bibc 100
1/91
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
92 Terms
fours types of weak interactions between water and biomolecules
hydrogen bonds
ionic interactions
phosphorylated aa
serine, threonine, tyrosine
histidine
regulated enzyme activity
protein/protein interactions
protein degradation
signal transduction
sulfanated aa
tyrosine
hormone receptor-ligand interactions
G protein signaling
Methylation
lysine, arginine
epigenetic regulation
histone function (charge is maintained)
Hydroxylation of aa
Proline, lysine
O-linked glycosylation
phosphorylation
Carboxylation
glutamate
affinity for Calcium ions
hydrogen bonds
between H2O due to electron structure across O-H
O more electronegative
electrons not equally distributed → dipole
due to tetrahedral arrangement of orbitals → interacts 4 neighbors
hold h-bonds with average of 3.4 water molecules
O-H h-bond length x2 the covalent
dissociation E is 23kJ/mol Hbond < 470kJ/mol covalent
lifetime between each other is 1 to 20 picoseconds
high cohesion → melting/boiling point are higher for water cus stick together
polarity and its affects on interactions
polar → large electronegative diff
all weak interactions except hydrophobic
non-polar → small electronegative diff
hydrophobic and van der Waals
amphipathic → part hydrophobic part hydrophilic (phenylalanine + phospholipids)
all weak interactions
ionic interactions
electrostatic interactions based on O-H bond dipole of water → interacts with chaarged ions
ionic bonds are tranfered interactions are not
hdipole+ with Cl- → attractive
ionic salts want to be disordered and not in Crystal structure
hydrophobic interactions
free moving water is favorable over caged water
less total surface area → smaller decrease in entropy → more energetically favorable
maximum possible entropy means more energetically favorable
aggregation
hydrophobic interactions do not occur because non-polar solutes attract one another
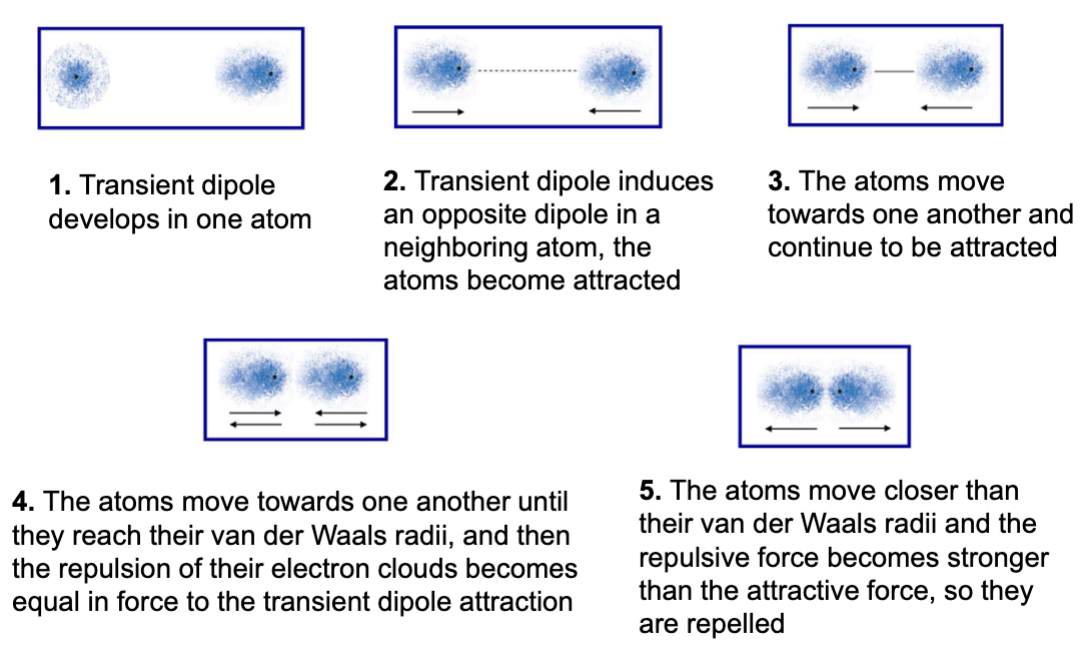
van de Waals interactions
very weak transient version of other electrostatic interactions
fluctuating dipole
random variations in electron positions around a nucleus causes transient dipole
distance is important
equilibrium is achieved when attractions and repulsion are balanced
lipid bilayers are a result of ___ effect on ___ solutes
hydrophobic + amphipathic
polar head on the amphipathic molecule
causes what look like ordered structures
__ molecular interactions drive protein folding
weak
