GCT
1/325
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
326 Terms
Contact dependent signaling
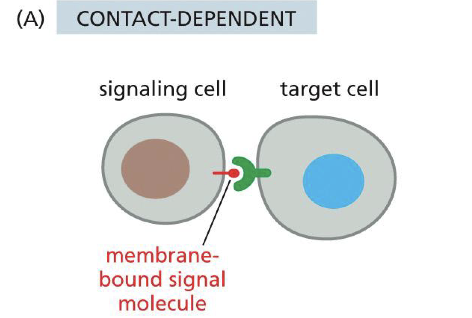
Paracrine signaling
Signaling cell --> secretes local mediator.
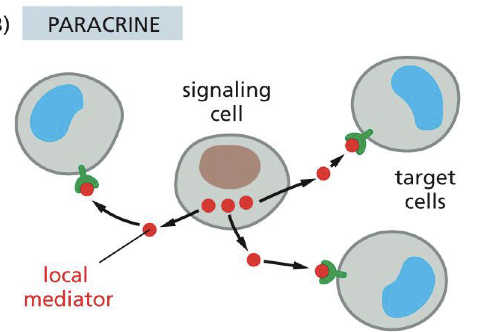
Endocrine signaling
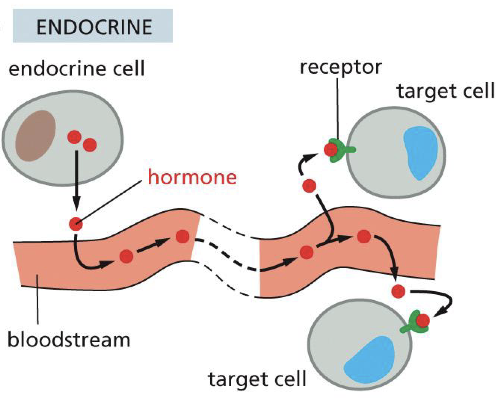
Synaptic signaling
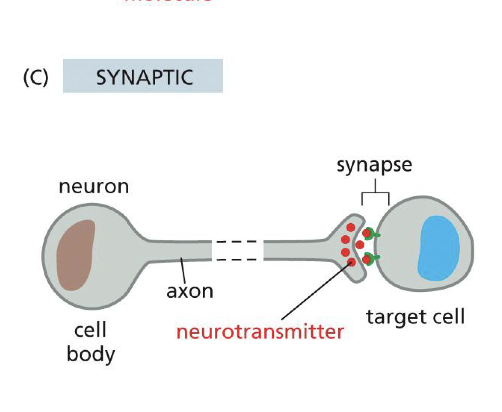
Name all Intracellular signaling On-Off switches
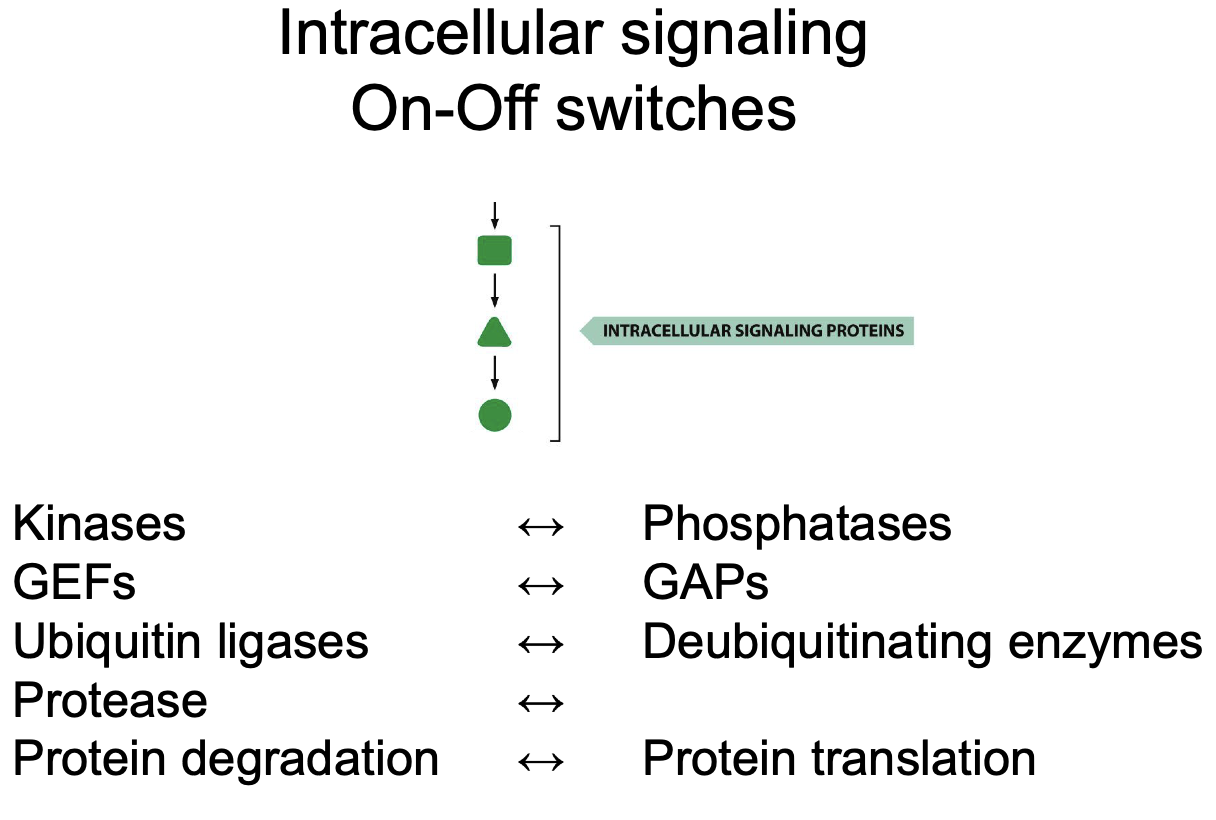
Switching on-off by G-proteins: Name all relevant players
