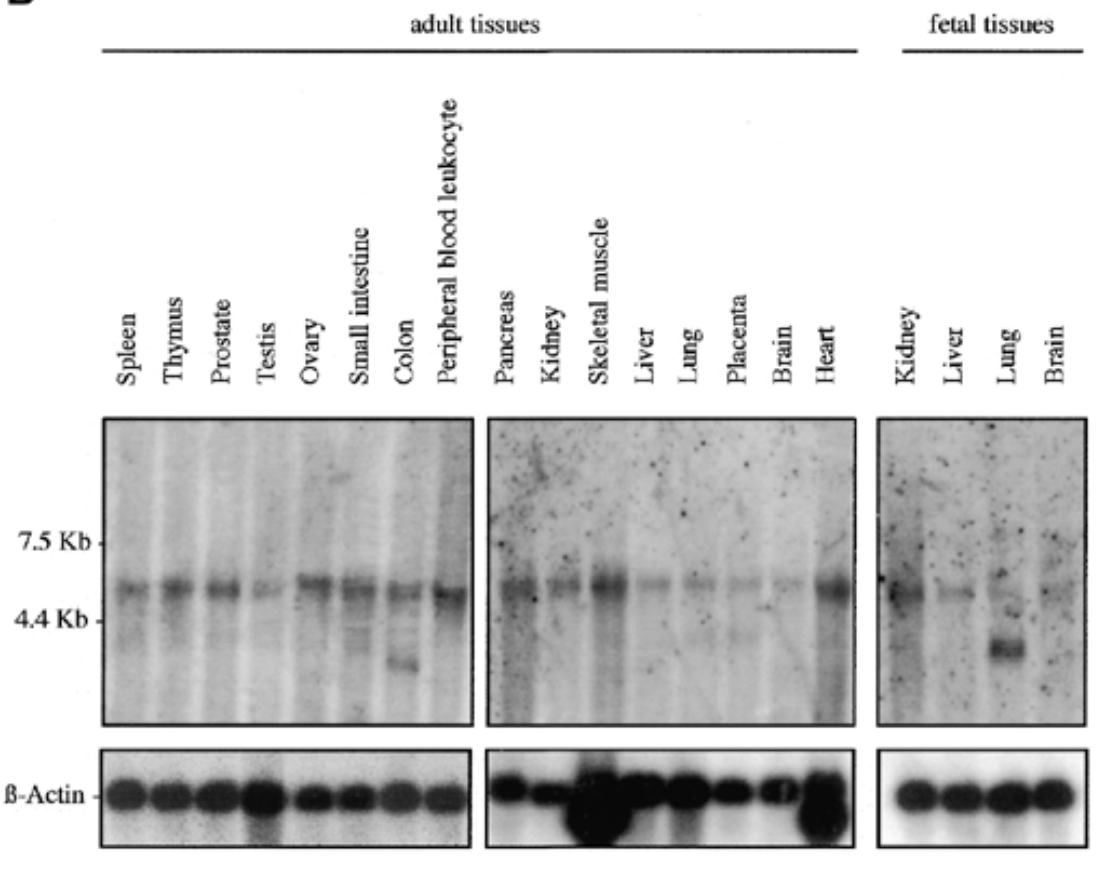
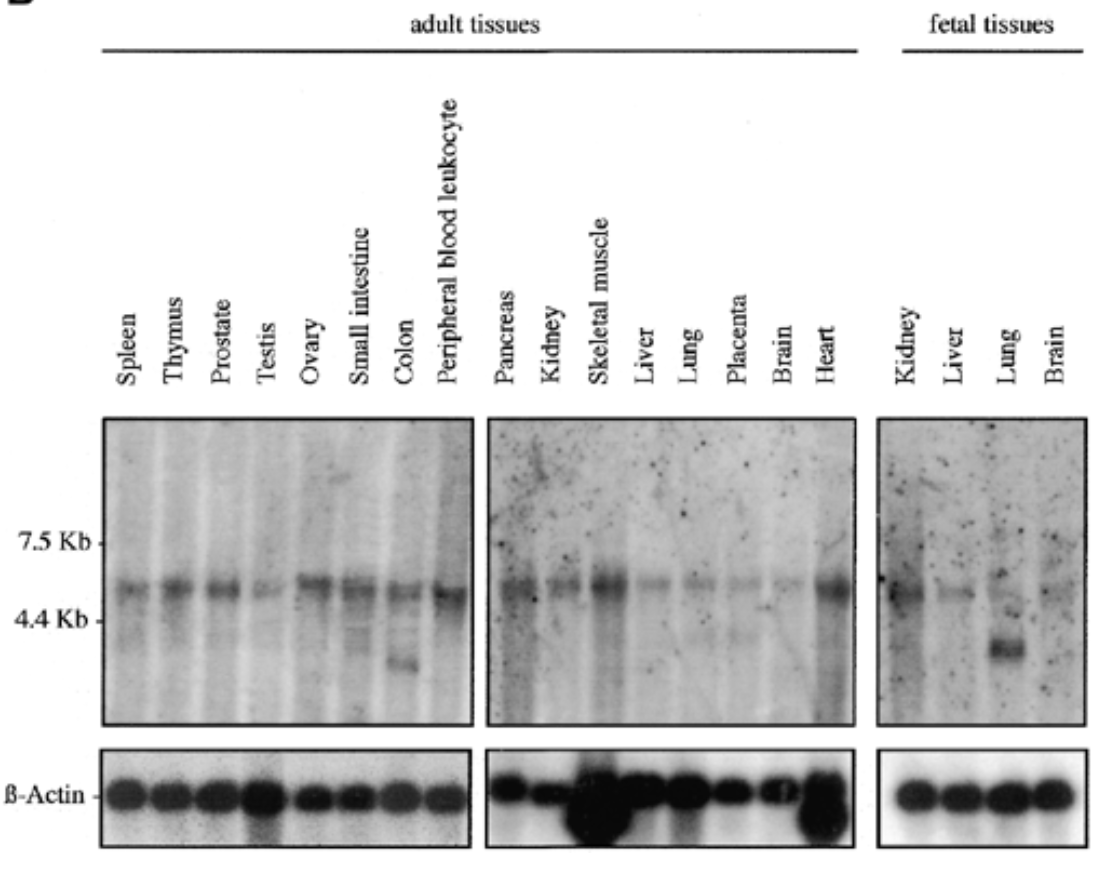
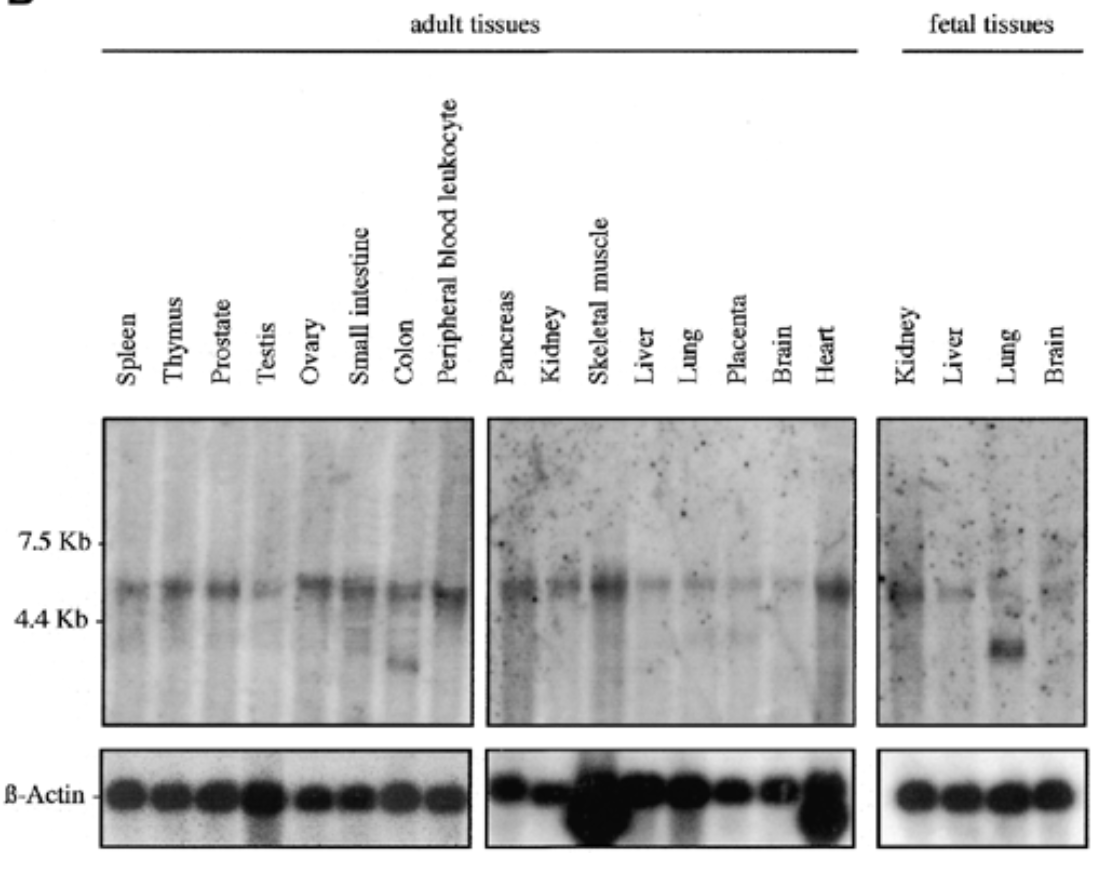
where is sox7 mrna exp in humans
method
northern blot
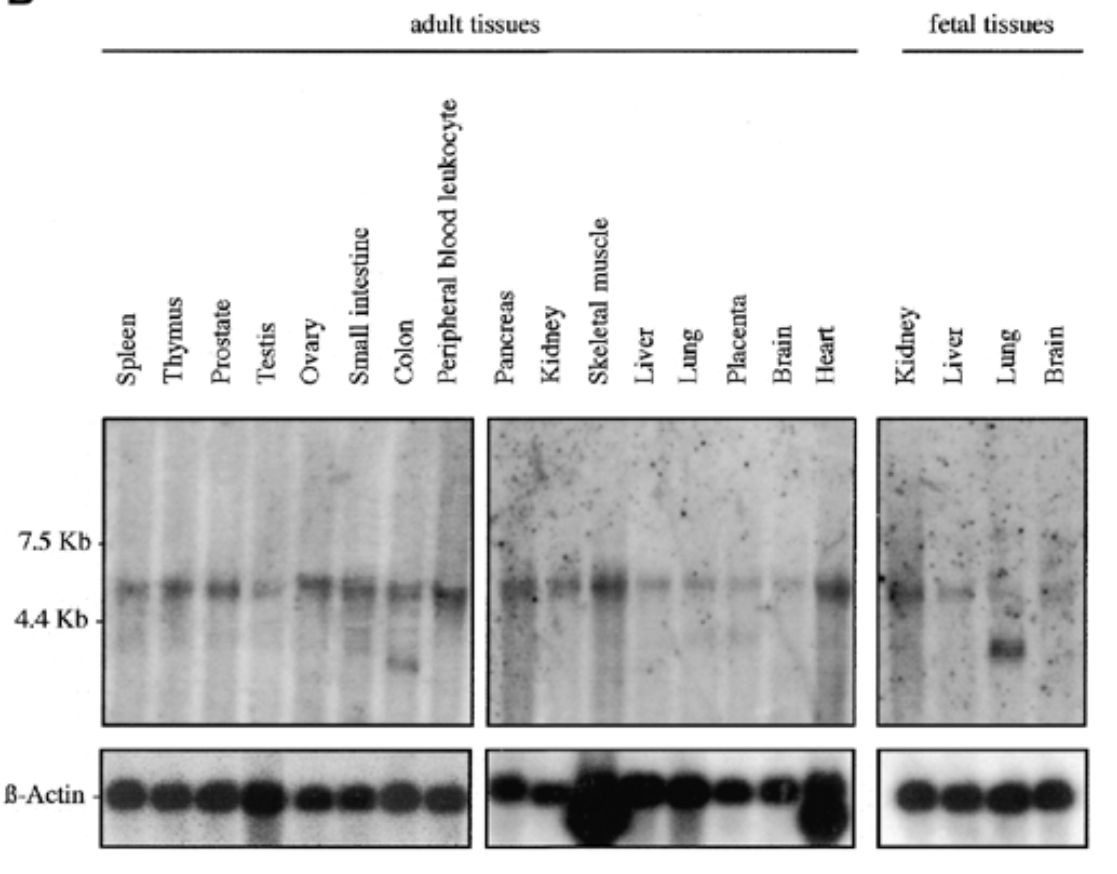
conditions
fetal and adult tissue
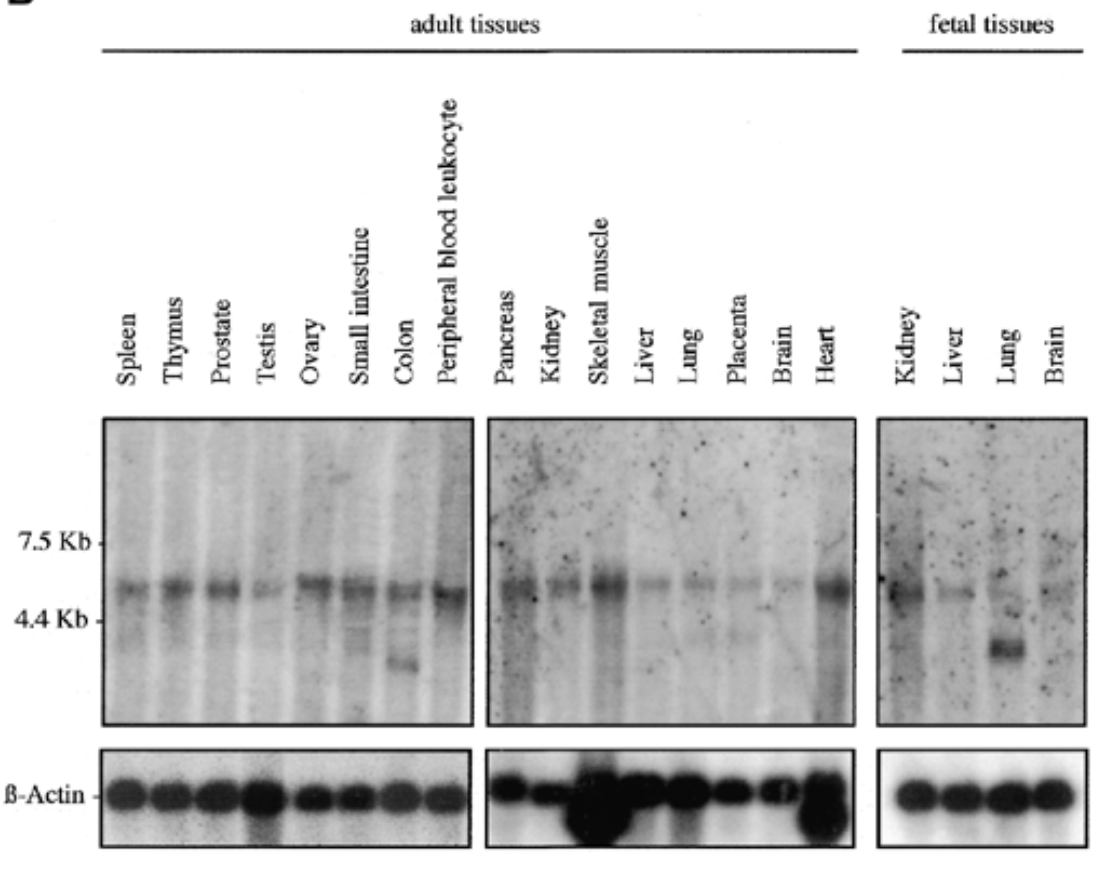
result
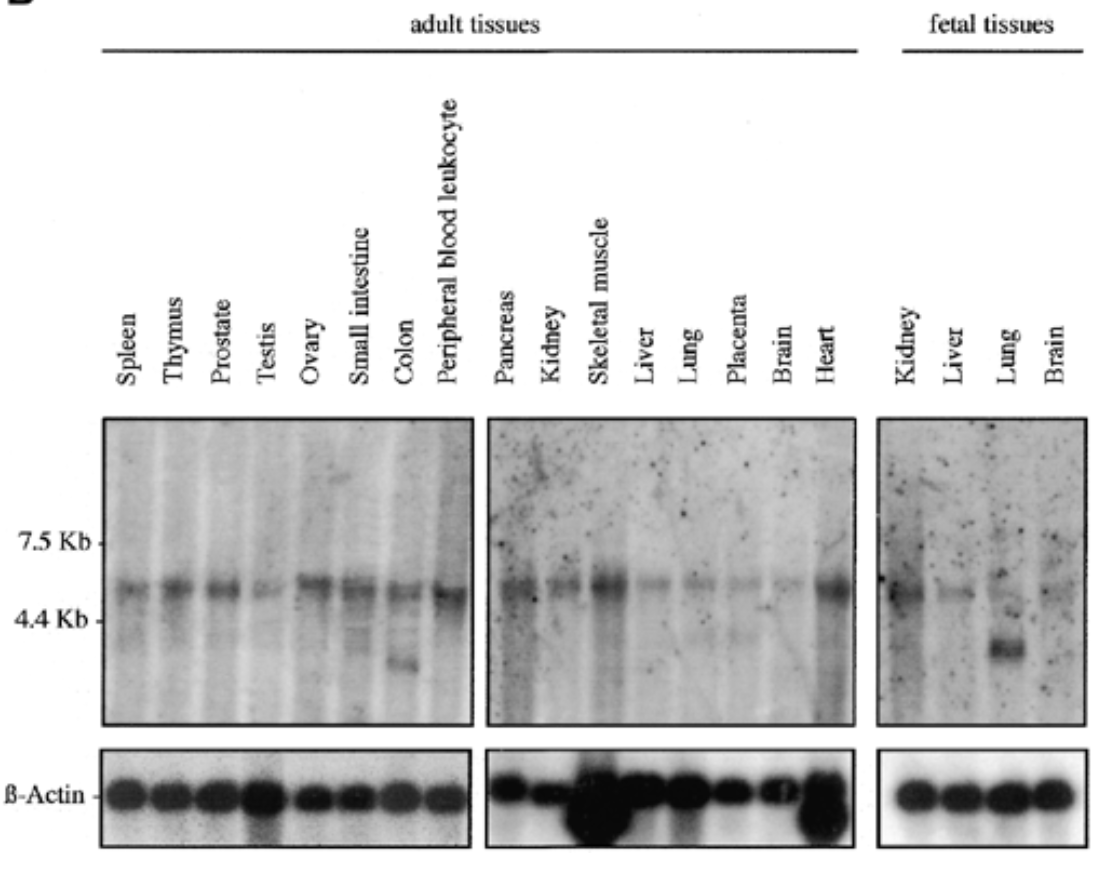
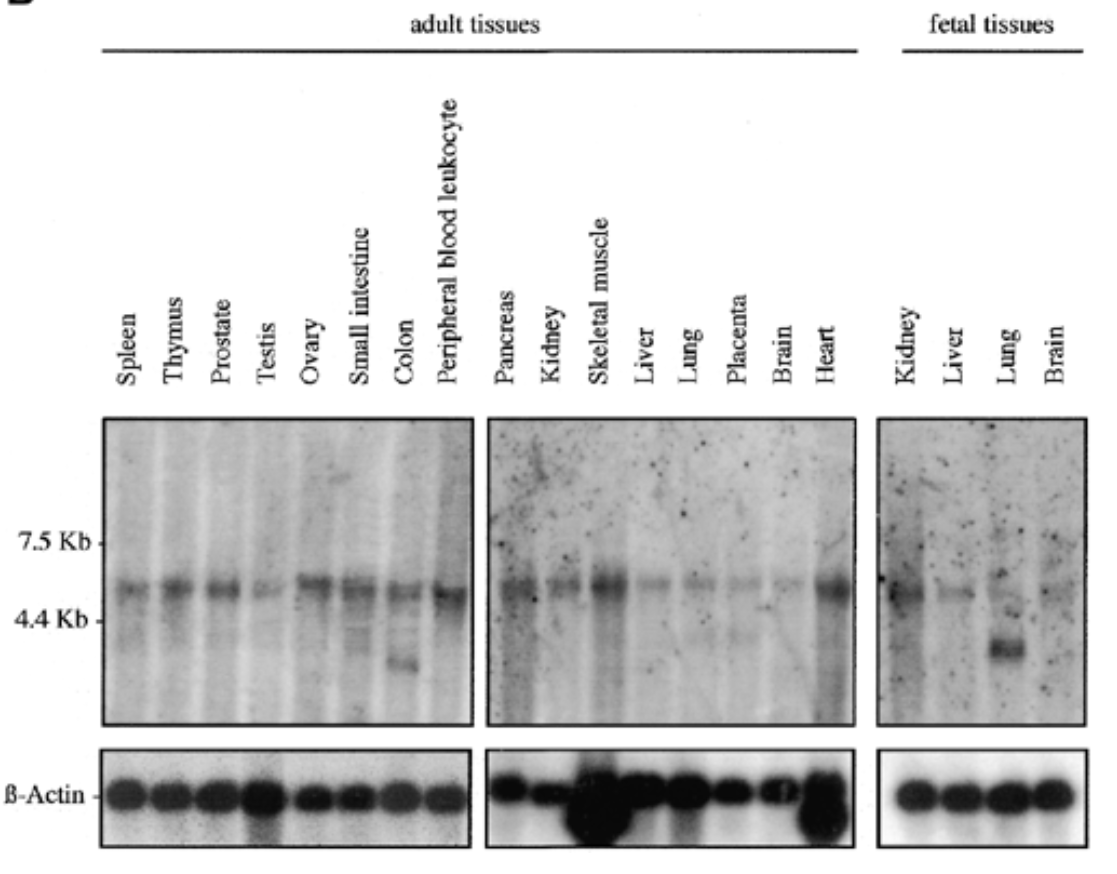
sox7 exp in all fetal and adult human tissue at 5kb. additional 2 bands at 2.5 kb for adult colon and fetal lung
control
b actin for equal loading to verify rna is present
conclude
sox7 plays role in all tissues. bands at 2.5 kb can be splice variants or unspliced messengers.
