Manipulating Genomes: General Exam Flashcards
1/41
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
42 Terms

Contraction of smooth muscle
Extra mucus production
Inflammation
any of the two

Conservation ;
Gene bank ;
Teaching ;
Leisure ;
any of the following
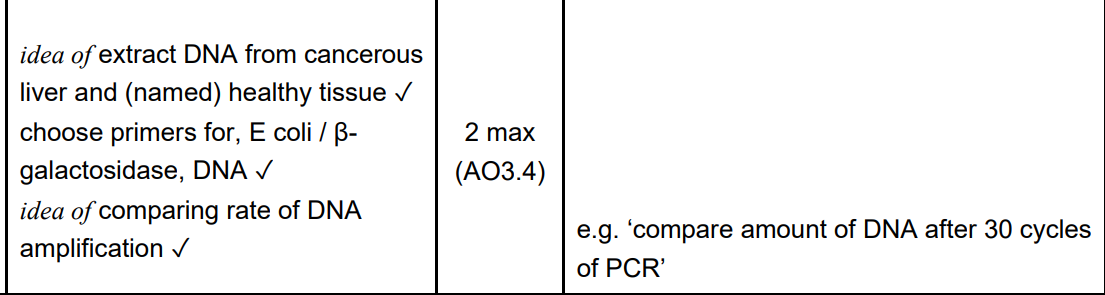
(b) DNA sequencing techniques have provided new information about plant relationships.
Outline the roles of each of the following procedures in sequencing a genome:
(i) the polymerase chain reaction (PCR) (2)
To amplify DNA
and produce a range of different lengths.
(iii) digestion of DNA by restriction enzymes. (2)
To cut the DNA into smaller fragments.
and to cut vectors.
(c) Suggest why a genome has to be fragmented before sequencing. (2)
Genome is very large
so better accuracy.
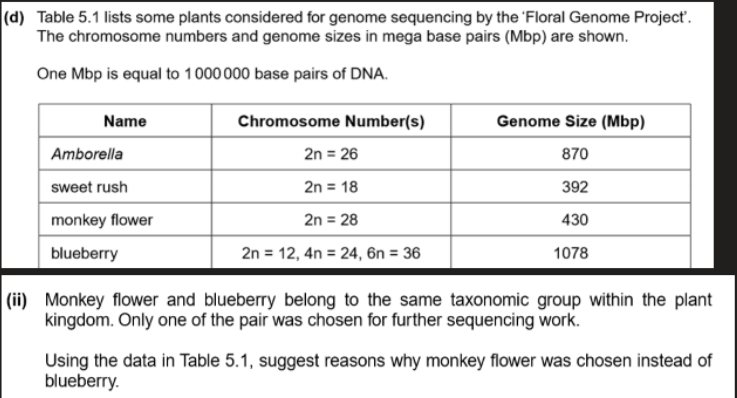
Monkey flower has fewer genome
So quicker and cheaper
* Try to avoid saying ‘easier’ as that isn’t proper biological terminology.