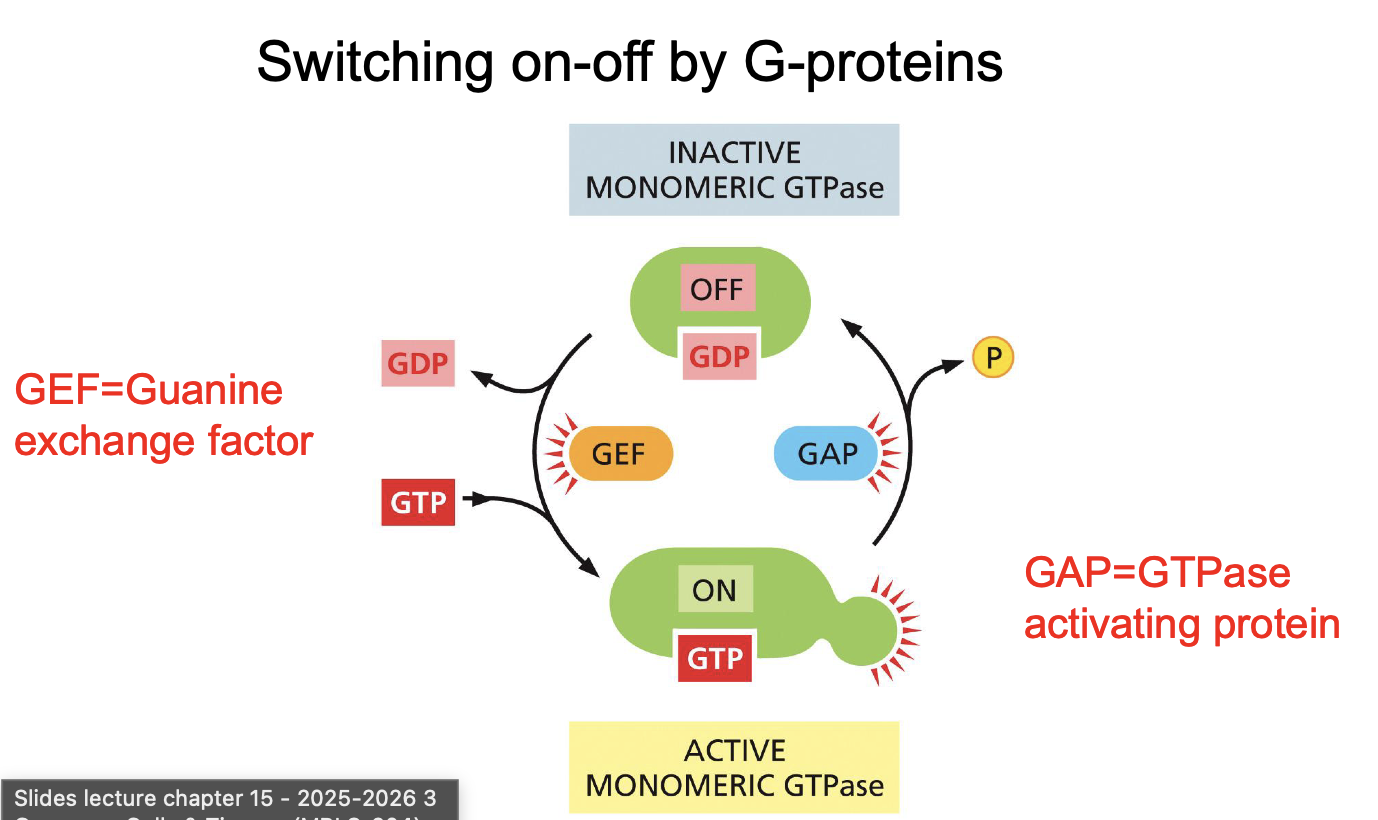
Switching on-off by ubiquitination: Name all relevant players and name all steps
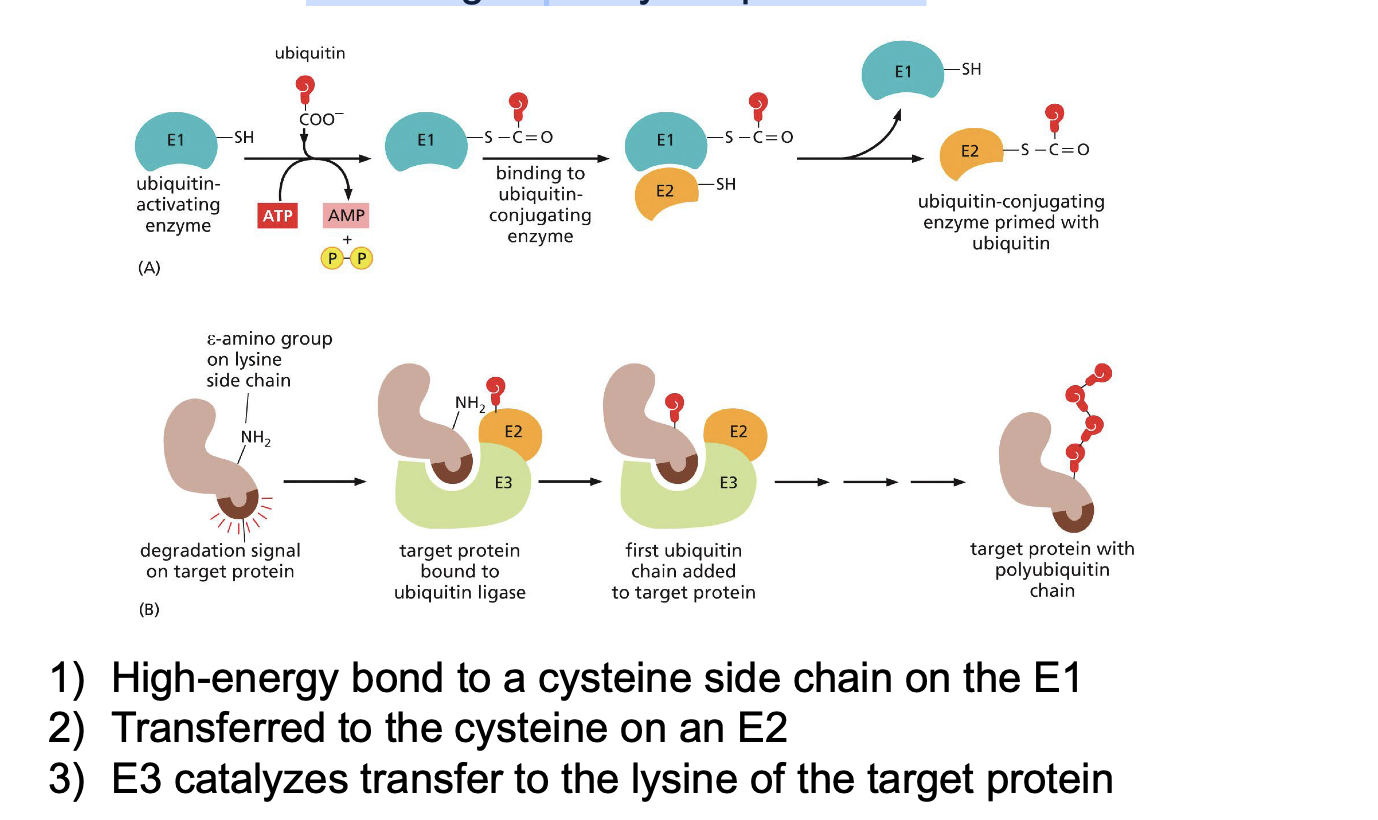
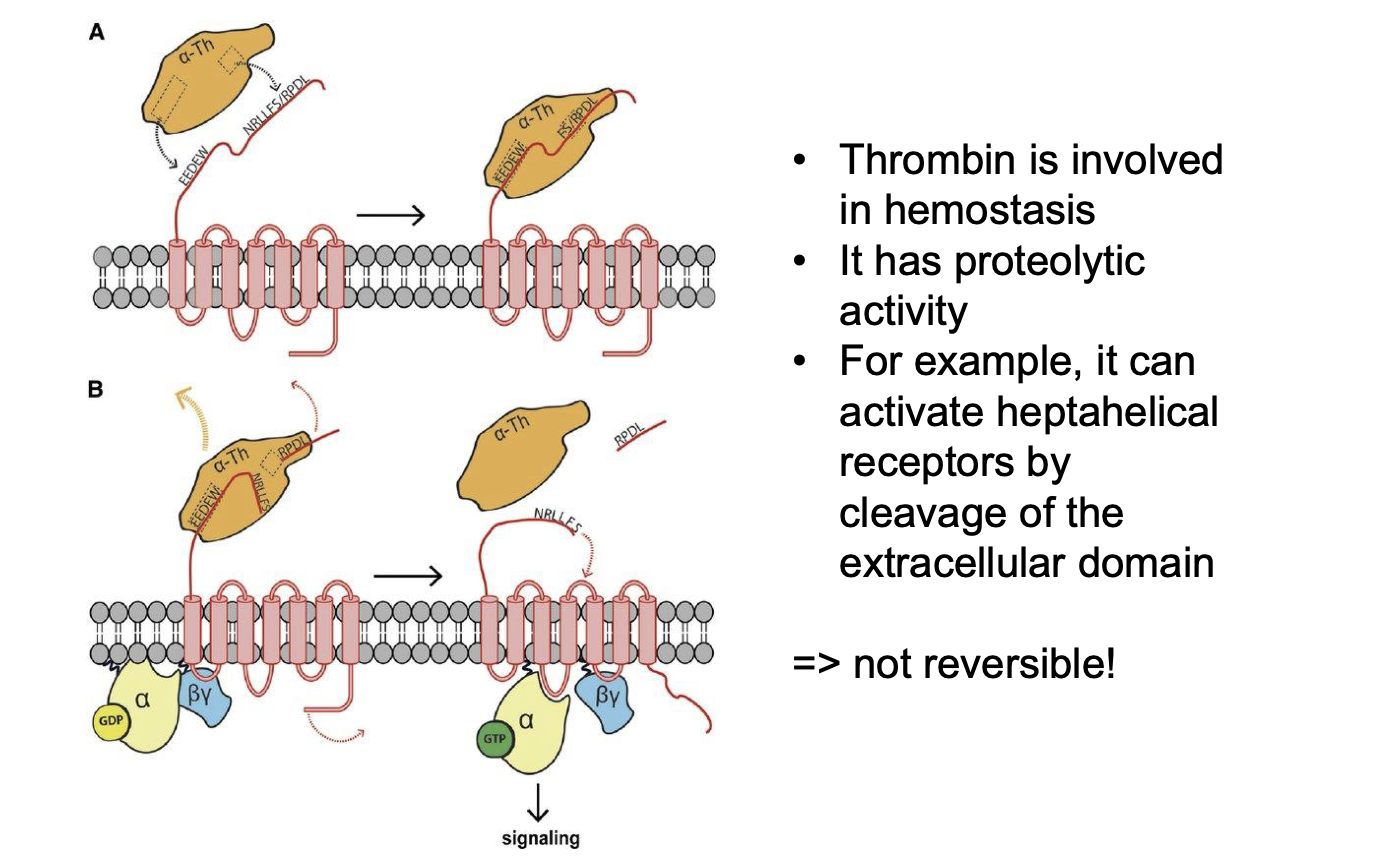
Switching on-off by proteolytic cleavage:
Name relevant proteins?