classify
hypothesis/establishing, expression
question
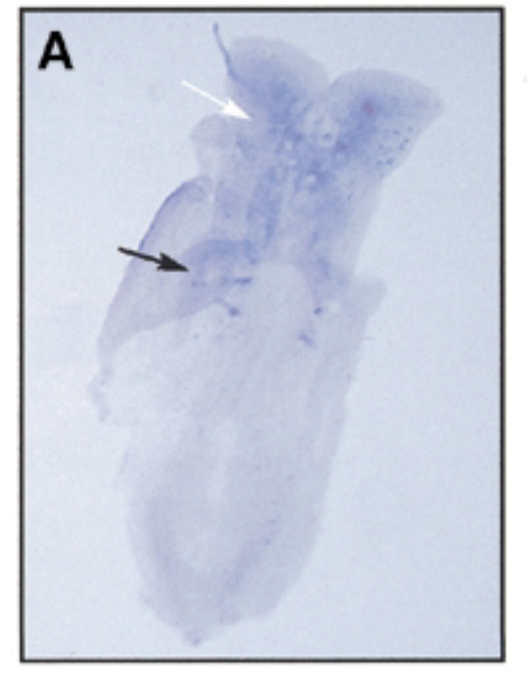
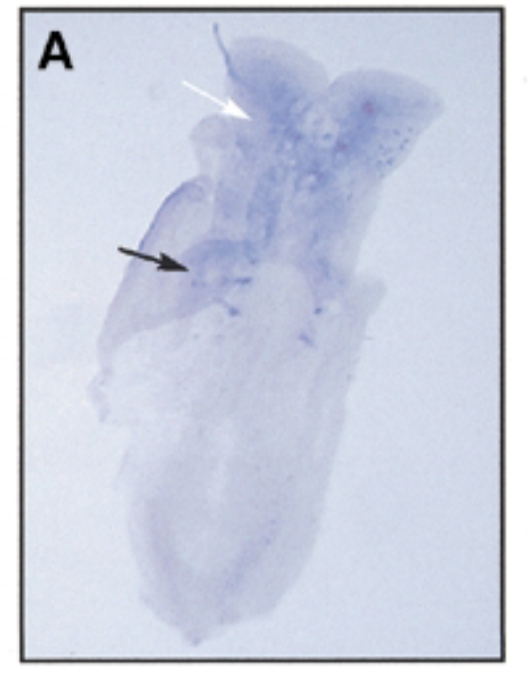
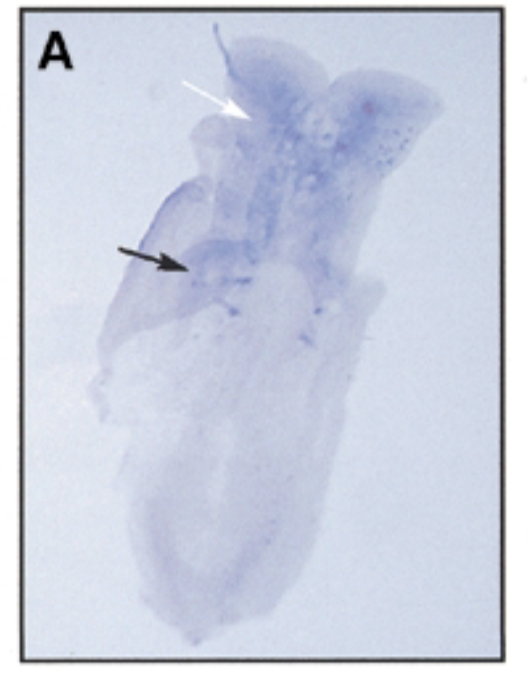
where sox7 is exp as mouse embryo devlop over time 8 dpc
method
whole mount in situ hybridisations
conditions
mouse embryo at 8 to 11.5 and 17.5 dpc
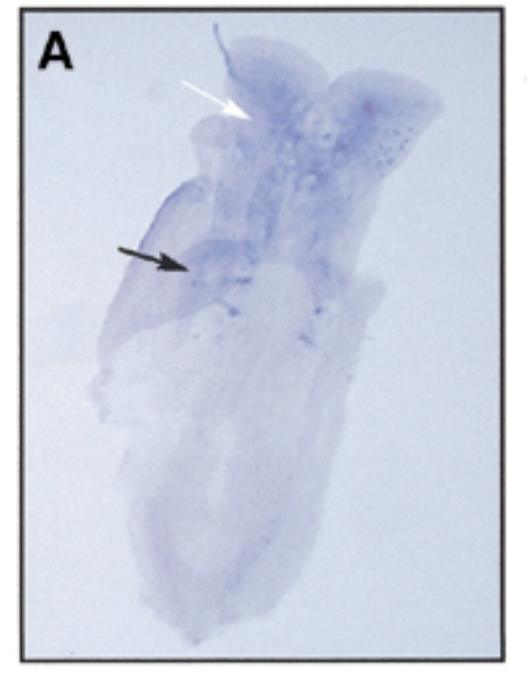
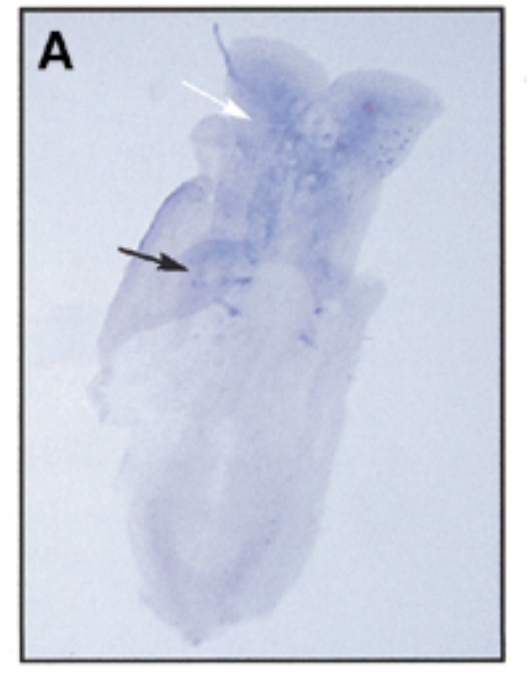
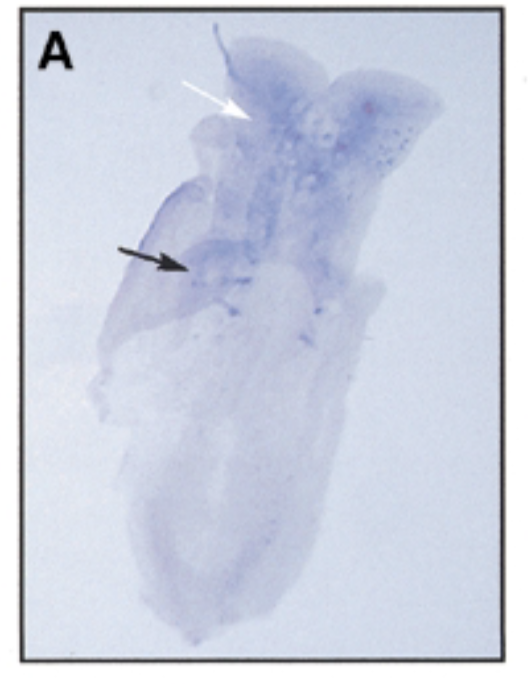
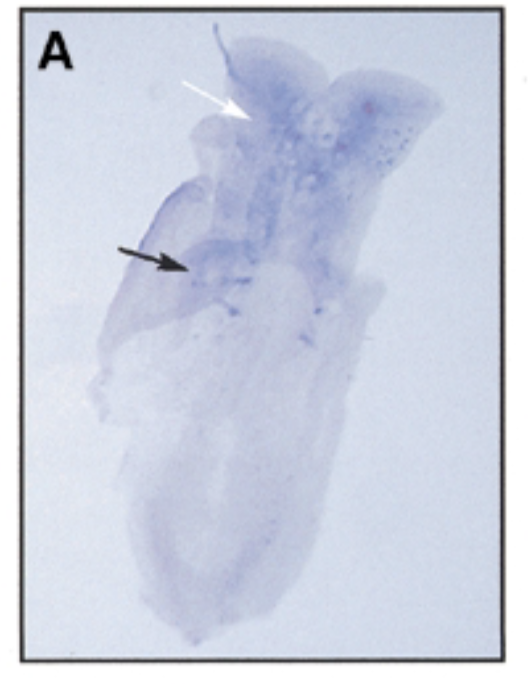
result
at 8 dpc, sox7 is exp in somites (black) and brain (white)
control
sense probe = negative control that would show no staining. rna probe is antisense to sox7 mrna. the sense probe wouldn’t bind to sox7 mrna.
conclude
sox7 is exp in the regions that need to be developed. there is early exp and restricted head dev.
classify
hypothesis, exp
question
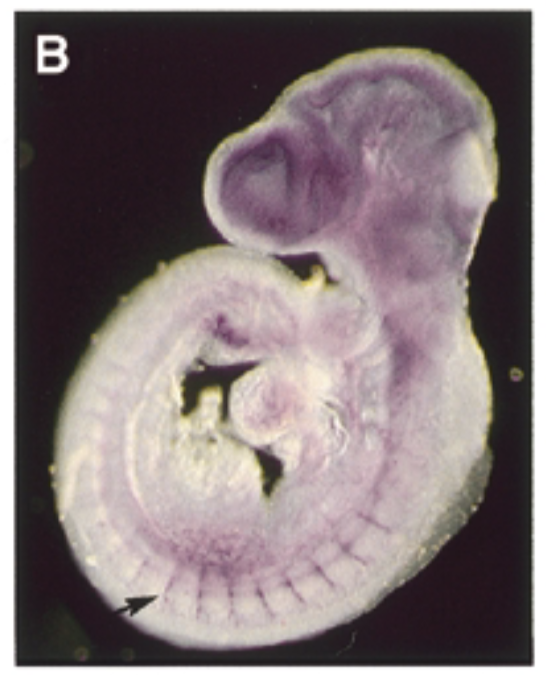
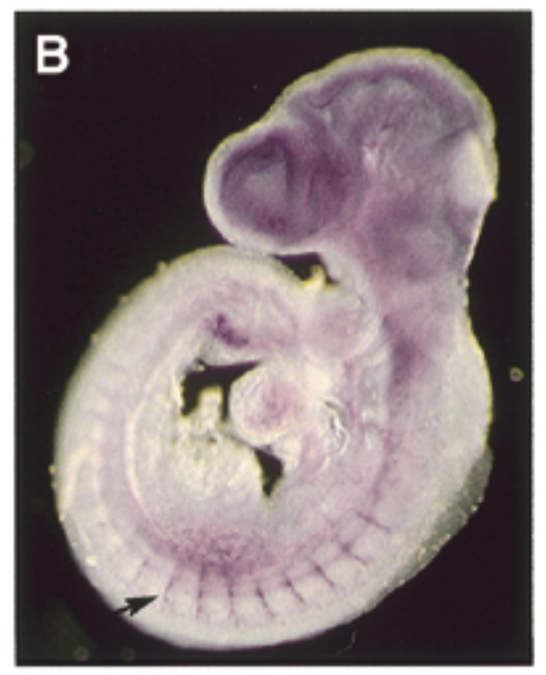
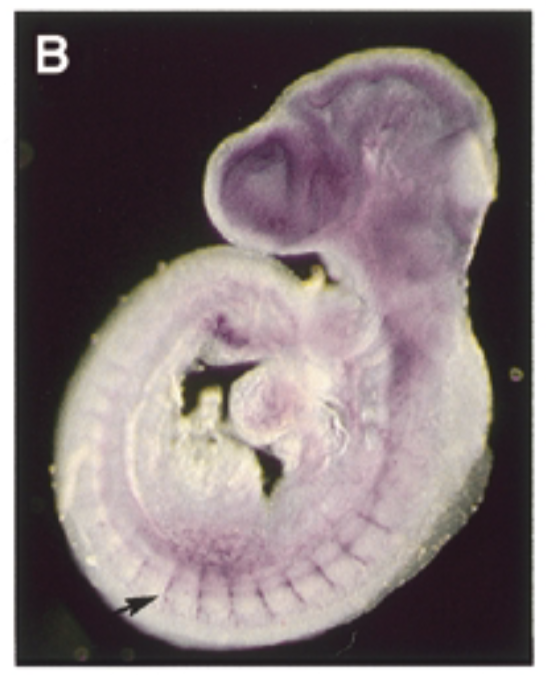
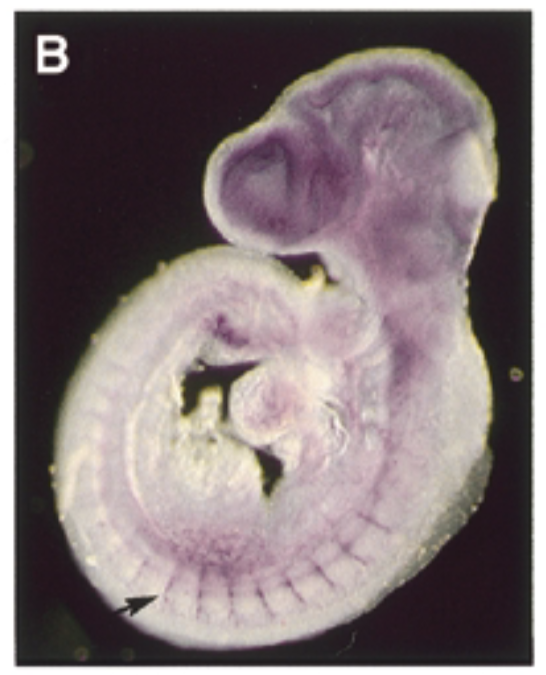
where sox7 is exp in a developing mouse embryo 9.5 dpc