Larger.
*This is because of the fact that the nucleus has to hold 6x the normal amount of genetic info, so the cells have to compensate by being larger.
DNA sequence information is most useful when used with the phylogenetic (cladistic)
approach to classification.
How does the phylogenetic approach to classifying species differ from the biological species concept?
No need to test for interbreeding
Can apply to organisms that reproduce asexually
the biological concept of identifying species is based on whether species can reproduce to have fertile offspring, whilst the phylogenetic concept is based on identification of evolutionary links between species.
(b) Discuss the potential benefits to mankind and the ethical concerns raised by the following
examples of genetically modified organisms:
• rice modified for increased vitamin A content (‘Golden RiceTM’)
• humans having somatic gene therapy treatment for a genetic disease.
In your answer you should give a balanced account of the benefits and concerns for
each example of genetic modification.
Golden Rice
Reduction of Vitamin A deficiency in Asia
However the genetic diversity of rice is reduced
clone may suffer from one disease
potential hybridisation with wild rice
the seeds are very expensive
Somatic Gene Therapy
Allows for a better quality of life
can help with conditions such as Cystic Fibrosis/ Parkinson’s
Hence expanding the lifespans of individuals undergoing treatment.
However the virus vector could potentially cause disease
Invasive procedure, which needs to be repeated
Potential rejection from the immune system
Unknown effects for both.
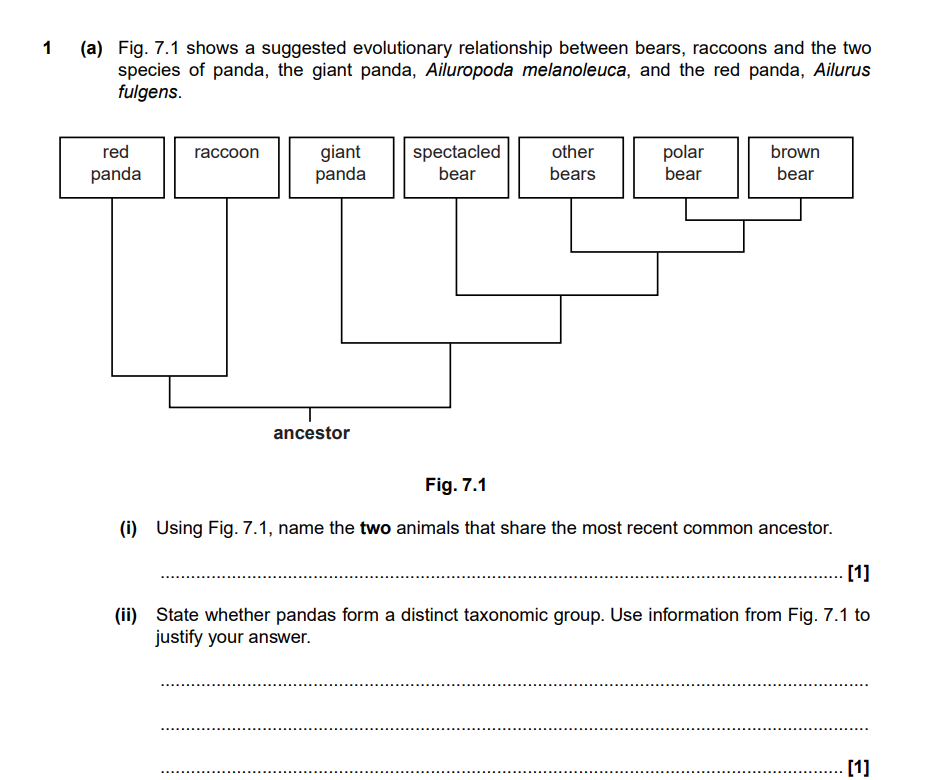
i) Polar bear and Brown Bear
ii) The giant panda is more closely related to the spectacled bear and other bears than it is to the red panda.
The red panda shares a more recent common ancestor with the raccoon.
So no.
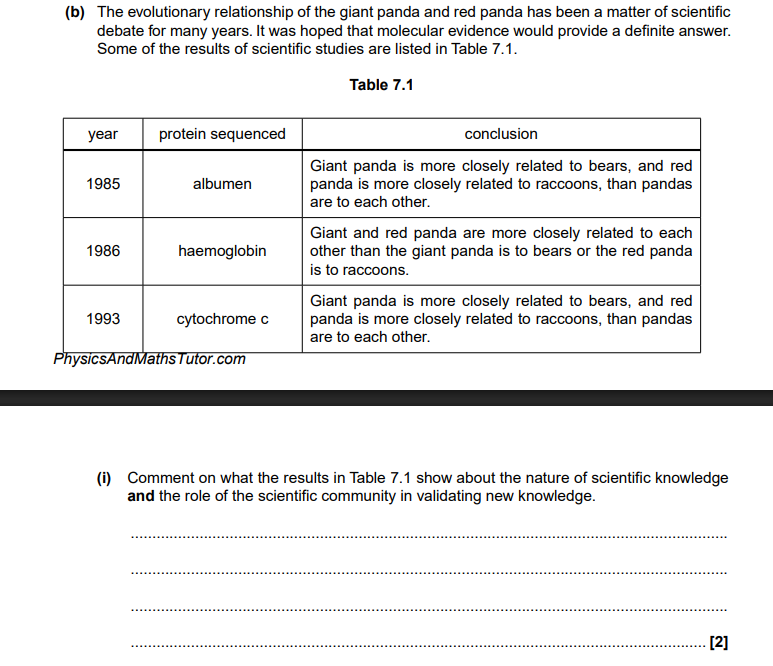
Scientific knowledge is always developing over time,
and the role of the scientific community is to validate new knowledge via peer review.
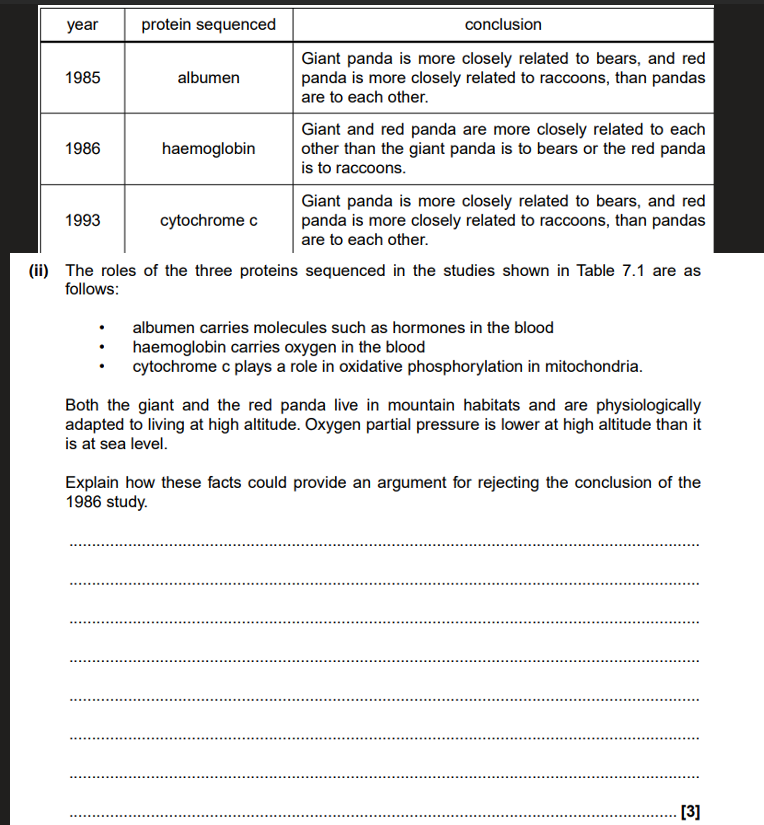
Both pandas share the same selection pressures,
So the haemoglobin may act as an adaptation,
and could be a result of convergent evolution.