hydrophobic inside hydrophilic outside
charged/polar amino acids at the surface to participate in electrostatic interactions
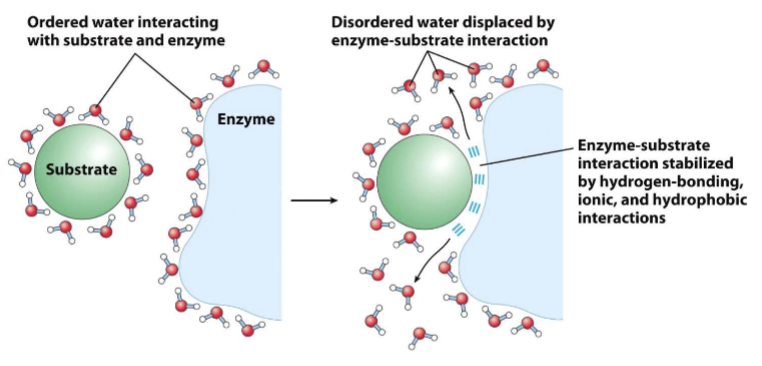
release of ___ water facilitates formation of___ complexes
ordered + enzyme-substrate
non polar solutes cause water to be constrained into more orderly orientations along interface
polar cause constrained water molecular motion to lesser degree bc compensatory electrostatic interactions
disruptions of ordered water drives interaction of polar substrates with the complementary polar surface of an enzyme
entropy increases as the water molecules associated with the substrate and enzyme are displaced
__ interactions are critical to __
weak + biomolecule stability
cumulative effects of interactions has significant strength
large # of weak interactions → more stable complex
to dissociate 2 biomolecules joined through multiple weak interactions → need disruption all at the same time
this is less probable so the stability of the weak interactions is greater than would be expected if they only had an additive effective
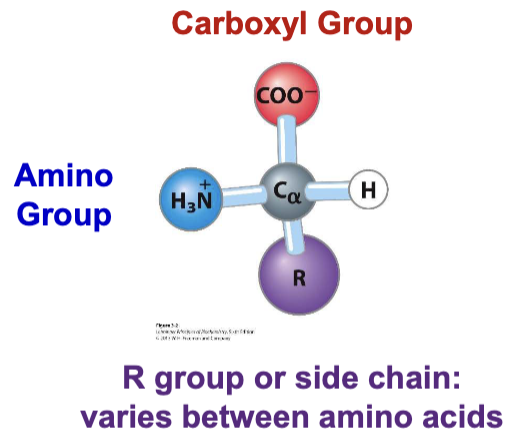
proteinogenic amino acid
alpha carbon - carboxyl and amino group bonded to same carbon
stereoisomers
L AA → amino group before carboxyl clockwise
D AA → carboxyl before amino clockwise
20 AA in 5 groups based on side chains
nonpolar or aliphatic (hydrophobic)
aromatic (varies)
polar uncharged (hydrophilic)
positive charged (basic)
negatively charged (acidic)
stacking of aromatic AA
pi-pi stacking
stabilizing protein structure
mediating interactions with other protein, DNA/RNA, small molecules
overlapping orbitals
interaction between TRP side chain in DNA polymerase eta subunit and adenine residue in DNA
positive charged AA
lysine
primary amino group , pka = 10.53
arginine
guanidinium group , pka = 12.48
histidine
aromatic imidazole , pka = 6
lysine and arginine stabilize protein binding to nucleic acids interacting with __ backbone
phosphate
positive charge on Lys/Arg with negative phosphate groups on backbones
involved in protein interactions w nucleic acids
ex: p53 DNA binding protein contacts the DNA backbone through K120, R248, R273 and R280
Mutations of these residues in p53 often result in cancer due to disrupted p53 function
histidine is — and — to small shifts in —
aromatic and sensitive to small shifts in pH
aromatic → charge is shared across bonds