method
in situ hybridisations
conditions
mouse embryo at 8 to 11.5 and 17.5 dpc
result
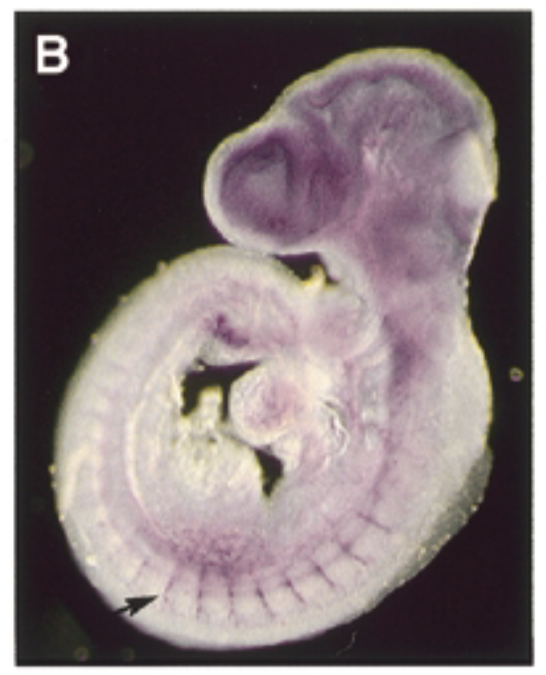
at 9.5 dpc, sox7 is exp thru vascular system in intersomitic vessels. sox7 helps create the endothelial cells which are the cells inside skin of vessels. exp of sox7 makes the endothelial cell the cell that it is
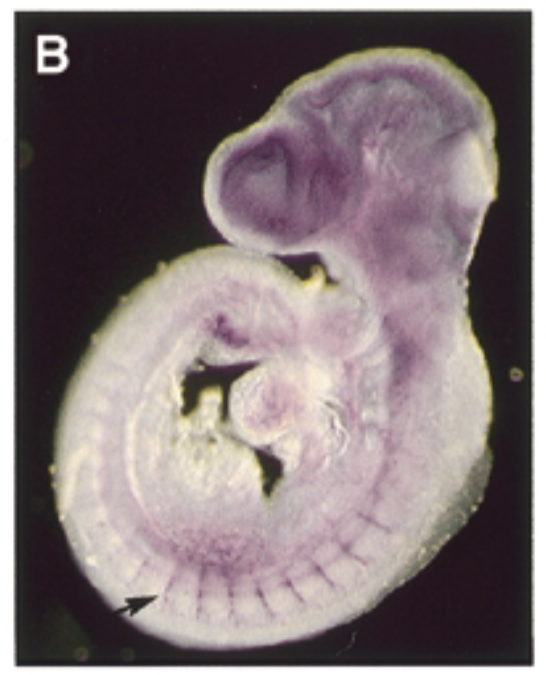
control
sense probe
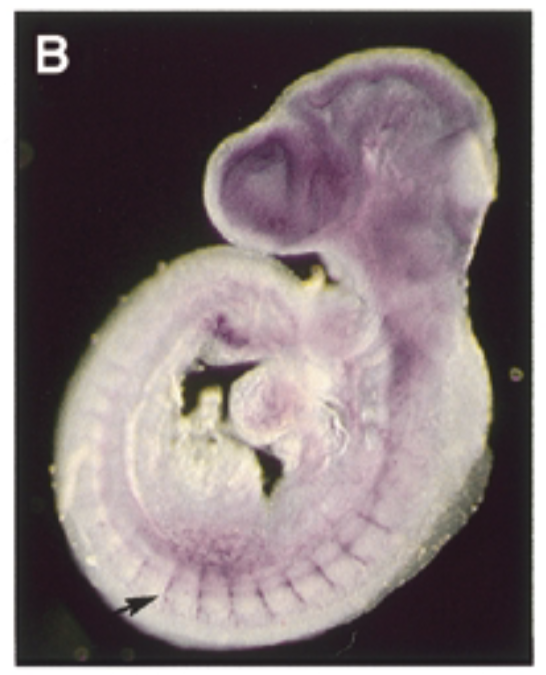
conclude
sox7 plays a role in vascular development
classify
hypothesis, exp
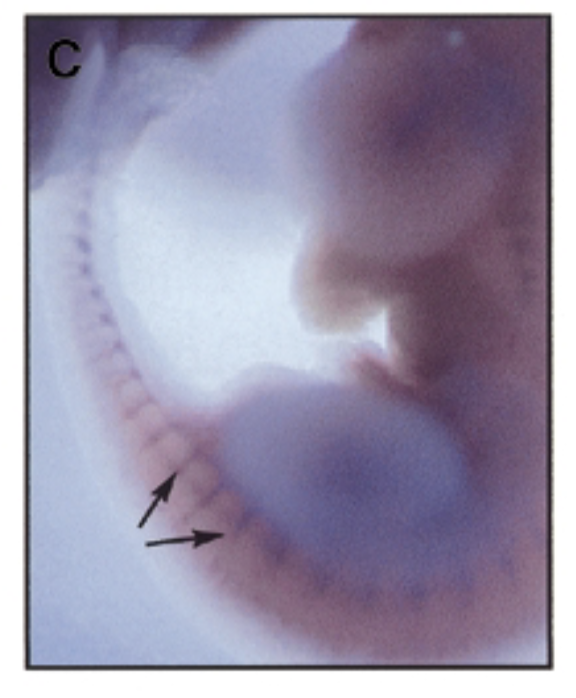
question
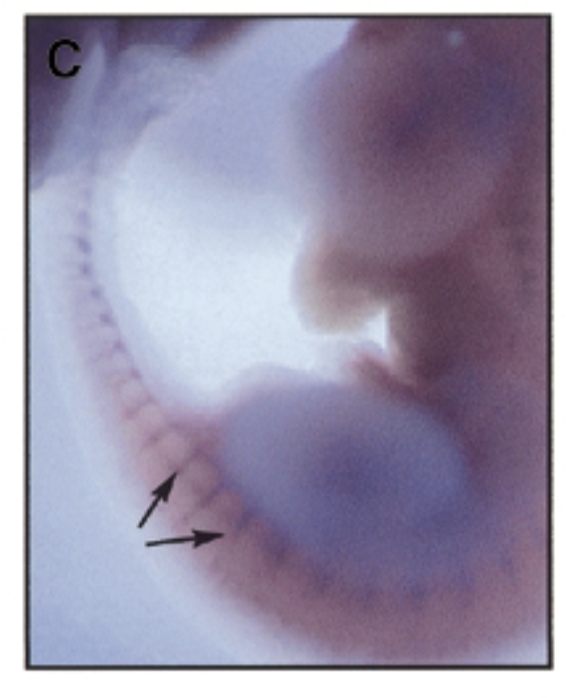
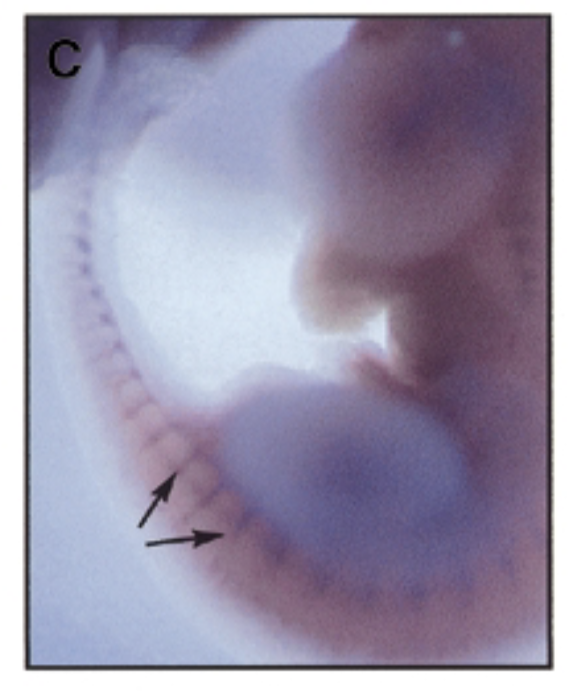
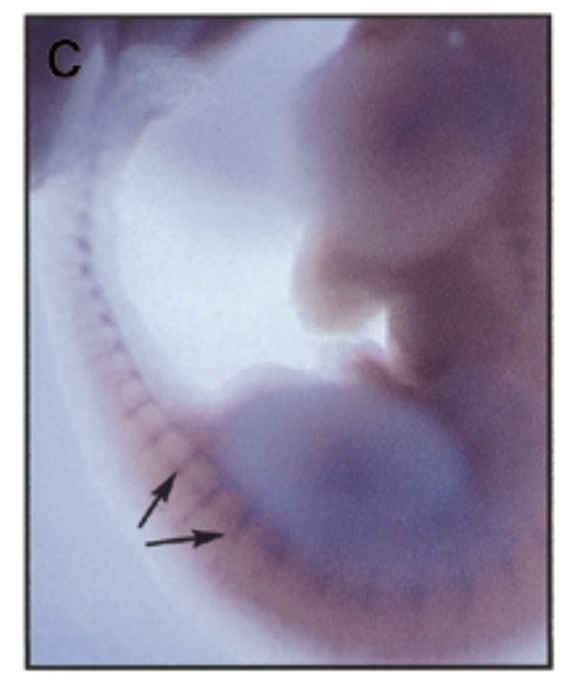
where is sox7 exp in developing mouse embryo 11.5 dpc
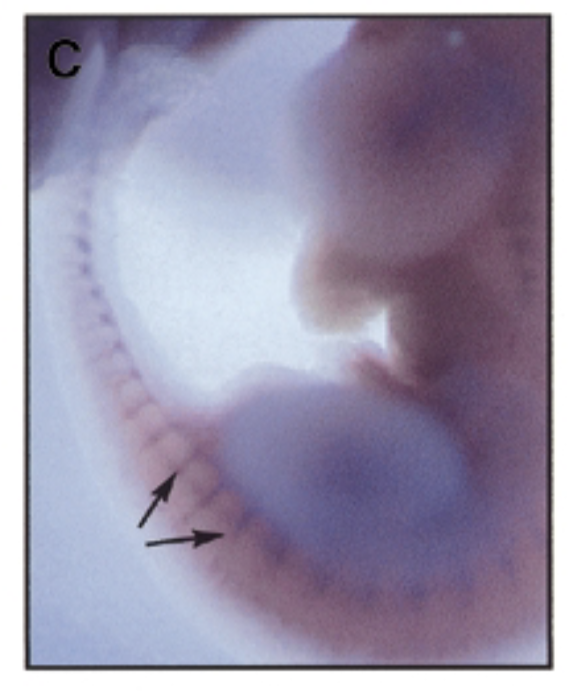
method
in situ hybridisations
conditions
mouse embryo at 8 to 11.5 and 17.5 dpc
result
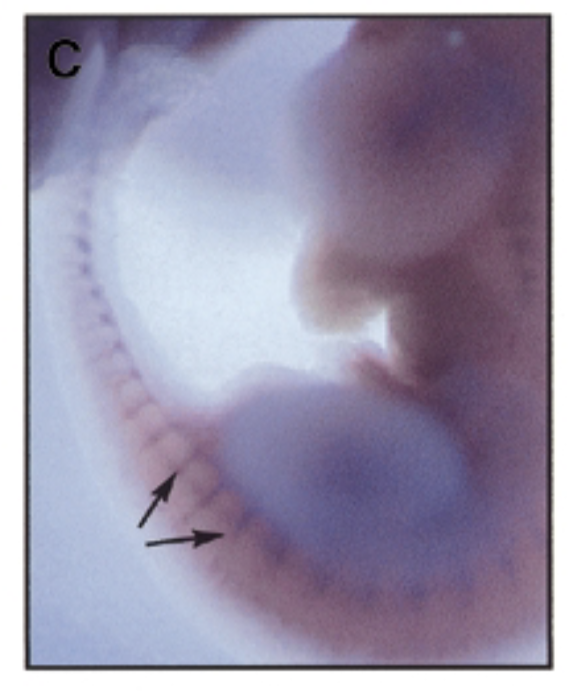
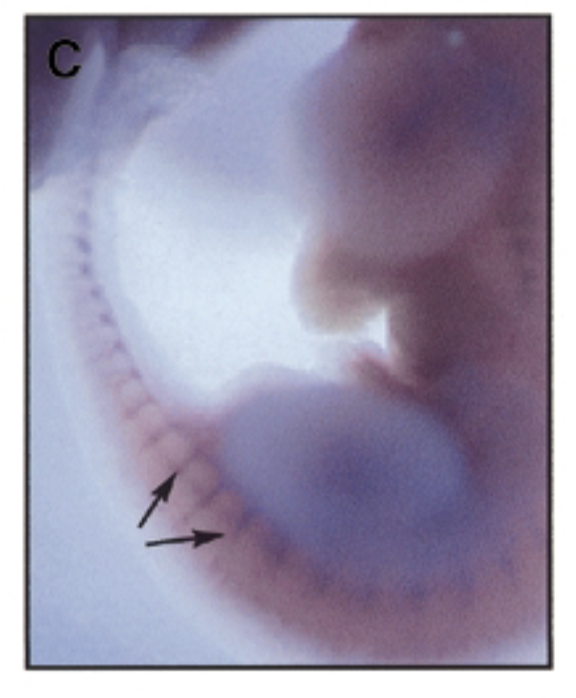
at 11.5 dpc, sox7 is mainly exp in intersomitic vessles
control