Model Answer: Humans are diploid organisms, meaning they possess two sets of chromosomes. Therefore, an individual inherits one copy (allele) of the DRD4 gene from their mother and one copy from their father.
(e) One allele of DRD4 has been found more frequently amongst individuals whose personality is described as ‘novelty-seeking’ and whose behaviour tends to be exploratory and impulsive. Suggest how this particular allele of the DRD4 receptor could have become common in the human population. (4)
Model Answer: 1. Selection Pressure: In the past, 'novelty-seeking' behavior provided a selective advantage.
2. Survival: Individuals with this allele were more likely to explore new habitats or find new food sources, increasing their chances of survival.
3. Reproduction: These individuals were more likely to survive to adulthood and reproduce successfully.
4. Inheritance: The advantageous allele was passed on to their offspring.
5. Frequency: Over many generations, the frequency of this allele increased within the population.
Describe how gel electrophoresis can be used to compare two samples of DNA (6 marks).
Mark 1: Preparation. Mentioning restriction enzymes (endonucleases) to cut DNA into fragments of different lengths.
Mark 2: Loading. DNA samples are placed into wells at the negative electrode (cathode) end of an agarose gel.
Mark 3: Potential Difference. An electric current is applied across the gel.
Mark 4: Charge/Movement. DNA is negatively charged (phosphate groups) and moves toward the positive electrode (anode).
Mark 5: Size Separation. The gel acts as a mesh; smaller fragments move faster and further than larger fragments.
Mark 6: Comparison. Use of a DNA ladder (standard) or comparing the final band patterns (profiles) to see if they match.
What must be used to see the DNA bands after electrophoresis?
A fluorescent dye (e.g., Ethidium Bromide) or a radioactive probe/UV light.
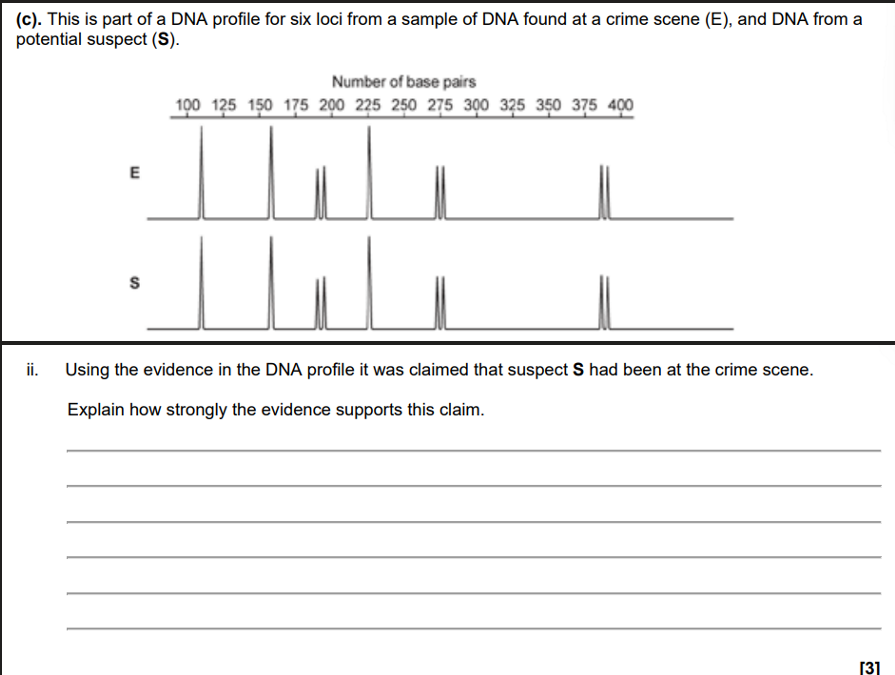

