9. Biotechnology: Principles and Processes
1/142
Earn XP
Description and Tags
Clear your concepts and then use these flashcards for revision! Suitable for revision for IAT, NEET, NEST, etc. Question mode: Flashcards only. Answer mode: Answer wtih definition. Recommended study mode: Spaced repetition. Good luck with your exams!
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
143 Terms
What is the primitive definition of biotechnology?
Biotechnology deals with techniques of using live organisms or enzymes from organisms to produce products and processes useful to humans.
What is the modern definition of biotechnology given by EFB (European Federation of Biotechnology)?
The integration of natural science and organisms, cells, parts thereof, and molecular analogues for products and services.
What are the two core techniques that enabled the birth of biotechnology?
Genetic engineering
Bioprocess engineering
What is genetic engineering?
Techniques to alter the chemistry of genetic material (DNA and RNA), to introduce these into host organisms and thus change the phenotype of the host organism.
What is bioprocess engineering?
Maintenance of sterile (microbial contamination-free) ambience in chemical engineering processes to enable growth of only the desired microbe/eukaryotic cell in large quantities for the manufacture of biotechnological products like antibiotics, vaccines, enzymes, etc.
_______ reproduction preserves the genetic information, while _______ reproduction permits variation.
(sexual / asexual)
Asexual reproduction preserves the genetic information, while sexual reproduction permits variation.
What is the disadvantage of using traditional hybridisation procedures in genetic engineering?
Traditional hybridisation procedures used in plant and animal breeding, very often lead to inclusion and multiplication of undesirable genes along with the desired genes.
What is the likely fate of a piece of DNA, which is somehow transferred into an alien organism?
Most likely, this piece of DNA would not be able to multiply itself in the progeny cells of the organism. But, when it gets integrated into the genome of the recipient, it may multiply and be inherited along with the host DNA.
What is “origin of replication”?
In a chromosome there is a specific DNA sequence called the origin of replication, which is responsible for initiating replication.
This is a sequence from where replication starts and any piece of DNA when linked to this sequence can be made to replicate within the host cells. This sequence is also responsible for controlling the copy number of the linked DNA.
[definitions from two parts of the NCERT chapter]
For the multiplication of any alien piece of DNA in an organism it needs to be a part of a chromosome(s) which has a specific sequence known as ___________________.
For the multiplication of any alien piece of DNA in an organism it needs to be a part of a chromosome(s) which has a specific sequence known as ‘origin of replication’.
In genetic engineering, why is an alien DNA is linked with the origin of replication in the host body?
So that this alien piece of DNA can replicate and multiply itself in the host organism.
The construction of the first recombinant DNA emerged from the possibility of linking a gene encoding antibiotic resistance with a native _______ of Salmonella typhimurium.
The construction of the first recombinant DNA emerged from the possibility of linking a gene encoding antibiotic resistance with a native plasmid of Salmonella typhimurium.
The construction of the first recombinant DNA emerged from the possibility of linking a gene encoding antibiotic resistance with a native plasmid of Salmonella typhimurium. Which two scientists accomplished this?
Stanley Cohen and Herbert Boyer accomplished this.
The construction of the first recombinant DNA emerged from the possibility of linking a gene encoding antibiotic resistance with a native plasmid of Salmonella typhimurium.
Stanley Cohen and Herbert Boyer accomplished this how?
Stanley Cohen and Herbert Boyer accomplished this in 1972 by isolating the antibiotic resistance gene by cutting out a piece of DNA from a plasmid which was responsible for conferring antibiotic resistance.
The cutting of DNA at specific locations became possible with the discovery of what?
‘molecular scissors’ – restriction enzymes
How do plasmids act as vectors?
During genetic engineering, the cut piece of DNA was then linked with the plasmid DNA. These plasmid DNA act as vectors to transfer the piece of DNA attached to it. You probably know that mosquito acts as an insect vector to transfer the malarial parasite into human body. In the same way, a plasmid can be used as vector to deliver an alien piece of DNA into the host organism.
The linking of antibiotic resistance gene in Salmonella typhimurium with the plasmid vector became possible with the enzyme _____________.
The linking of antibiotic resistance gene in Salmonella typhimurium with the plasmid vector became possible with the enzyme DNA ligase.
The linking of antibiotic resistance gene in Salmonella typhimurium with the plasmid vector became possible with the enzyme DNA ligase.
What does this enzyme do?
It acts on cut DNA molecules and joins their ends.
What is recombinant DNA?
The linking of antibiotic resistance gene with the plasmid vector became possible with the enzyme DNA ligase. This makes a new combination of circular autonomously replicating DNA created in vitro and is known as recombinant DNA.
The linking of antibiotic resistance gene with the plasmid vector became possible with the enzyme DNA ligase. This makes a new combination of circular autonomously replicating DNA created in vitro and is known as recombinant DNA.
What happens when this recombinant DNA is transferred into Escherichia coli?
When this DNA is transferred into Escherichia coli, a bacterium closely related to Salmonella, it could replicate using the new host’s DNA polymerase enzyme and make multiple copies.
What is “cloning” of a gene? (example)
The ability to multiply copies of antibiotic resistance gene in E. coli was called cloning of antibiotic resistance gene in E. coli.
What are the three basic steps in genetically modifying an organism?
identification of DNA with desirable genes;
introduction of the identified DNA into the host;
maintenance of introduced DNA in the host and transfer of the DNA to its progeny.
In the year 1963, the two enzymes responsible for restricting the growth of bacteriophage in Escherichia coli were isolated. One of these added methyl groups to DNA, while the other cut DNA.
What was the latter called?
restriction endonuclease
What is the first restriction endonuclease?
restriction endonuclease–Hind II
The first restriction endonuclease (Hind II)‘s functioning depends on what?
a specific DNA nucleotide sequence
It was found that restriction endonuclease Hind II always cut DNA molecules at a particular point by recognising a specific sequence of how many base pairs?
6
It was found that restriction endonuclease Hind II always cut DNA molecules at a particular point by recognising a specific sequence of 6 base pairs.
This specific base sequence is known as ___________________ for Hind II.
recognition sequence
Besides Hind II, today we know more than ________ restriction enzymes that have been isolated from over 230 strains of bacteria each of which recognise different recognition sequences.
(how many?)
Besides Hind II, today we know more than 900 restriction enzymes that have been isolated from over 230 strains of bacteria each of which recognise different recognition sequences.
What is the convention for naming of restriction enzymes?
The first letter of the name comes from the genus and the second two letters come from the species of the prokaryotic cell from which they were isolated.
Roman numbers following the names indicate the order in which the enzymes were isolated from that strain of bacteria.
Restriction enzyme EcoRI comes from which organism? (including strain)
Escherichia coli RY 13
Restriction enzymes belong to a larger class of enzymes called __________.
Restriction enzymes belong to a larger class of enzymes called nucleases.
Restriction enzymes belong to a larger class of enzymes called nucleases. What are the two kinds of nucleases?
exonuc
Restriction enzymes belong to a larger class of enzymes called nucleases. These are of two kinds; exonucleases and endonucleases.
What do exonucleases do?
Exonucleases remove nucleotides from the ends of the DNA.
Restriction enzymes belong to a larger class of enzymes called nucleases. These are of two kinds; exonucleases and endonucleases.
What do endonucleases do?
Endonucleases make cuts at specific positions within the DNA.
How do restriction endonucleases function?
Each restriction endonuclease functions by ‘inspecting’ the length of a DNA sequence. Once it finds its specific recognition sequence, it will bind to the DNA and cut each of the two strands of the double helix at specific points in their sugar -phosphate backbones.
Each restriction endonuclease recognises a specific palindromic nucleotide sequences in the DNA.