only ionizable amino acid with a side chain pka near physiological pH
small shifts in pH can alter charge
common enzyme active sites and participates in many catalysis reactions
negatively charged AA side chains
acidic
found at the external face of proteins where they participate in electrostatic interactions
(-) charge of asp/glu complementary to the charge of lys/arg can mediate protein interactions
asp/gly involed in coordination of metal ions
general pkas of side chain
lys/arg → 10-12
his → 6
asp/glu →3/4
Two other amino acids have been found to be genetically encoded and incorporated into proteins
selenocysteine
TGA-codon
pka = 5.2
cysteine with sleneium instead of sulfur
pyrrolysine
found in methanogenic archaea and bacteria
lysine with pyrroline ring attached
bc selenocysteine is an effective — it is important in the active site of many —
nucleophile + redox enzymes
selenium is more efficient nucleophilic attack
ex: required active site of gluthione peroxidase (antioxdant protect cell’s lipids)
seleneocysteine sec is encoded by an —- —- —- —- in some proteins
opal stop codom UGA
to encorpoart sec into proteind uring translation mRNA transcirpt must contain 2 structural elemetns
opan stop codon complementary to tRNA carrying sec
secondary structural element in 3’ untrans region of mRNA formed by SECIS
60 nucleotide in length & forms stem loop
secondary structure of secis
The translational machinery pauses at the opal stop codon
The stem-loop structure of the SECIS physically interacts with the translational machinery allowing the Sec tRNA to be recruited to the mRNA being translated
The Sec tRNA delivers Sec to the opal stop codon
The 10 bp tRNA acceptor stem of Sec tRNA is longer than most (most are 7-9 bp) to facilitate interaction with the paused translational machinery
pyrrolysine
present in active site of methyl transferase
ring coordinates positioning and displaying the methyl group
incroporated at a stop codon Amber
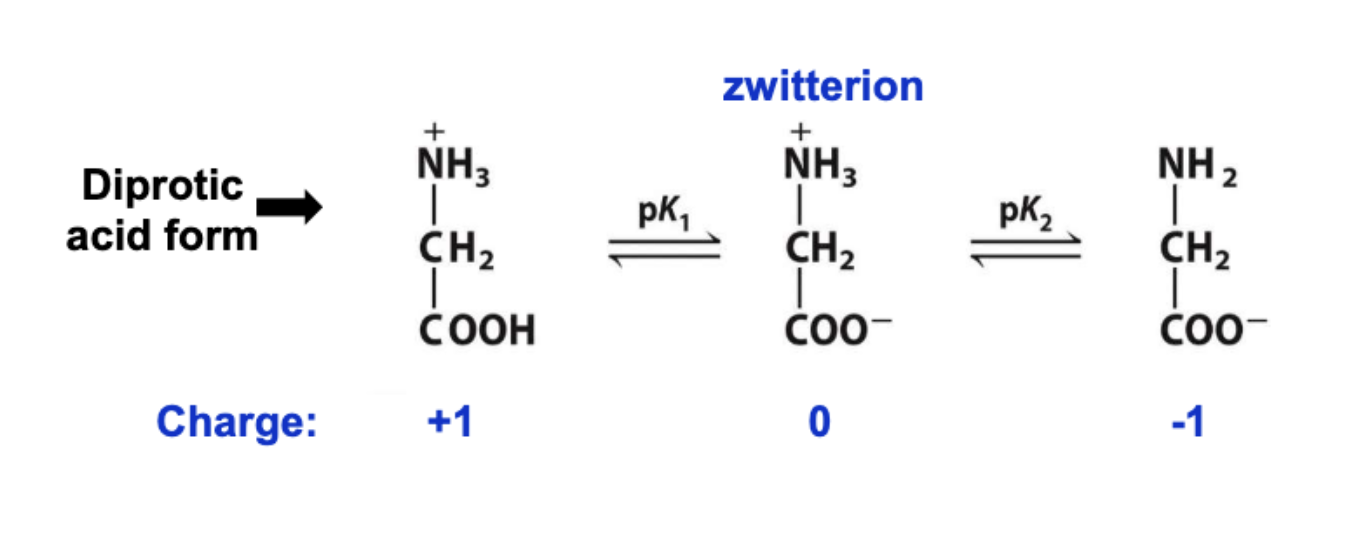
zwitterions
dipolar ions acts as acid or base
at neurtral pH → aa are present as zwitterions
pk1 = dissociate of H+ from carboxylic acid group
pk2 = amino group
titration curve for glycine
2 buffering areas
dissociation of H+ CA is pk1 = 2.3
isoelectric point: 5.97 (charge is 0)
dissocaition from amino acid = 9.6
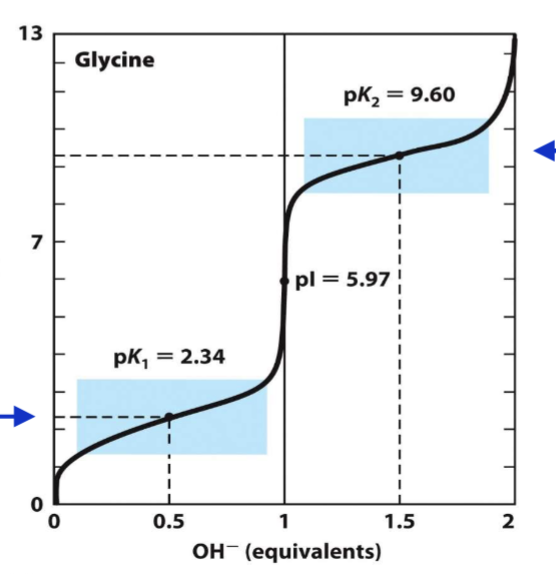
structure of aa affect pka
pka of carboxyl is lower than other CO
→ opposite charge amino group stabilizes the zwitterion