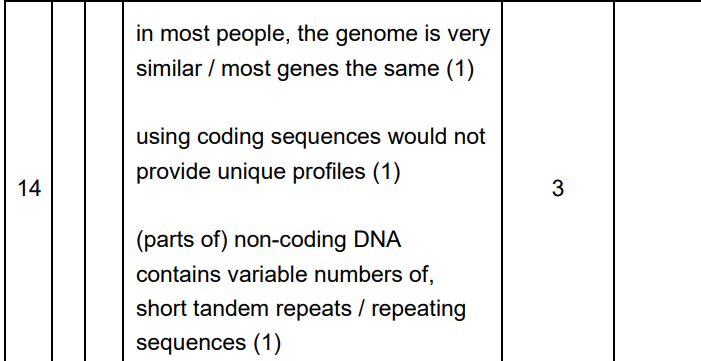
DNA Profiles are identical
Probability of two people having identical profiles very low
BUT
6 is a low number of peaks
Individuals may be closely related
Could be identical twins

Gene sequencing
• determines order of bases • order of bases linked to order of amino acids • order of amino acids is protein primary structure • DNA sequence can be inferred from, and implies, protein primary structure
Bioinformatics
• stores and organises large amounts of data • databases of o DNA and amino acid sequences o protein structures o metabolic pathways • facilitates fast retrieval and sharing of information • algorithms and statistical tests
Computational biology
• needed for analysis of large amounts of data • rapid processing of data • prediction of amino acid sequences • modelling of protein structure or function • algorithms and statistical tests

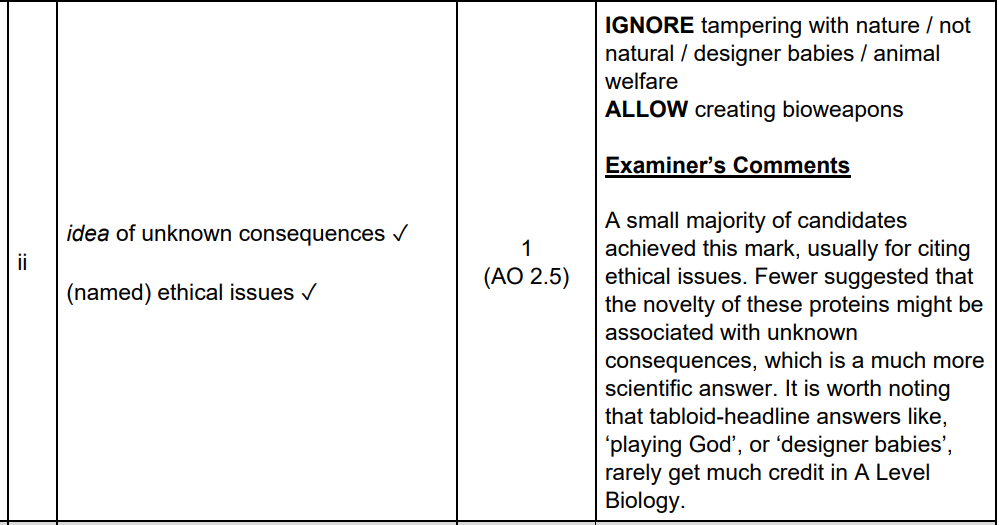

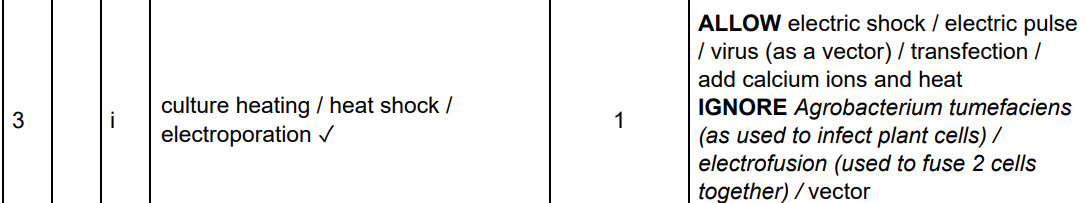
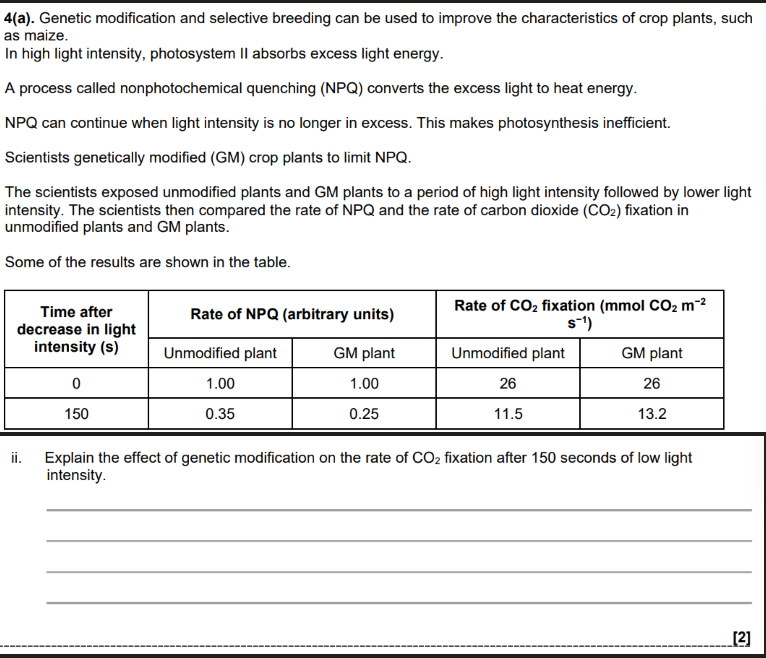
Don’t compare the wrong parts from the table.
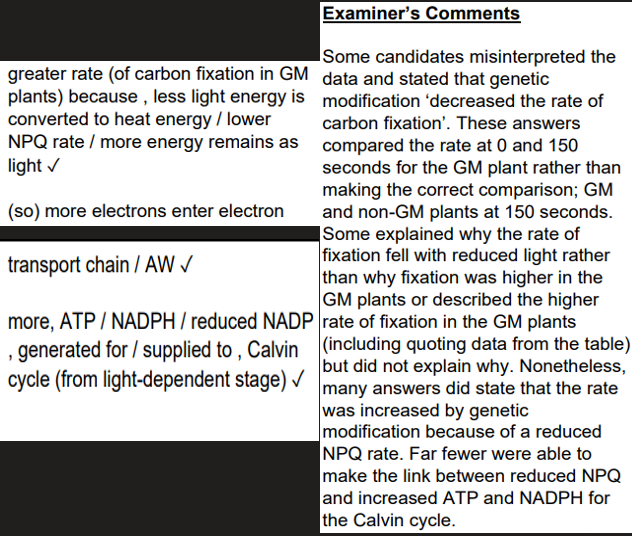