What part is cleaved?
Reversible?
Thrombin
Switching on-off by post translational modifications:
What 4 types of ranslational modifications:
(Same as intercellular pathway)
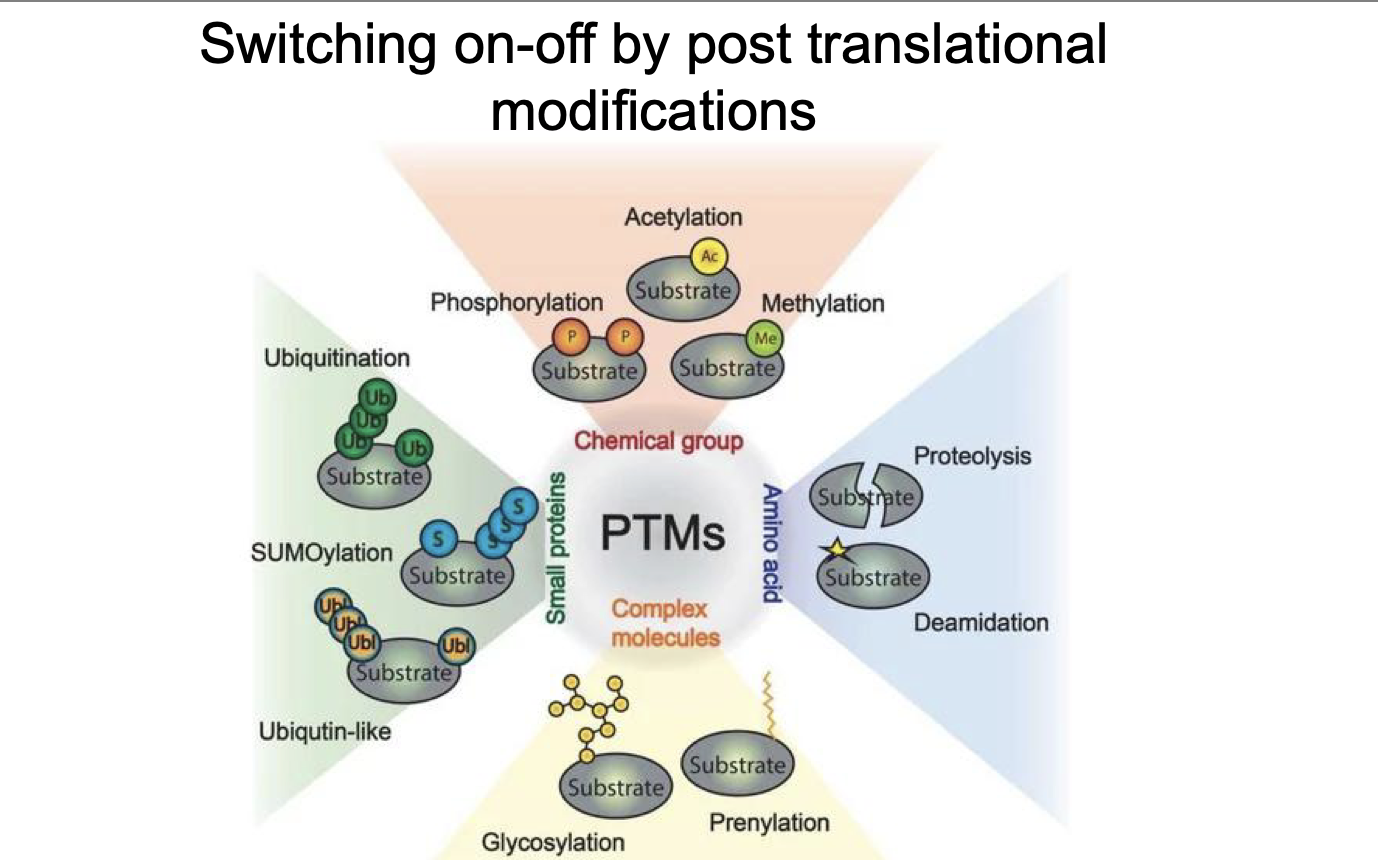
GTPase function:
Ras?
Rho?
Rab?
Ran?
