M4L3 - Co-translational Targeting
1/27
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
28 Terms
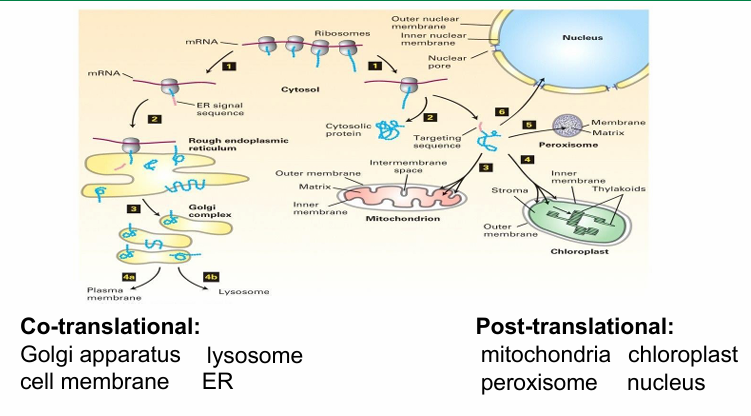
Secretory Pathway
Proteins transported to ER is co-translational instead of post-translational
Secreted pathways are synthesized in ER before vesicle-based transport carries them out of the cell
RER Characteristics
Bound by single membrane (it’s also continous with outer nucleus membrane
No genome (RER proteins all encoded in nuclear genome
Site of protein synthesis and modification
First destination for secreted proteins
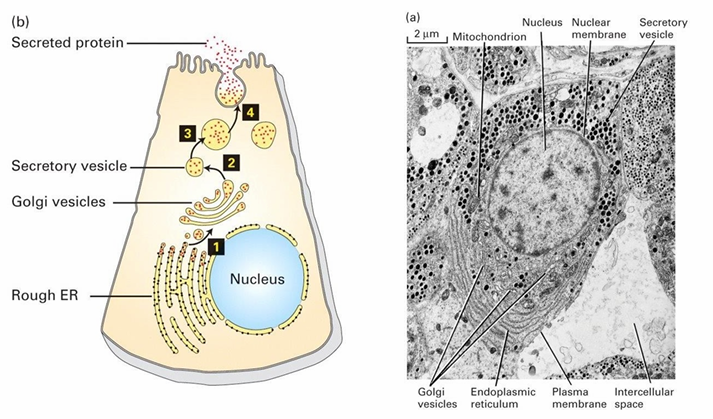
TEM of ER
RER is continous with nucleus membrane
Punctate black dots is membrane-bound ribosomes
They synthesize secreted proteins
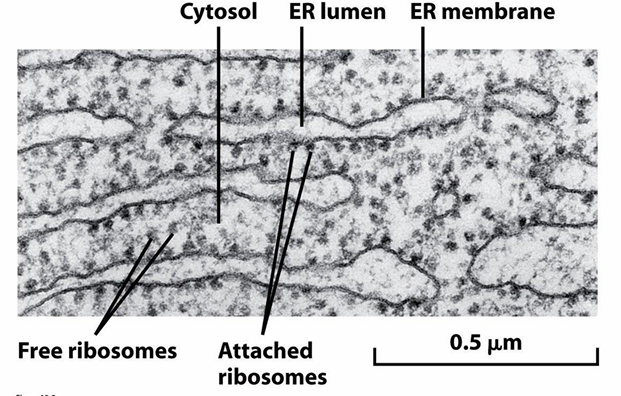
TEM of RER
ER associated ribosomes are on cytosolic side of the ER membrane
At any given time, there will be free and ER-associated ribosomes
Both are translationally active and come from the same common pool of ribosomes
They just cycle on and off the ER membrane
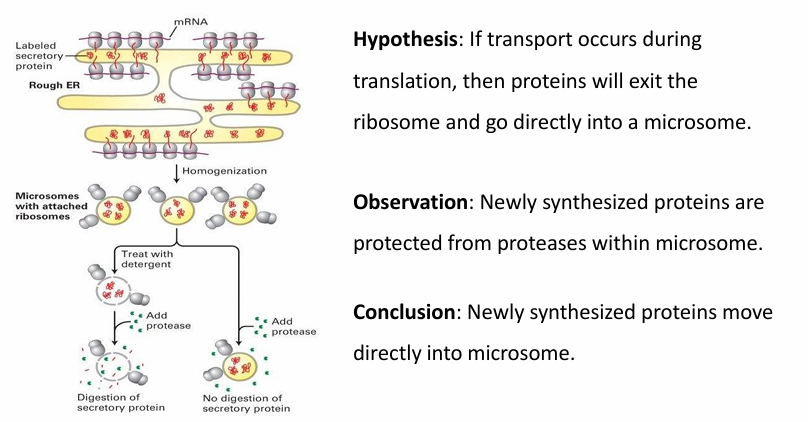
Proteins Targeted while Translation is Active Experiment
Microsomes used to mimic ER membrane
Made by disrupting the ER membrane by homogenization
They spontaneously assemble into spheres mimicing normal ER behaviour
Two experiments were conducted
Treat microsomes with detergent
This releases the proteins inside
Protease is added to digest the proteins
This shows that the protease can digest proteins