use , algorithms / statistical tests / (mathematical) models ✓
store , data / information (from different DNA sequences) ✓
analyse / identify, differences / similarities, in DNA (sequences) / alleles ✓
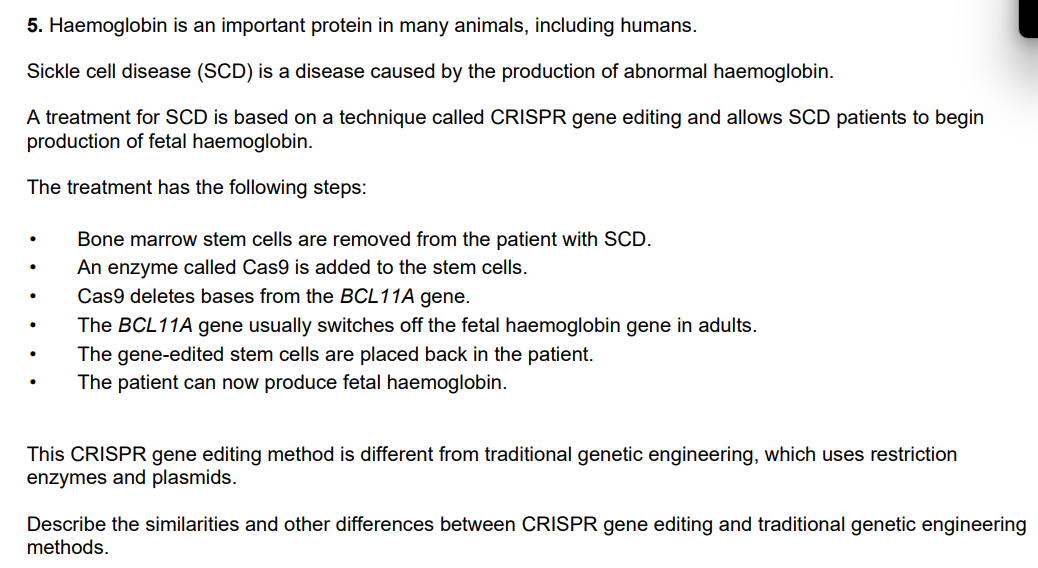


(4 marks)
Similarities
(both) use enzyme that cuts DNA ✓
(both) change base, sequence / order, in the organism ✓
idea that they both result in the production of a new polypeptide ✓
Differences
no , gene / DNA , insertion in (this method of) CRISPR ✓
no, marker genes / ligase / bacterial cells / vector, used in CRISPR ✓
idea that traditional genetic engineering is illegal in humans ✓
What are the three specific enzymes/tools required for traditional genetic engineering mentioned in the OCR specification?
Restriction enzymes (to cut DNA), plasmids (as vectors), and DNA ligase (to join DNA fragments/form recombinant DNA).
What is 'Electroporation'?
A technique used in genetic engineering where an electrical current is applied to cells to increase membrane permeability, allowing DNA (like plasmids) to enter.

i) virus / viral vector ✓ liposome ✓ (any one)
ii) Sharing knowledge.

DNA sequencing determines the order of bases (nucleotides) in a gene.
Each triplet of bases (codon) codes for a specific amino acid.

Sequencing (1 Mark): High-throughput sequencing allows scientists to monitor the rapid rate of mutation in the Ebola genome over time.
Sequencing (1 Mark): It can be used to identify changes in antigen/surface protein structure that might render current vaccines ineffective.
Bioinformatics (1 Mark): Facilitates genome-wide comparisons between different viral strains to identify highly conserved (unchanging) regions of the genome to target with vaccines.
Bioinformatics (1 Mark): Can be used in epidemiology to track the spread of specific strains through a population, allowing for targeted "ring vaccination" in high-risk areas.