pka of amino is lower
→ electronegative carboxyl pulls e- from amino group
titration for charged aa
2 zone of buffering + zone of buffering for loss of H+ for side chain
histidine = 6
lys/arg = 10-12
asp/glu = 3-4
peptide form through condensation
joined through amide group and hydroxyl
amino acids in peptide chain are called residues
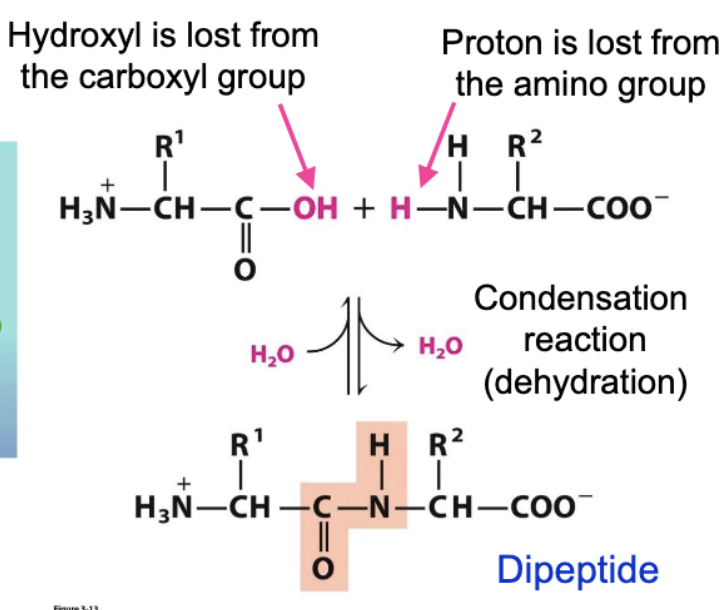
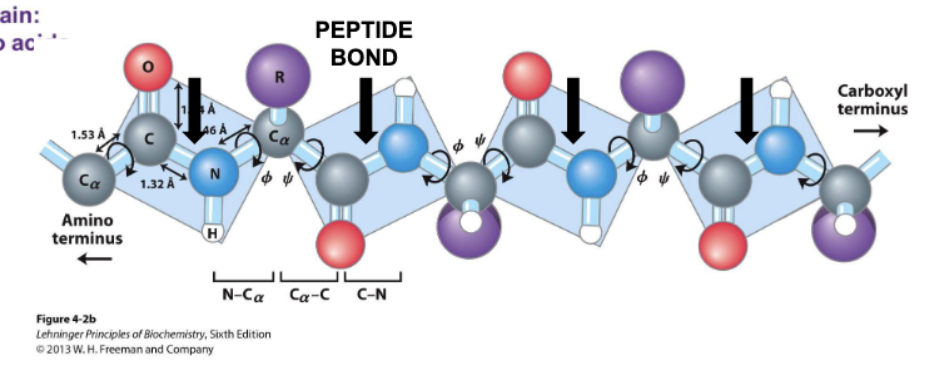
where/what are peptide chains
composed of repeating amino nitrogen, alpha carbon, and carbonyl
have amino and carboxyl terminus
oligopeptide: >2 residues
polypeptide: <10,000 kiloDaltons
protein >10,000
examples of peptides
small are important
aspartame, oxytocin, amantin (toxin from mushrooms inhibits RNA polymerase
aa side chain can coordinate metal cofactors that enhance enzyme activity
aa: his, cys, asp, glu, selenocysteine
metals: iron, zinc, magnesium, calcium
ex: rebonucleotide reductase → iron atoms coordinated in the active sire by glu, his, asp
triggers free radical production in tyr to deoxygenate ribonucleotides
alcohol dehydrogenase
zinc atom is coordinated in active side by cys and his residues
zinc binds an alcohol mol in position to facilitate hydride transfer to coenzyme nicotinamide
oxygen binding proteins contain — — groups to bind to oxygen gas
heme prosthetic
none of aa side chaine in proteina re suitable for reversible binding oxygen
solutions: coordination of heme prosthetic groups containing iron to reversibly bind oxygen
modifcations to aa
add/remove charge groups
add functional groups
local changes → protein structure → leading to changes in protein function
enzymatic function
protein localization in cell
ability to form complexes
post transitional modifications
phosphorylation (ser, thr, tyr - eukaryotes) (his prokaryotes)
sulfation - tyr
methylation (lys, arg)
phosphorylation importance
transcirptional activity
proetin stabilty
subcellualr localization
creat confomartipnal change -> ATP
hydroxylation
proline and lysine
permit other modifications such as O-linked glycosylation or phosphorylation
carboxylation
glutamate for clotting
increases affinity for Ca+
glycosylation (linked)
O-linked: to hydroxyl oxygen of Ser, Thr, Tyr, Hydroxyproline, hydroxylysine
N-linked: to nitrogen of Asn (or very rarely to Arg, only known in bacteria)
C-linked: very rare, to Trp
critical to protein stability, at least 40 human diseases are a result of mis-glycosylation, none of which have a known cure!