Protease added with intact microsome
The newly synthesized proteins are protected inside microsome
Conclusion: Newly synthesized proteins move directly into the microsome without being exposed to external env
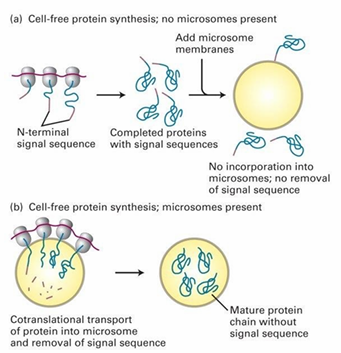
Translation and Translocation Co-occurance Experiment
Can protein transport to ER occur post-translationally?
Hypothesis: If proteins have completed translation, import won’t occur into ER
Control: Figure b
ER-targeted proteins are synthesized in vitro in presence of microsomes
Mature proteins are found inside microsomes
Experiment: Figure a
ER proteins synthesized in vitro in absence of microsomes
The translated proteins are added to a solution with microsomes
They’re unable to enter after full translation
Conclusion: Import must occur co-translationally
Co-Translational Protein Transport Rules
Signal sequence
Receptor for signal sequence
Translocation channels
E requirement (GTP hydrolysis)
Method of targeting proteins to different organelle locations
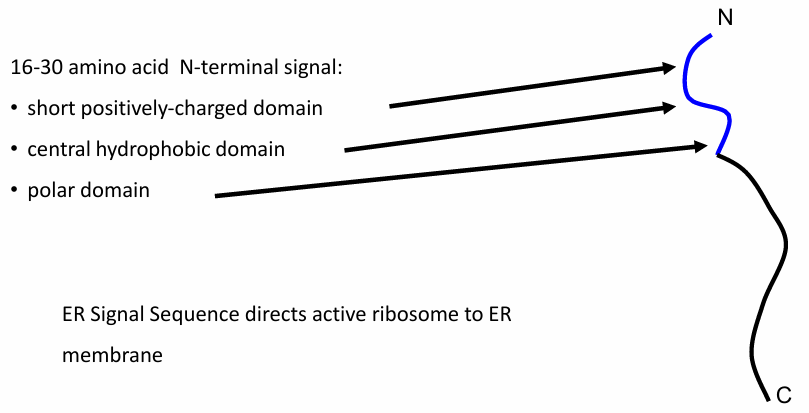
Rule 1: ER protein Transport
Domain of 16-30 amino acids needed
Sequence located at N-terminus carrying
A short (+) domain
Hydrophobic domain
Polar domain
Signal Sequence is produced first bc N-terminus
First part of nascent protein translated
2 roles:
Targets protein to ER
Guides ribosome translating the protein to ER membrane
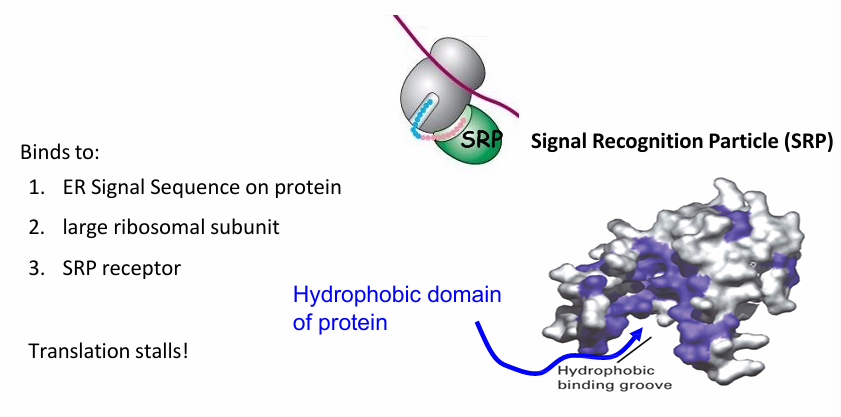
Rule 2: ER protein Transport
Signal Recognition Particle (SRP)
Made of 6 proteins and functional RNA (300 nucleotides long)
