Biochem lecture 22: translation
1/122
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
123 Terms
Characteristics of the genetic code
a nonoverlapping triplet code
Nonoverlapping code
Each three-nucleotide triplet is distinct.
Overlapping code (ex. reading ABC, BCD, CDE) (a single base can…)
a single base can be part of multiple consecutive codons
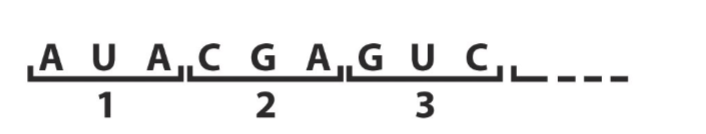
nonoverlapping code
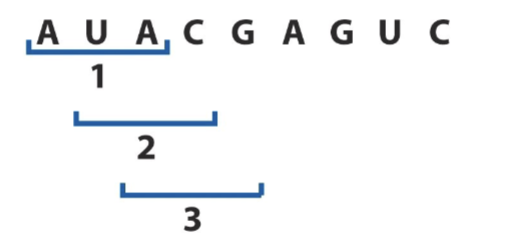
overlapping code
Codes examples with possible # of AAs for each
single base -> 4 (4¹), two bases -> 16 (4²), three bases -> 64 (4³)
Possible # of AA’s
4, 16, 64
For 20 amino acids at least __ nucleotides are necessary per codon
3
____ reading frames are possible because the code is nonoverlapping
3
Each frame gives a different set of _____
triplets
Selection and insertion result in
frame shifts
Insertion and deletion or three insertions can result in
returning to the reading frame
Deciphering the code-using trinucleotides to bind ____ ____ to ____
charged tRNA to ribosomes
Deciphering the code- 1) using trinucleotides to bind charged tRNAs to ribosomes ex.
UUU-> tRNA (Phe), AAA -> tRNA (lys), CCC -> tRNA (pro)
Deciphering the code- 2) using defined ….
polynucleotides in a cell-free translation system
How are polyribonucleotides synthesized
chemically or with polynucleotide phosphorylase
Deciphering the code- resulted in the
genetic code table
The genetic code is _____
degenerate
Degenerate
some amino acids have more than 1 codon associated with it
Degeneracy can minimize
the deleterious effects of mutations
3rd position mutations can be ____
silent
____ is the adapter
tRNA
tRNA molecule can do… and this is what type of process
base pairing, selective process
Anticodon and codon are in _____ direction
opposite direction
Codon and anticodon are inherently antiparallel due to what property?
polarity
Watson-Crick base pairs
A-U, G-C
What is a base pair that is also possible that Watson-Crick did not know about
G-U
Wobble hypothesis
the third base of a mRNA codon (at the 3’ end) can pair flexibly with the first base of a tRNA anticodon (at the 5' end), allowing one tRNA to recognize multiple synonymous codons
___ codons possible, ___ codons for stop leaves ___ codons to be recognized only up to __ unique tRNA anticodons discovered
64; 3; 61; 45
we know wobble hypothesis exists because…
there are more codons than anticodons
Wobble pairing allows some ….
tRNA (anticodons) to recognize more than one codon
The wobble hypothesis 1
The 1st two 5' bases of an mRNA codon always form strong base pairs with the tRNA anticodon, conferring most of the specificity.
I pairs with
A, U, or C
The wobble hypothesis 2
The 5’ base of the anticodon determines the number of codons recognized by the tRNA
when the anticodon 5’ base is ___ or ___ only 1 codon is recognized
C or A
When the anticodon 5’ base is ___ or ___ 2 different codons may be recognized
U or G
when the anticodon 5’ base ___ 3 different codons can be recognized
I
The wobble hypothesis 3
When an amino acid is specified by several different codons, the codons that differ in either of the 1st two bases require different tRNAs
The wobble hypothesis 4
minimum of 32 tRNAs (4^2 X 2) are required to translate all 61 codons.
The translation machinery
mRNA, tRNA, ribosomes, soluble protein factors (initiation factors, elongation factors, termination factors)
tRNA is composed of
amino acid arm, Tψ (psi) C arm, Anticodon Arm, Anticodon, D Arm
uridine is found in what part of tRNA
anticodon arm
dihydrouridine is found in what part of tRNA
D arm
pseudouridine (ψ) is found in what part of tRNA?