sense probe
conclude
exp of sox7 plays a role in development of mouse embryo
classify
hypothesis, exp
question
where is sox7 exp in developed embryo 17.5
method
in situ hybridisations
conditions
mouse embryo at 17.5 dpc vs 8-11.5
result
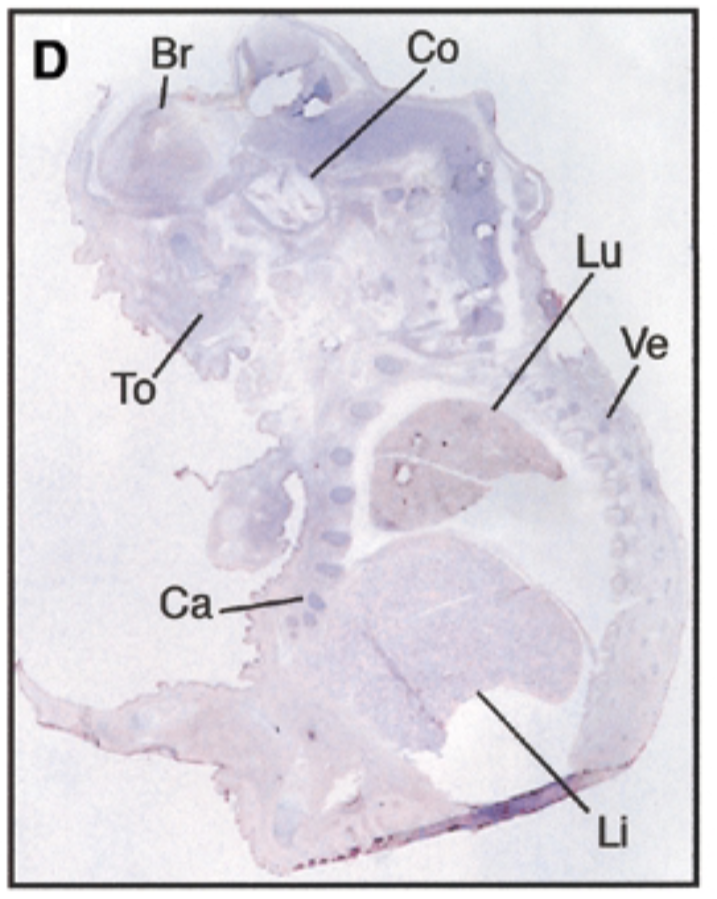
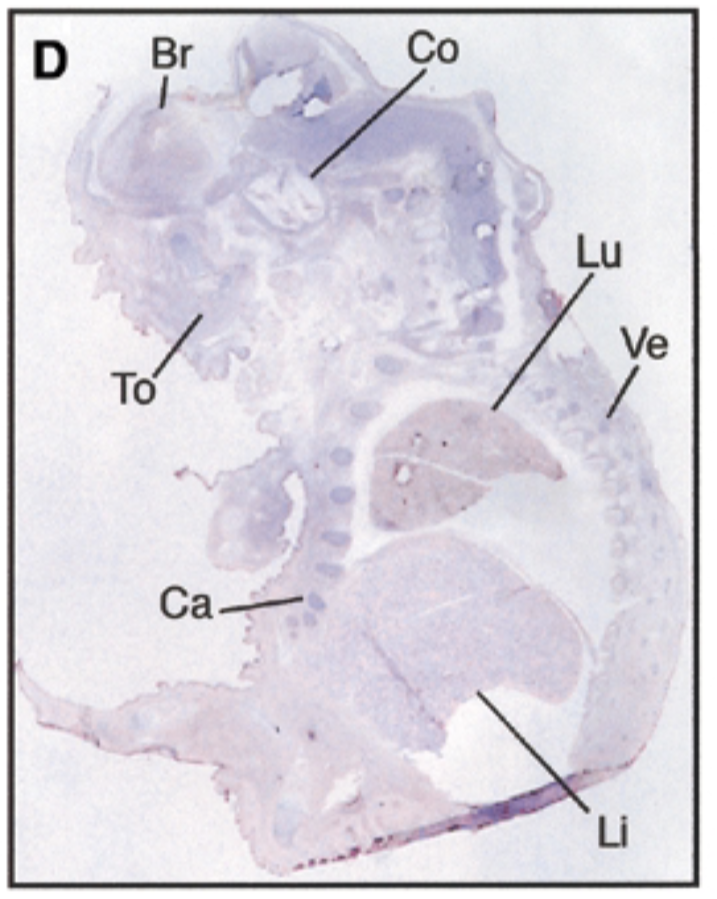
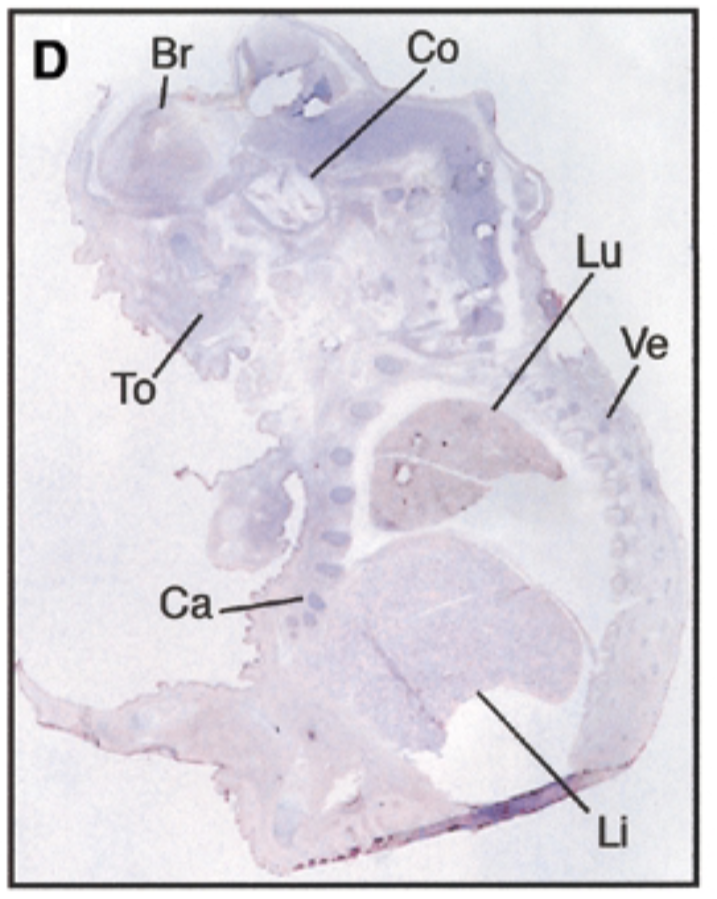
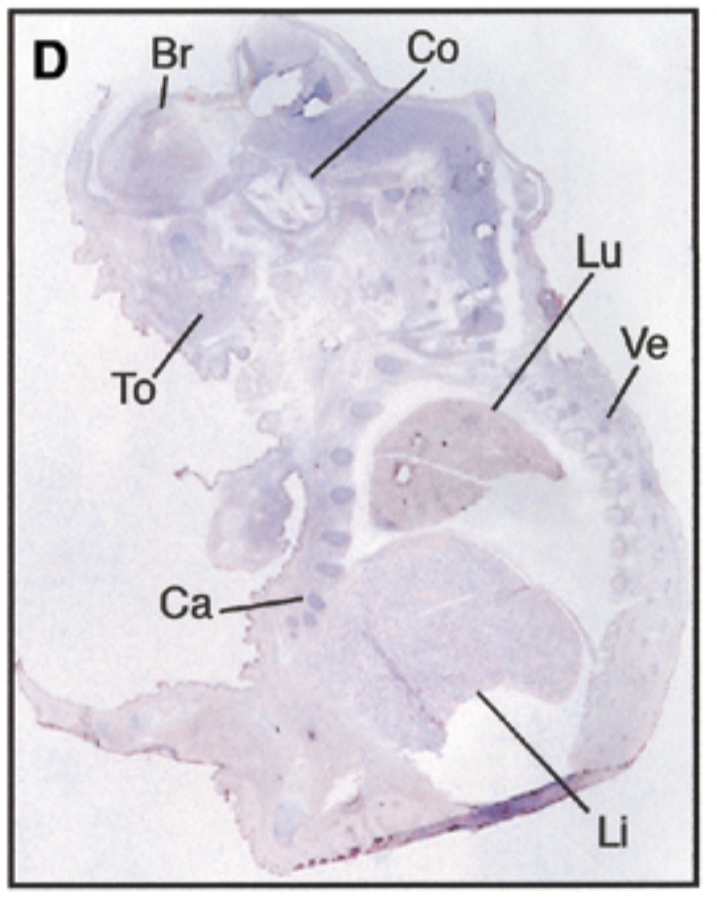
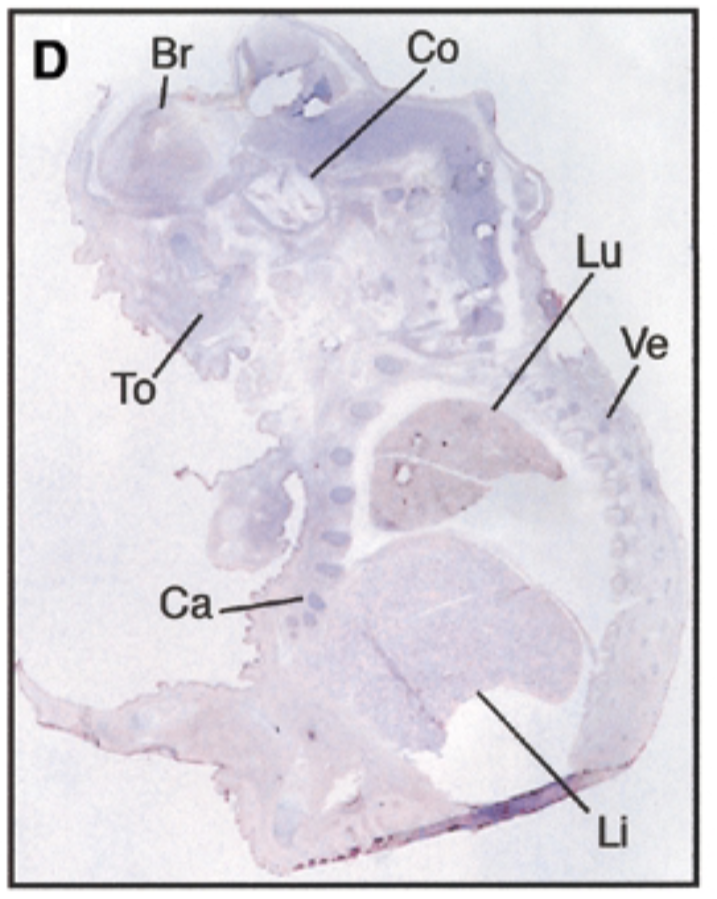
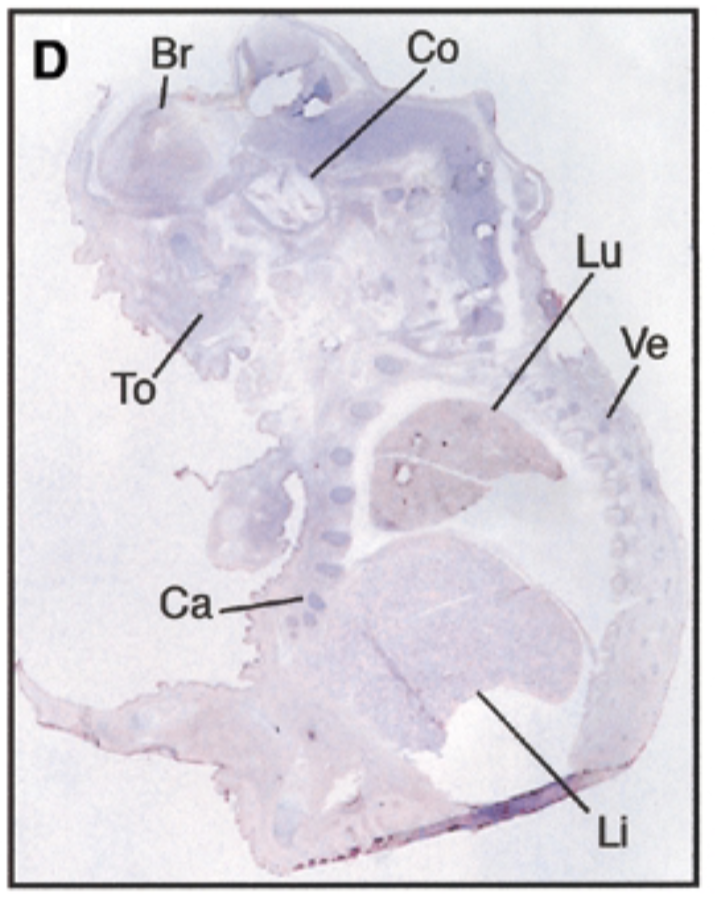
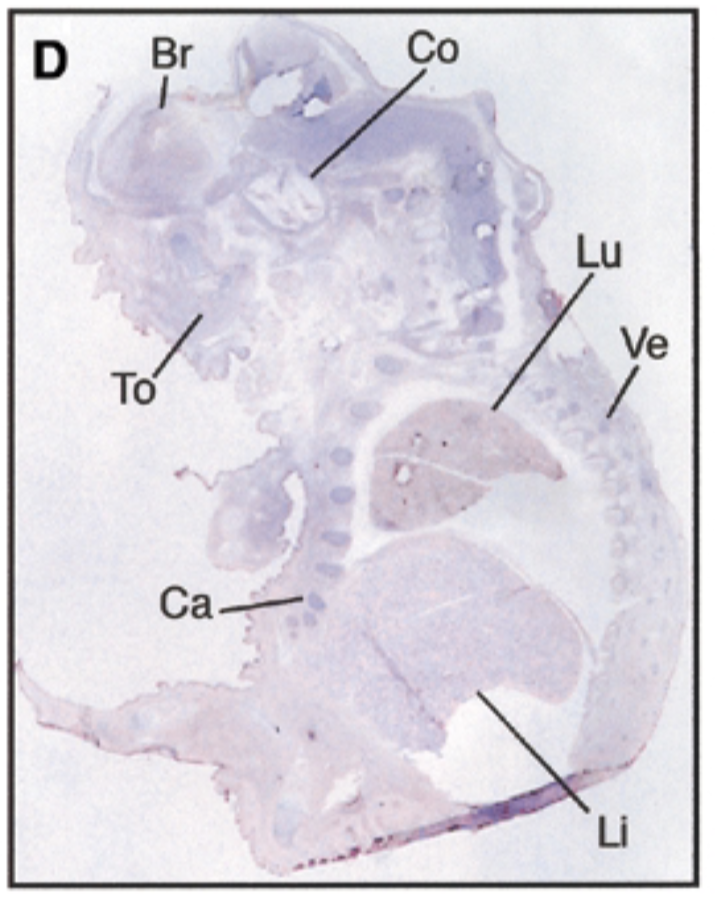
at 17.5 dpc, sox7 is seen in brain, cochlea, tongue, cartilage, lung, liver, vertebrae. starts being restricted
control
sense probe=negative control & positive control=nuclear fast red staining=counterstain that stains all structures + probe show where mrna is exp
conclude
after threshold of 17.5 dpc, exp becomes restricted to certain site
classify
hypothesis, exp
question
where is sox7 exp in human and if conserved btn mouse and human
conditions
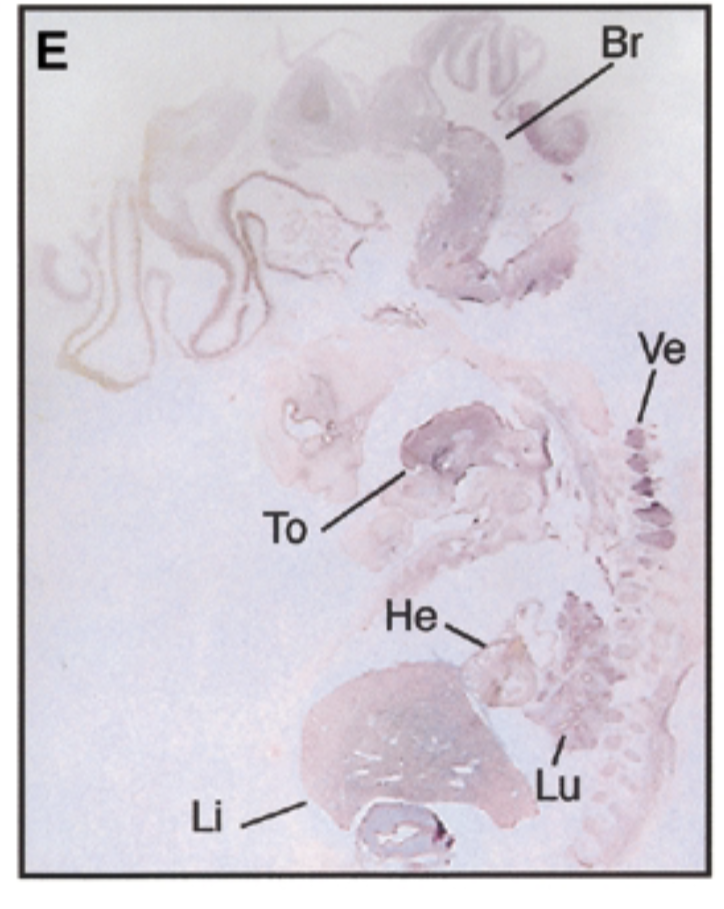
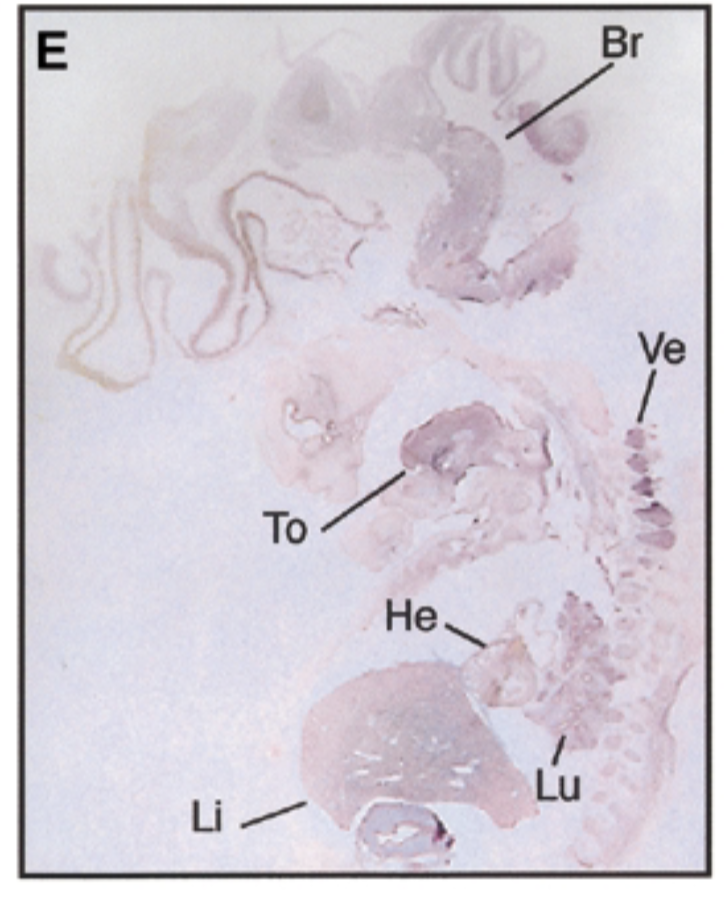