It binds to
N terminal ER sequence
Large subunit of Ribosome
This pauses translation for a bit
If the sequence was on the C-terminus, translation would be complete before SRP binding
No ribosome = no transport bc this is co-translational
The SRP receptor has hydrophobic bindng groove that recognizes the hydrophobic domain of ER signal sequence
SRP Cytosolic Complex
Requires:
SRP
SRP receptor on ER membrane
SRP Receptor
transmembrane dimer with an alpha and beta subunit
SRP and alpha subunit of SRP receptor are GTP-binding
SRP receptor is associated with ER translocon
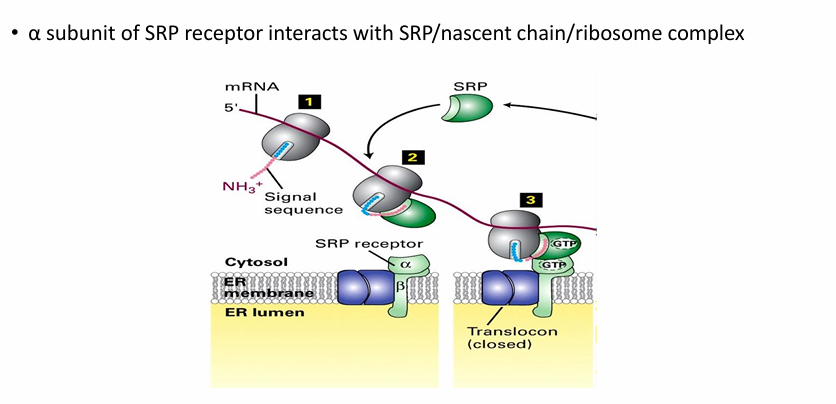
Components Associated with ER Membrane Surface During Co-Translational Protein Transport
Ribosome
mRNA
N-terminus of nascent protein
SRP
SRP receptor
closed translocon
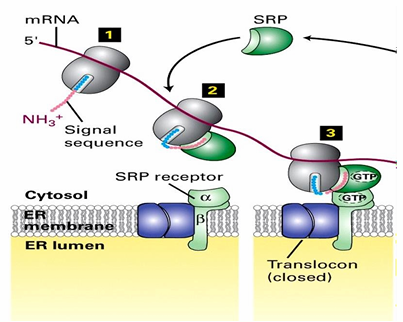
Rule 3: ER protein Transport
The translocon is initially closed
Opens as the nascent peptide is transferred to interior
SRP dissociates from ER signal sequence during translation
Translation then continues
Ribosome remains associated with ER membrane
As translation continues, the nascent peptide is pushed through translocon
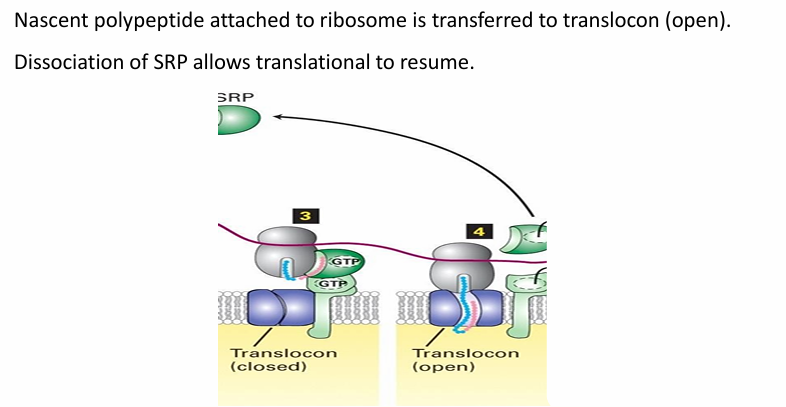
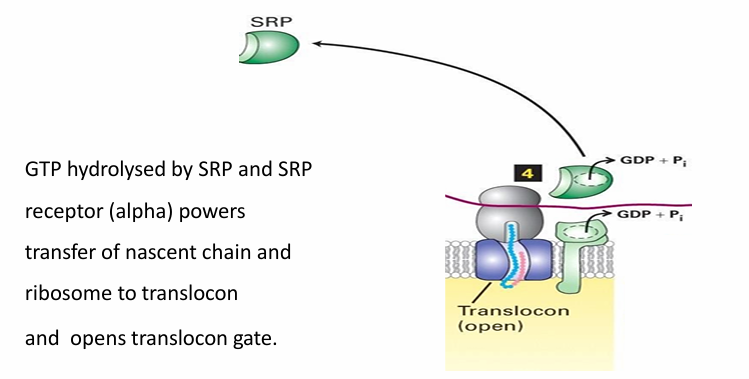
Rule 4: ER protein Transport
SRP and SRP receptor have intrinsic GTPase activity
Both hydrolyze GTP for E
Powers the transfer of nascent peptide into translocon and opens it
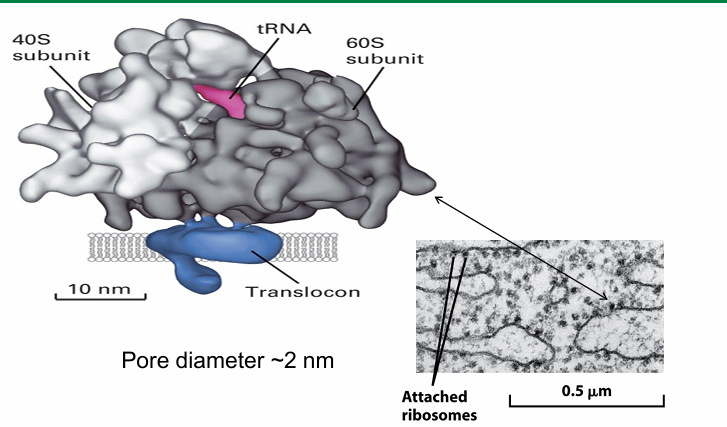
Ribosome-Translocon Complex: Co-Translational Protein Transport