acetylation
lys
epigenetic regulation
histone structure
interaction with DNA
antibody glycosylation
stabilizes them
AB for immunity
directly block viral entry into cells
AB tagged molecules or organism recognized by other immune cells
— groups can be attached to amino acids
to facilitate —-
hydrophobic + protein localization to membranes
Myristoylation: Myristic acid (C14) tail covalently attached by an amide bond to the alpha-amino group of an N-terminal glycine residue, important for signaling and apoptosis
Palmitoylation: covalent attachment of palmitic acid (C16), to cysteine and less frequently to serine and threonine residues of proteins. Can be reversibly removed, important for hemagglutinin-mediated influenza infection
Prenylation: transfer of either a farnesyl or a geranyl-geranyl moiety to C-terminal cysteine(s) of the target protein, important for protein-protein interactions
Hydrophobic groups can be attached to amino acids to facilitate protein localization to membranes
Hydrophobic groups insert into the cell membranes – this allows proteins to localize within the cytosol to the face of specific membranes
Palmitoylation is readily reversible, and can regulate function of a protein through changes in localization
i.e. Some signaling proteins are not functional unless they are present at the plasma membrane, and their function can be negatively regulated by removing them from the membrane (depalmitoylation)
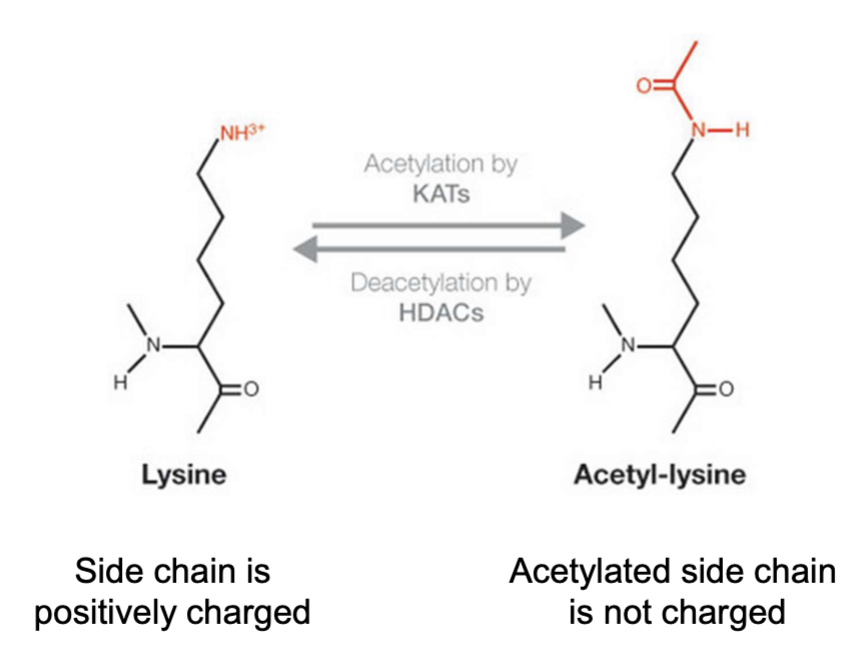
— acetylation is important for regulation
of DNA accessibility and gene transcription
Lysine
DNA is packed around histone proteins in our cells
Histone proteins are spherical-shaped octamers made from 5
types of subunits, named H1-H5
DNA is wrapped around the exterior of the histone octamer
Lysine acetylation weakens histone interactions with DNA
positive lysine residue interact wiht negative phophaste on DNA
K64 can be acetylated which weaken interaction with DNA cus there is less charge
DNA falls off histone
disruption of electrostatic interaction
loosed DNA → opens up interact w transcriptional machinery
regulate whether genes are turned on or off
chracteristic of protein
size, charge, polarity
chromatography → protein run through column with beads/matrix
separate from one another depending on structure
charge → ion exchange chromatography
beads have negative charge + proteins that are negative move fast
positive take the longest
anion change is the opposite
size → gel filtration
beads w channels → large protein goes fast
small protein go slow
polarity → hydrophobic interaction (Reversed phase)
hydrophobicity
nonpolar stationary phase
acivity assay
try to find specific protein
sequence multple seperations
peptide bonds and rotation
dipole and bond is shorter and rigid
no fre rotation around bond
phi (ϕ) Rotation is around the N-Cα bond
psi (ψ) Rotation is around the Cα-C bond
large side chain aa have fewer possible angles of rotation