Why is monitoring 'rapid mutation rates' critical for vaccine effectiveness?
Mutations can change the shape of antigens (proteins) on the pathogen's surface, meaning memory cells from previous vaccinations may no longer recognize them.
Define 'Epidemiology' in the context of DNA sequencing.
Using genomic data to track the source, spread, and transmission patterns of a disease within a population.
What is the advantage of 'high-throughput' or 'nanopore' sequencing over traditional Sanger sequencing?
It is significantly faster and can sequence entire genomes rapidly, which is essential during a fast-moving viral outbreak.

Inbreeding, which reduces genetic diversity.
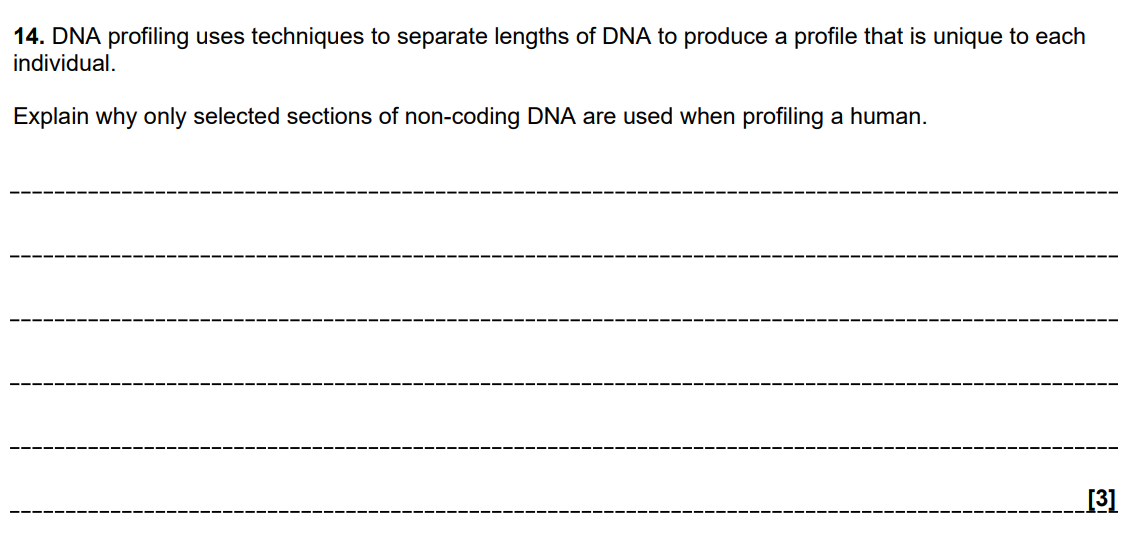