The large ribosomal subunit directly interacts with translocon
Minimal space exposes the nascent polypeptide emerging from ribosome to cytosol
So not susceptible to protease in external env
This close association gives the rough ER speckled appearance
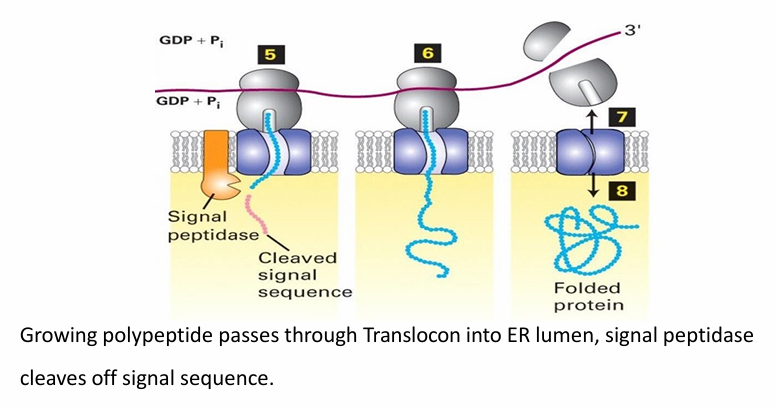
What Happens when the N-terminus of the nascent protein reaches the lumen of RER
ER signal sequence is cleaved by signal peptidase
Peptide is pushed through translocon as translation continues
After translation, it’s released into the lumen where folding occurs
Translocon closes and ribosome dissociates
Depending on the mRNA the ribosome binds to next, it can be a free ribosome or bound
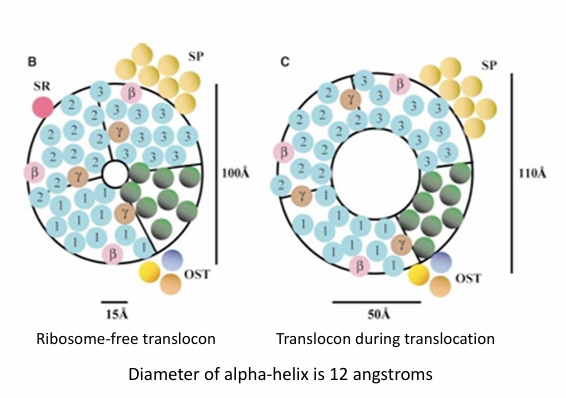
Co-Translational Protein Transport Translocon Structure
Comprised of many transmembrane a-helices forming the wall
Inner circle is the inner diameter of the translocon
When no ribosome present, the opening is narrow (15Å)
When ribosome is attached, the open conformation is adopted
Nascent protein goes through
opening widens to 50Å in diameter
Rule 5: ER protein Transport
The proteins here are soluble
They either
Remain in ER
Transported via secretory vesciles to GA
Transported to lysosomes
Transported out of the cell
Needs a second signal sequence
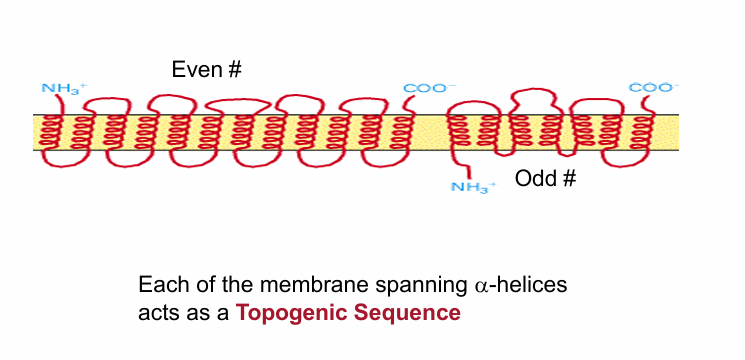
Topogenic Sequence
Embeds proteins in the ER membrane during co-translational transport
25 amino acid long sequence forming a hydrophobic a-helix with ability to embed in membrane