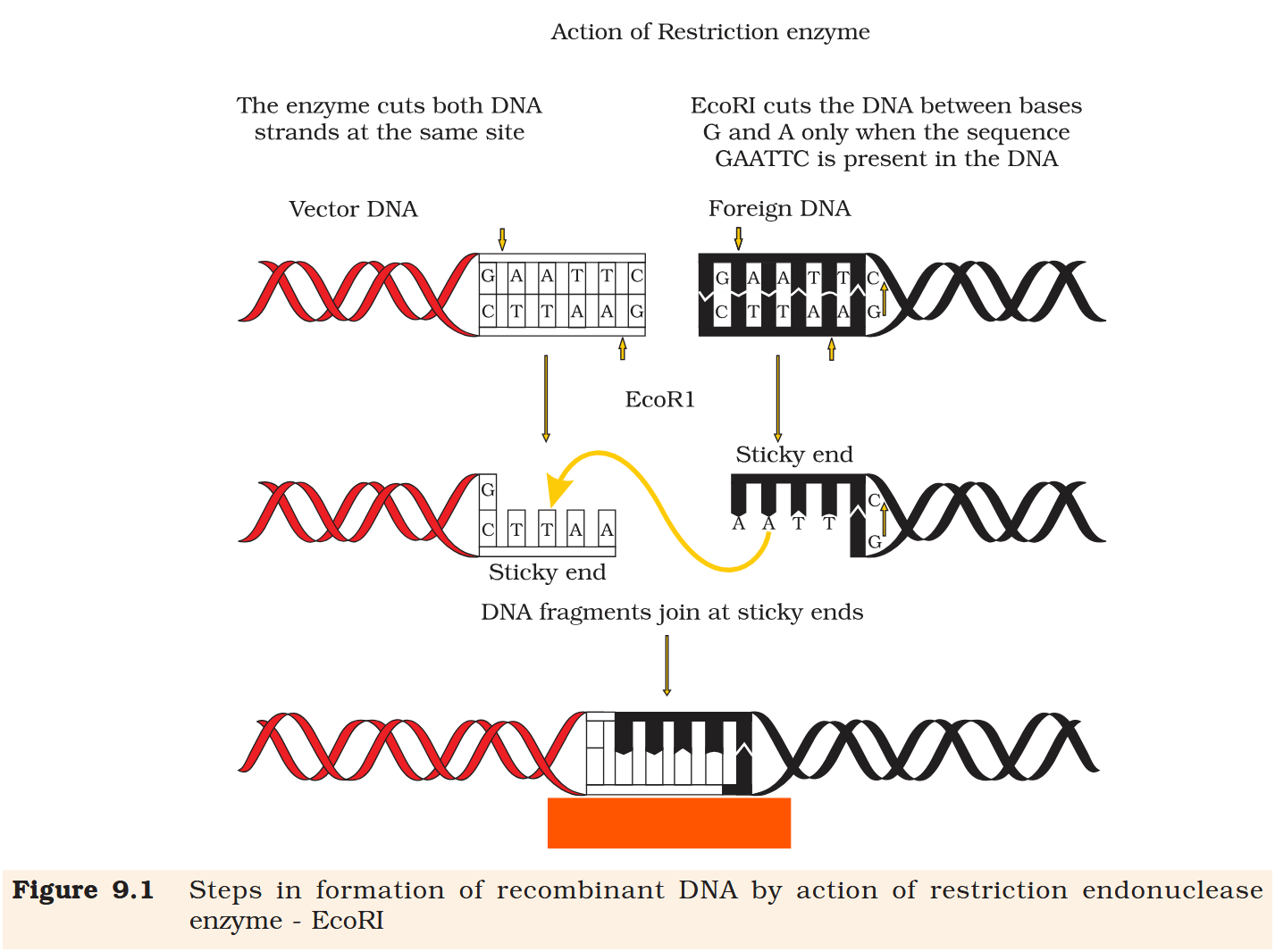
Recombinant DNA
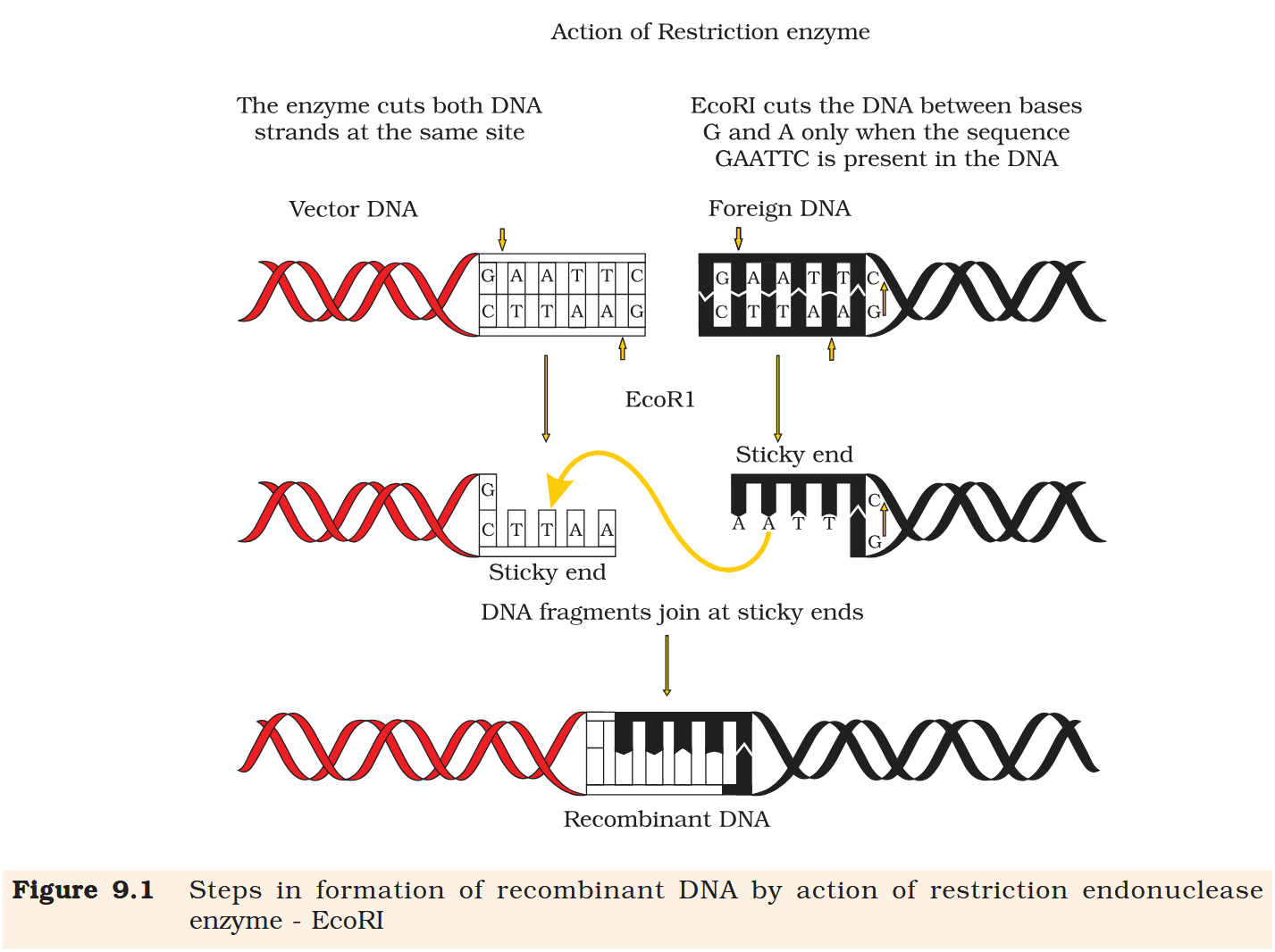
What is a palindrome in DNA?
A palindrome in DNA is a sequence of base pairs that reads same on the two strands when orientation of reading is kept the same.

Is this palindromic DNA?
Yes
At which position do restriction endonucleases cut DNA strands?
Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands.
Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands. This leaves single stranded portions at the ends. There are overhanging stretches called _____ _____ on each strand.
Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands. This leaves single stranded portions at the ends. There are overhanging stretches called sticky ends on each strand.
Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands. This leaves single stranded portions at the ends. There are overhanging stretches called sticky ends on each strand.
Why are they called sticky ends?
These are named so because they form hydrogen bonds with their complementary cut counterparts.
Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands. This leaves single stranded portions at the ends. There are overhanging stretches called sticky ends on each strand.
This stickiness of the ends facilitates the action of the enzyme _____________.
DNA ligase
Restriction endonucleases are used in genetic engineering to form ‘___________’ molecules of DNA, which are composed of DNA from different sources/genomes.
Restriction endonucleases are used in genetic engineering to form ‘recombinant’ molecules of DNA, which are composed of DNA from different sources/genomes.
To form recombinant molecules of DNA, why must the vector DNA and foreign DNA be cut with the same restriction endonucleases?
When cut by the same restriction enzyme, the resultant DNA fragments have the same kind of ‘sticky-ends’ and, these can be joined together (end-to-end) using DNA ligases.
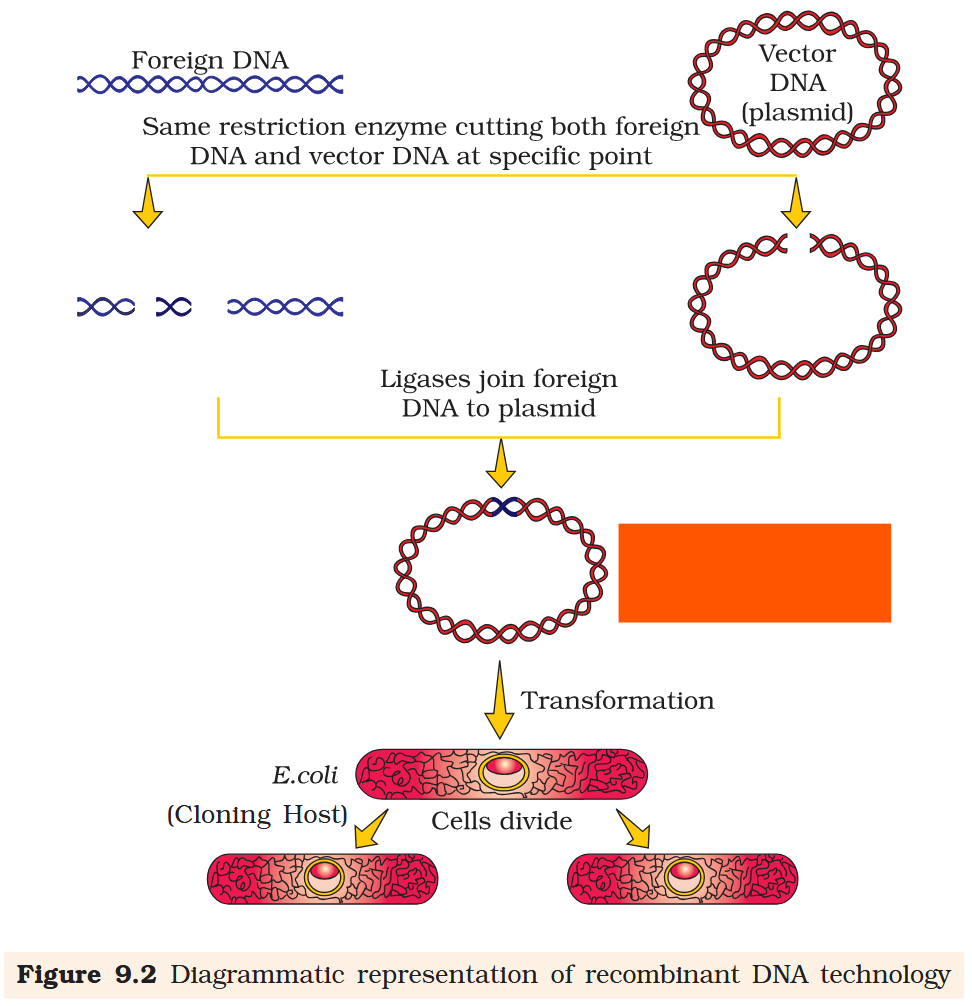
Recombinant DNA molecule
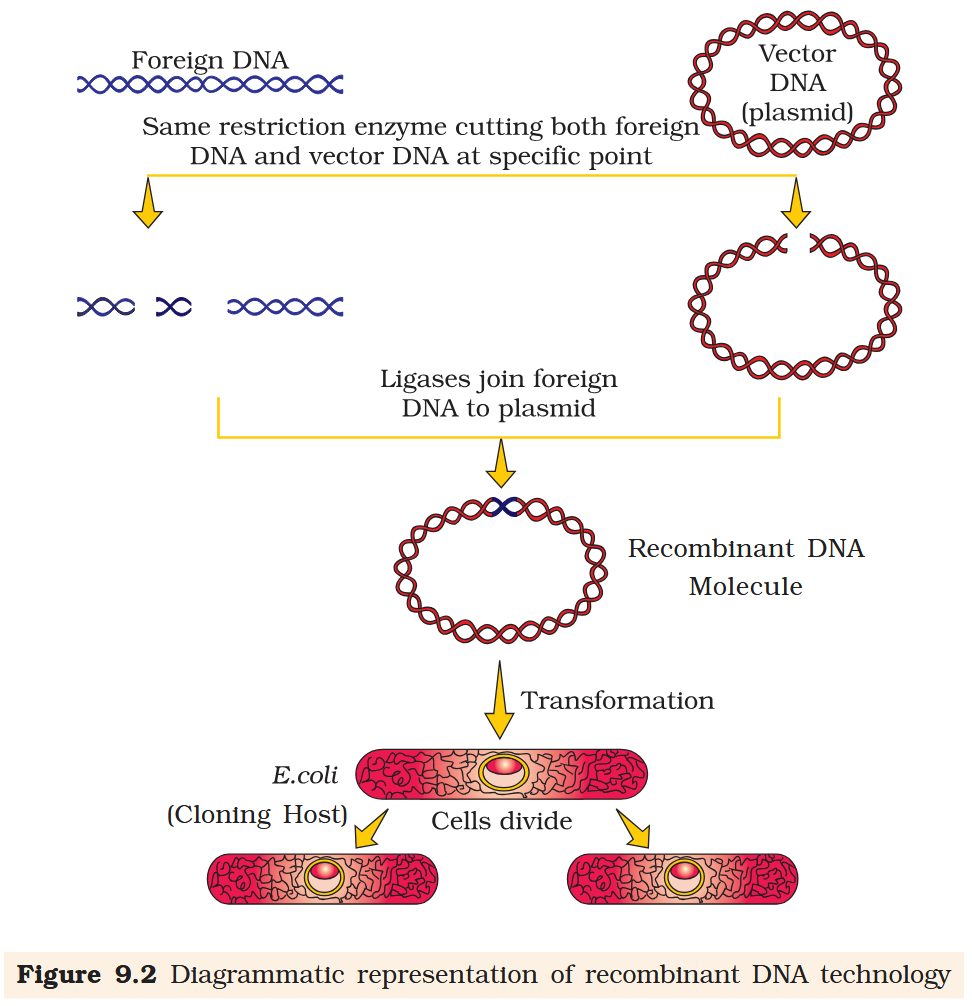
The cutting of DNA by restriction endonucleases results in the fragments of DNA. These fragments can be separated by a technique known as ____ _____________.