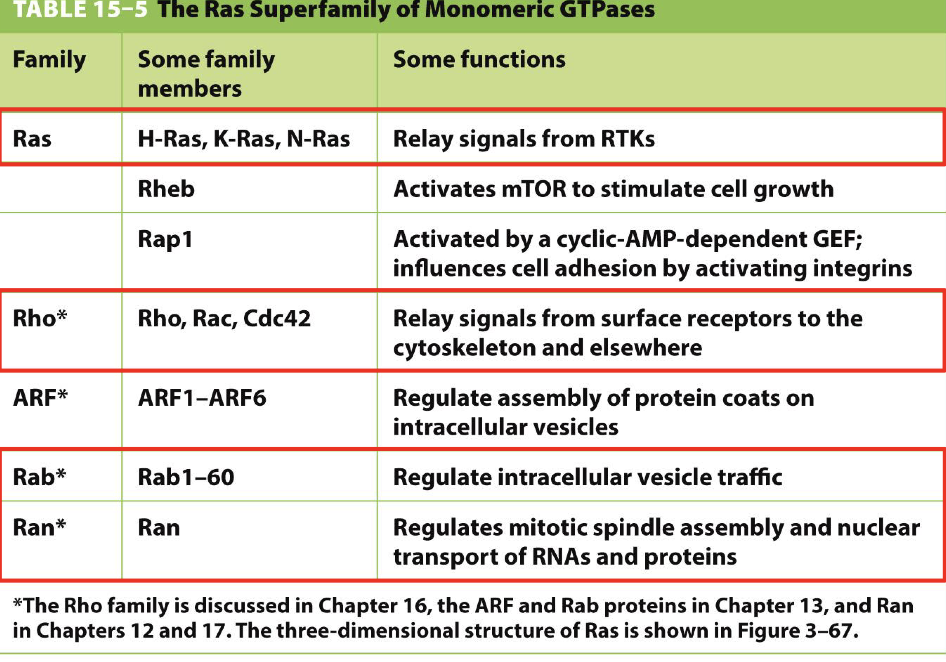
Receptor desensitization (pathways for destroying receptor) describe the following pathways:
Negative feedback?
Delayed feedforward?
Receptor inactivation?
Receptor sequestration?
Receptor destruction?
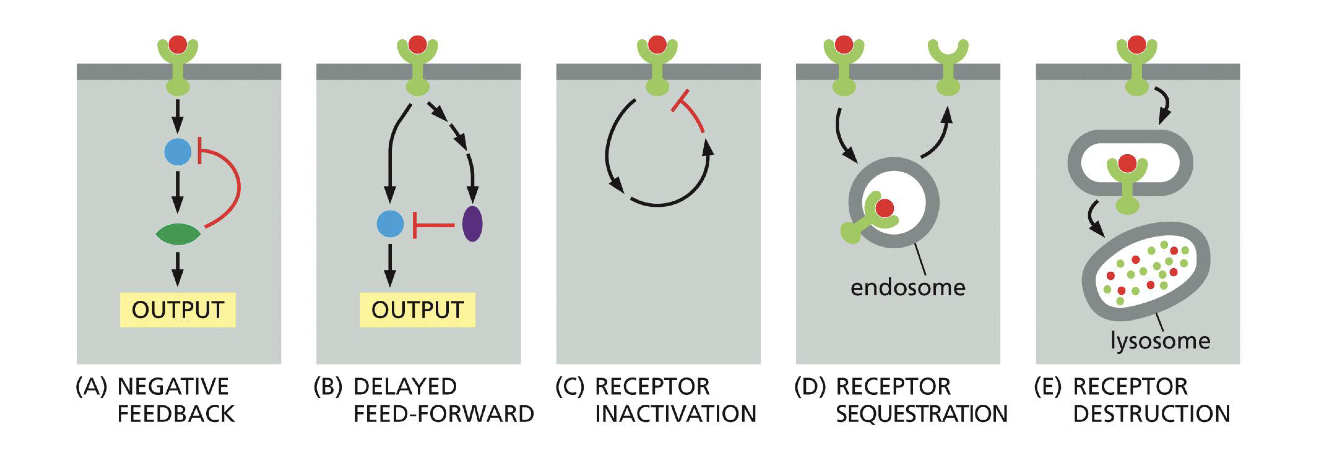
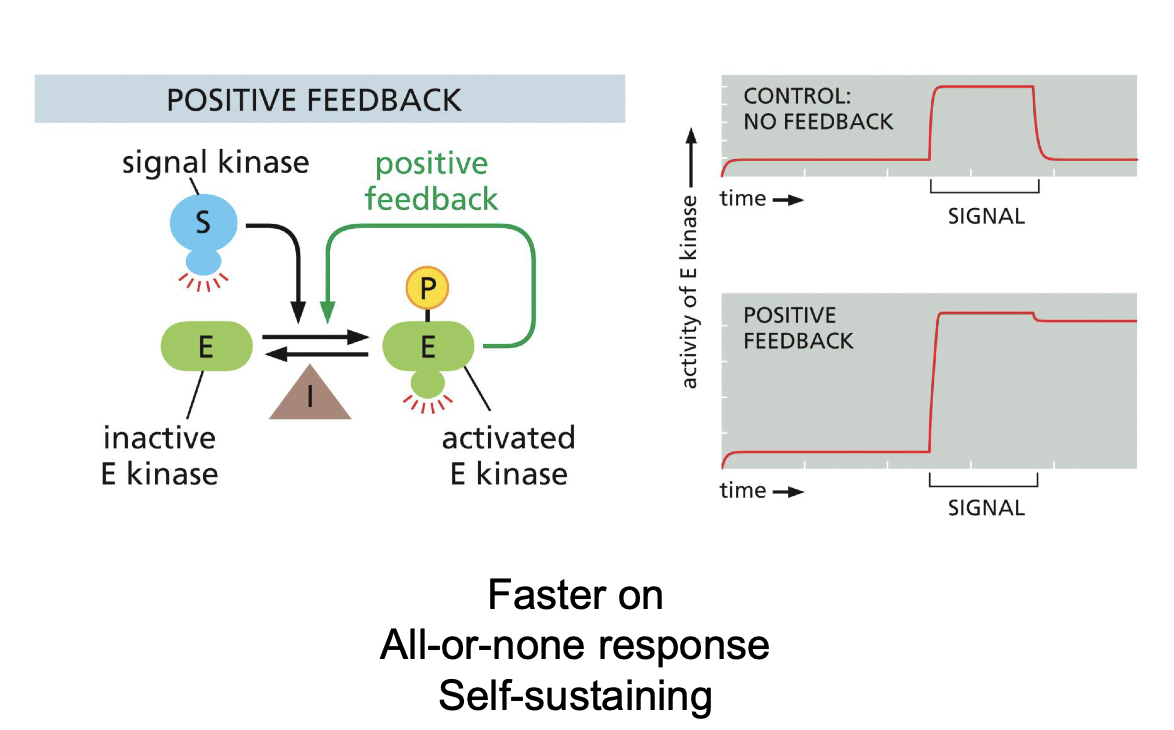
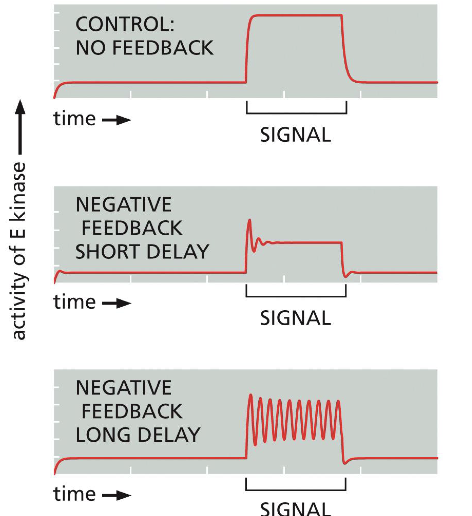