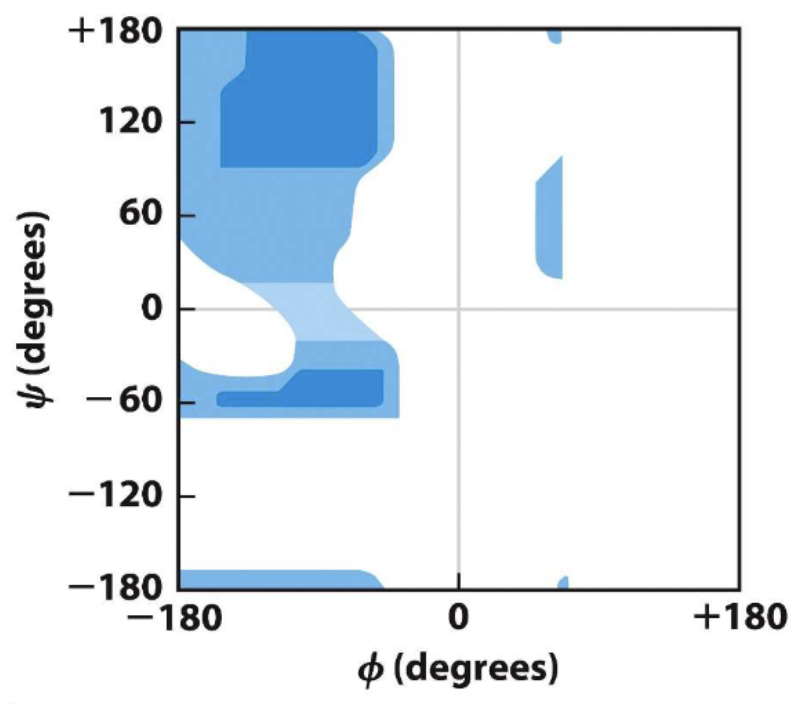
phi psi angles plot
alanine
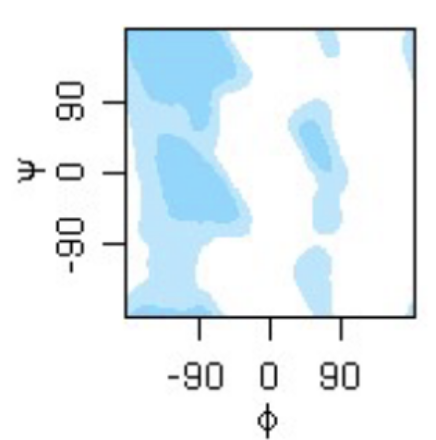
phi psi angles plot
general
glycine
Proline
Before proline in the peptide chain
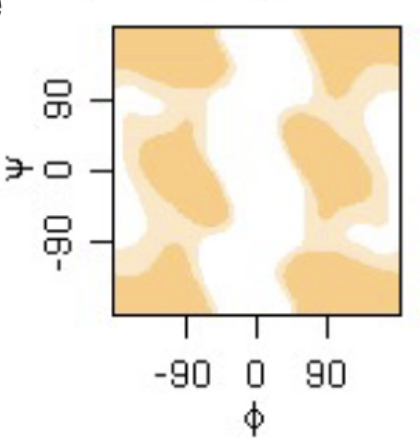
phi psi angles plot
glycine.
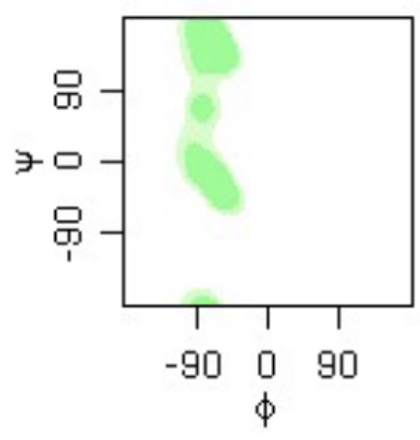
phi psi angles plot
proline
right handed alpha helix, beta sheets, left ahnded beta sheets
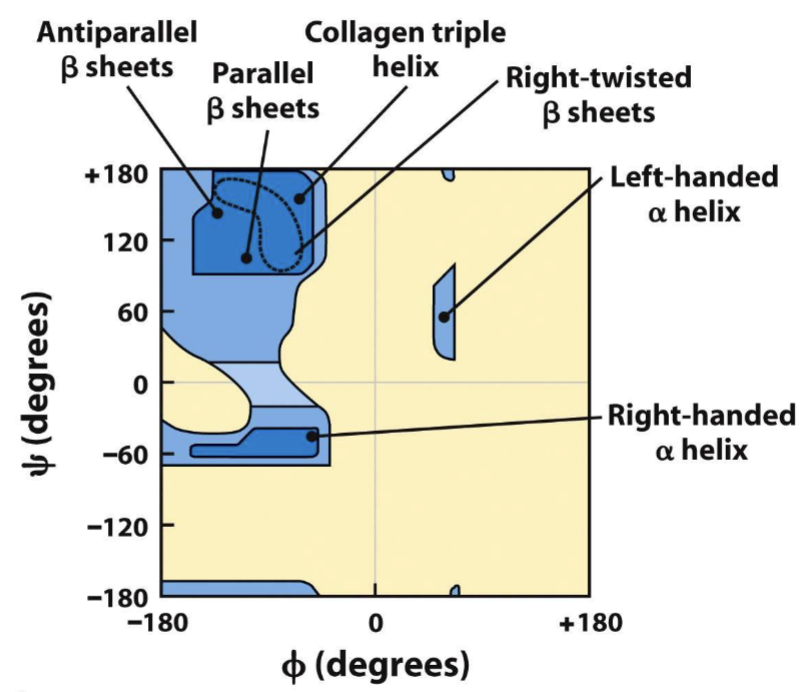
alpha helices
stable bc hydrogen bonds
H attached to N
electronegative carbonyl O attached to peptide bond C
net dipole : amino has partial positive, carboxyl has negative
right handed are more stable for L amino acids
less likely to find glycine and proline in a helix
stability effected by:
phi/psi bond angle
proximal r groups (opp charges stabilize)
bulky R group destabilize
gly too much rotation, pro no enough
same aa can destabilize
helical wheel projection
Amphipathic helix
(one side polar, one side nonpolar)
one side of the helix exposed and one side packed against the interior of the protein
Nonpolar residues - interior protein, hydrophobic
polar residues - helix is hydrophilic - fully exposed
HGH is amphipathic helices
beta shrands
antiparallel - amino to carboxyl, next is carboxyl amino
most stable - hydrogen bonds
parallel - carboxyl to amino for all
carbonyl dont match up as easily, not 180, less stable
beta barrel
common
hydrophilic exterior, hydrophobic
each strand has one face in our face out
alpha/beta structure
repeats of alpha beta strand
frequent domains
ex: leucine rich repeats
inner beta sheet, out alpha helices
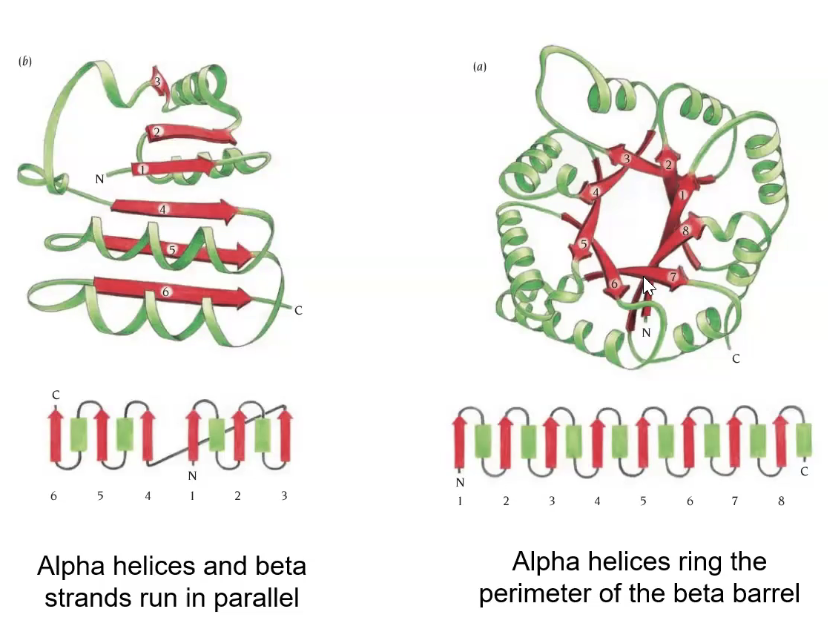
alpha+beta
any combination of a and b
SH2 domain twisted beta flanked by alpha helices
coiled-coil
long alpha helices intertwined between each other
strong and flexible
3.5 aa per turn not 3.6
repeat period of 7 aa - heptad repeat
hydrophobic interactions stabilize
salt bridge can stabilize ionic electrostatic interaction
keratin
protonfilament: 2 cc in staggered fashion (4 h)
protofibril: 2 staggered protonfilament (4cc, 8 ah)
filament: 4 protonfibrils twisted in right hand conformation (16cc, 32 ah)
keratin stabilized by
helices bundled in staggered conformation
disulfide bonds btwn protofilament and filament
collagen
repeats of pro-hyp-gly
loose left handed helix
triple helix bundled
can hbond to each other
staggered super helices
fibroin
ala and gly
tightly packed
hydrophobic side of each stack against one another
crystalline beta sheets of fibroin are interspersed with amorphous regions
amorphous is what can stretch to allow for flexibility
beta barrel and carrier protein
retinol binding protien
has a hydrophobic cavity where VA can go to where its needed
green fluorescent protein
protects against aqueous environment
gfp
ef hands
antigens
retinol binding
rubisco
enzyme active sites
beta propeller
4 beta strand twisted
neuraminidase: flu virus protein