The cutting of DNA by restriction endonucleases results in the fragments of DNA. These fragments can be separated by a technique known as gel electrophoresis.
What is gel electrophoresis?
Since DNA fragments are negatively charged molecules they can be separated by forcing them to move towards the anode under an electric field through a medium/matrix.
The DNA fragments separate (resolve) according to their size through sieving effect provided by the gel. Hence, the smaller the fragment size, the farther it moves.
Nowadays the most commonly used matrix in gel electrophoresis is _______ which is a natural polymer extracted from sea weeds.
Nowadays the most commonly used matrix in gel electrophoresis is agarose which is a natural polymer extracted from sea weeds.
During gel electrophoresis, the separated DNA fragments can be visualised only after staining the DNA with a compound known as _________ _________ followed by exposure to UV radiation.
During gel electrophoresis, the separated DNA fragments can be visualised only after staining the DNA with a compound known as ethidium bromide followed by exposure to UV radiation.
During gel electrophoresis, the separated DNA fragments can be visualised only after staining the DNA with a compound known as ethidium bromide followed by exposure to _______________.
(which radiation?)
During gel electrophoresis, the separated DNA fragments can be visualised only after staining the DNA with a compound known as ethidium bromide followed by exposure to UV radiation.
What is the process of elution (in gel electrophoresis)?
The separated bands of DNA are cut out from the agarose gel and extracted from the gel piece. This step is known as elution.
If we are able to link an alien piece of DNA with bacteriophage or plasmid DNA, we can multiply its numbers equal to the ______ _________ of the plasmid or bacteriophage.
If we are able to link an alien piece of DNA with bacteriophage or plasmid DNA, we can multiply its numbers equal to the copy number of the plasmid or bacteriophage.
Bacteriophages because of their ____ number per cell, have very ____ copy numbers of their genome within the bacterial cells.
(high / low)
Bacteriophages because of their high number per cell, have very high copy numbers of their genome within the bacterial cells.
How many copies per cell do plasmids have generally?
Some plasmids may have only one or two copies per cell whereas others may have 15-100 copies per cell. Their numbers can go even higher.
