B7- DNA and Protein synthesis ✅
1/19
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
Compare and contrast DNA in eukaryotic cells with DNA in prokaryotic cells
Similarities:
Nucleotide structure is identical - deoxyribose attached to phosphate and a base
Adjacent nucleotides joined by phosphodiester bonds, complementary bases joined by hydrogen bonds
DNA in mitochondria / chloroplasts have similar structure to DNA in prokaryotes → Short, circular, not associated with proteins
Differences:
Eukaryotic DNA is longer
Eukaryotic DNA is linear, prokaryotic DNA is circular
Eukaryotic DNA is associated with histone proteins, prokaryotic DNA is not
Eukaryotic DNA contain introns, prokaryotic DNA does not
what is a homologous chromosome
pair of chromosomes
with the same genes
in same loci
define the term gene
A sequence of DNA (nucleotide) bases that codes for:
The amino acid sequence of a polypeptide
Or a functional RNA (eg. ribosomal RNA or tRNA)
define the term allele
different versions of the same gene
define the term degenerate
more than one triplet codes for the same amino acid
what is a chromosome ?
Long, linear DNA + its associated histone proteins
In the nucleus of eukaryotic cells
what is a locus?
Fixed position a gene occupies on a particular DNA molecule.
what’s the difference between a:
triplet
codon
anticodon
3 bases on strand on DNA
3 bases on a strand of RNA ( complementary to DNA base sequence )
3 bases on a strand of tRNA
Describe the nature of the genetic code
Triplet code→ A sequence of 3 DNA bases, called a triplet, codes for a specific amino acid
Universal → The same base triplets code for the same amino acids in all organisms
Non-overlapping→ Each base is part of only one triplet so each triplet is read as a discrete unit
Degenerate → An amino acid can be coded for by more than one base triplet
What are ‘non-coding base sequences’ and where are they found?
Non-coding base sequence - DNA that does not code for amino acid sequences / polypeptides:
1. Between genes - eg. non-coding multiple repeats
2. Within genes - introns
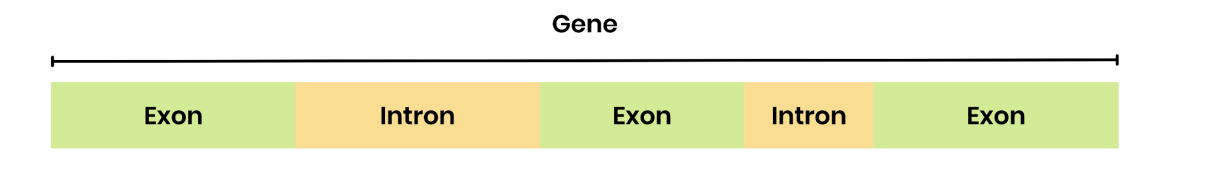
What are introns and exons?
introns→ base sequence of a gene that doesn’t code for amino acids, in eukaryotic cells
exons→ base sequence of a gene coding for amino acid sequences (in a polypeptide)
define genome