Determines topology (# of times the protein crosses membrane and orientation of it)
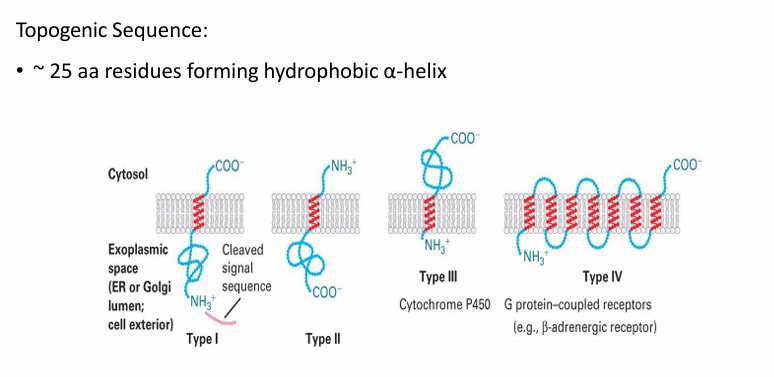
There are 4 classes of proteins based on their topology and sequences used
Type I / II / III: Span the membrane once
Type IV: Span the membrane multiple times
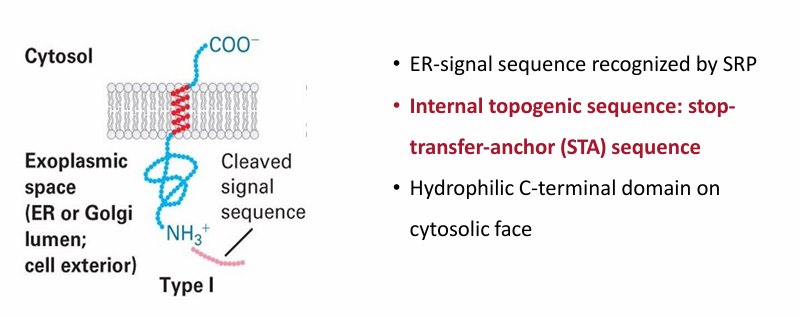
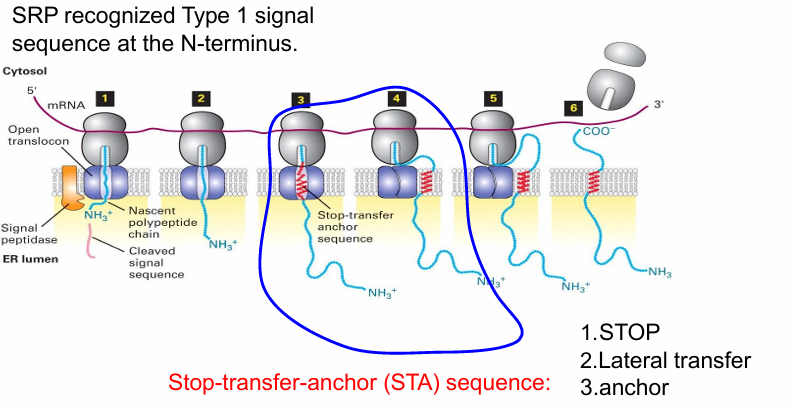
Type 1 Proteins
Same N-terminal signal sequence in soluble ER proteins
STA: Stop-Transfer-Anchor internal togogenic sequence
Forms hydrophobic alpha helix embedding the protein in the ER membrane
Stops translocation through translocon to transfer the protein to membrane
Anchors protein in place
The protein maintains this topology even when moving to other locations via vesicle transport
N-terminus faces exoplasmic space (inside ER or GA lumen)
C terminus always faces cytosolic space
Transport and Insertion of Type I Protein
N terminal sequence recognized by SRP and brought to SRP receptor
N-terminal sequence threaded into translocon
Protein synethesis pushes the protein through
When STA sequence is translated, it folds into an a-helix
Recognized by interior wall of translocon
Stops translocon and causes it to open laterally
Allows topogenic sequence to diffuse into surrounding membrane
Keeps protein anchored in place
After translation/translocation is complete the C and N terminal domains fold within their envs
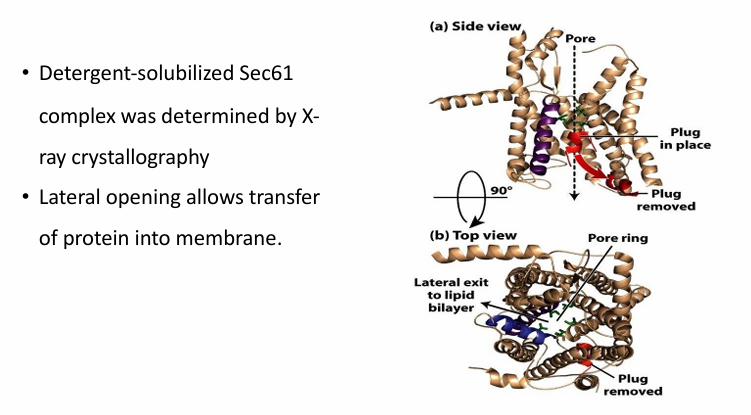
Sec61 Complex
Part of traanslocon in co-transloation protein transport