Vectors used are present are engineered in such a way that two processes become easier, which are?
Vectors used at present, are engineered in such a way that;
they help easy linking of foreign DNA
they help in easy selection of recombinants from non-recombinants.
What are the three features that must be present in a vector to facilitate cloning in that vector?
origin of replication (ori)
selectable marker
cloning sites
If one wants to recover many copies of the target DNA it should be cloned in a vector whose origin of replication supports ____ copy number.
(high / low)
If one wants to recover many copies of the target DNA it should be cloned in a vector whose origin of replication supports high copy number.
What is the function of a selectable marker in a cloning vector?
It helps in identifying and eliminating non-transformants and selectively permitting the growth of the transformants.
What is transformation as a procedure under genetic engineering?
Transformation is a procedure through which a piece of DNA is introduced in a host bacterium.
Normally, the genes encoding resistance to antibiotics such as ampicillin, chloramphenicol, tetracycline or kanamycin, etc., are considered useful ________ ________ for E. coli.
Normally, the genes encoding resistance to antibiotics such as ampicillin, chloramphenicol, tetracycline or kanamycin, etc., are considered useful selectable markers for E. coli.
Normal E. coli cells carry resistance to ampicillin.
True or false?
False. The normal E. coli cells do not carry resistance against any antibiotics.
In order to link the alien DNA, the vector needs to have very few, preferably single, recognition sites for the commonly used restriction enzymes.
Why are single sites preferable?
If the restriction enzymes act only on one site in the vector, then it avoids having multiple fragments of the vector (thus less risk of loss of ori).
Presence of more than one recognition sites within the vector will generate several fragments, which will complicate the gene cloning.
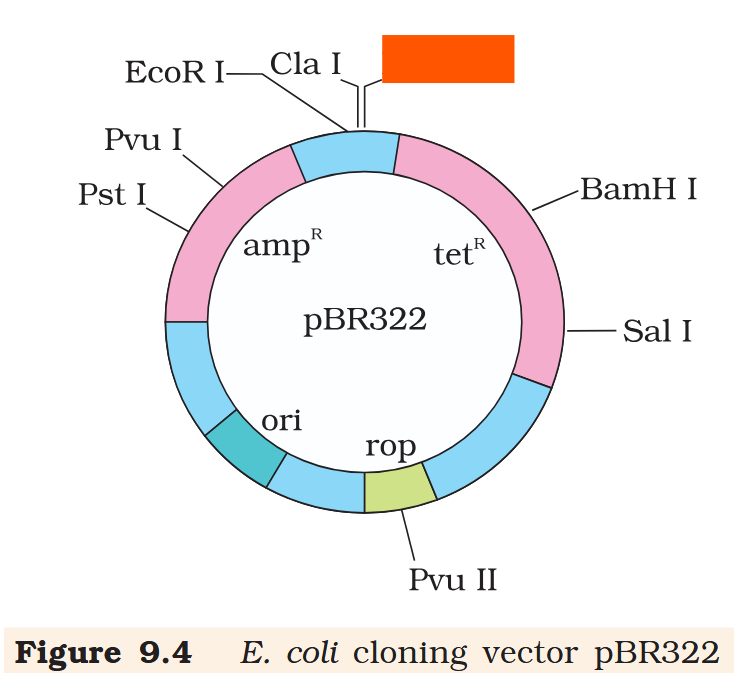
Hind III
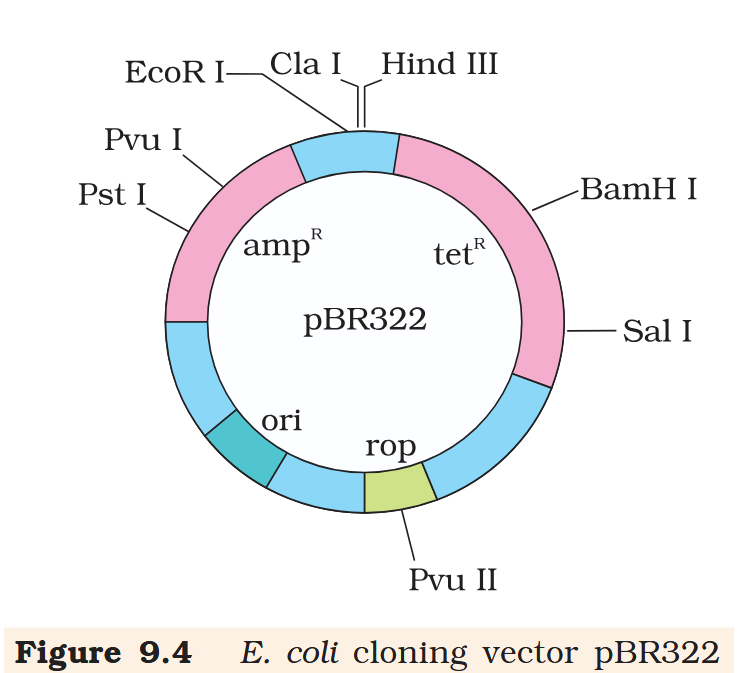
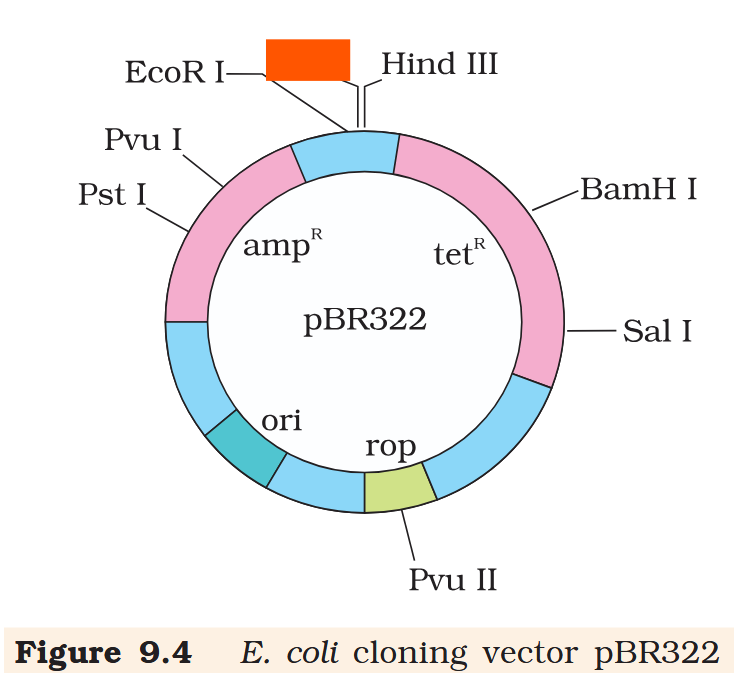
Cla I
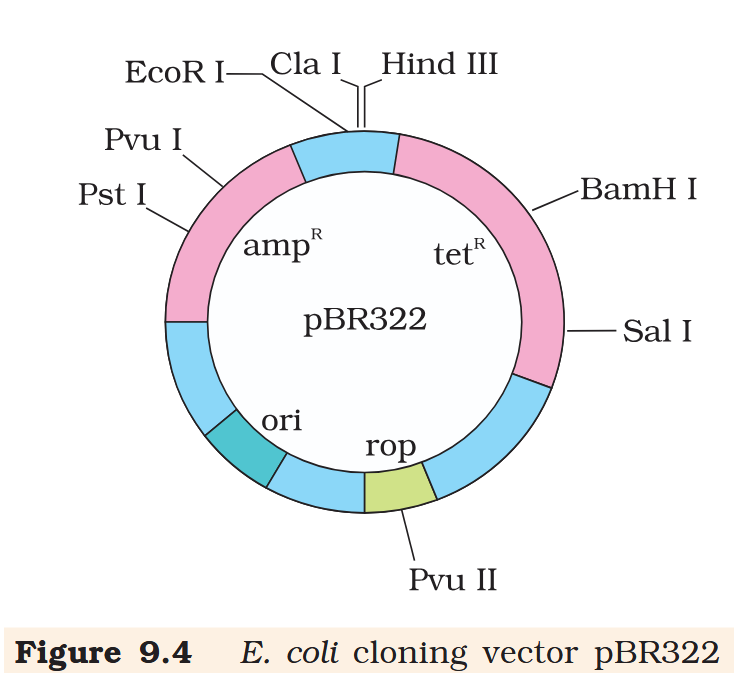
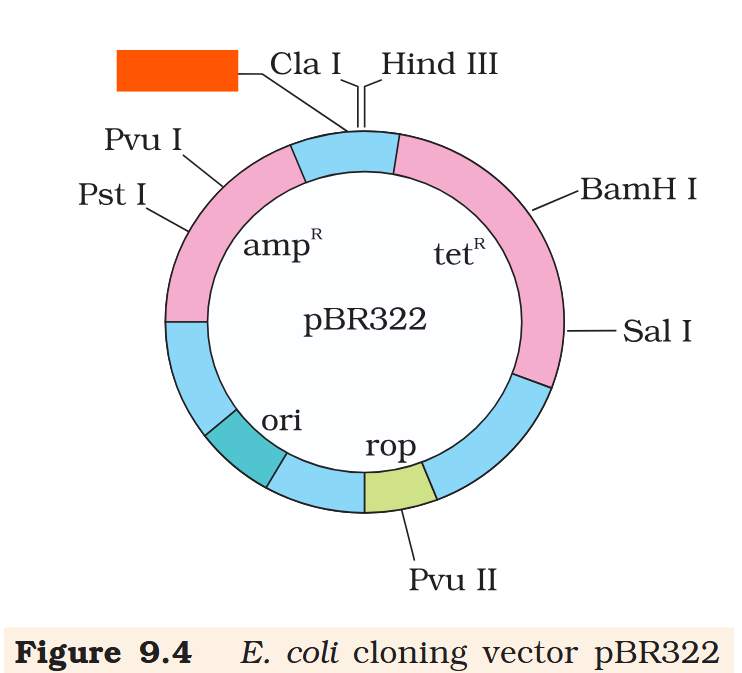
EcoR I
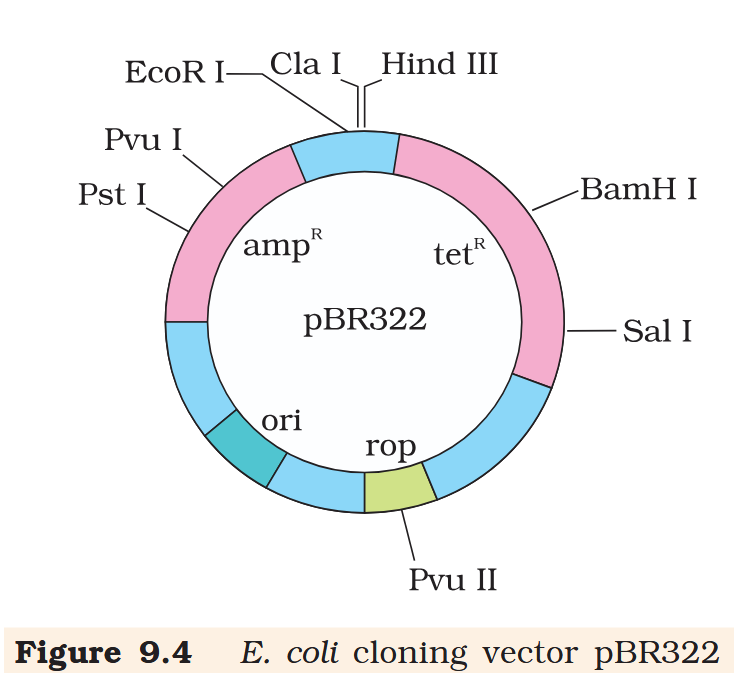
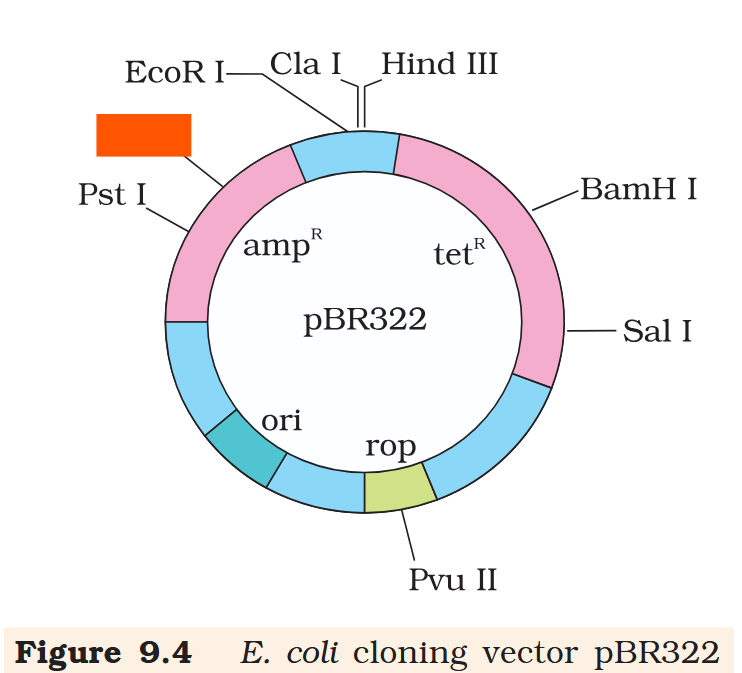
Pvu I
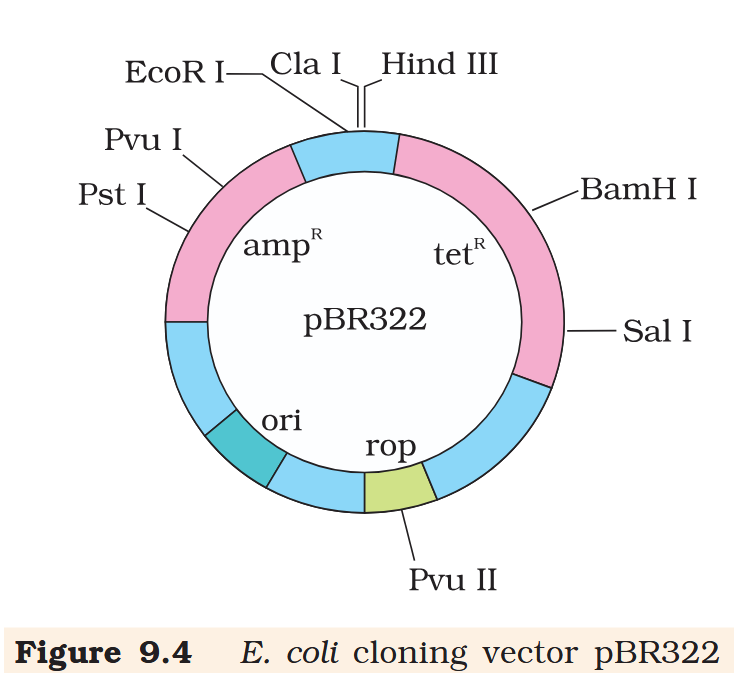