TψC arm
tRNA 2D structure
cloverleaf
Why do we have tRNA instead of just RNA? (delete this)
because tRNA adds a benefit, regular RNA gets stiff, tRNA is more flexible (Temperature decrease -> motion decrease -> need tRNA)
Difference between uridine vs dihydrouridine
uridine has an aromatic ring (double bond, planar)
Dihydrouridine is more ____ than uridine
flexible
tRNA 3D structure
an L in 3D
Ribosomes are ….
RNA-protein machines
S =
svedberg (sedimentation rate), correlates with size
Eukaryotic ribosome
two subunits, larger than bacterial ribosomes (proteins are larger)
Bacterial ribosome
two subunits, smaller than eukaryotic ribosome
rRNA are key for ________ function
ribosomal
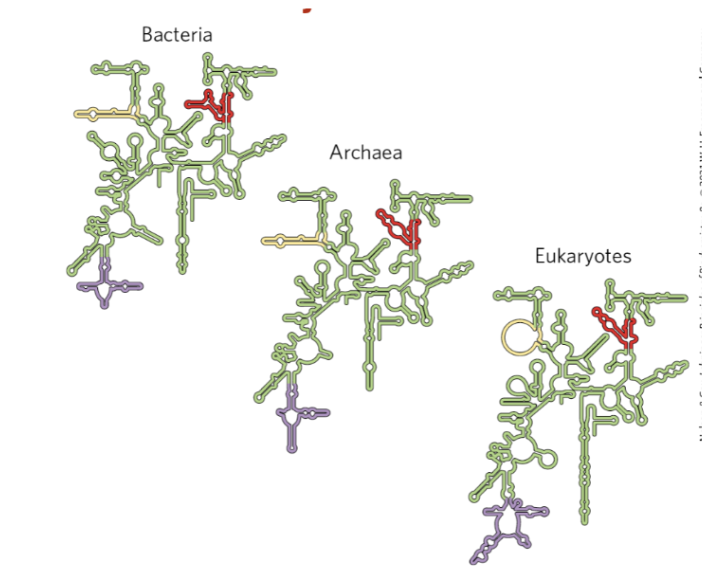
secondary structure of rRNA
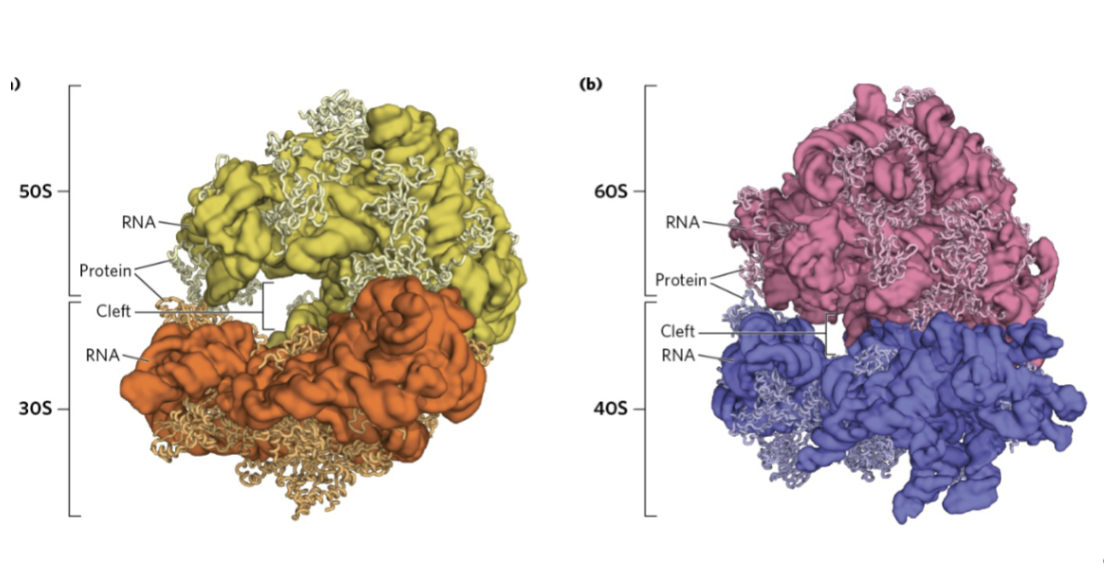
folded ribosome structure
Ribosome structure- Ribosomes are made of…
RNA, protein, and a cleft
Assembly of ribosomes in eukaryotes begins in the ____ (location)
nucleus
Assembly of ribosomes in eukaryotes-location goes from where to where
nucleolus -> nucleoplasm -> cytoplasm
Protein synthesis 5 major steps
activation of amino acids, initiation, elongation, termination and ribosome recycling, folding and posttranslational processing
Protein synthesis- stage 1
aminoacylation reaction or tRNA charging
Protein synthesis- stage 1 overall
amino acid + ATP + tRNA <-> aminoacyl-tRNA + AMP + PPi
Protein synthesis- stage 1 Nomenclature
Ser-tRNA^ser charged; tRNA^ser uncharged
Aminoacylation is carried out by
aminoacyl-tRNA synthetase
Aminoacylation-____ here is essential
accuracy
Protein synthesis- stage 2
initiation
Protein synthesis- stage 2 steps
30S binds IF-1 and IF-3, mRNA binds positioned by 16S rRNA, if IF-2 GTP binds 30S subunit, fMet-tRNA^fmet binds, which base-pairs with start codon, 50S binds IF-2 GTP hydrolyzed, IF-1, IF-2, IF-3 released, 70S initiation complex
An RNA-RNA interaction with ____ ____ positions ____ on the _____ in prokaryotic mRNA
16S rRNA positions the mRNA on the ribosome
The eukaryotic initiation process
scanning to 1st AUG from 5’
The eukaryotic initiation process
factors binding to subunit , propagate along the message until you fit AUG
Protein Synthesis- Stage 3
elongation
Protein Synthesis- Stage 3 steps
1) binding of aminoacyl-tRNA to A site 2) peptide bond formation 3) translocation
Protein Synthesis- Stage 3.1 steps