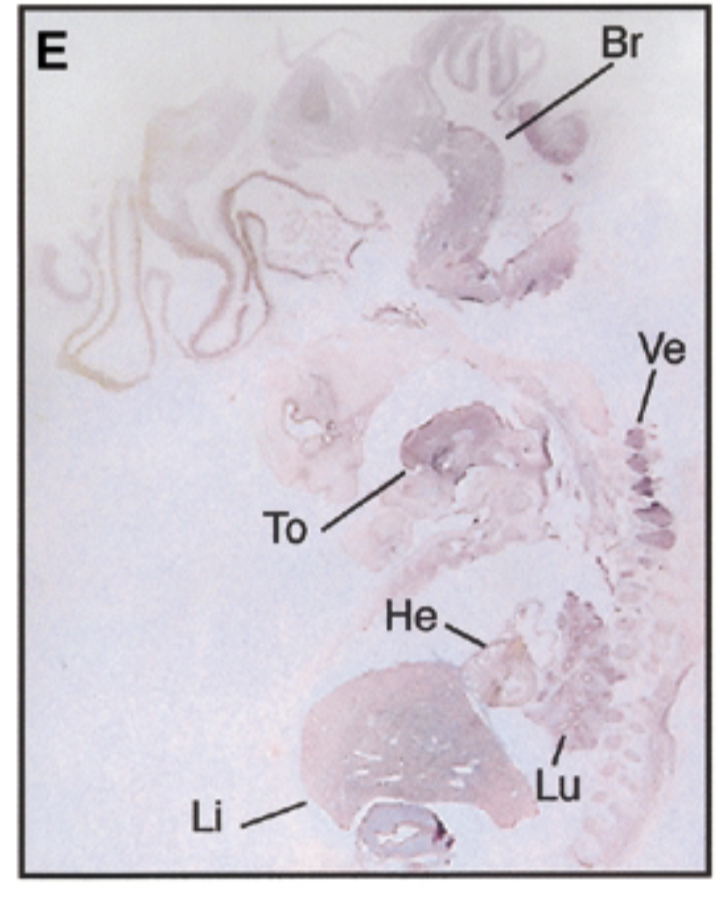
8 week human embryo at carnegie stage 19 vs 17.5 dpc mouse
result
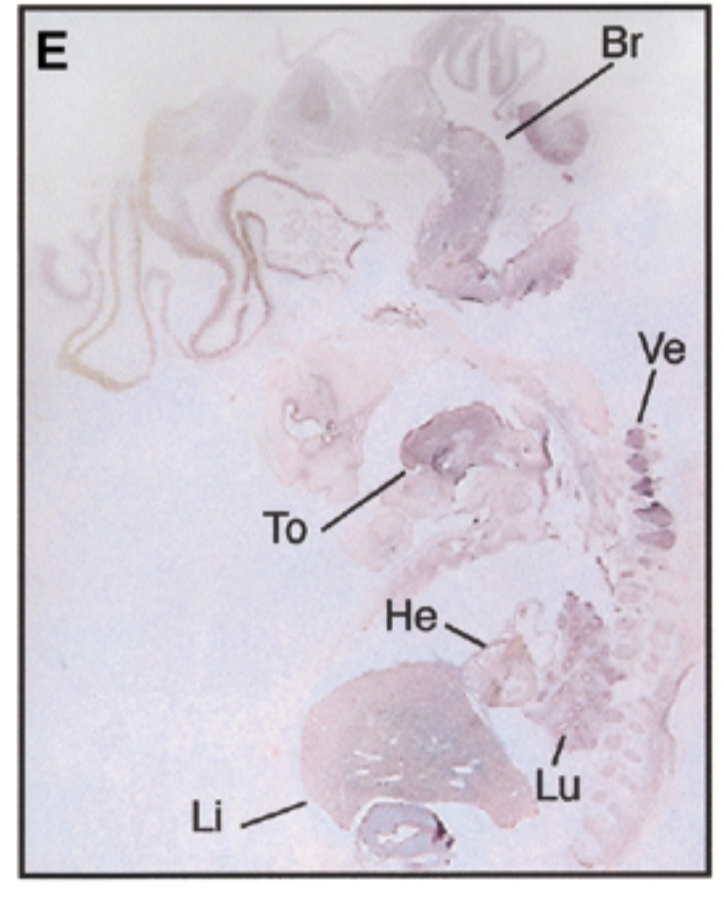
8 week embryo, sox7 is in brain tongue heart liver lung vertebrae in pattern similar to mouse
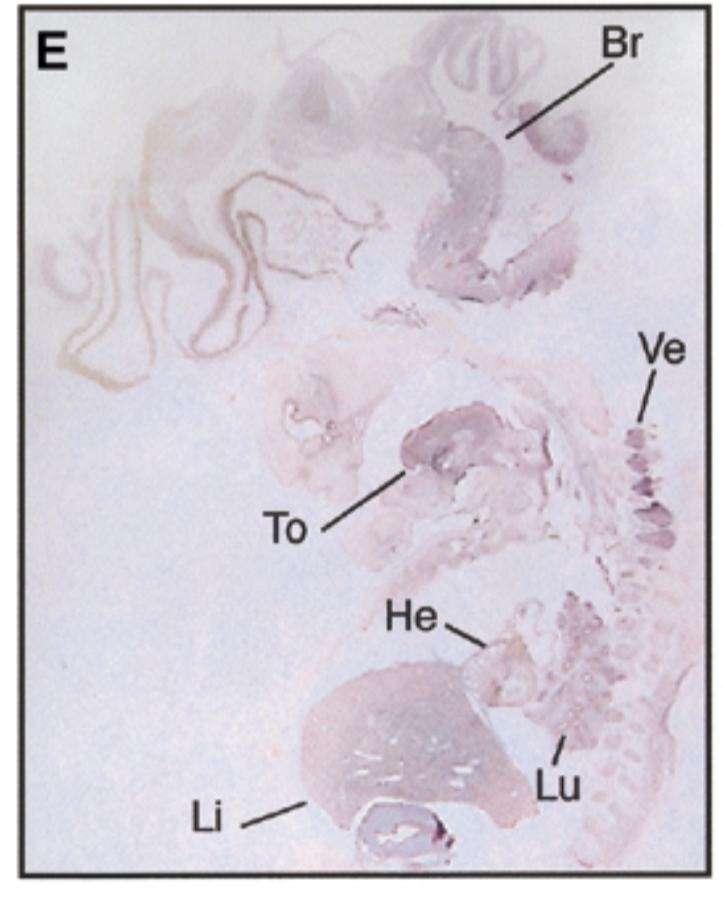
control
negative control=sense probe, positive control=nuclear fast red stain=counterstain that stains everything + probe that shows where mrna exp
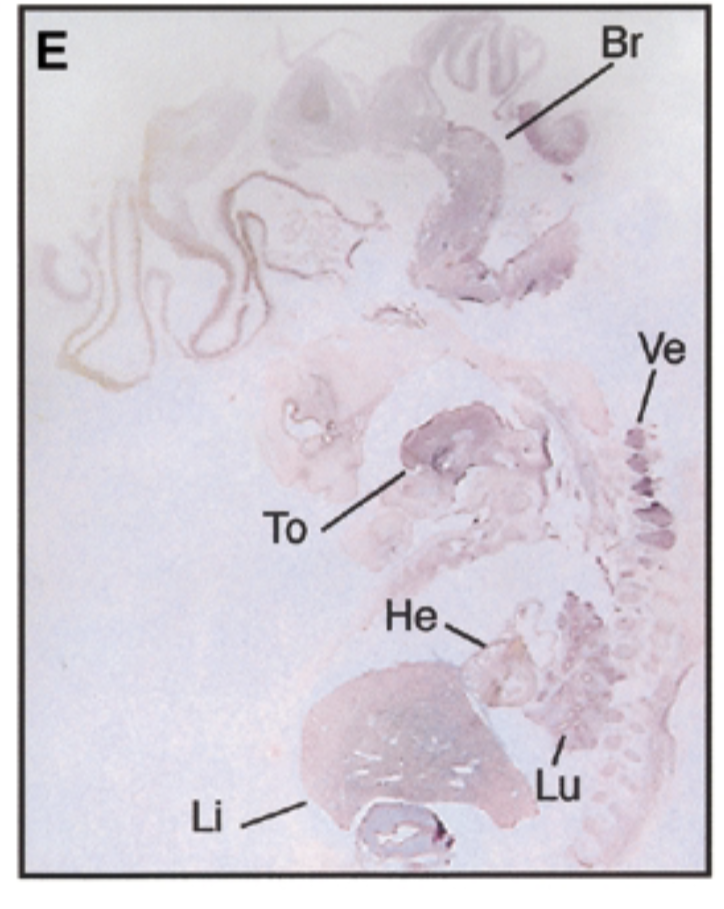
conclude
sox7 plays a role in development AND differention and exp gets restricted like mice → sox7 is conserved btn mouse and embryo. those organs are where epithelial mesencymal transition happen, so sox7 is involved in emt
classify
hypothesis, exp
question
how is sox18 expressed compared to sox7
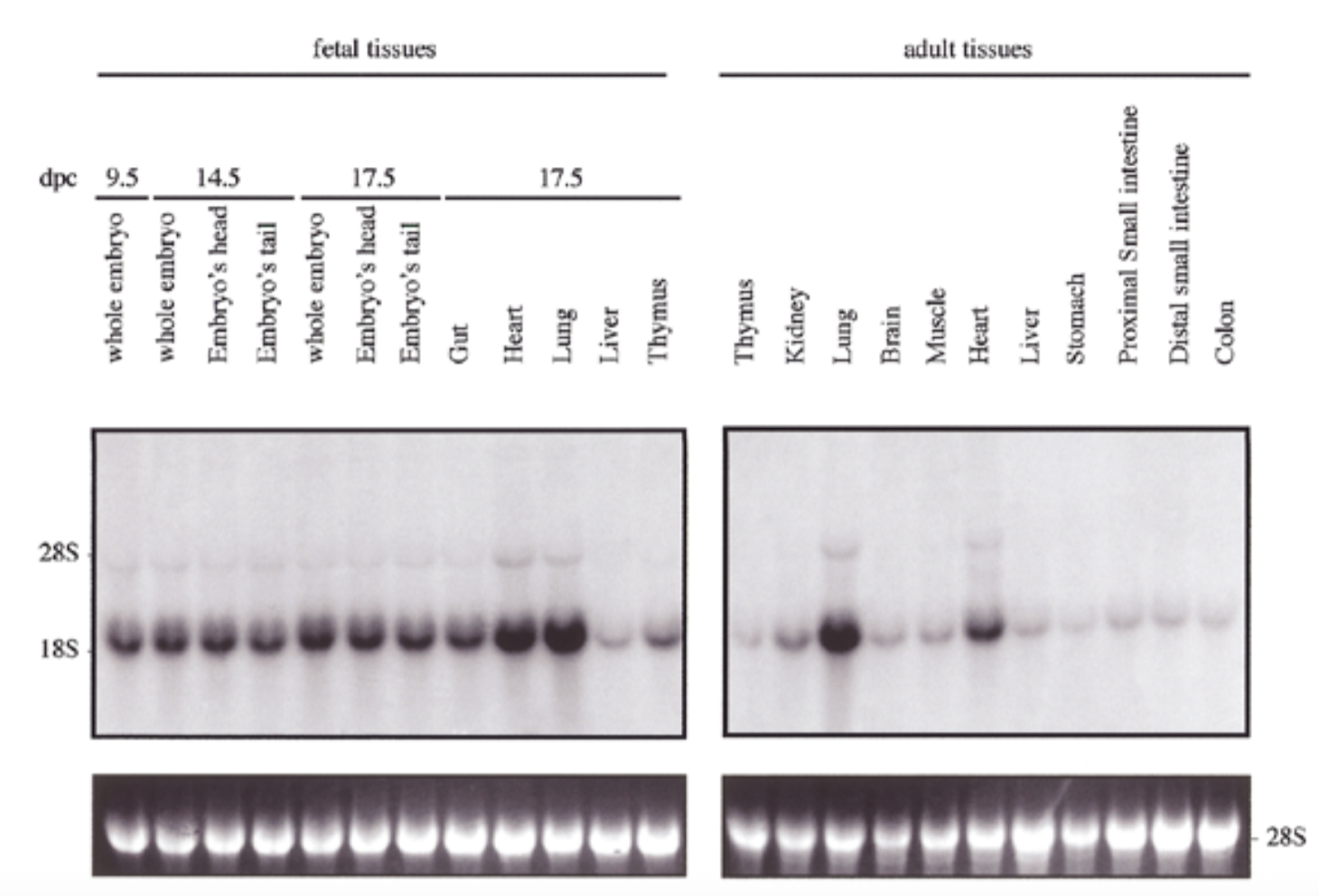
method
northern blot
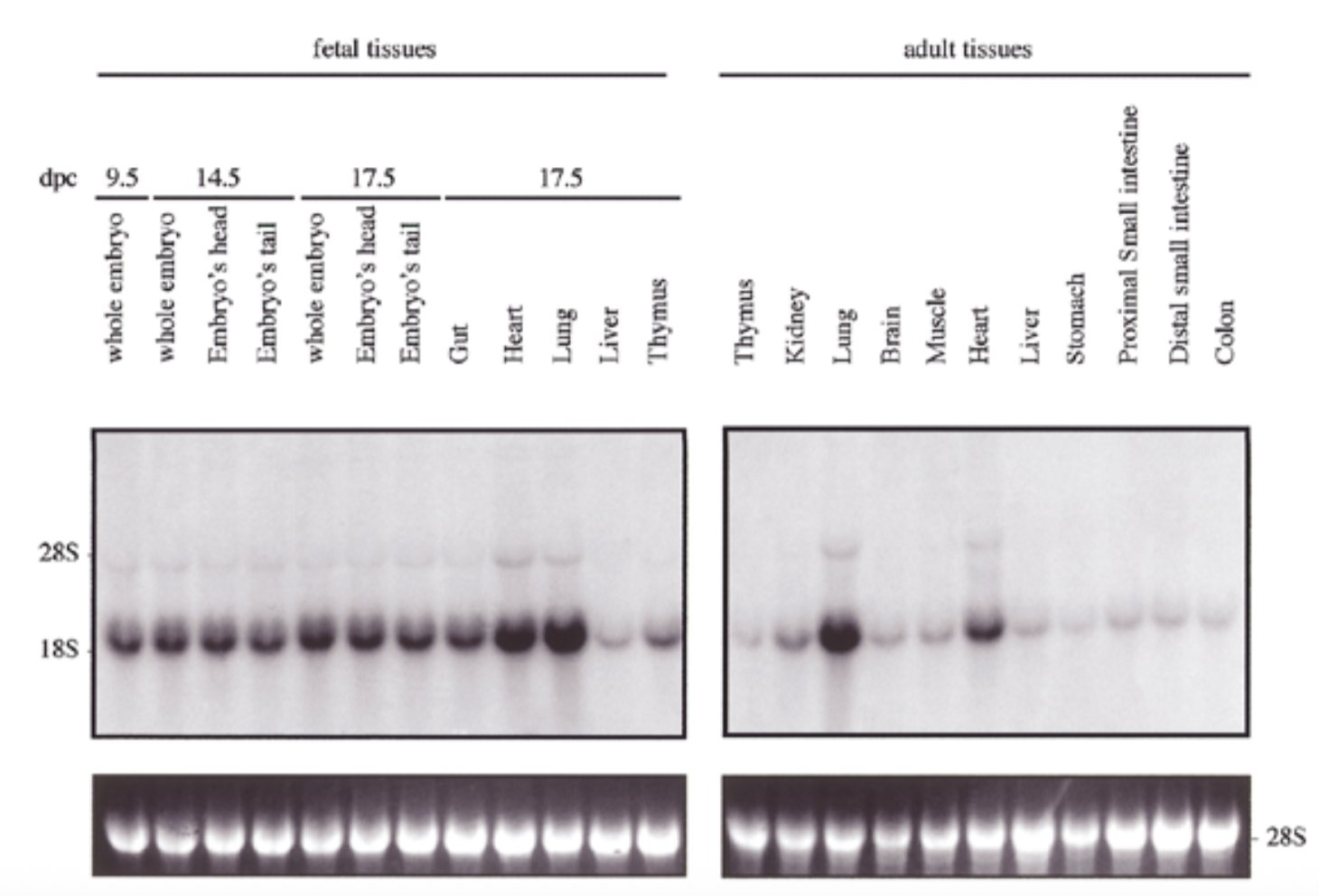
conditions
sox18 probe + embryonic vs adult
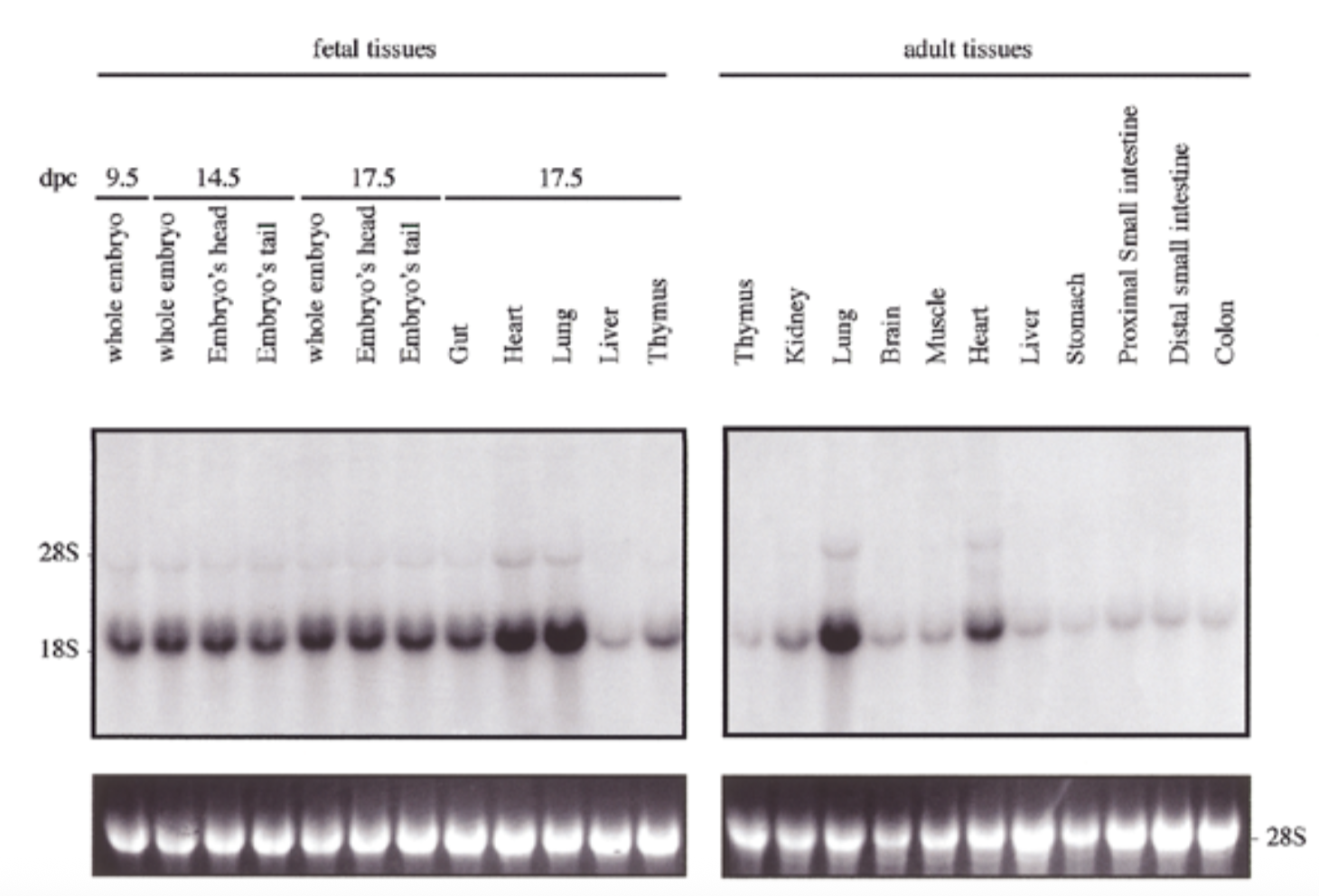
result
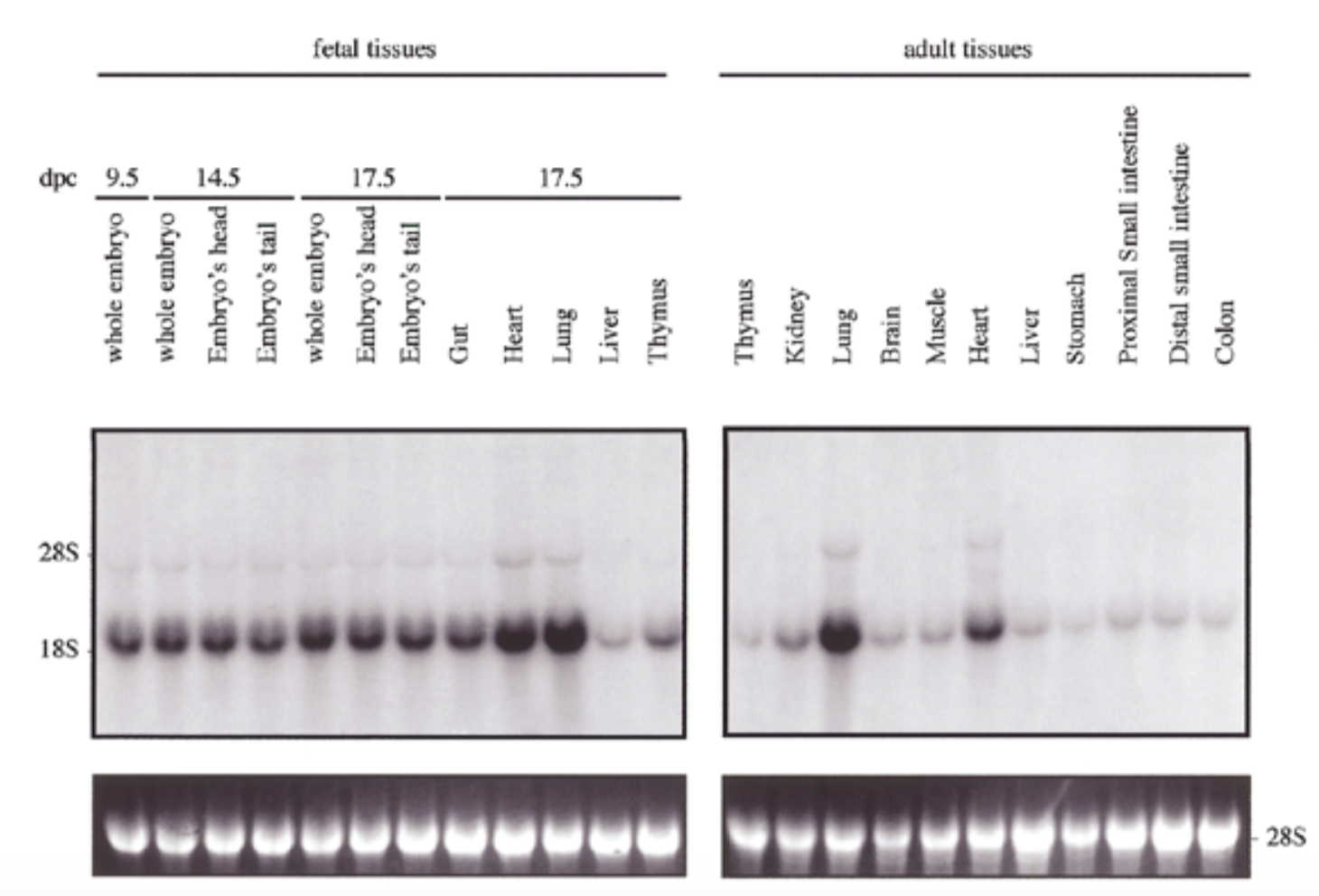