uses hemagglutinin HA to being to receptor protein glycosylated with sialic acid
active site made by loops stuck to beta strands (kinda flexible is important)
TIM structure
stable core barrel strucutre
rubisco: catalyzes carboxylation
active sire on top between loops
bind DNA and calcium
ef hand long loop - binds calcium
short - binds DNA
charged polar residues asp and glu
CDPKs: conformational change after Ca binds to EF hands
important for signal transduction
allows for site to be open
diseases like malaria causes it → block activation of the kinase so it cant get it
antigen binding on AB
beta barrels bind to the antigens
loops bind the antigen
bind to organism for them to be degraded
toll-like receptors (TLR) are embeded into extracellular membranes
TLRs ar the security guards of immune system
made of the leucine rich
prediction of tertiary structure: levinthal’s paradox
in vivo porteinf old in 10^-1 → 10^-3 sec >98 orders
folding is too fast to be random
folding is due to collapse into most thermodynamically stable state
native protein vs denatured
folded - native (weak IM interactions maximized)
Native is tightly packed, a compact hydrophobic core
Determined by the amino acid sequence and how it interacts with the aqueous environment
Each protein sequence yields a unique native structure
The native folding state is thermodynamically stable (it represents the lowest achievable energy state for that protein)
unfolded - denatured (structure open side chains loosened)
step 1 of protein folding
hydrophobic collapse forms molten globule
into semi molten state
reduce in # protein conformations possible
peptide backbones hydrogen bond w one another and alpha helices and beta sheets emerge while protein collapse
unfolded → bury core (hydrophobic effect) → 2° → molten globule (loose 3° structure)→ 3° → 4°
molten globules & native fold
stable
next step is slower → arrangment of all side chains
lower free energy not as low as native fold
native fold:
weak IM interactions w aa are maximized
released energy w bonds → this more thermodynamically stable
thermodymics
∆G = ∆H - T∆S
change in free energy = change in enthalpy - change in entropy
∆G is negative → E is released more spontaneous
large ∆S → more spontaneous decrease in entropy
smaller ∆H → more spontaneous
enthalpy for folding
folded (native) → low enthalpy/entropy
unfolded (denatured) → high enthalpy/entropy
system w folding → decrease enthalpy, increased entropy
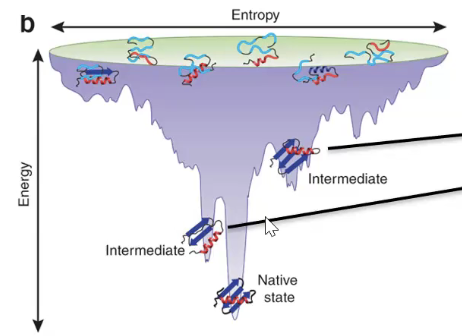
energy funnel diagram
troughs represent intermediates
wide funnel → lots of folding paths
anfinsens
ribonuclease a (degrades RNA) treated with B-mercapethanol and urea to denature protein
urea removed and reformation of Hbonds occur 1% activity regain
b-mercapethanol removed reform disulfide bonds 100% activity
denaturation
took active native folded enzymes and disrupt them
denature
pH change - alter ionation of aaa side chains disrupts Hbonds