Red a-helix is the plug swinging down to open the pore during translocation
Blue helix illustrates the lateral opening of the translocon
This complex was determined by x-ray crystallography
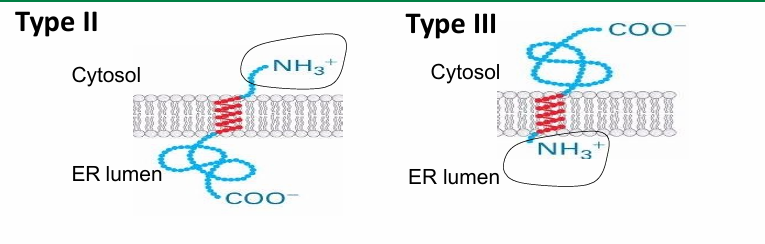
Type II / III Proteins
Single-pass proteins
Lack N-terminal signal sequence
They have a single signal sequence
SA Sequence: Signal Anchor
Signal for both SRP and topogenic sequence
Type II:
Oriented with N-terminus on cytosolic side
C-terminus on luminal side
Type III:
Oriented same way as Type I
N terminus on luminal side
C terminus on cytosolic side
Have very short N-terminus (allows it to be threaded through)
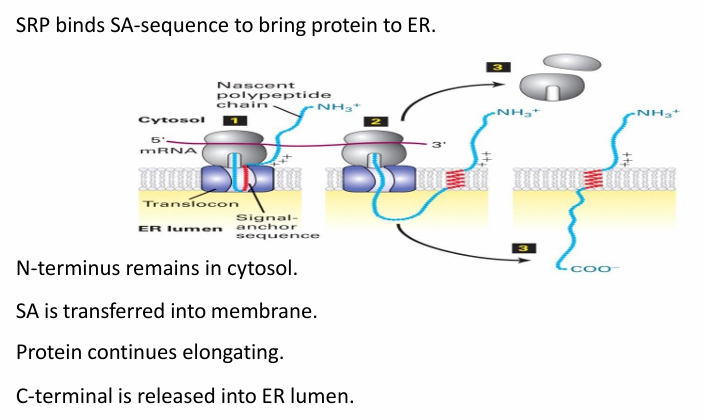
Type II Membrane Protein Transport
The SA sequence (red) is recognized by SRP to bring the protein and ribosome to ER membrane
SA sequence is then transferred to translocon
(+) residues prevent transfer of N-terminal portion of protein into translocon
Translocon opens laterally to allow diffusion of SA sequence into membrane
C-terminus pushed through translocon into ER lumen
Type III Membrane Protein Transport
SA sequence (red) is recognized by SRP to bring to ER membrane
A short N-terminal sequence threaded into translocon
SA sequence enters allowing translocon to open laterally
(+) residues ensure the sequences neighbouring SA sequence stay in cytosol
Newly synthesized protein is pushed away from ribosome-translocon complex while being translated
The C-terminus stays on cytosol side and folds there