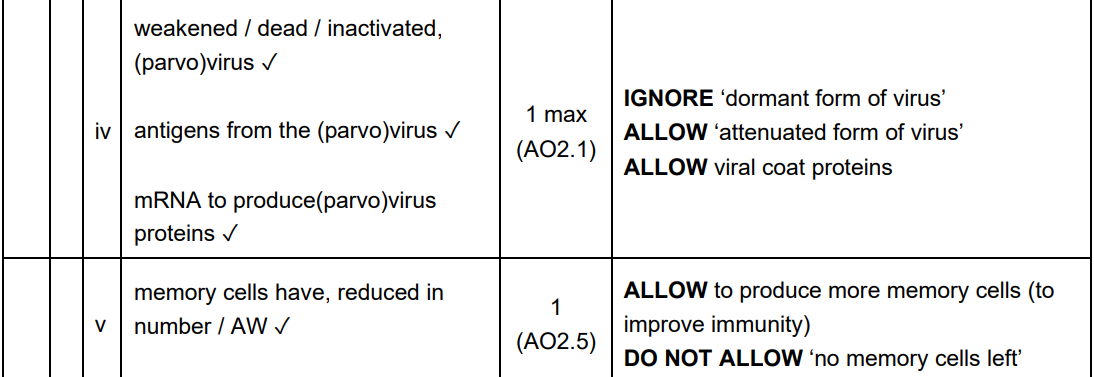



The Trap: Many students describe "putting DNA in a plant" but fail to mention the specific enzymes or the method of cell delivery.
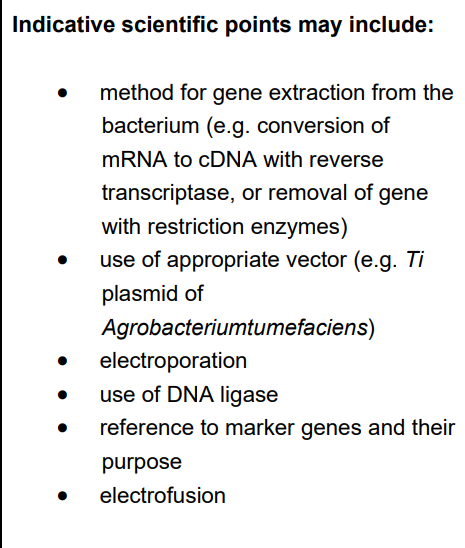
Marking Criteria: You need to cover three distinct stages: Isolation, Insertion into Vector, and Transformation/Cloning.
Isolation of the Bt Gene:
Identify the specific gene for moth larvae resistance in Bacillus thuringiensis.
Use restriction endonucleases to cut the gene from the bacterial DNA.
These enzymes leave "sticky ends" (short sections of single-stranded DNA).
Creation of Recombinant DNA:
Cut a plasmid (vector) using the same restriction enzyme so the sticky ends are complementary.
Mix the Bt gene and the opened plasmid together.
Use DNA ligase to join the sugar-phosphate backbones, forming recombinant DNA.
Transformation and Plant Growth:
Insert the recombinant plasmid into the plant cells (transformation). This can be done via electroporation or using a secondary bacterial vector like Agrobacterium tumefaciens.
Use tissue culture (micropropagation) to grow the transformed cells into adult, pest-resistant aubergine plants.

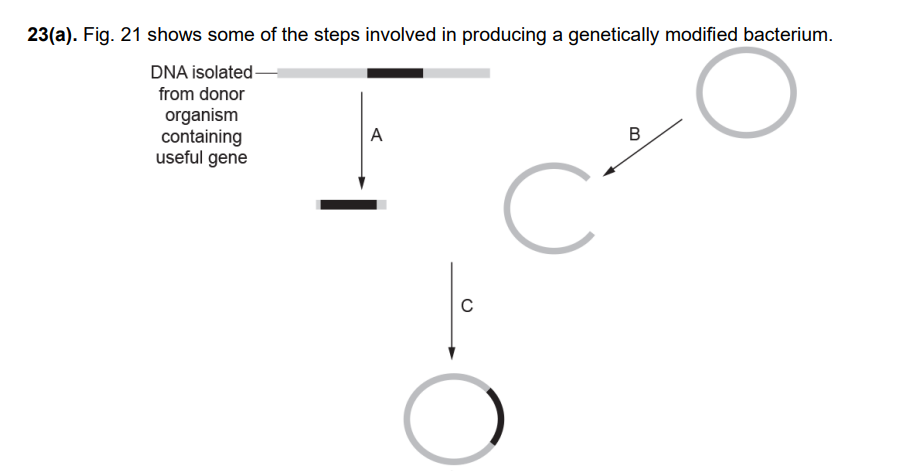
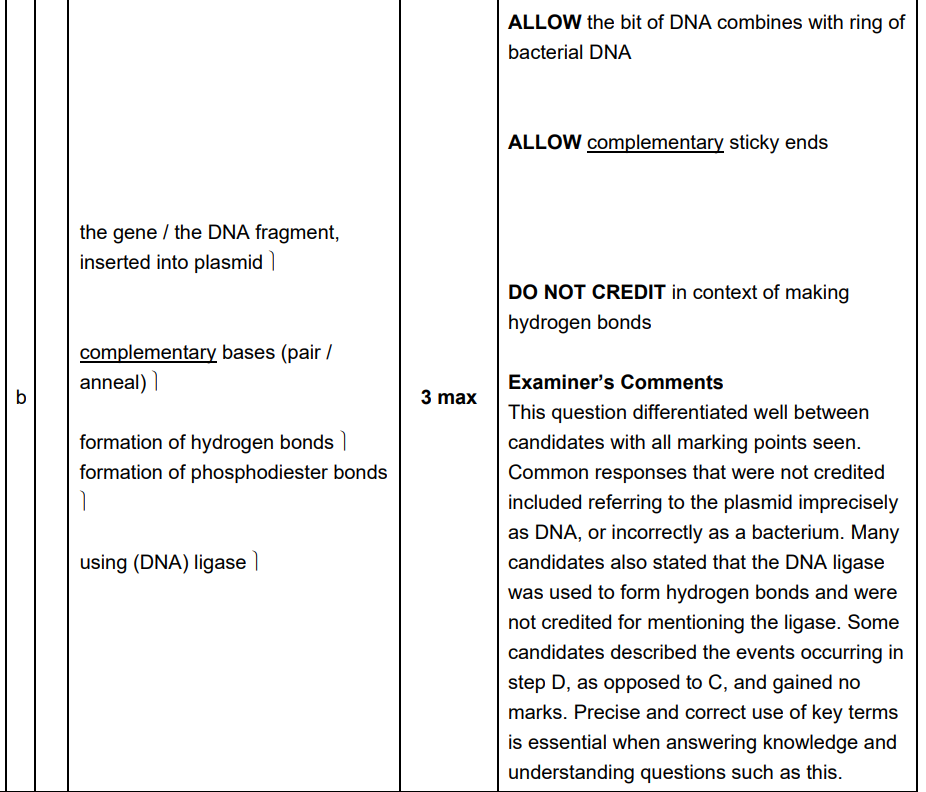
Describe the events which are happening at C. (3 marks)
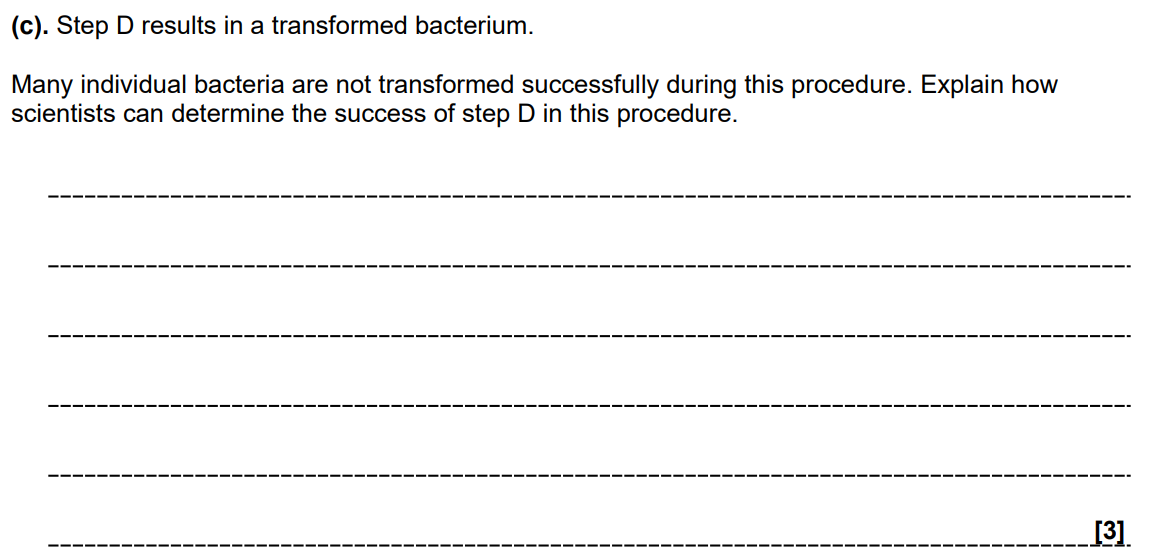


