BIS101 Midterm 2: Protein, translation, and the genetic code
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
What do proteins do?
biochemical catalysis
provide structure for the cell
control movement
Transport of materials around the body
Signaling
Protection of the body and of individual cells
Storage functions
Proteome
The entire complement of proteins present in a cell
Protein Structure
Proteins are made of polypeptides
A polypeptide is a long chain of amino acids
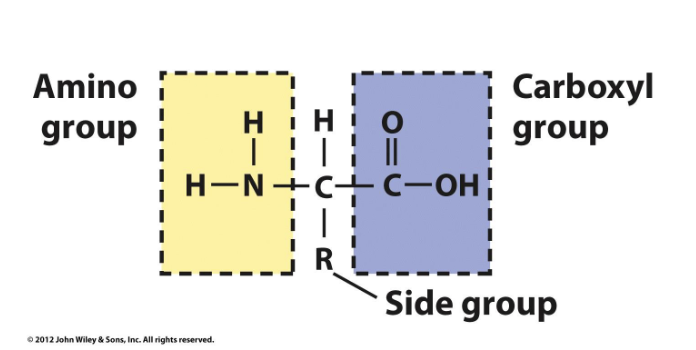
Amino acids have a free amino group, a free carboxyl group, and a side group (R)
Amino acids are then joined by peptide bonds
-COOH of one AA is attached to the -NH2 of another
Levels of protein structure
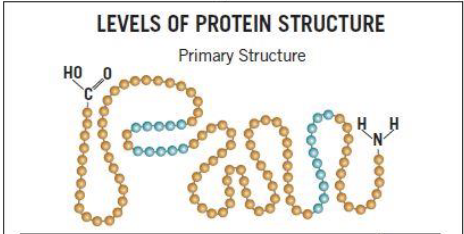
Primary structure
Secondary structure
Tertiary Structure
quaternary structure
Primary Structure
amino acid sequence
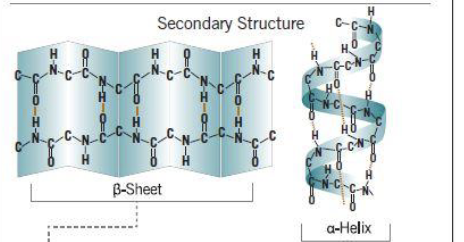
Secondary structure
local structures involving H bonds of amino acids near one another, including alpha helices and beta sheets
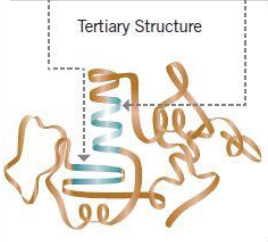
Tertiary Structure
Overall 3D shape of polypeptide
Quaternary Structure
Shape of a protein containing more than one polypeptide chain

What do we need for translation?
mRNA template
Ribosomes, composed of polypeptide and RNA molecules
tRNAs
Amino acids, and Amino-acid activating enzymes
Soluble proteins involved in polypeptide chain initiation, elongation, and termination
Ribosomes
“machines” that contain multiple ribosomal RNAs and proteins
translate mRNA in the 5’ to 3’ direction
Read each triplet (3 bps) codon and assemble the amino acids in the order specified by the codons
tRNA
adapters between amino acids and the codons in mRNA molecule
Aminoacyl-tRNA synthetase
Add amino acids to the tRNAs
Add the correct amino acid based on the anticodon the tRNA carries
Highly accurate with an error rate of ~1:100,000
Specificity of tRNAs
tRNA molecules must have the correct anticodon sequence
tRNA molecules must be recognized by the correct aminoacyl-tRNA synthetase carrying the correct amino acid
tRNA molecules must bind to the appropriate sites on the ribosome
A site
binds incoming tRNA carrying next amino acid to be added to the chain
P Site
binds tRNA to which growing peptide is attached
E site
binds departing uncharged tRNA
Prokaryotic Translation Initiation
Have a small subunit
Once small subunit binds, can build initiation complex
Find start coden then find initiator tRNA (fMet) and bring it into P site
only time that you start off in the P site
Then large subunit can assemble
Then translation can begin
How does prokaryotic ribosome know where to start?
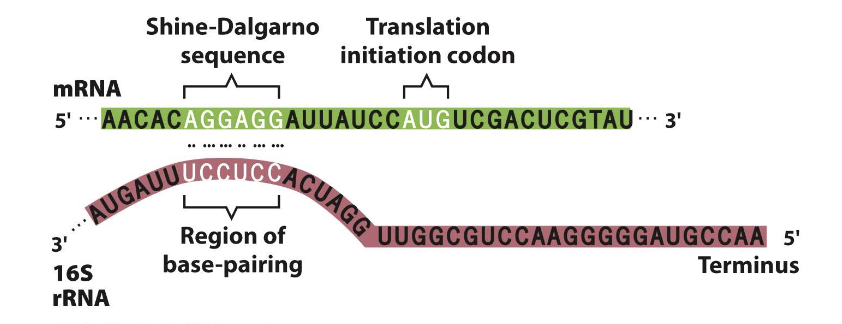
Shine-Dalgarno Sequence
neaer 5’ end of mRNA, upstream of start codon
This sequence is complementary to the 16s rRNA component of the small ribosomal subunit
The ribosome P-site is positioned opposite the AUG start codon via this RNA: RNA pairing
Eukaryotic Translation Initiation
Very similar but now have the 5’ cap so we know where to start
A complex of the small subunit plus IF bind to the 5’ cap of mRNAs
Scans for the first start odon from 5’ end
initiation tRNA interacts with in IF and binds start codon at ribosome P site
Large subunit binds, initiation is complete
How does eukaryotic ribosome know where to start?