EF-Tu GTP binds to aa-tRNA^aa, aa-tRNA^aa-EF-Tu GTP binds to A site, GTP is hydrolyzed EF-Tu GDP dissociates, EF-Tu GTP is regenerated
Protein Synthesis- Stage 3.2
peptide bond formation
Protein Synthesis- Stage 3.2 steps
A-site amino acid’s NH2 attacks P-site amino acid -> peptide bond forms -> chain transfers to A site
Protein Synthesis- Stage 3.3
translocation
Protein Synthesis- Stage 3.3 steps
amino acids shift from A binding site to P binding site catalyzed by EF-G-GTP, AUG moves from P to E site
Protein Synthesis- Stage 4
termination
Protein Synthesis- Stage 4 steps
stop codon enters A site -> release factor binds to UAG -> peptidyl-tRNA link hydrolyzed -> peptide leaves -> components dissociate
Protein synthesis is energetically
expensive!
AMP -> AMP + PPi occurs in what step/s
aminoacylation
GTP -> GDP + Pi occurs in what step/s
proofreading and translocation
___ high energy phosphate bonds per peptide bond
4
___ ____ for initiation per protein
1 GTP
Coupling of _____ and ____ in bacteria
transcription and translation
In bacteria, messenger RNA (mRNA) is protected by
ribosomes, acting as polysomes
Bacterial protein synthesis is _________, but eukaryotic protein synthesis is not
co-transcriptional
Protein synthesis-Stage 5
post-translational processing
Stage 5: Post translational processing examples
N- and C- terminal modification, loss of signal sequences, amino acid modification, disulfide bond formation, glycosylation, isolation, addition of prosthetic groups, proteolytic processing
Addition of carbohydrate side chains function and location
plays a key role in protein targeting, occurs in ER
Addition of isoprenyl groups
adds aliphatic chain, anchor otherwise soluble proteins to the membrane
Many antibiotics target
protein synthesis
Ribosomes have three binding sites
A (aminoacyl) site, P (peptidyl) site, and E (exit) site
Antibiotic examples
chloramphenicol, Cycloheximide, Erythromycin, Fusidic acid, Puromycin, Streptomycin, Tetracycline, Diphtheria toxin, and Ricin
Chloramphenicol
Inhibits peptidyl transferase on the prokaryotic large subunit
Cycloheximide
Inhibits peptidyl transferase on the eukaryotic large subunit
Erythromycin
Inhibits translocation by the prokaryotic large subunit
Fusidic acid
inhibits elongation in prokaryotes by binding to EF-G GTP in a way that prevents its dissociation from the large subunit