heat - add E
detergents - amphipathic molecules associate with nonpolar residues
chaotropic agent - ion/small organic molecule at high conc. disturb IM interactions
Urea - interferes with H bonding
Beta-mercaptoethanol breaks disulfide bonds between cysteine
problems to folding
aggregation
non native disulfide bonds
isomerization of proline
removal of misfolded proteins
endoplasmic reticulum
Partially folded or misfolded →
PDI, PPI, HSP70, HSP 10/60 chaperones: assist folding
26S Proteosome: degrade misfolded protein
Misfolded and aggregation
IRE1 signaling in unfolded protein response (trigger cell death)
HSP100 break up aggregate
Protein Disulfide Isomerase (PDI)
Non-native disulfide bridge formation
hydrophobic inside of PDI
serve as point of contact for unfolded protein
oxidized or reduced forms
oxidize helps forms disulfide bonds
reduced PDI helps break and reform
Peptidyl Prolyl Isomerases (PPI)
proline is only one in cis conformation (trans 4x stable)
PPI switch proline form cis to trans
immunophilin recognize substrates through 4 aa motif
beta barrels
parvulins (PIN1)
target substrates proteins where serine or threonine precede pro
only recognizes substrates after phosphorylation
too much PIN1 is around, these proteins are more readily activated and sometimes active when they shouldn’t be
HSP 60/10
GroEL cylinder
unfoled protein in one half and then the other
HSP10 (GroES) is the lid
Parts:
Apical (alpha beta motif)
forms opening whern unfoled enters
flexible
hydrophobic
intermediate
allow ATP/ADP diffusions
flexible hinges
Equatorial
ATP bindings
stabilise chambers
GroES: 7 subunit structures
β sheet
β hairpin (roof)
Mobile loop (interacts with GroEL)
hsp 70
nucleotide binding domain - binds ATP
substrate binding domain - clamping to substrate binding domain
closing and opening domain when ADP→ATP
take on a new substrate
hsp 100
detangles misfolded protein
each subunit has ATP bound which helps tug misfolded protein through
how does accumulation of misfolded proteins get recognized to undergo cell death
ER-localized HSP70 called BIP
PERK1 kinase that stops translation of new proteins
ATF6 contains transciption factor that is released to turn on genes to help clean up
golgi
IRE1
causes splicing of mRNA transcript that activates expression of genes needed for the UPR
causes degradation of most other mRNA transcripts
induces apoptosis if it remains active too long Iif BIP does not re-associate)
How do aggregated and misfolded proteins get degraded?
1. If they can be disaggregated they are degraded by the 26S proteasome
2. If they remain as an aggregate they are disposed of through autophagy
misfolded go to cytoplasms through
ubiquitylated (targets them to 26S for degradation)
small lysine residues
disease and folidng
form very small stable beta structures
amyloid beta - alzeimers
tau protein - parkisons
native fold can misfold into beta strands
large aggregated beta sheets