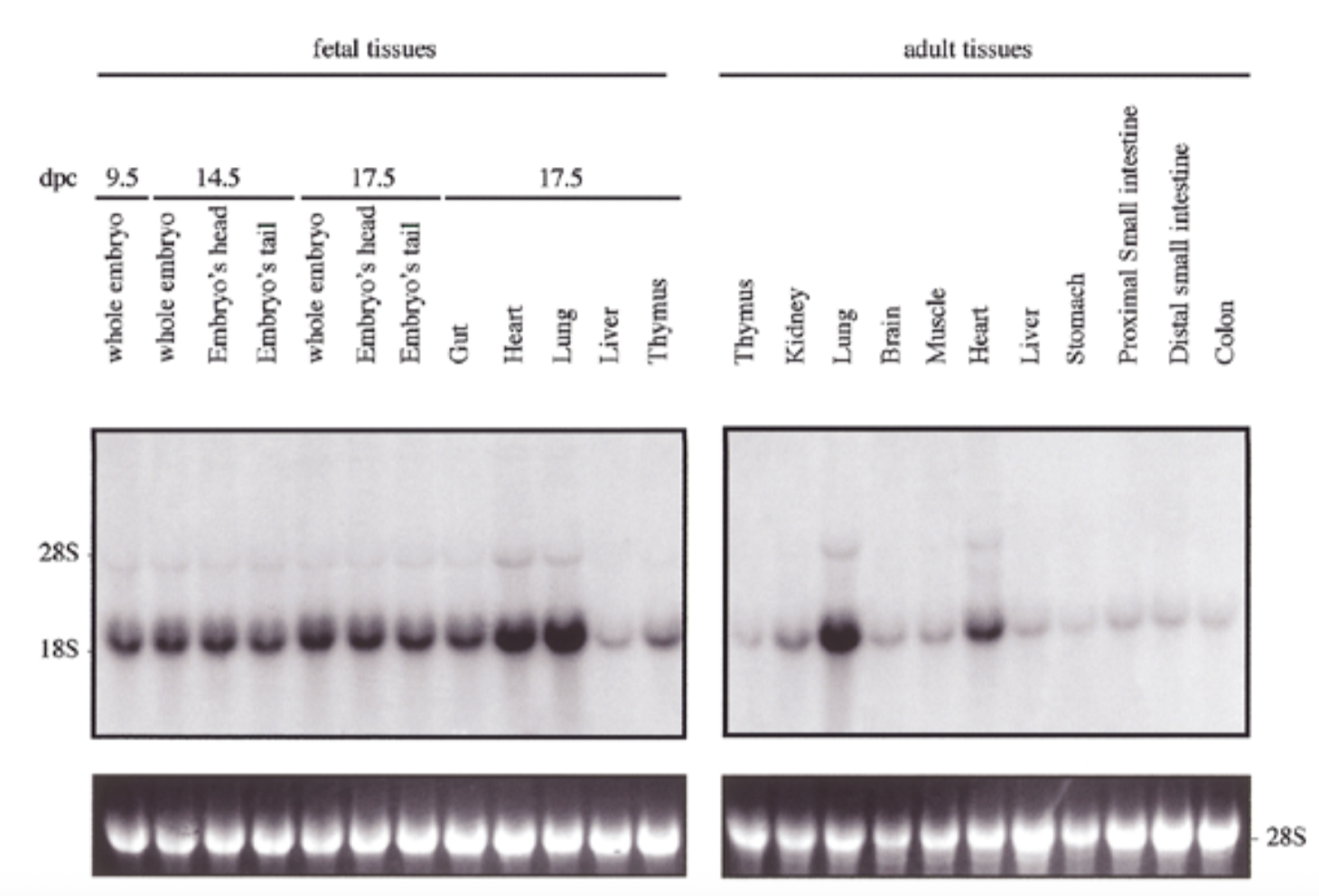
exp of sox18 had similar pattern to sox7 for both embryo and adult: exp in all tissues but particularly high in heart and lung
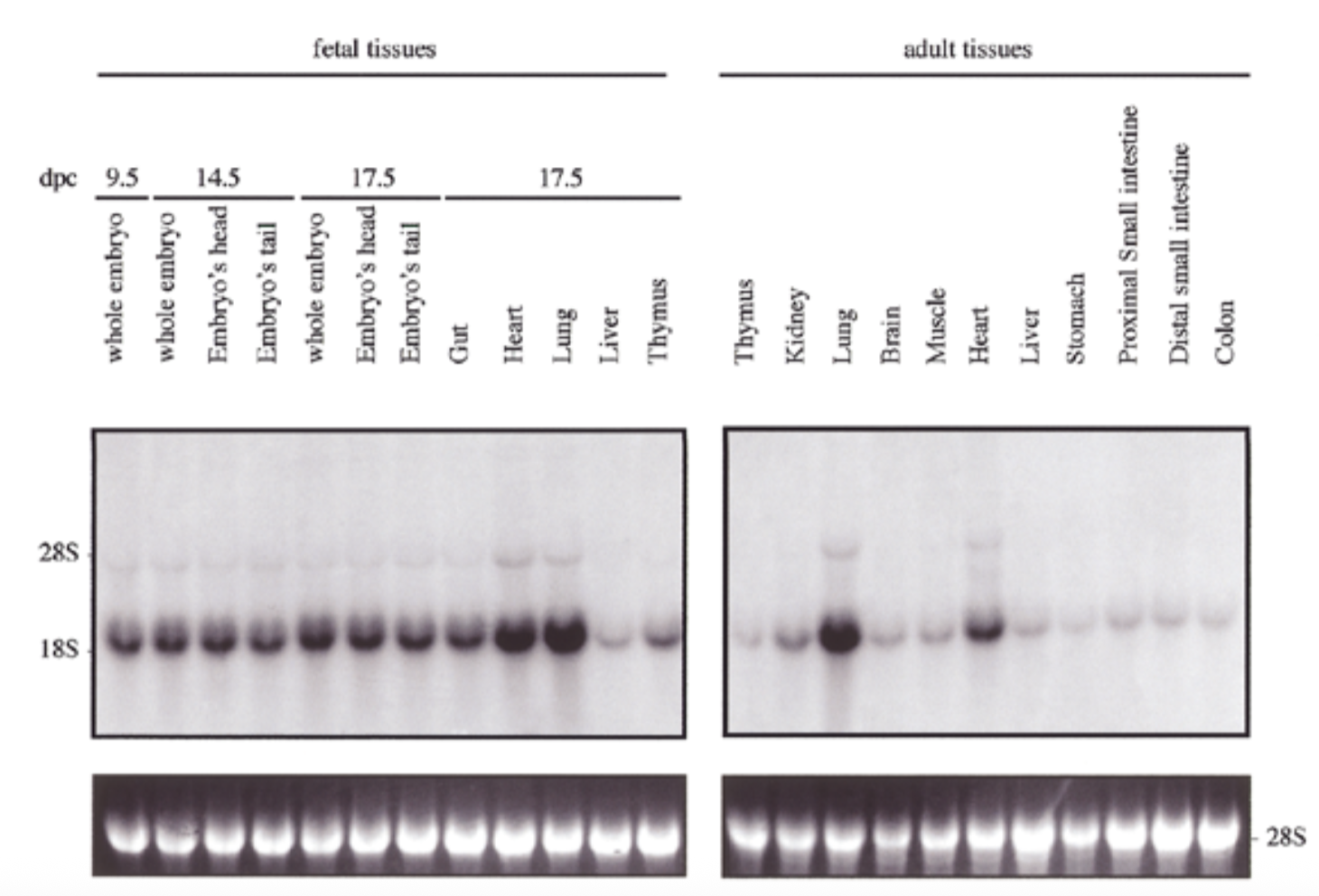
control
28s is rrna that is always exp at same levels → shows equal loading + ethidium bromide staining to show if equal amt of rna in each lane
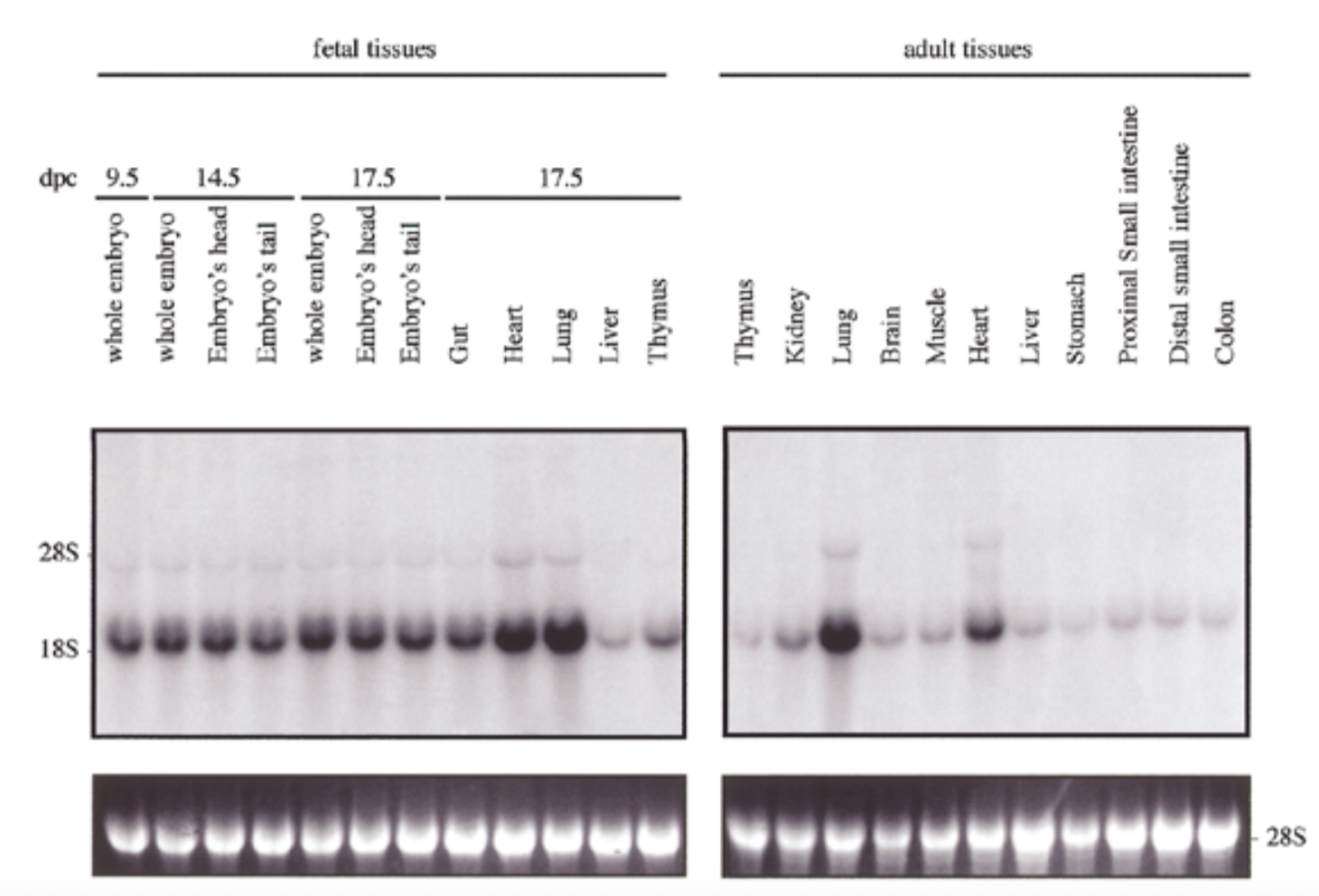
conclude
sox7 is comparable to sox18 → fxs overlap → genes come from same ancestor
classify
hypothesis/establishing, exp
question
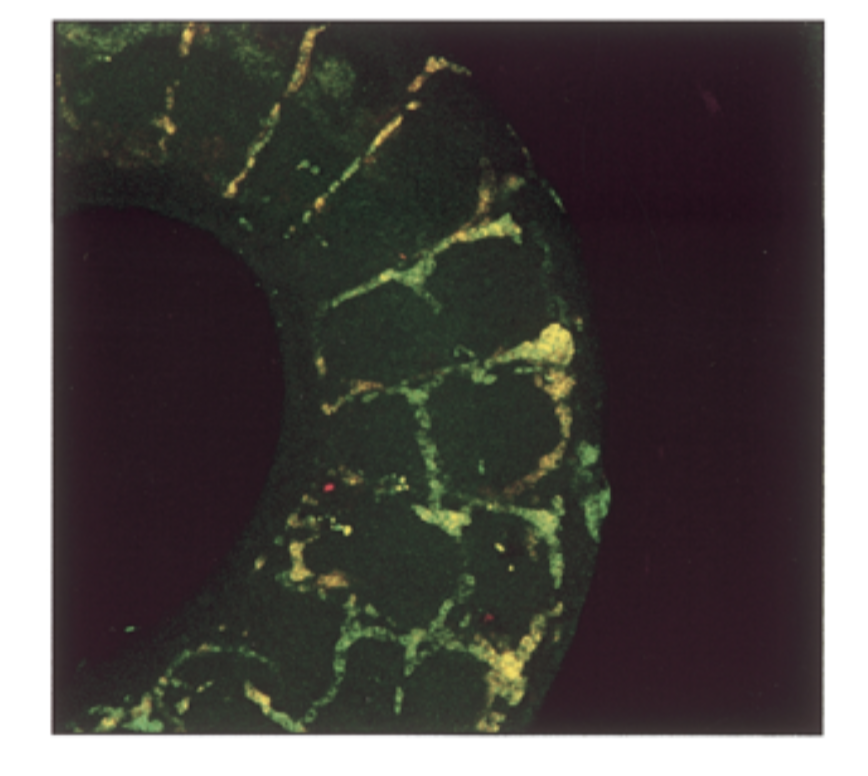
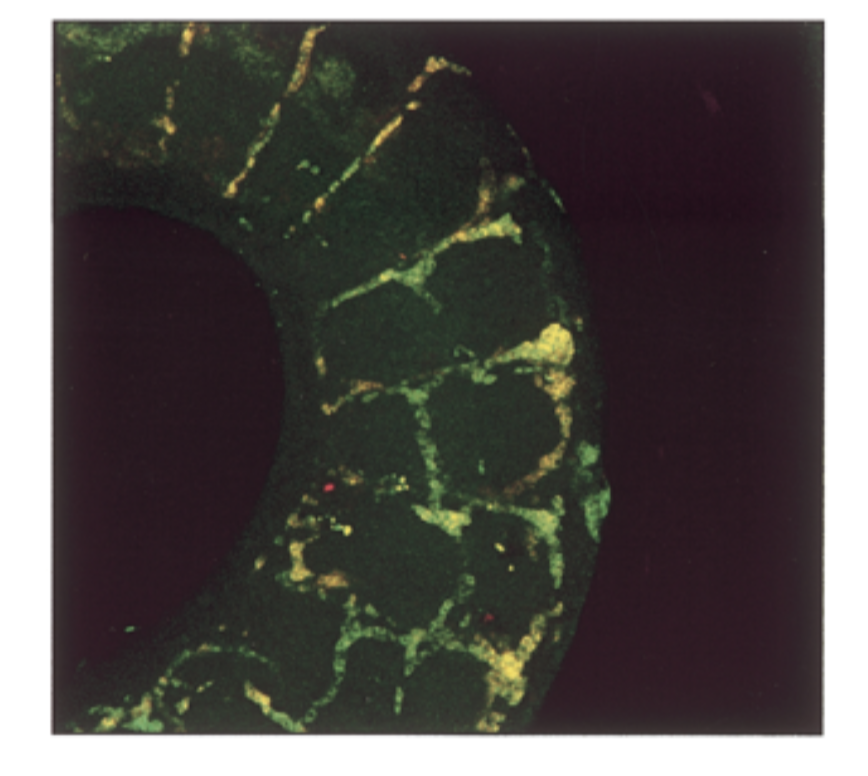
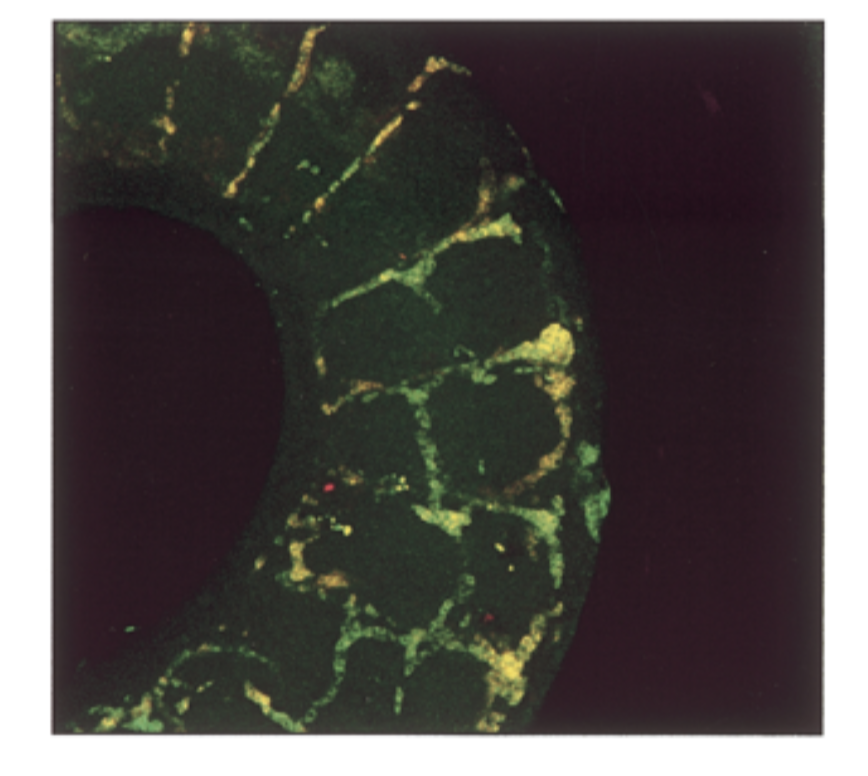
where is sox18 exp compared to sox7. do they colocalize in intersomitic vessels of 11.5 dpc mouse
method
fish
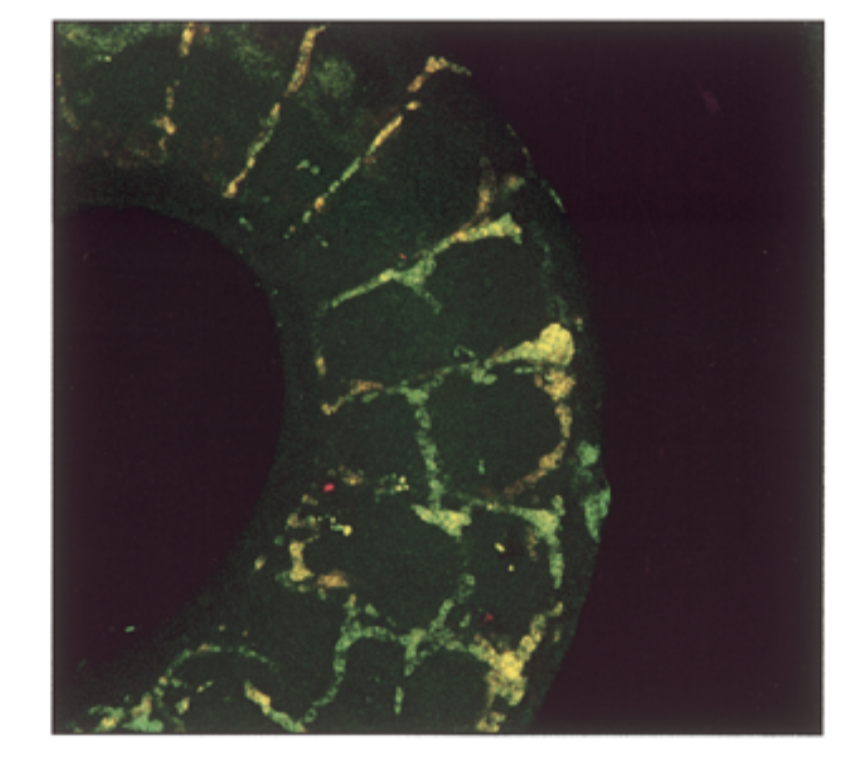
conditions
sox7 antisense prob, sox18 antisense probe
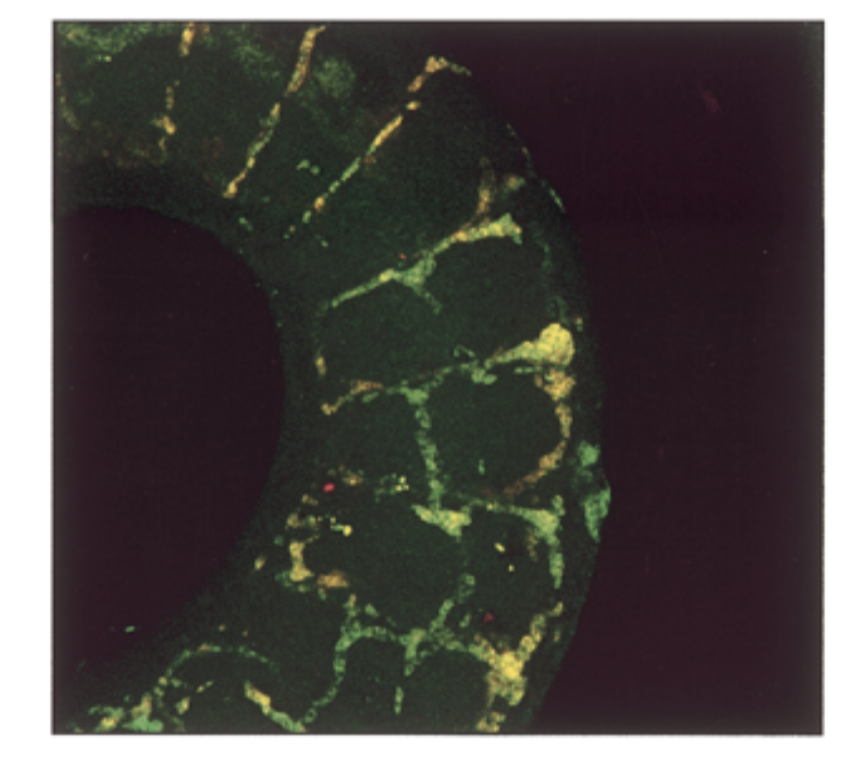
result
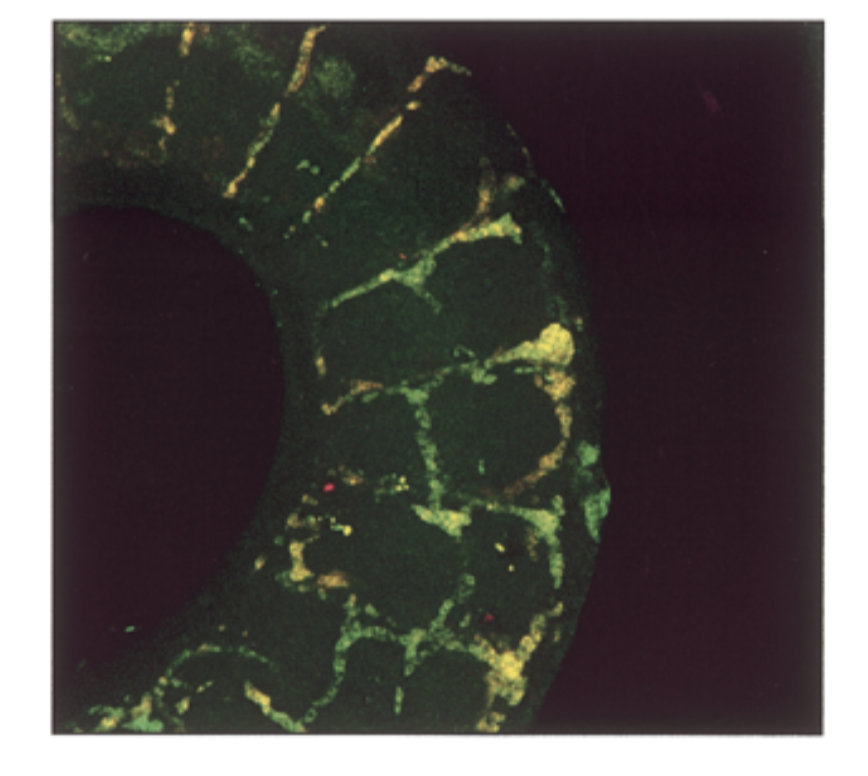
mrna exp had a lot of overlap = yellow
control
sense probe
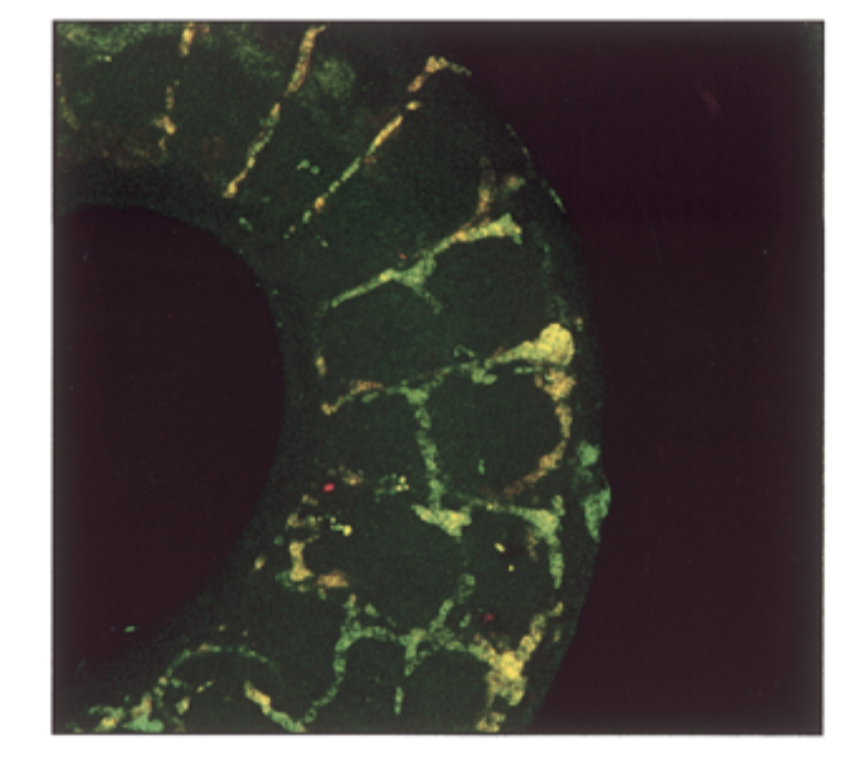
conclude