The complete set of genes in a cell (including those in mitochondria and /or chloroplasts)
define proteome
The full range of proteins that a cell can produce (coded for by the cell’s DNA / genome)
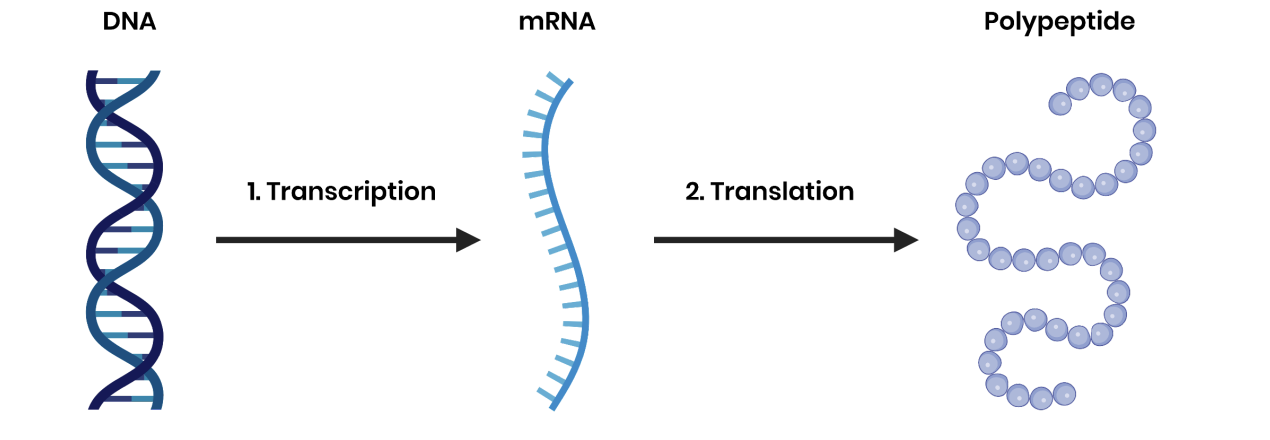
Describe the two stages of protein synthesis
transcription- Production of messenger RNA (mRNA) from DNA, in the nucleus
translation- Production of polypeptides from the sequence of codons carried by mRNA, at ribosomes
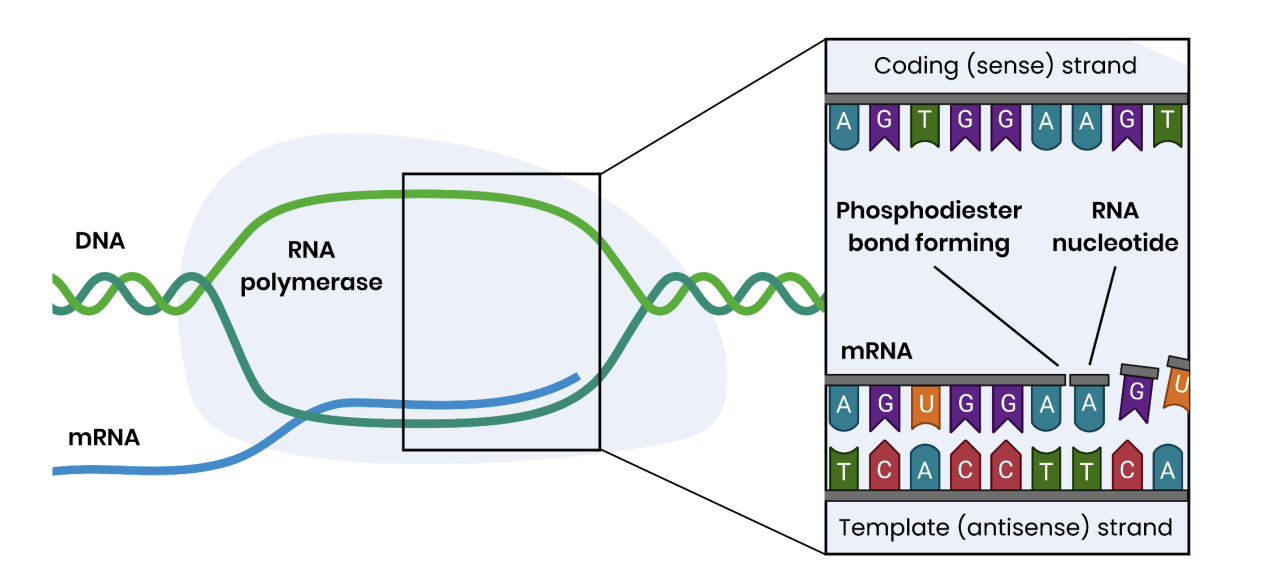
transcription process
DNA helicase and RNA polymerase breaks weak hydrogen bonds between DNA bases and strands separate
Only one DNA strand acts as a template
Free RNA nucleotides align next to their complementary bases on the template strand
In RNA, uracil is used in place of thymine (pairing with adenine in DNA)
RNA polymerase joins adjacent RNA nucleotides
This forms phosphodiester bonds via condensation reactions
Pre-mRNA is formed and this is spliced to remove introns, forming (mature) mRNA
how many sites are there for tRNA anticodons in a singular ribosome and why?
2 sites for 2 tRNA anticodons
ensuring the 2 tRNA molecules are close enough so a peptide bond can form between 2 amino acids
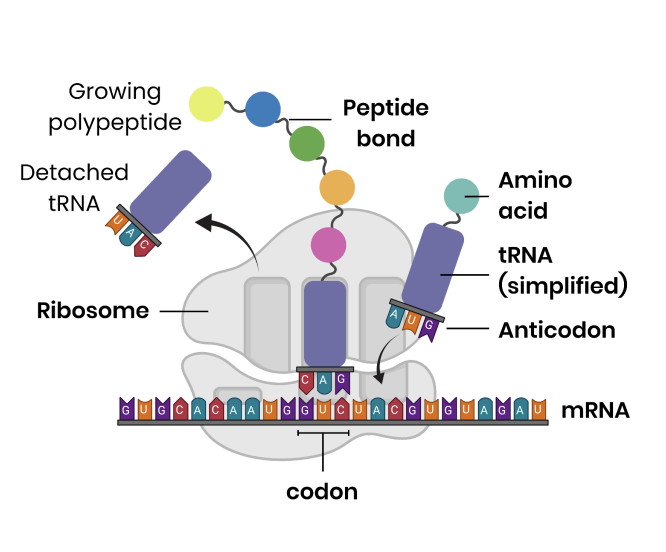
translation process
mRNA attaches to a ribosome and the ribosome moves to a start codon
tRNA brings a specific amino acid
tRNA anticodon binds to complementary mRNA codon
Ribosome moves along to next codon and another tRNA binds so 2 amino acids can be joined by a condensation reaction forming a peptide bond → Using energy from hydrolysis of ATP
tRNA released after amino acid joined polypeptide
Ribosome moves along mRNA to form the polypeptide, until a stop codon is reached
describe the role of ATP in translation
Hydrolysis of ATP to ADP + Pi releases energy
So amino acids join to tRNAs and peptide bonds form between amino acids
describe the role of tRNA in translation
● transports a specific amino acid, in relation to its anticodon
● tRNA anticodon complementary base pairs to mRNA codon, forming hydrogen bonds
● 2 tRNAs bring amino acids together so peptide bond can form
describe the role of ribosomes in translation
mRNA binds to ribosome, with space for 2 codons
Allows tRNA with anticodons to bind
Catalyses formation of peptide bond between amino acids (held by tRNA molecules)
Moves along (mRNA to the next codon) / translocation