Predict graph of protein after Positive feedback in intracellular signaling VS normal graph.
Predict graph of protein after Negative feedback in intracellular signaling VS normal graph:
Negative feedback short delay (plus term of signal)?
Negative feedback long delay (plus term of signal)?
Short delay -> adaptation
Long delay -> oscillations
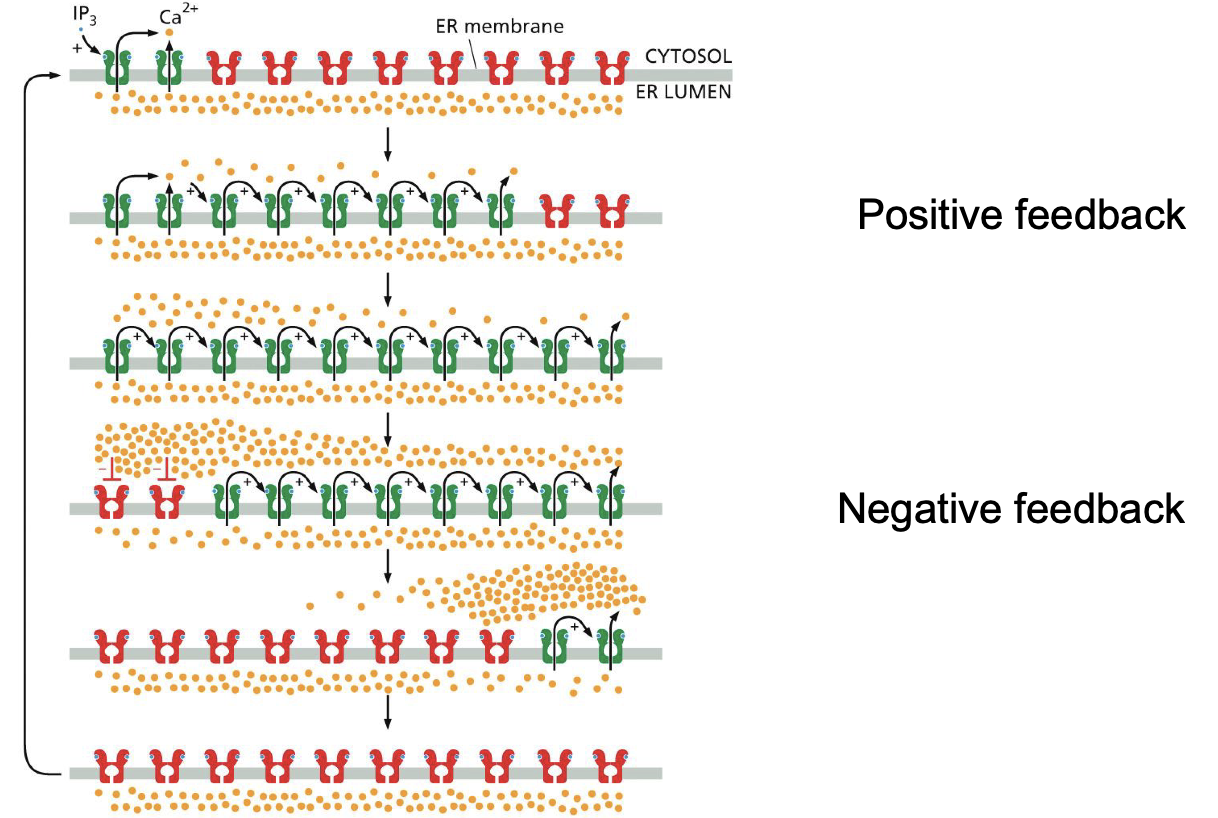
PLC signaling:
What does it induce
How does the feedback look like
Calcium oscillations
CamKII
Positive feedback
Negative feedback
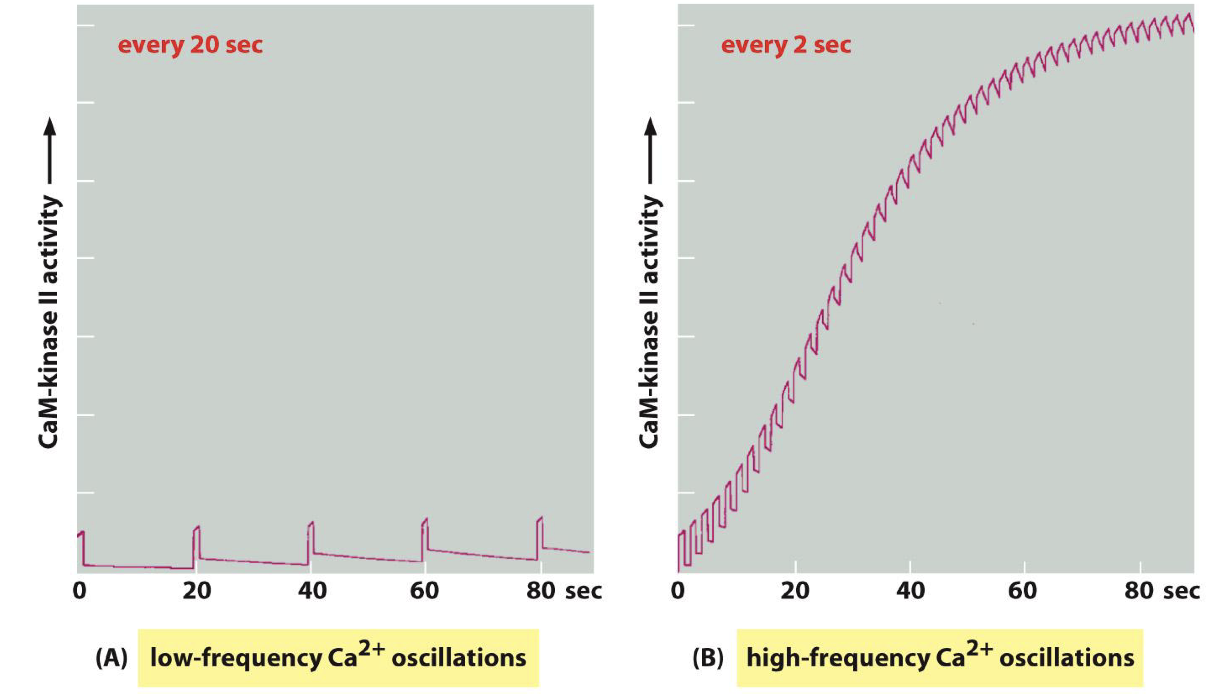
CamII
Low-frequency Ca²+ oscillation (every 20s)
High-frequency Ca²+ oscillation (every 2s)
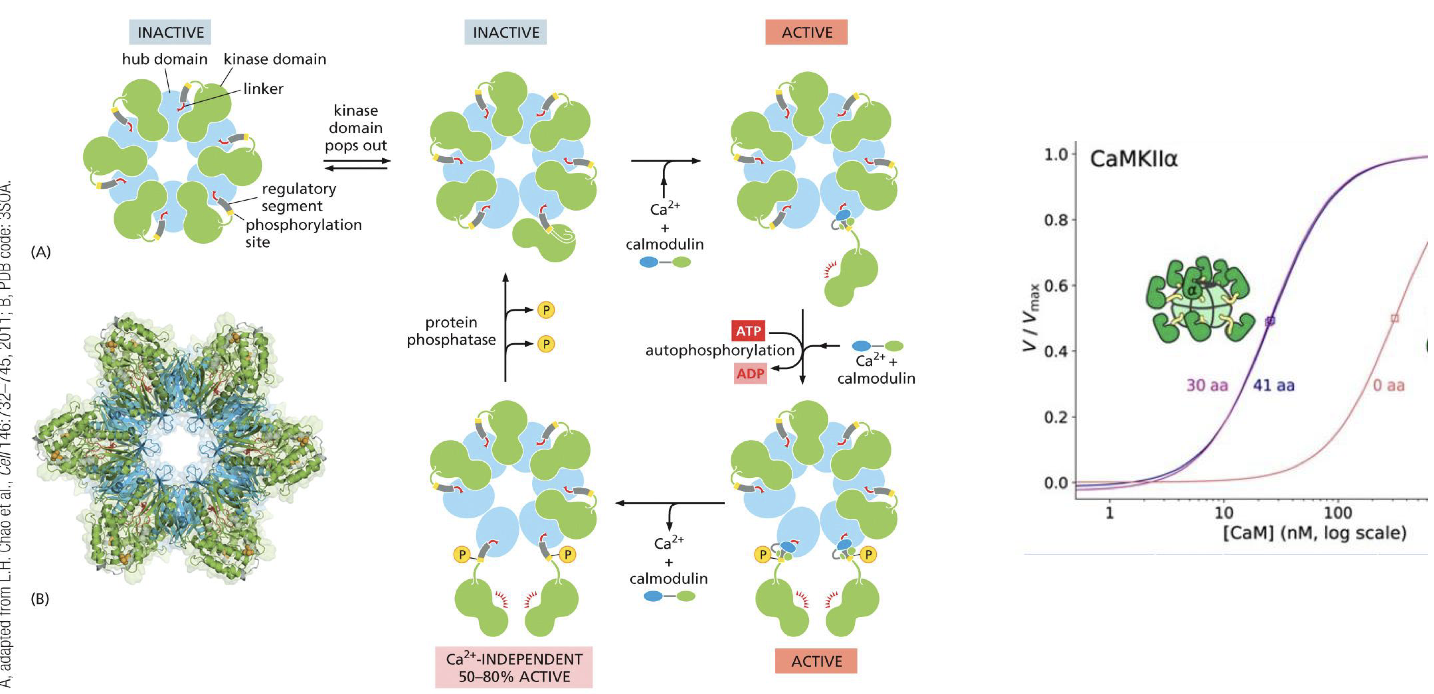
CamII:
Workings?