Marker Genes (1 Mark): The plasmid used as a vector contains a marker gene, such as a gene for antibiotic resistance or a fluorescent protein.
Selective Growth (1 Mark): The bacteria are grown on an agar medium containing the specific antibiotic.
Survival/Identification (1 Mark): Only the transformed bacteria that have successfully taken up the plasmid will possess the resistance gene and survive to form colonies; those not transformed will die.
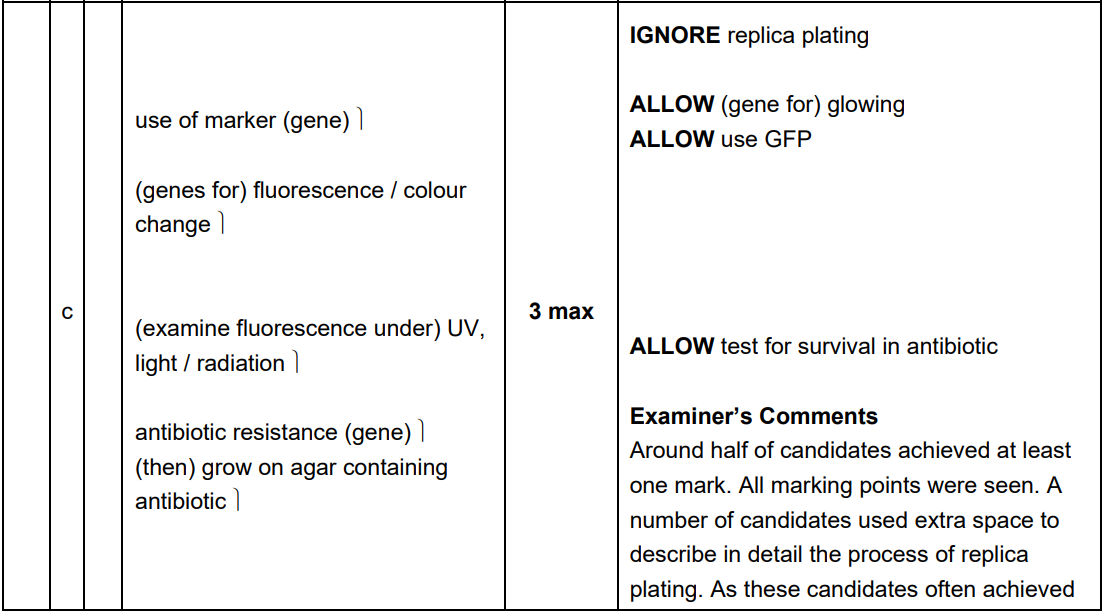
Why is a 'marker gene' necessary in a plasmid vector?
Because bacterial transformation is inefficient; the marker gene allows scientists to identify and select only the few cells that successfully took up the recombinant DNA.