sox7=sox18 in seq and exp → fxs overlap → genes come from same ancestor
classify
hypothesis, subcell loc
question
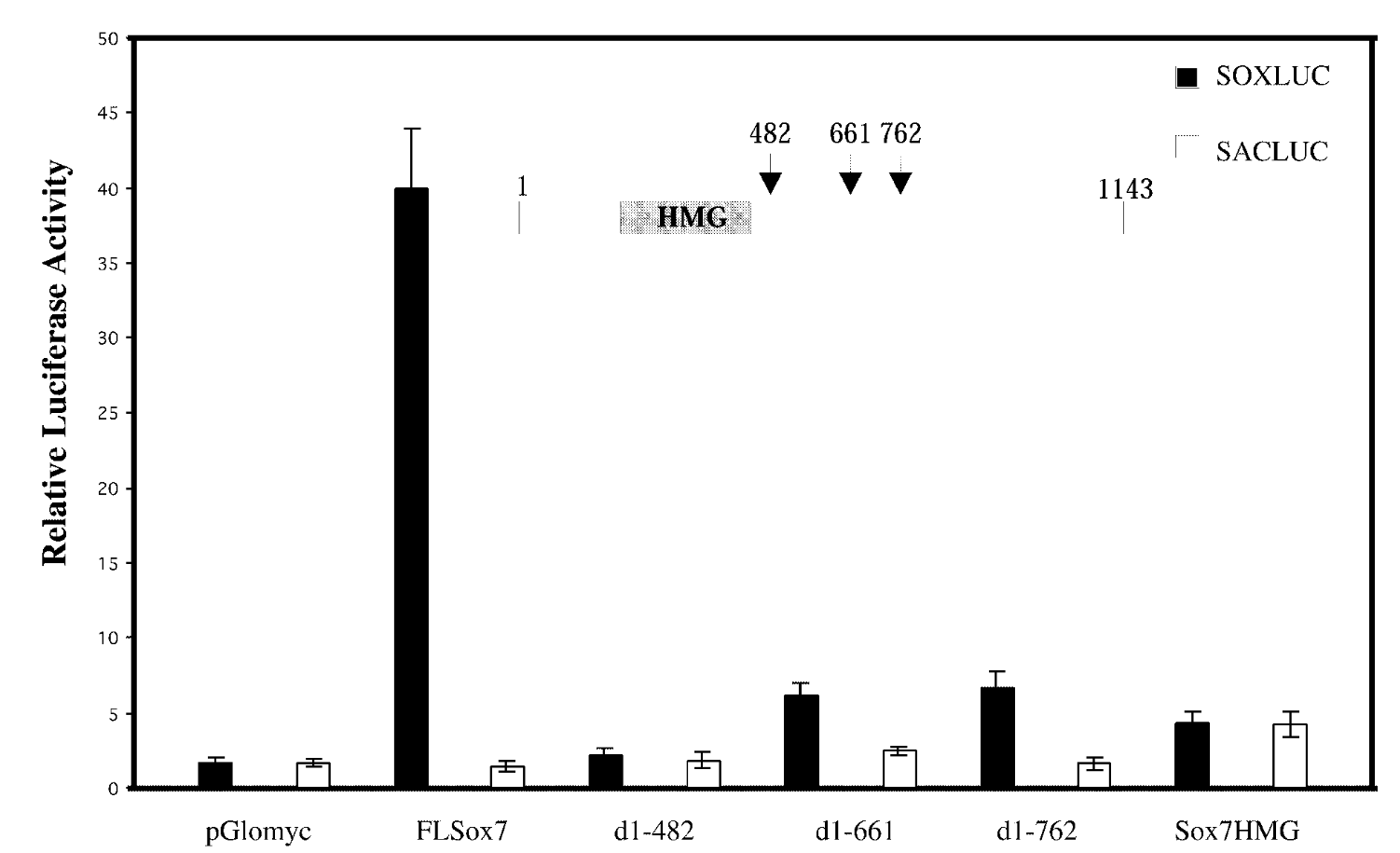
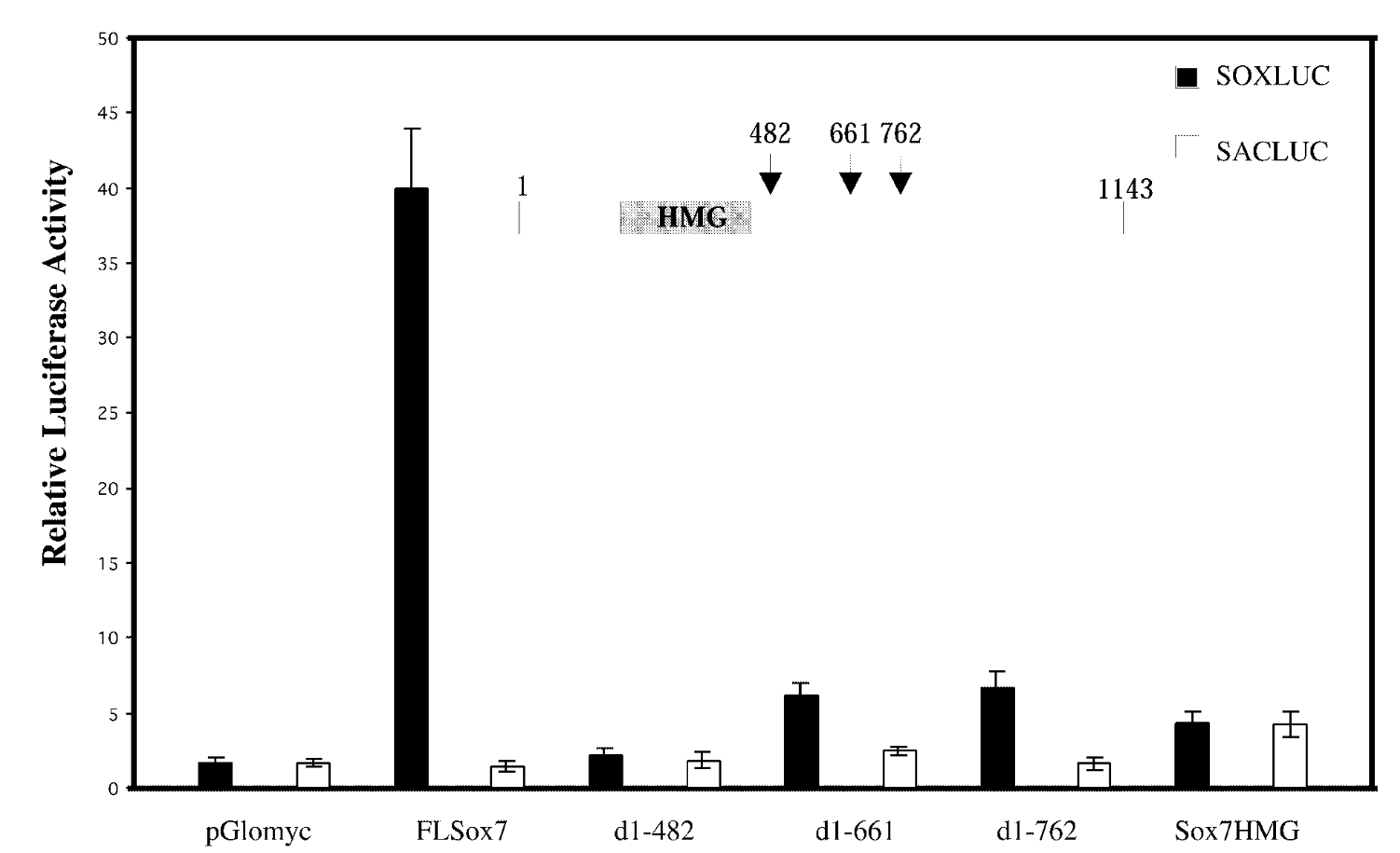
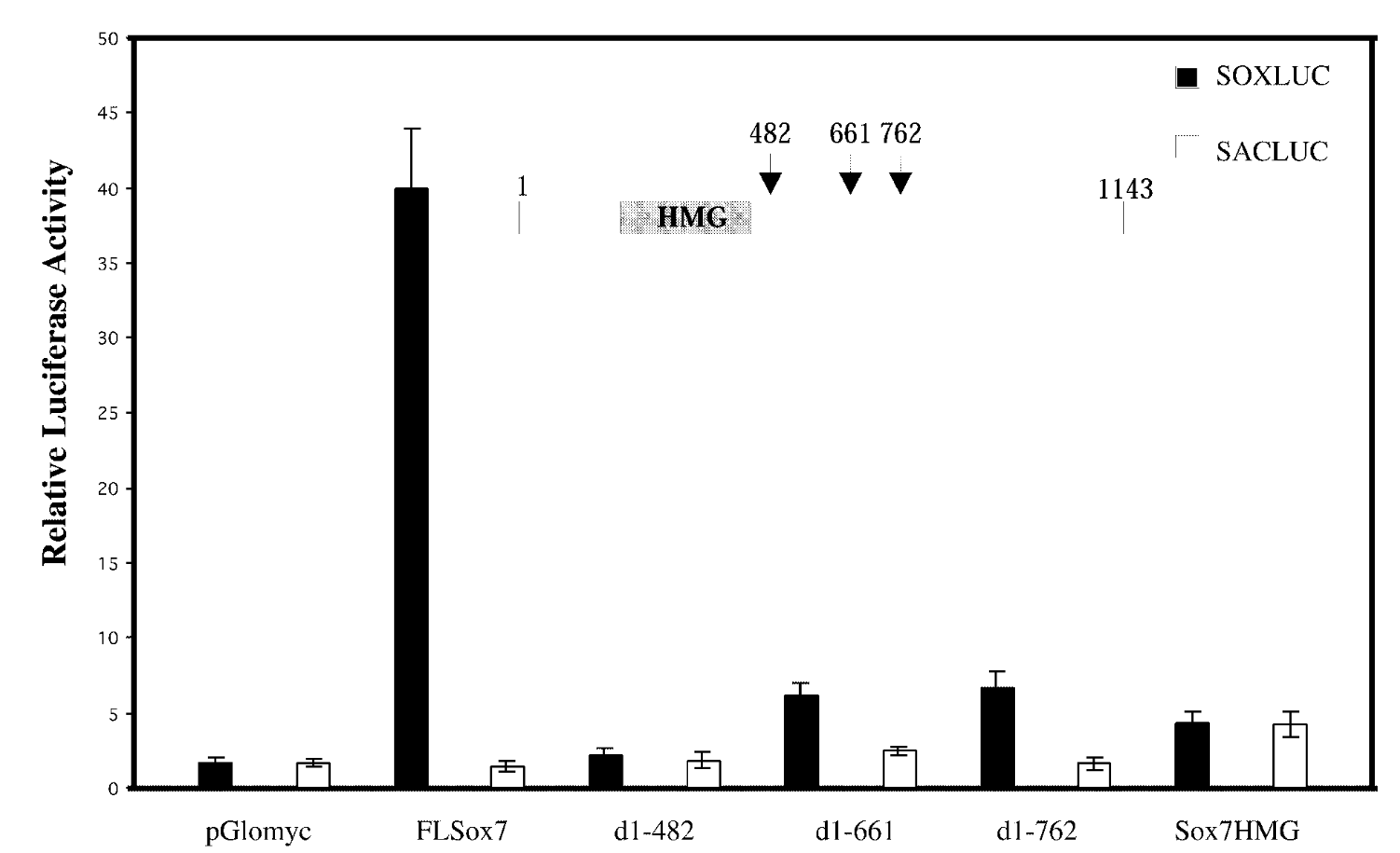
is sox7 a transcription activator or repressor and what section is responsible for activation
method
relative luciferase assay
conditions
full length sox7/sox7 deletion mutants; soxluc reporter plamsid that has motif/sacluc vector with no motif
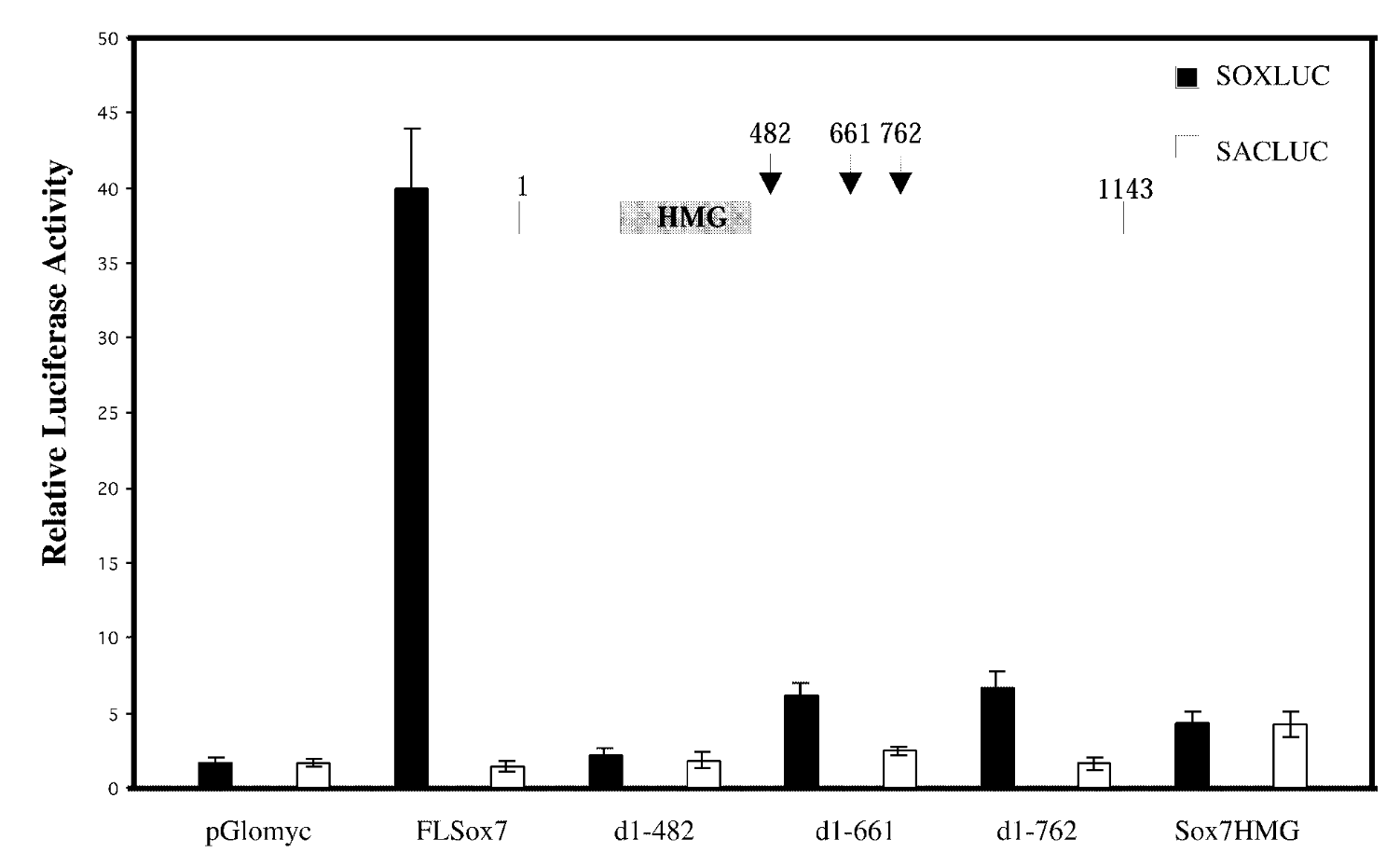
result
FL sox7 had high soxluc activity → sox7 binds to the dna binding site in reporter plasmid → sox7 is a transcription activator; mutants were not activators
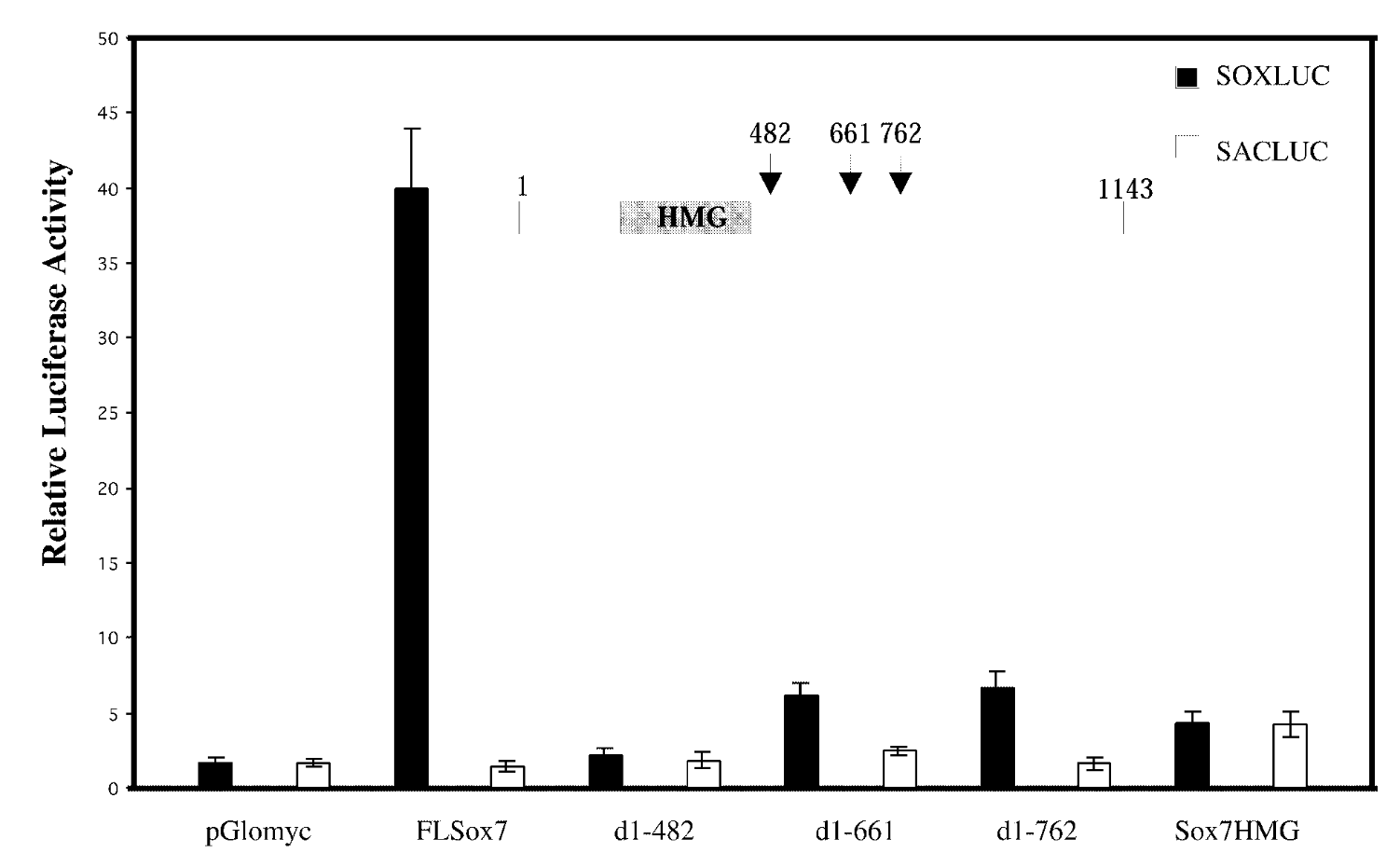
control
mutated sox7 with the c terminus cut off → baseline fx of sox7 with just hmg domain. sacluc → baseline activity of sox7. pglomyc=negative control=has nothing in it so won’t have any activity
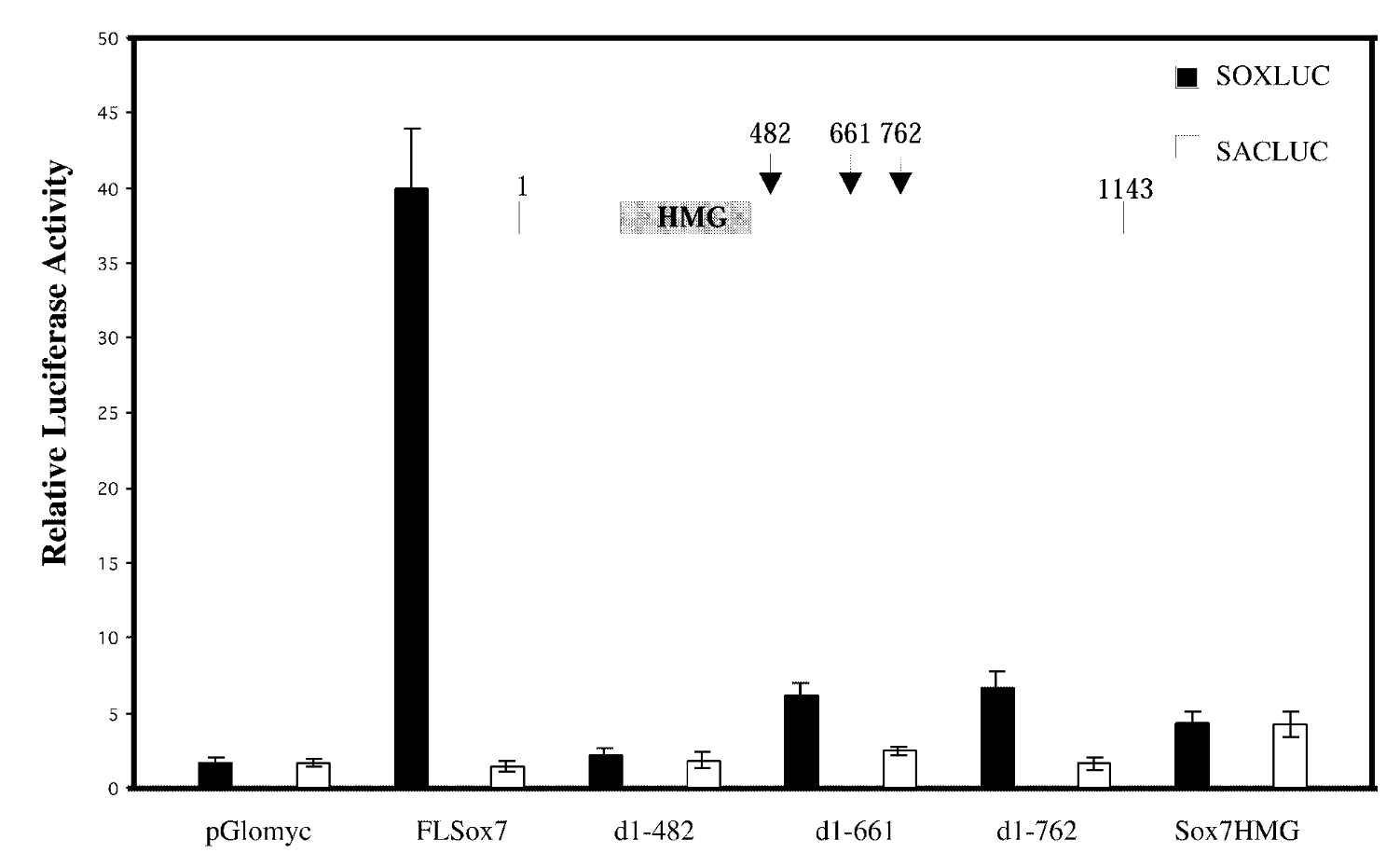
conclude
transcription activation is loc in cterminus of sox7.
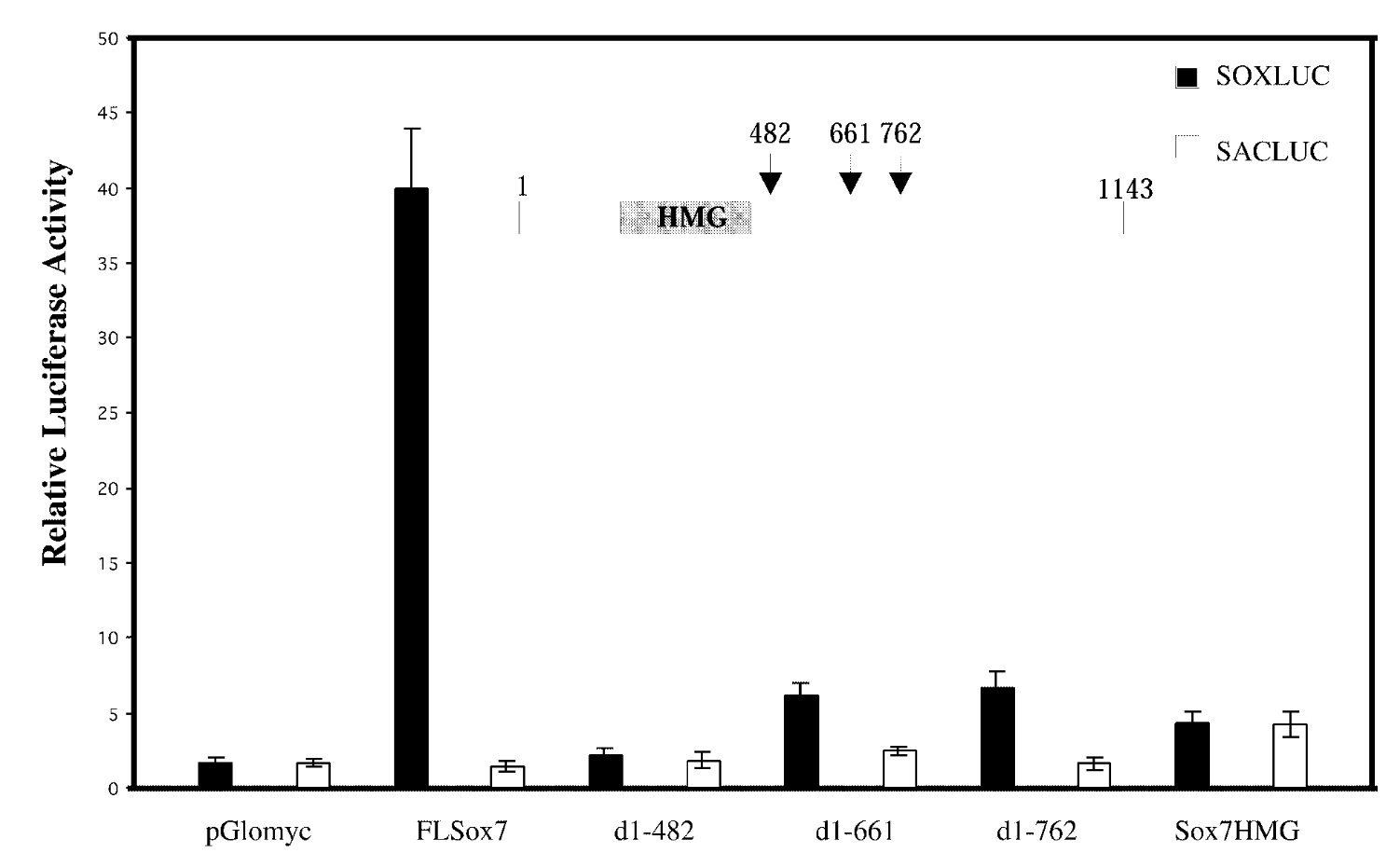