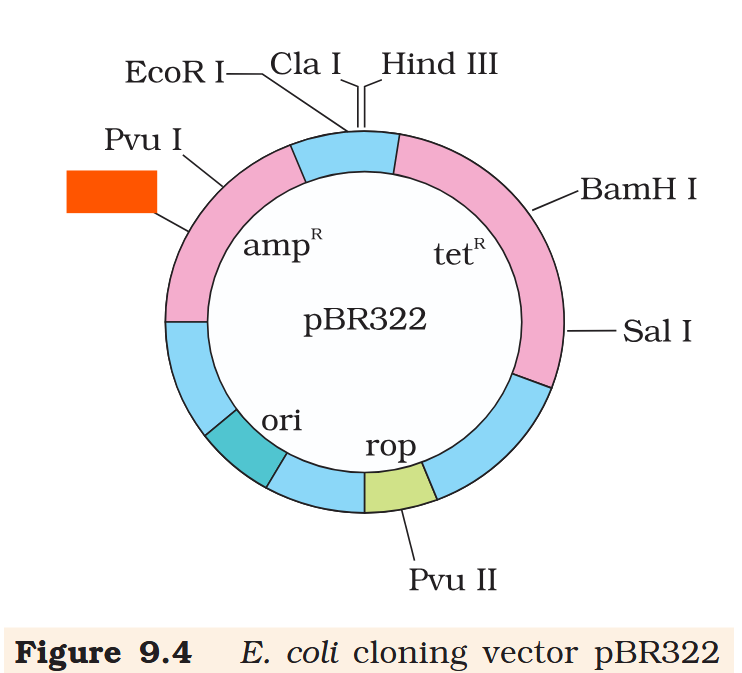
Pst I
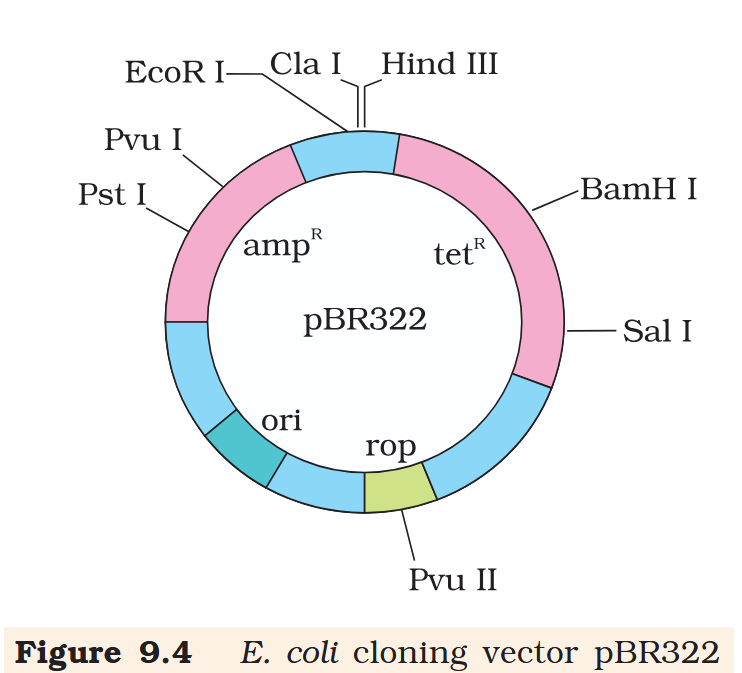
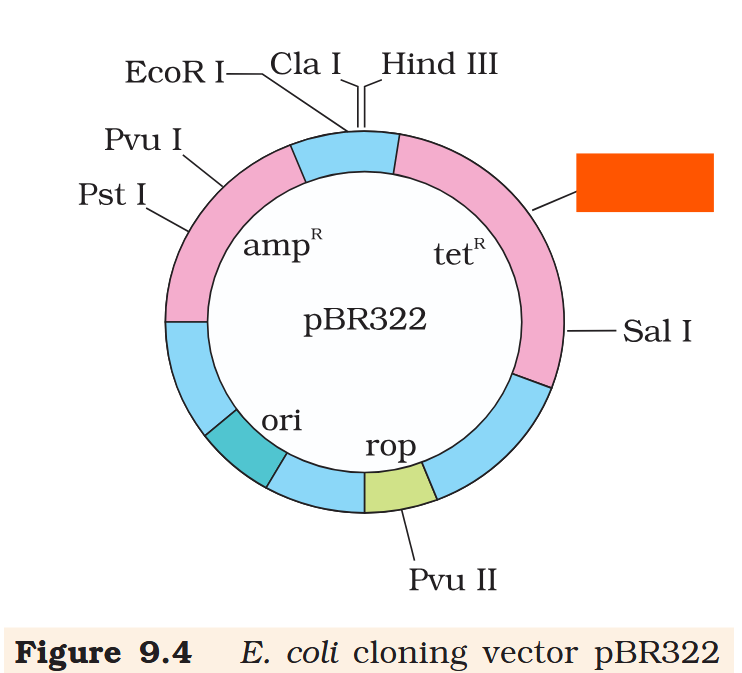
BamH I
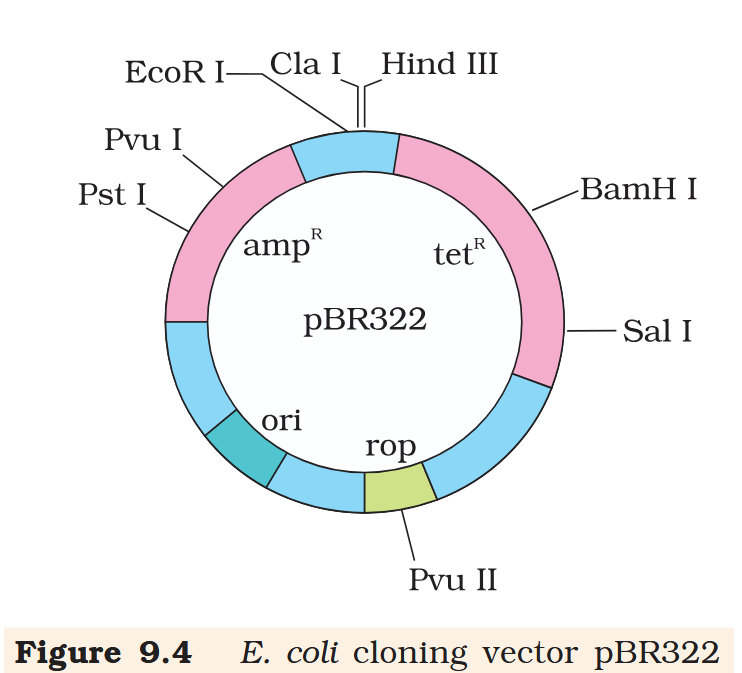
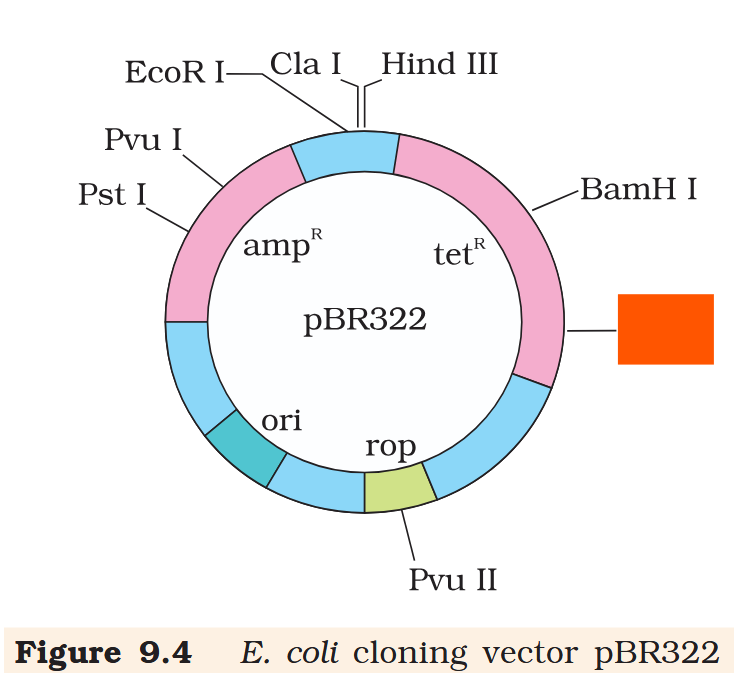
Sal I
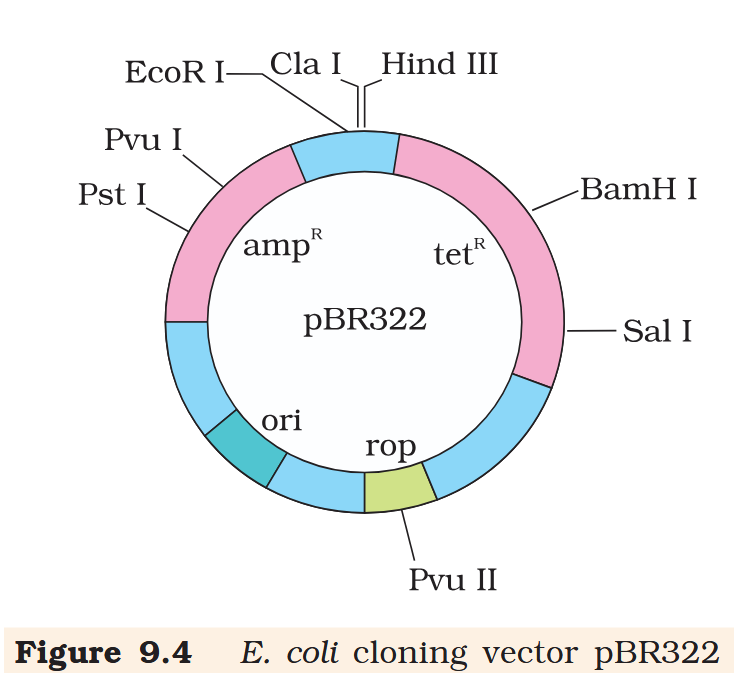
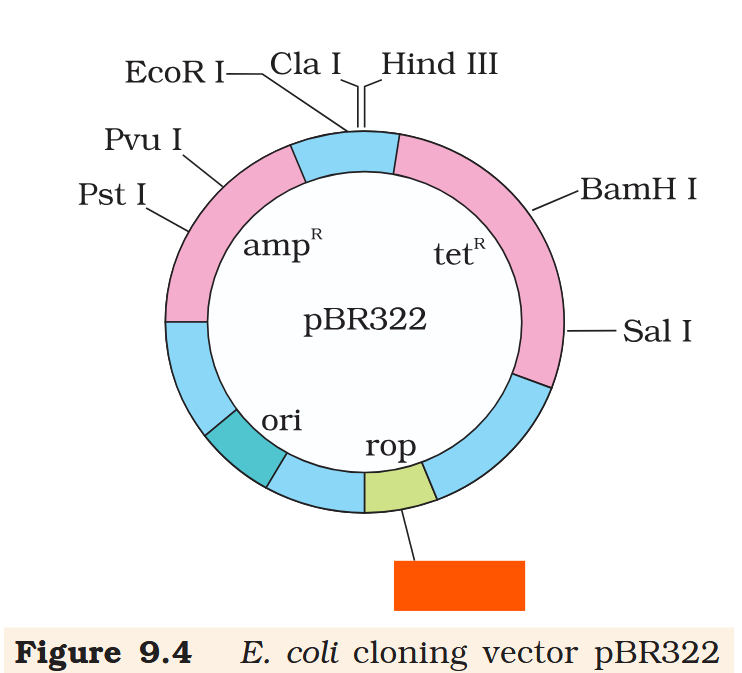
Pvu II
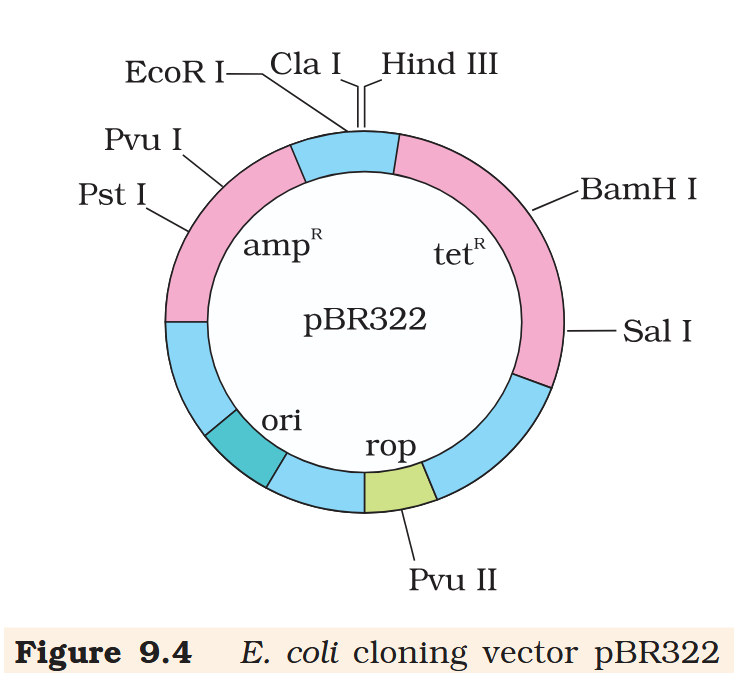
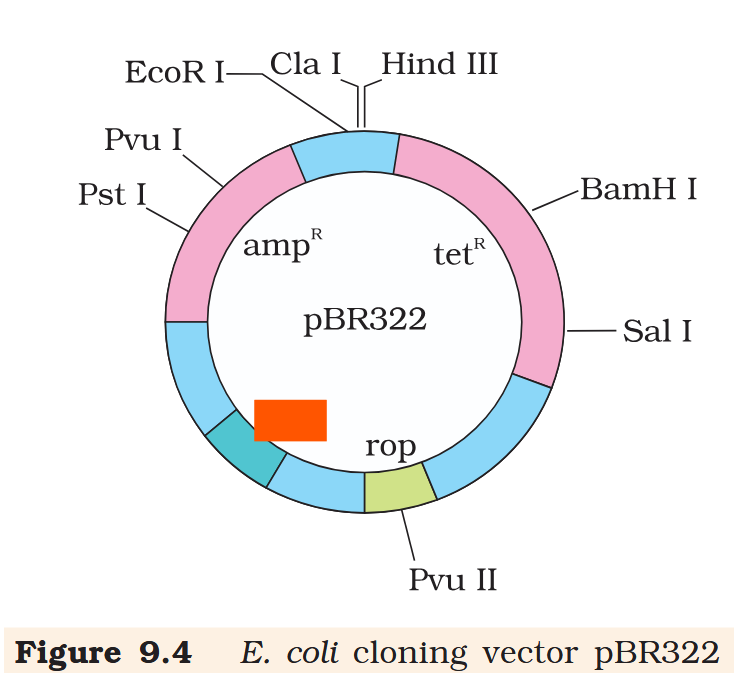
ori
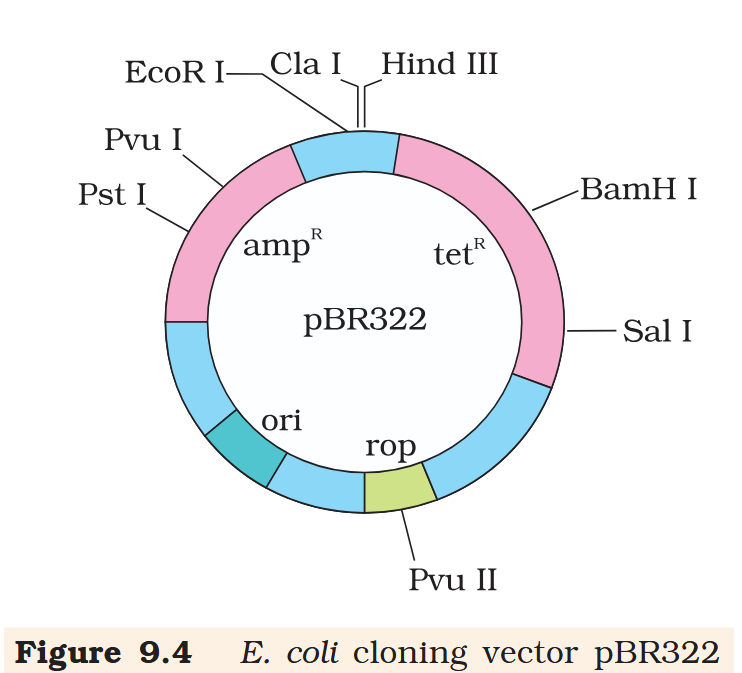
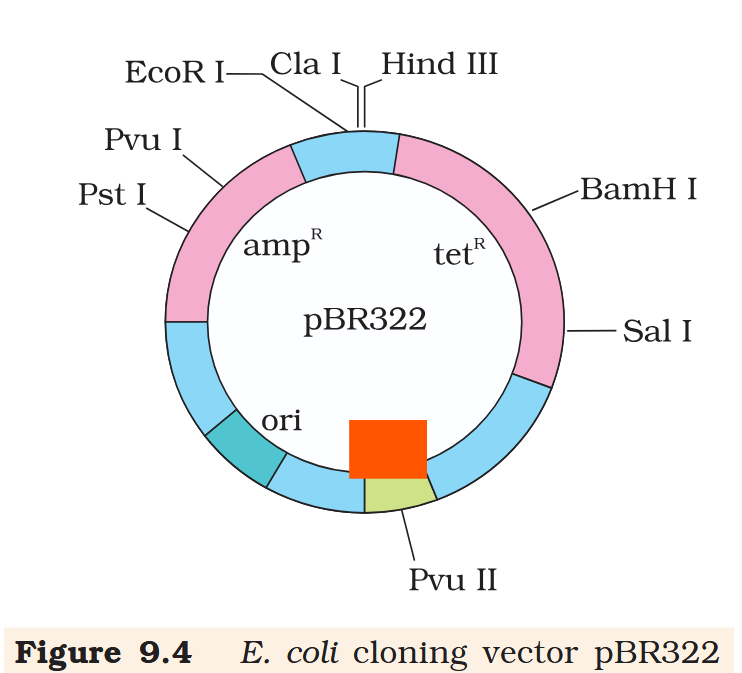
rop
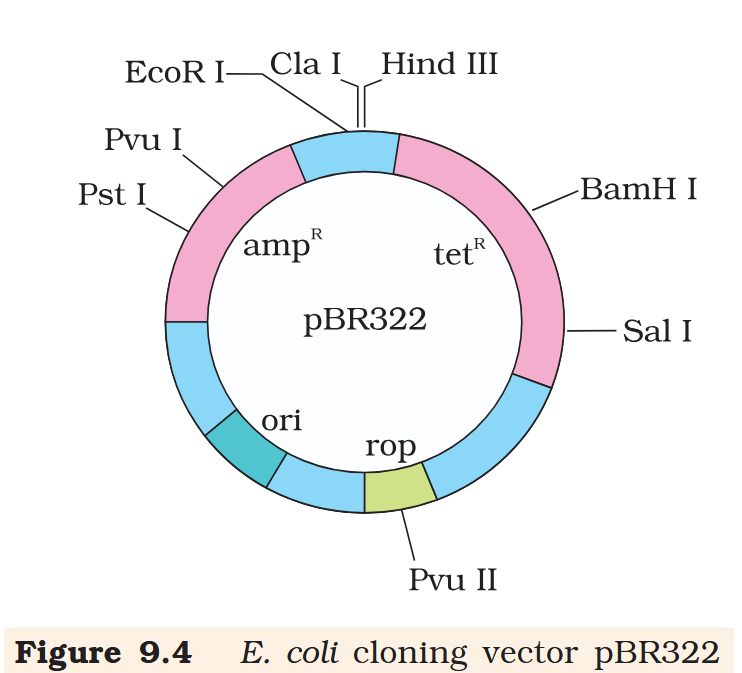
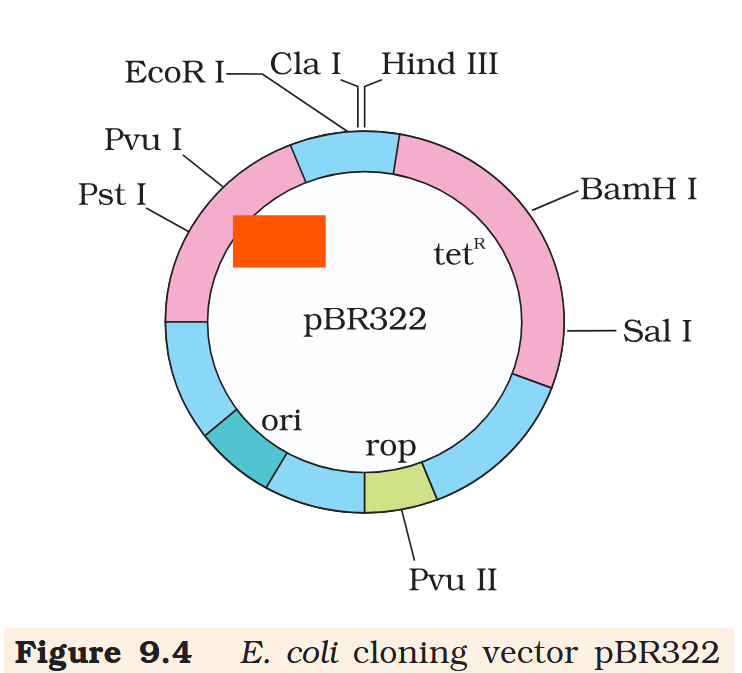
ampR
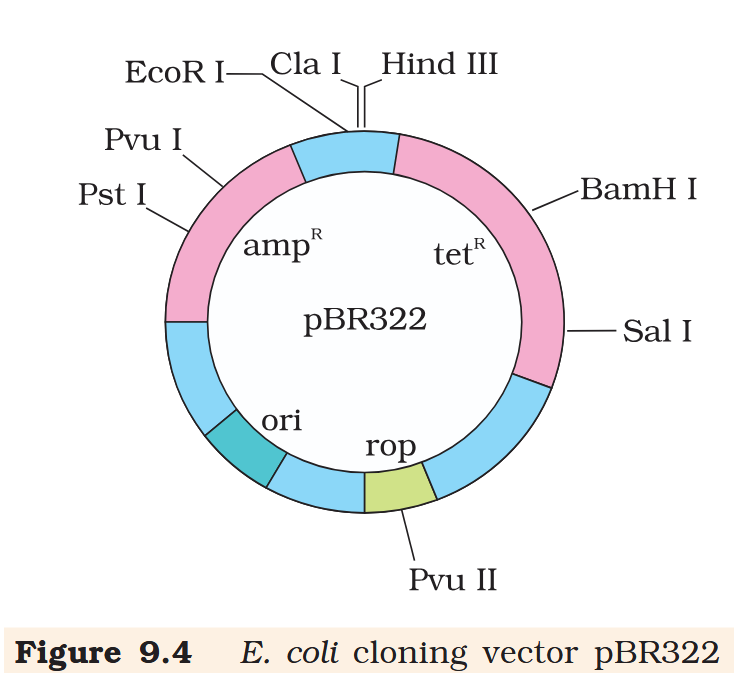
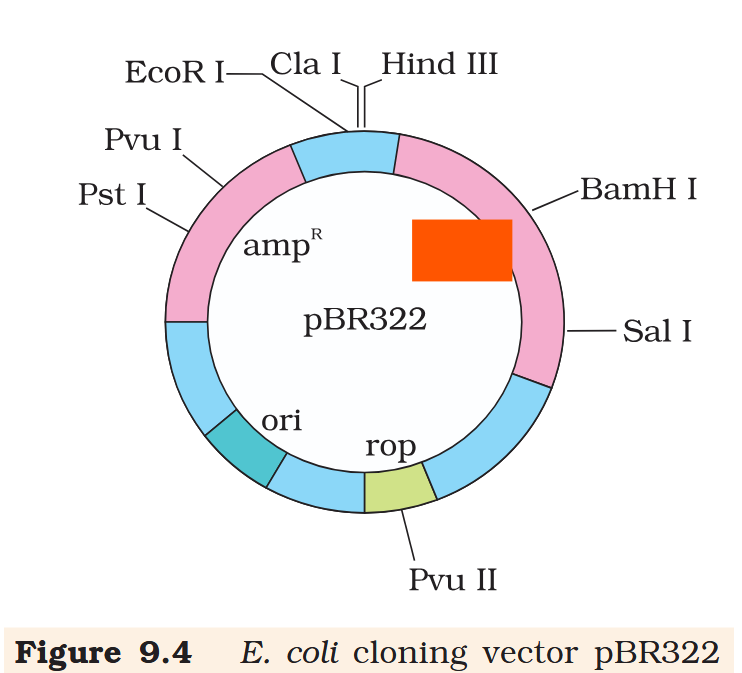
tetR
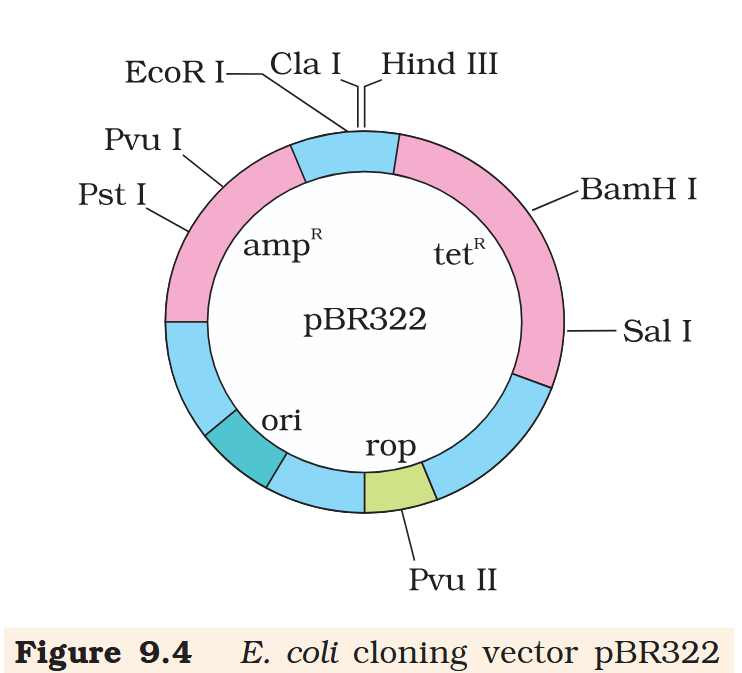
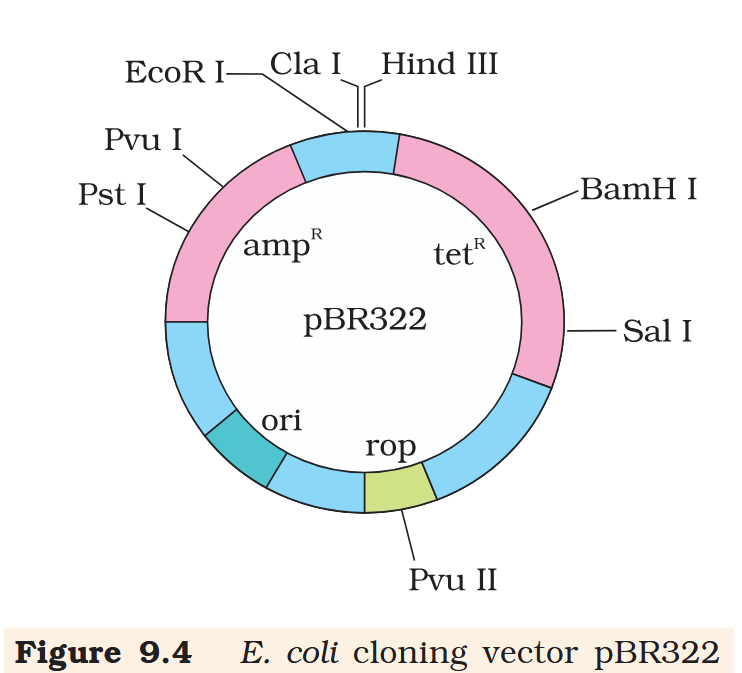
What does “rop” do?
rop codes for the proteins involved in the replication of the plasmid.
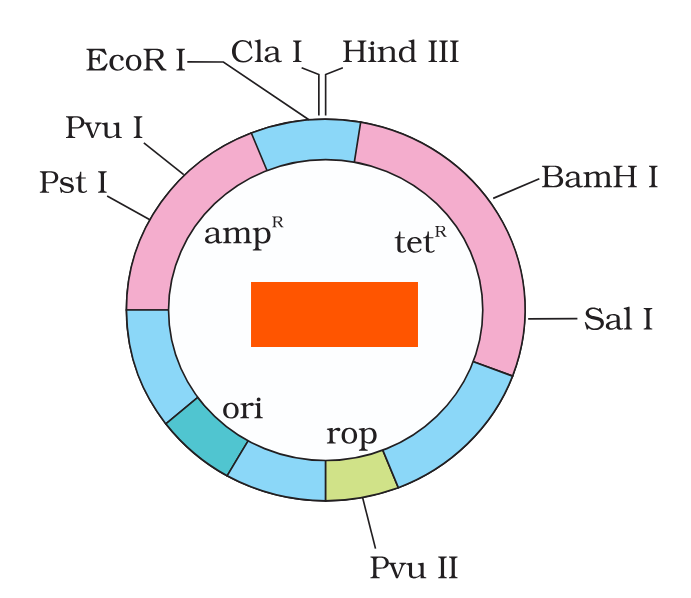
What is the name of this cloning vector?
pBRE322
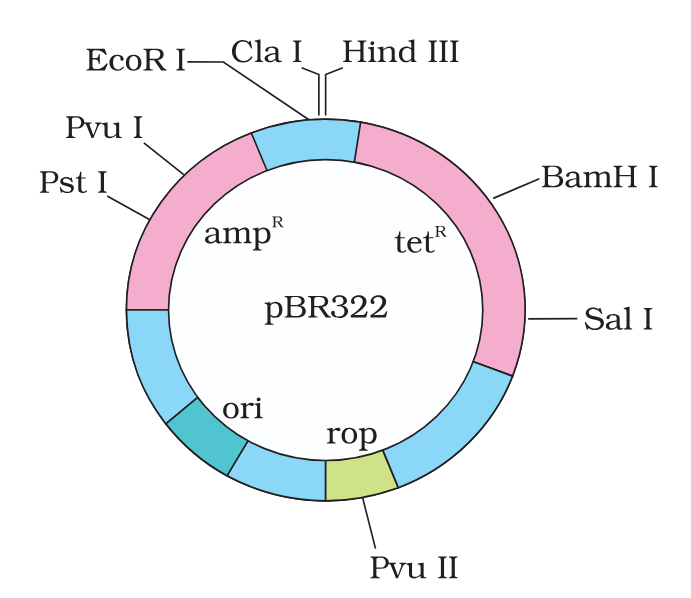
in E. coli cloning vector pBRE322, the ligation of alien DNA is carried out at a restriction site present where?
in E. coli cloning vector pBRE322, the ligation of alien DNA is carried out at a restriction site present in one of the two antibiotic resistance genes.