find 5’ cap
scan along transcript
most of the time use first AUG from 5’ end
but higher efficiency of using AUG is when it’s located in Kozak sequence
even if there’s another AUG closer to 5’ cap
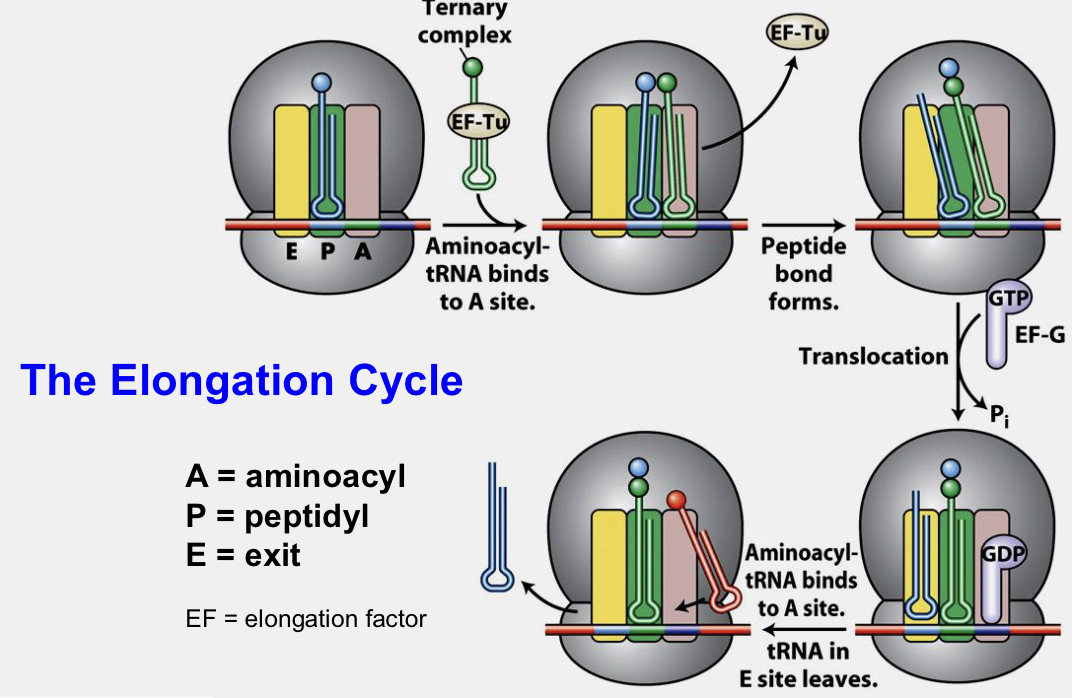
Elongation (for both prokaryotes and eukaryotes)
Aminoacyl-tRNA enters A site of ribosome, anticodon of tRNA must pair with codon in A site
Transfer of growing polypeptide from tRNA in P site to tRNA in A site
Translocate ribosome along 3 nucleotides toward 3’ end of transcript
Growing chain is in P site, uncharged tRNA in E site can then depart
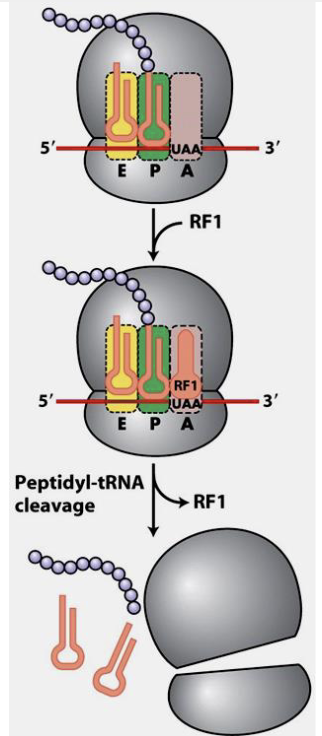
Polypeptide Chain Termination
Polypeptide chain termination occurs when stop codon enters A site of ribosome
UAA, UAG, UGA
when stop codon is encountered, a release factors binds to the A site, and the discharged tRNA in the E site departs
ribosome-mRNA-tRNA complex disassembles
Reversion Mutation
a genetic change that restores the original wild-type phenotype (appearance or function) in a mutant organism that previously acquired a forward mutation
Properties of the Genetic Code
It is composed of nucleotides triplets (codons)
It is nonoverlapping
It is comma-free
there are no pauses, punctuation, or gaps between codons
It is degenerate (more than one codon codes for the same amino acid)
It is ordered
It contains start and stop codons
It is nearly universal
Third Base Wobble
There are 61 codons that specify amino acids, but only 30-50 tRNAs in most genomes
This means there aren’t enough tRNAs for each to have a unique codon
Relaxation of the strict complementary base-pairing rules at the third base of the codon is called third base wobble
this permits a single tRNA to bind more than one codon in the mRNA
Post-Translational Processes
Protein Folding
Chemical modification of amino acids
Cleavage of polypeptides
Protein Folding
Primary, Secondary, etc structures
Proteins called chaperones help other proteins fold correctly
can help refold if they unfold
can sequester misfolded proteins
if a protein cant fold, its a loss of function
if misfolded proteins build up, they cause problems
Chemical modification of amino acids
Adding a new chemical group to a particular AA
Cleavage of Polypeptides
removal of fMet from N-terminus in bacteria
Polypeptides may be cleaved into multiple segments that have separate functions or that aggregate to form a functional protein