Type IV Integral Membrane Protein
Pass through multiple times
May be even/Odd # of topogenic sequences
Every time it passes through, there must be a sequence
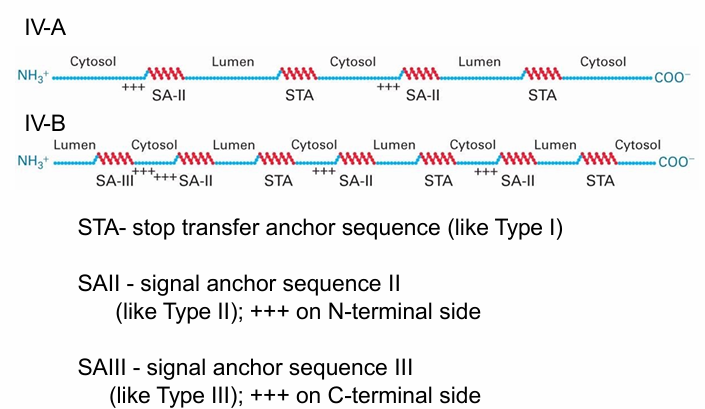
Type IV Integral Membrane Protein: 2 Examples
They differ at N-terminus
After N-terminus location decided, the rest is threaded through the membrane by alternating between STA sequence (Type I protein) and SA-II Sequence
Type IV-A
N-terminus is on cytosolic side
SA-II Sequence keeps N-terminus on cytosolic side
Type IV-B
N-terminus is on luminal side
SA-III Sequence keeps N-terminus on luminal side
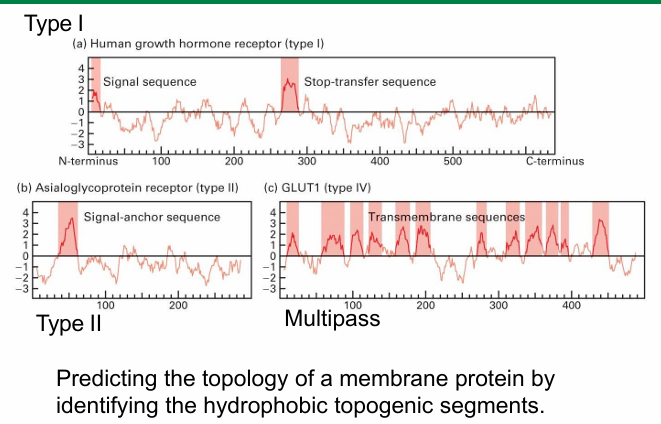
Hydropathy Profile
Graphic representation of all amino acids on polypeptide
Left —> Right on X axis
Amino acids from N to C-terminus
y axis: hydrophobicity of each amino acid
Cluster of hydrophobic residues (peaks) represent topogenic sequences
Type I
Hydrophobic peak at N-terminus
Typical for an ER signal sequence
Middle is another strong hydrophobic peak
Represents STA sequence
Type II
No hydrophobic sequence at N-terminus
Single hydrophobic peak representing SA sequence
Differentiating between Type II / III
Type II have longer N-terminus
Amino acids before SA sequence is also an indicator
(+) on N-terminal side suggest type II
(+) on C-terminal side suggests type III
Multipass Protein
Multiple hydrophobic peaks throughout
Suggests multiple topogenic sequences
Amount of peaks = amount of sequences = amount of passes through membrane
Overview Diagram