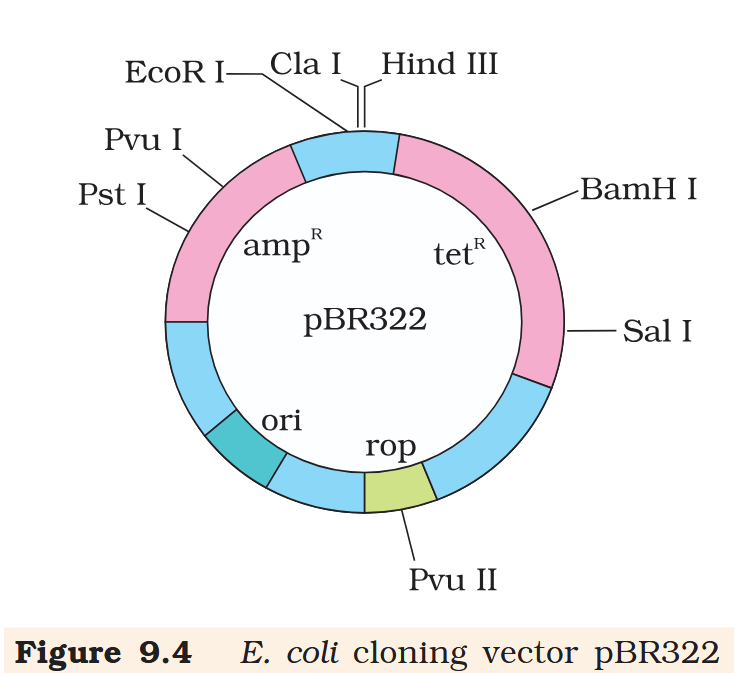
What is tetR for?
tetR is the tetracycline resistance gene
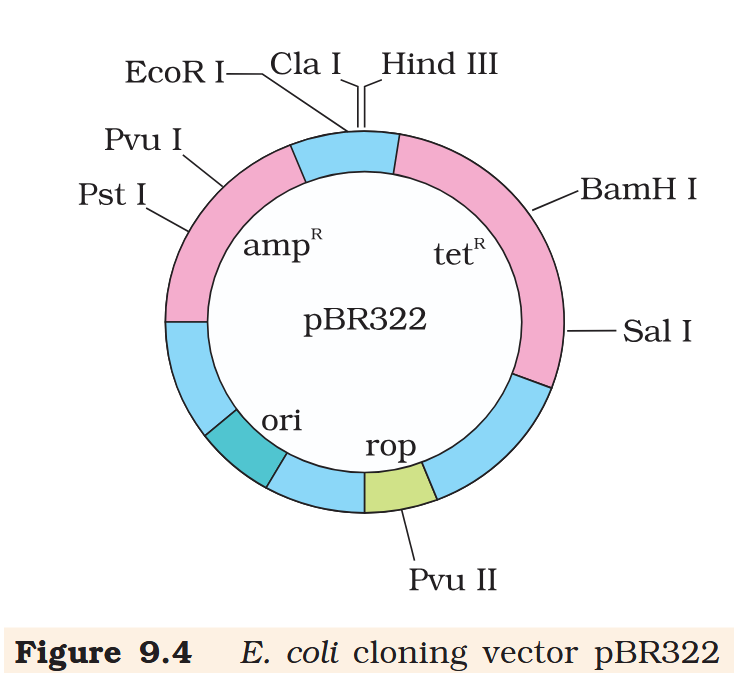
What is ampR for?
ampR is the ampicillin resistance gene
If you ligate a foreign DNA at the BamH I site of tetracycline resistance gene in the vector pBR322, what antibiotic resistance characters can be observed in the transformed bacteria?
Resistance to ampicillin is present, resistance to tetracycline is absent.
If you ligate a foreign DNA at the BamH I site of tetracycline resistance gene in the vector pBR322, how can you then differentiate between the recombinant and non-recombinant bacteria?
The recombinant plasmids will lose tetracycline resistance due to insertion of foreign DNA but can still be selected out from non-recombinant ones by plating the transformants on tetracycline containing medium. The transformants growing on ampicillin containing medium are then transferred on a medium containing tetracycline. The recombinants will grow in ampicillin containing medium but not on that containing tetracycline. But, non- recombinants will grow on the medium containing both the antibiotics.
Selection of recombinants due to inactivation of antibiotics is a cumbersome procedure. Why?
Because it requires simultaneous plating on two plates having different antibiotics.
Selection of recombinants due to inactivation of antibiotics is a cumbersome procedure.
Therefore, alternative selectable markers have been developed which differentiate recombinants from non-recombinants on the basis of what?
On the basis of their ability to produce colour in the presence of a chromogenic substrate.
Alternative selectable markers have been developed which differentiate recombinants from non-recombinants on the basis of their ability to produce colour in the presence of a chromogenic substrate.
How does this work?
In this, a recombinant DNA is inserted within the coding sequence of an enzyme, β-galactosidase. This results into inactivation of the gene for synthesis of this enzyme. The presence of a chromogenic substrate gives blue coloured colonies if the plasmid in the bacteria does not have an insert. Presence of insert results into insertional inactivation of the β-galactosidase gene and the colonies do not produce any colour, these are identified as recombinant colonies.
What is “insertional activation” under biotechnology?
In this, a recombinant DNA is inserted within the coding sequence of an enzyme, β-galactosidase. This results into inactivation of the gene for synthesis of this enzyme, which is referred to as insertional inactivation.
____________ _____________, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen.
Agrobacterium tumifaciens, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen.
Agrobacterium tumifaciens, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘T-DNA’ to transform normal plant cells how?
Agrobacterium tumifaciens, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen.
Agrobacterium tumifaciens, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘______’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen.
Agrobacterium tumifaciens, a pathogen of several dicot plants is able to deliver a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen.
__________ in animals have the ability to transform normal cells into cancerous cells.
retroviruses in animals have the ability to transform normal cells into cancerous cells.
retroviruses in animals have the ability to transform normal cells into ____________ cells.
retroviruses in animals have the ability to transform normal cells into cancerous cells.
A better understanding of the art of delivering genes by pathogens in their eukaryotic hosts has generated knowledge to transform these tools of pathogens into useful vectors for delivering genes of interest to humans.
Cool, huh?
Goated even
The tumor inducing (Ti) plasmid of ____________ ___________ has now been modified into a cloning vector which is no more pathogenic to the plants but is still able to use the mechanisms to deliver genes of our interest into a variety of plants. Similarly, ___________ have also been disarmed and are now used to deliver desirable genes into animal cells. So, once a gene or a DNA fragment has been ligated into a suitable vector it is transferred into a bacterial, plant or animal host (where it multiplies).
[NCERT se chhapa hai]
The tumor inducing (Ti) plasmid of Agrobacterium tumifaciens has now been modified into a cloning vector which is no more pathogenic to the plants but is still able to use the mechanisms to deliver genes of our interest into a variety of plants. Similarly, retroviruses have also been disarmed and are now used to deliver desirable genes into animal cells. So, once a gene or a DNA fragment has been ligated into a suitable vector it is transferred into a bacterial, plant or animal host (where it multiplies).
In order to force bacteria to take up the plasmid, the bacterial cells must first be made ‘competent’ to take up DNA.
Why?
Since DNA is a hydrophilic molecule, it cannot pass through cell membranes.
In order to force bacteria to take up the plasmid, the bacterial cells must first be made ‘competent’ to take up DNA.
How can this be done with the use of divalent cations?
This is done by treating them with a specific concentration of a divalent cation, such as calcium, which increases the efficiency with which DNA enters the bacterium through pores in its cell wall. Recombinant DNA can then be forced into such cells by incubating the aforementioned cells (along with with recombinant DNA) on ice, followed by placing them briefly at 42℃ (heat shock), and then putting them back on ice. This enables the bacteria to take up the recombinant DNA that it was incubated with.
In order to force bacteria to take up the plasmid, the bacterial cells must first be made ‘competent’ to take up DNA.
How can this be done with the “microinjection” method?
In a method known as micro-injection, recombinant DNA is directly injected into the nucleus of an animal cell.
[That’s all NCERT says sorry]
In order to force bacteria to take up the plasmid, the bacterial cells must first be made ‘competent’ to take up DNA.
How can this be done for plant cells?
In a method, suitable for plants, cells are bombarded with high velocity micro-particles of gold or tungsten coated with DNA in a method known as biolistics or gene gun.
In order to force bacteria to take up the plasmid, the bacterial cells must first be made ‘competent’ to take up DNA.
How can this be done using pathogens?
This method uses ‘disarmed pathogen’ vectors, which when allowed to infect the cell, transfer the recombinant DNA into the host.
What are the 6 steps in the processes of Recombinant DNA Technology?
Isolation of the genetic material (DNA)
Cutting of DNA at specific locations
Amplification of Gene of Interest (PCR)
Insertion of Recombinant DNA into the Host Cell/Organism
Obtaining the foreign gene product
Downstream processing
nucleic acid is the genetic material of all organisms without exception.
True or false?
True. Nucleic acid comprises of RNA and DNA both.
In order to cut the DNA with restriction enzymes, it needs to be in pure form, free from other macro-molecules.
True or false?
True.