class
hypothesis, functional (does that seq work and what is it doing)
question
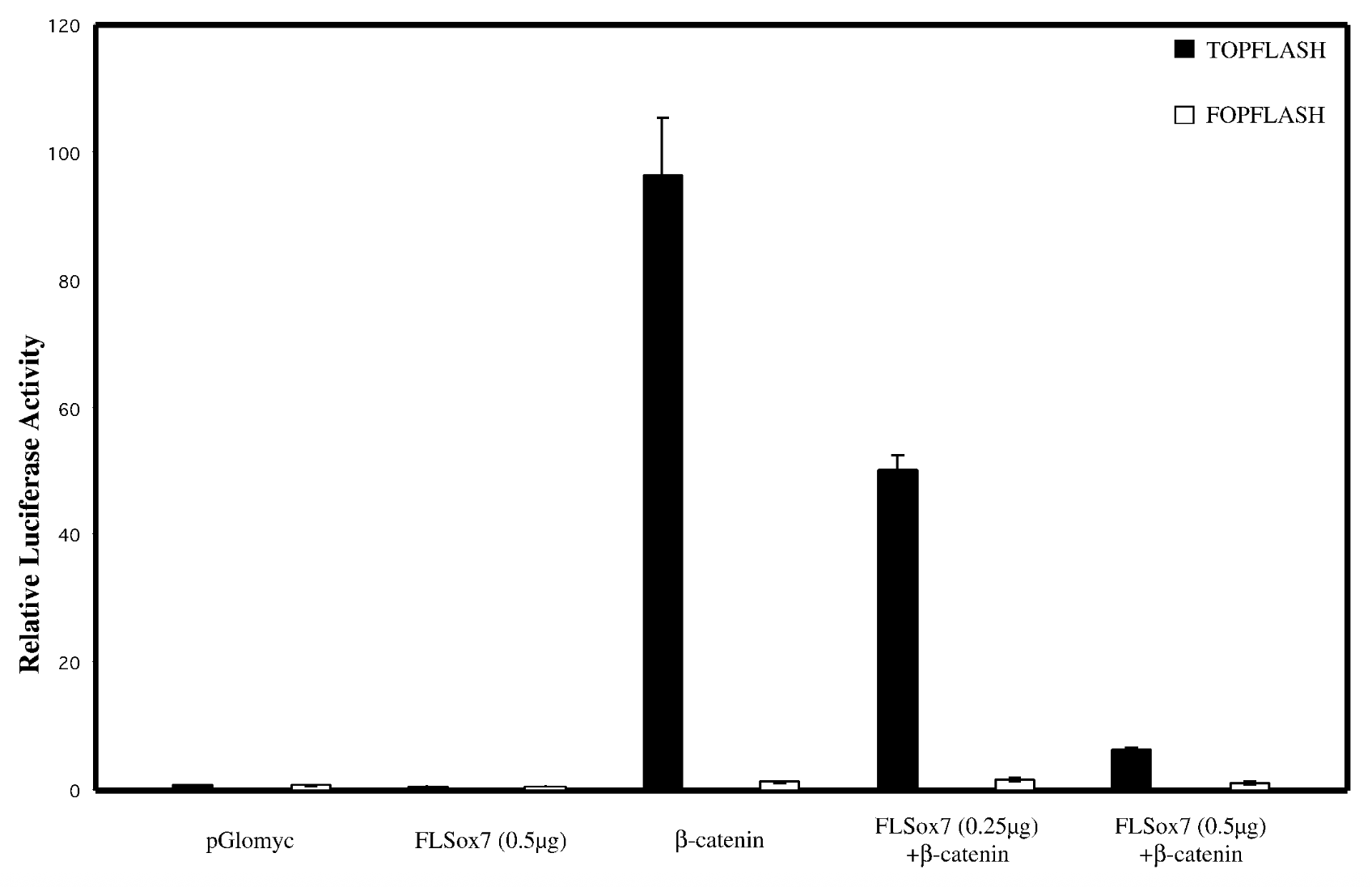
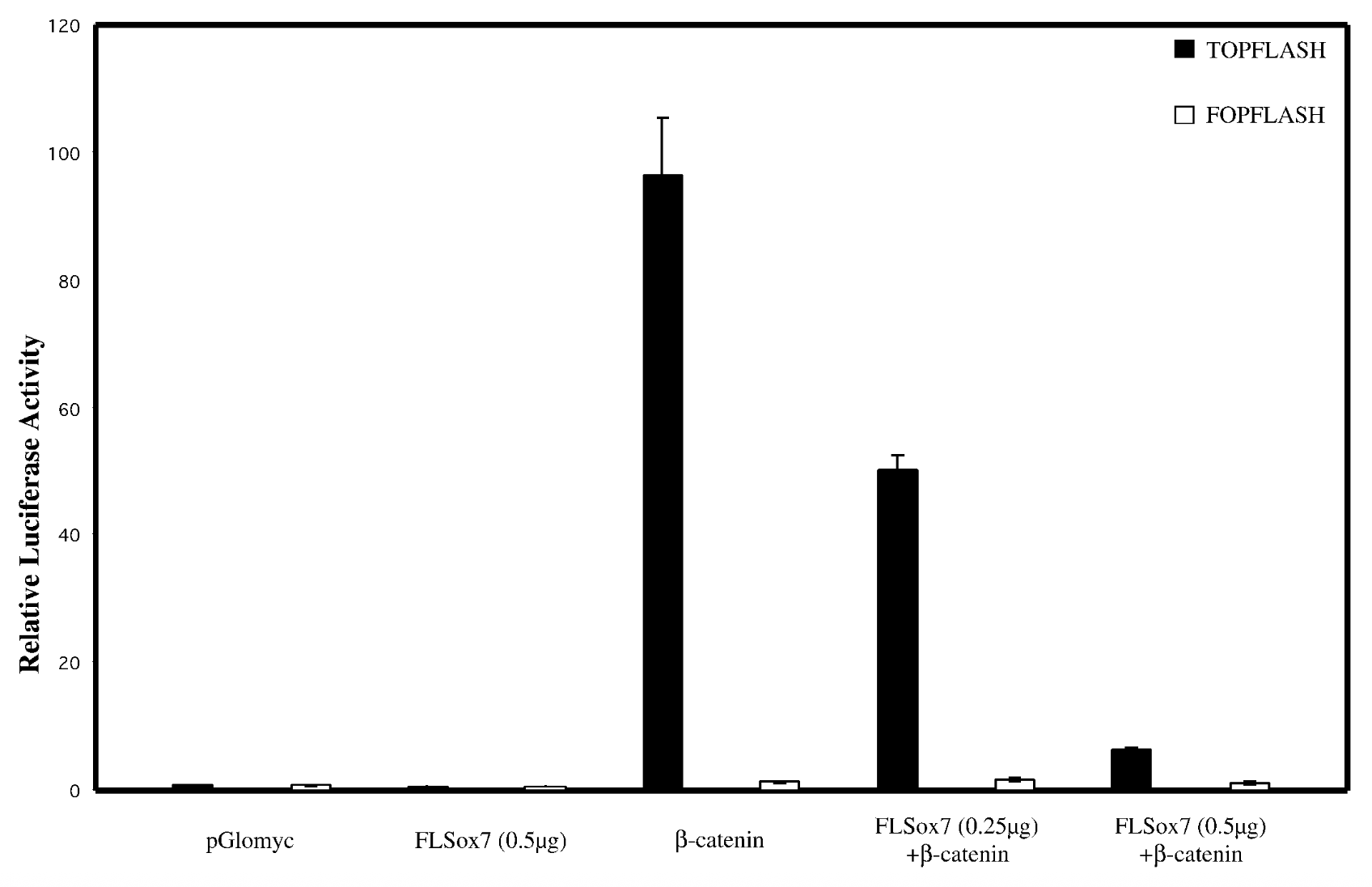
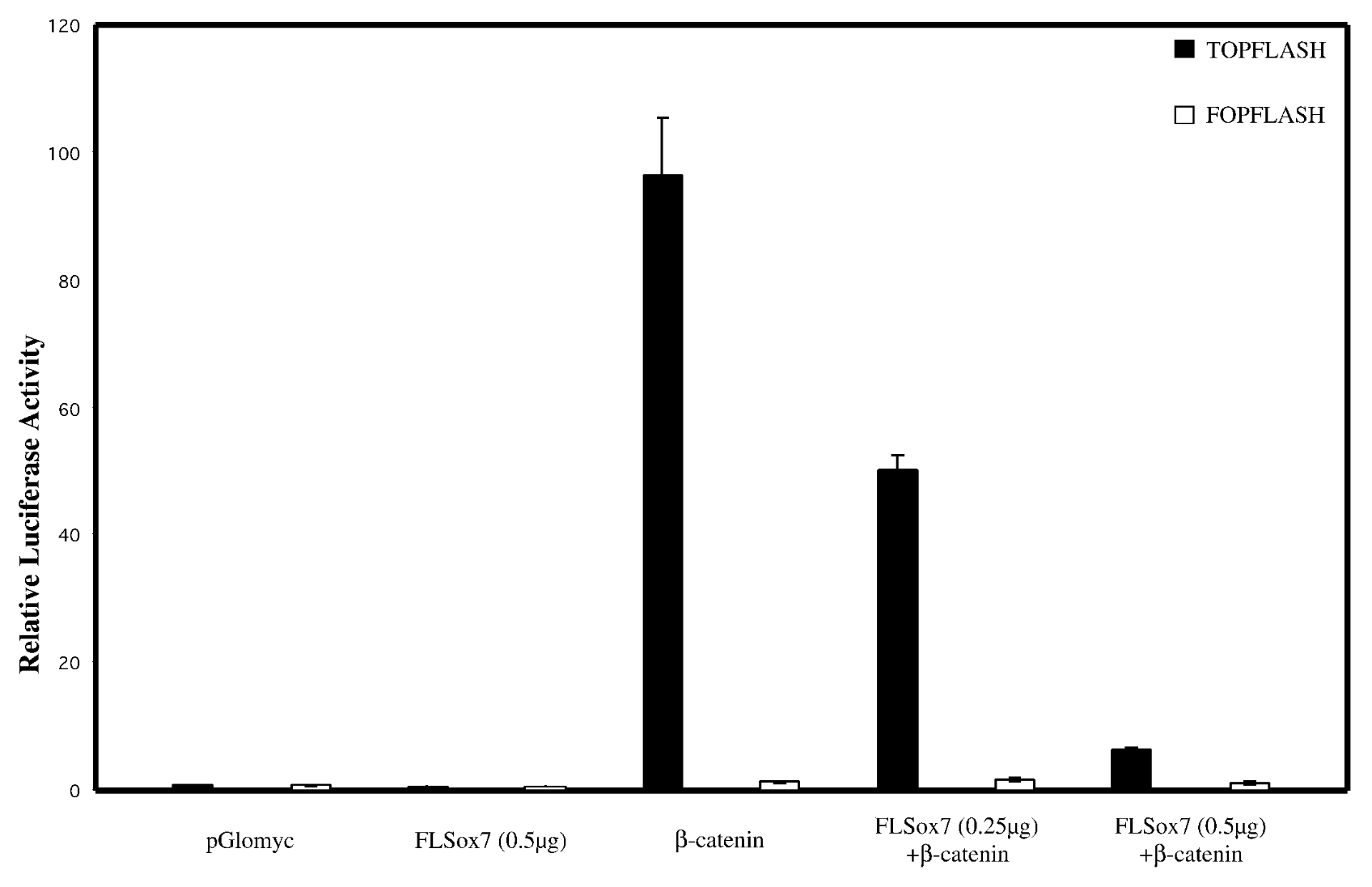
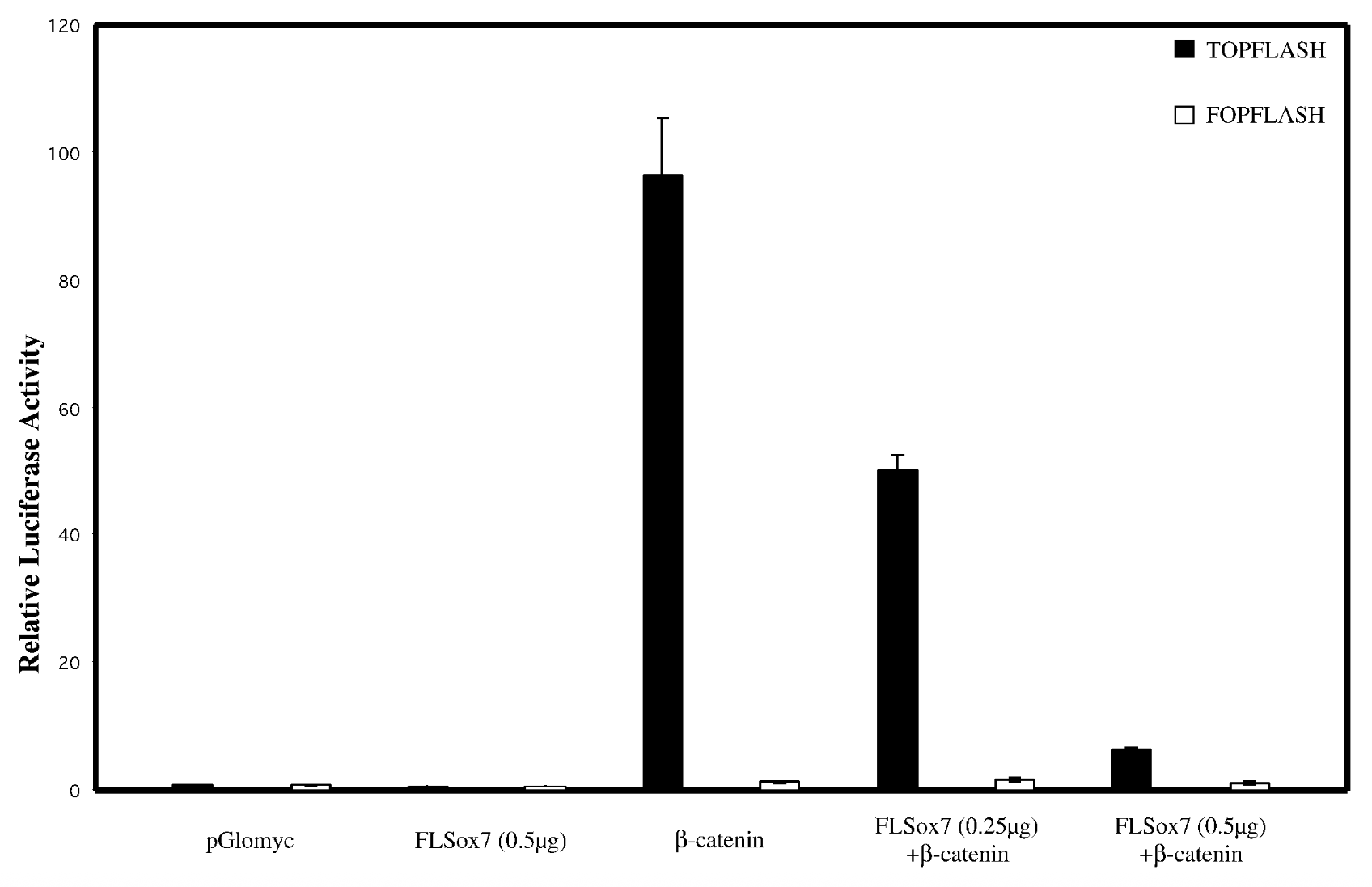
does sox7 affect wnt singaling pathway
method
relative luciferase assay
conditions
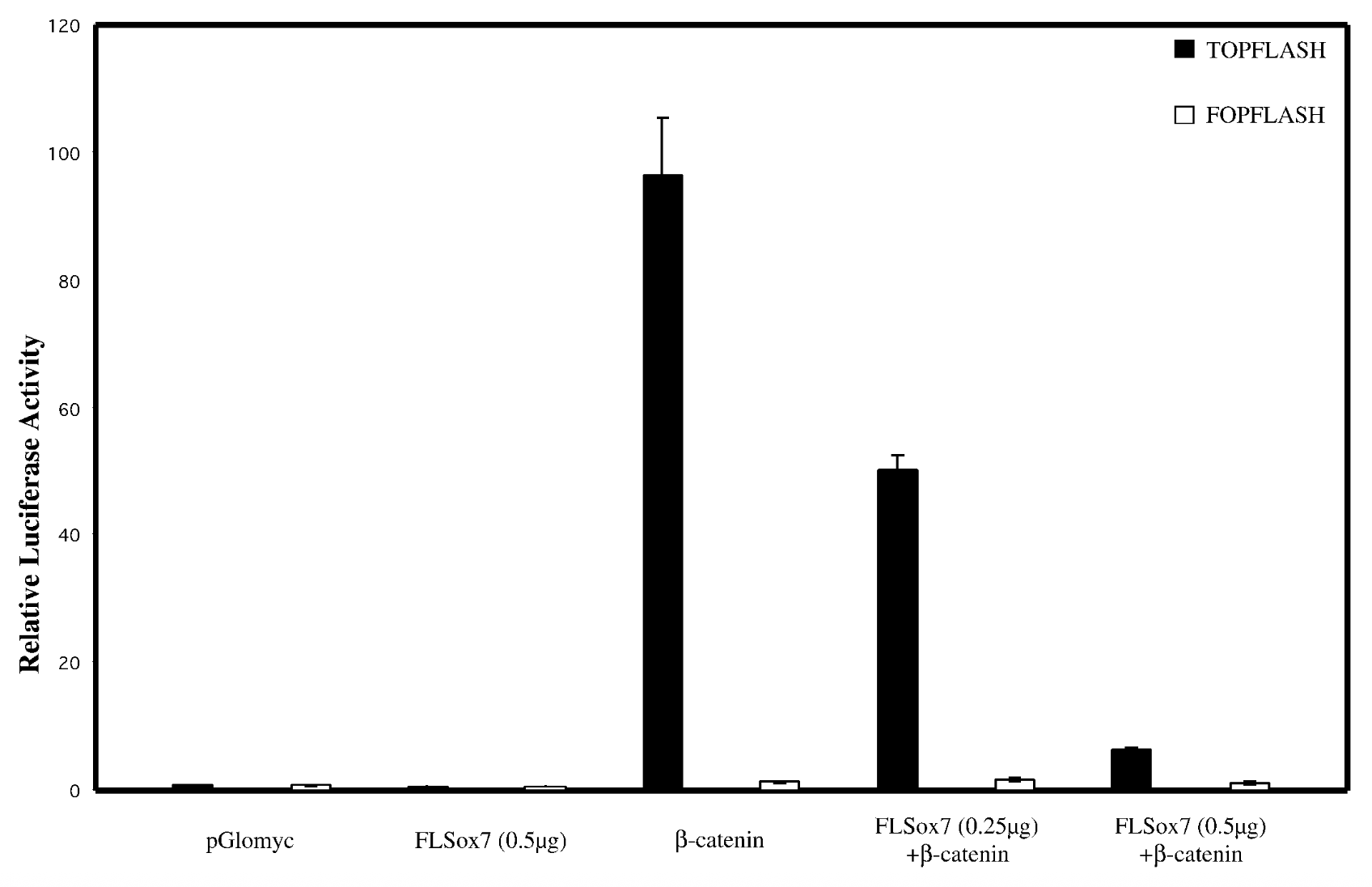
everything on x axis, bcatenins33 not regular bcatenin bc its the active stable ver + topflash=reporter plasmid that has bindnig sites, fopflash=plasmid with mutated bcatenin
result
flsox7+bcatenin < bcatenin alone → flsox7 represses luciferase reporter’s ability to activate b catenin
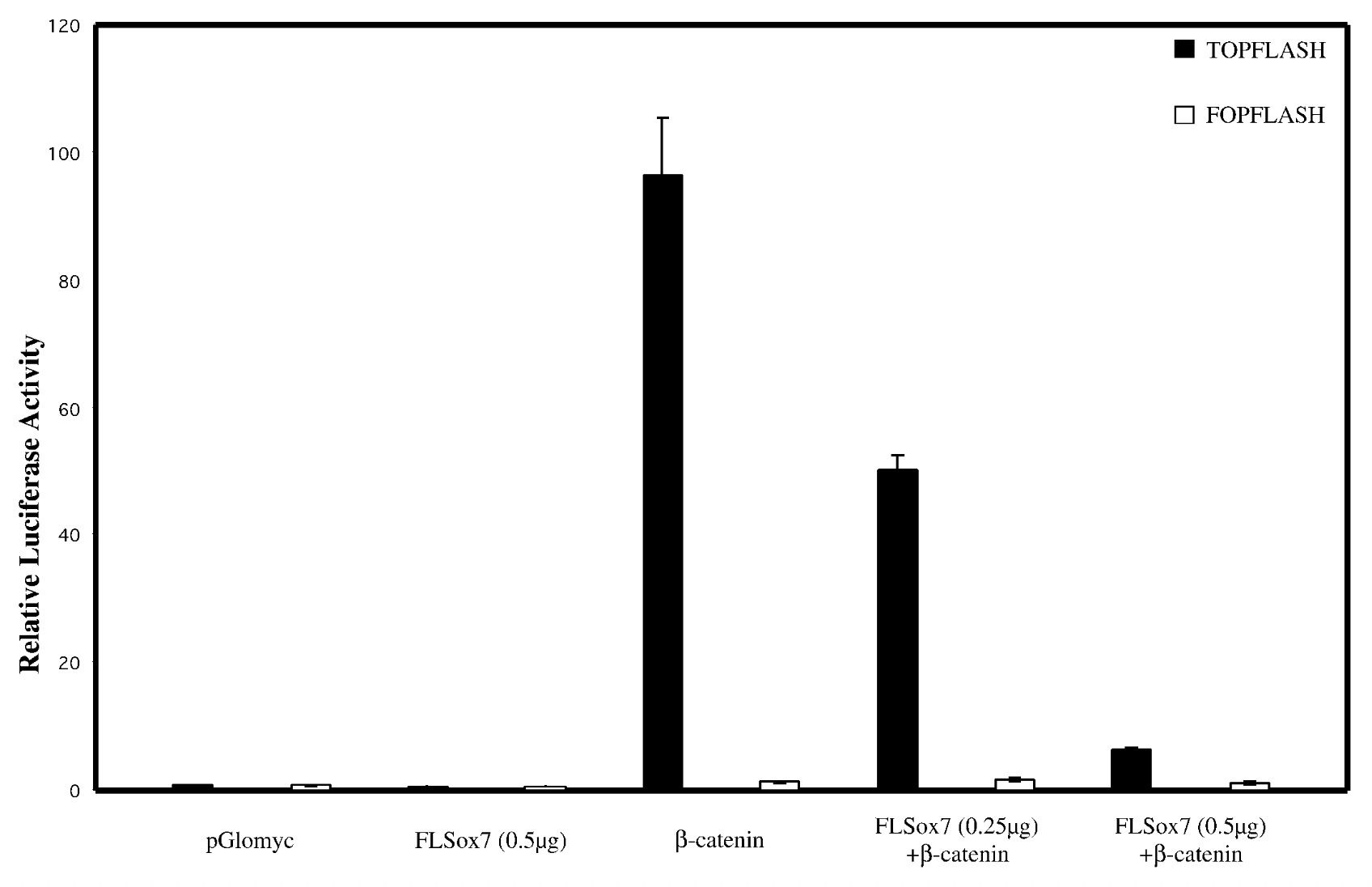
control
empty pglomyc = wild type seq with no b catenin → no bcatenin=nothing to activate. fopflash=mutated so nothing activates it
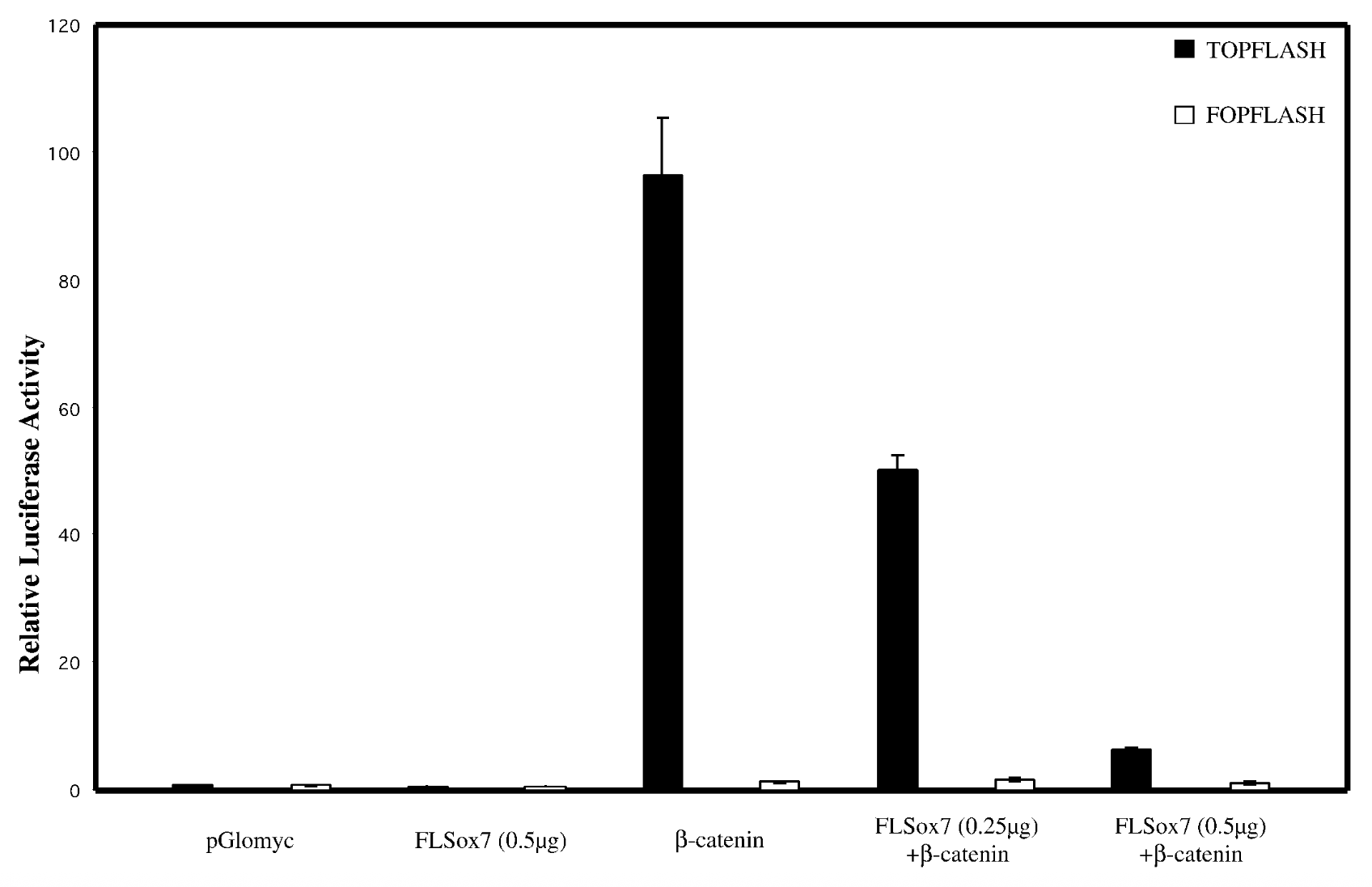
conclude
yes, sox7 impacts wnt singalling pathway. reg genes of wnt pathway get mutated in cancer → sox7 is tumor suppresor gene
class
hypothesis, functional