Key working enzymes?
What can CamII create?
High frequency calcium oscillations induce CamKII signaling
Calmodulin
Ca2+ activates
Phosphate active
Can create an extra synapse
Name the 3 types of formation/locations of intracellular signaling complexes.
And draw them. On vs Off version
Signaling complex on scaffolding protein.
Signaling complex on activated receptor.
Signaling complex on phosphoinositides
Check with powerpoint…
PI3 signaling:
For what?
Whole cascade, with protein names and domains.
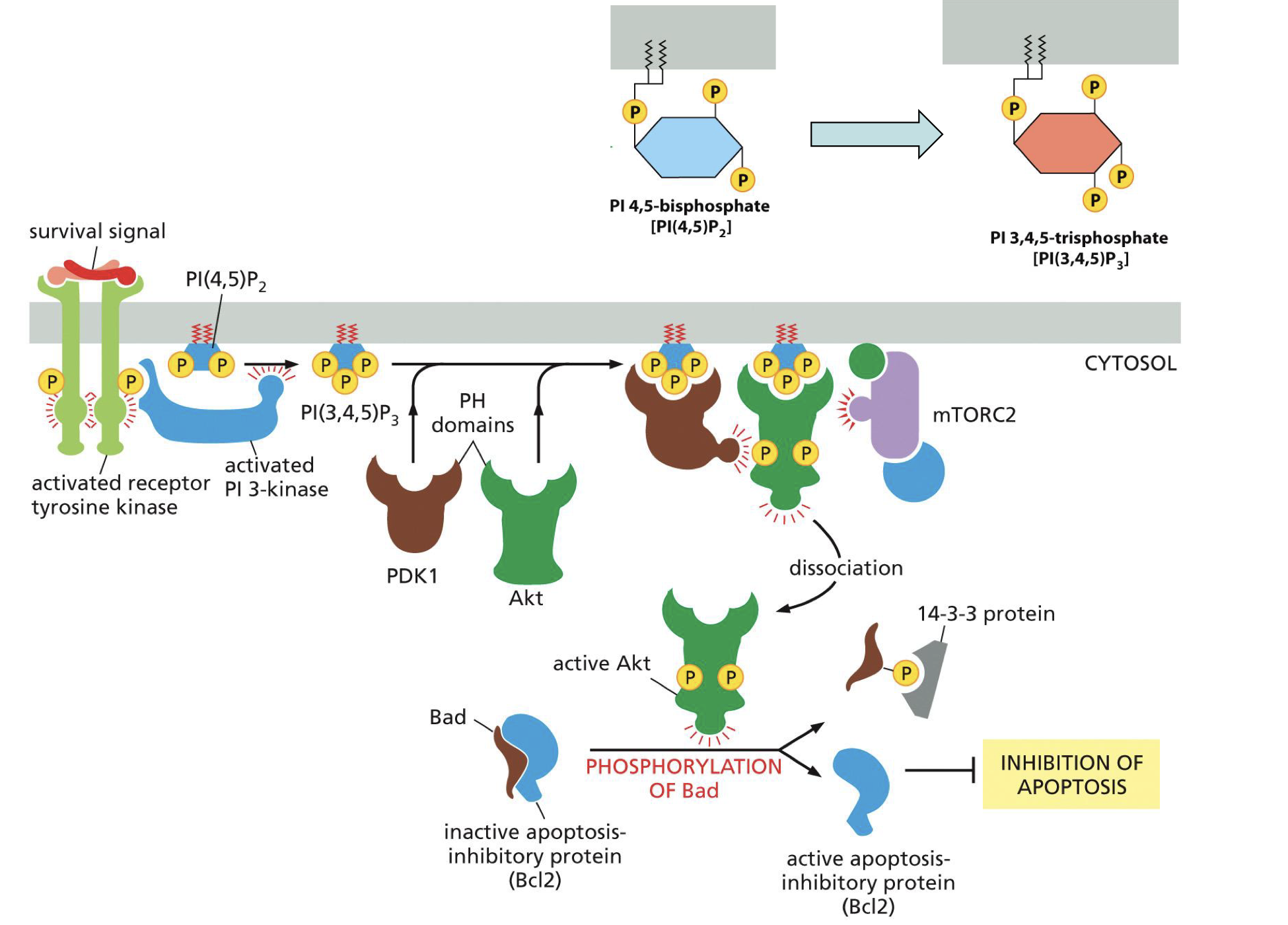
For cell survival
Survival signal, activated receptor tyrosine kinase,
activated PI 3 kinase,
PI(4,5)P2 → PI(3,4,5)P3,
PDK1 + Akt,
mTORC2,
active Akt,
phosphorylation of Bad on Bcl2 complex,
active Bcl2.
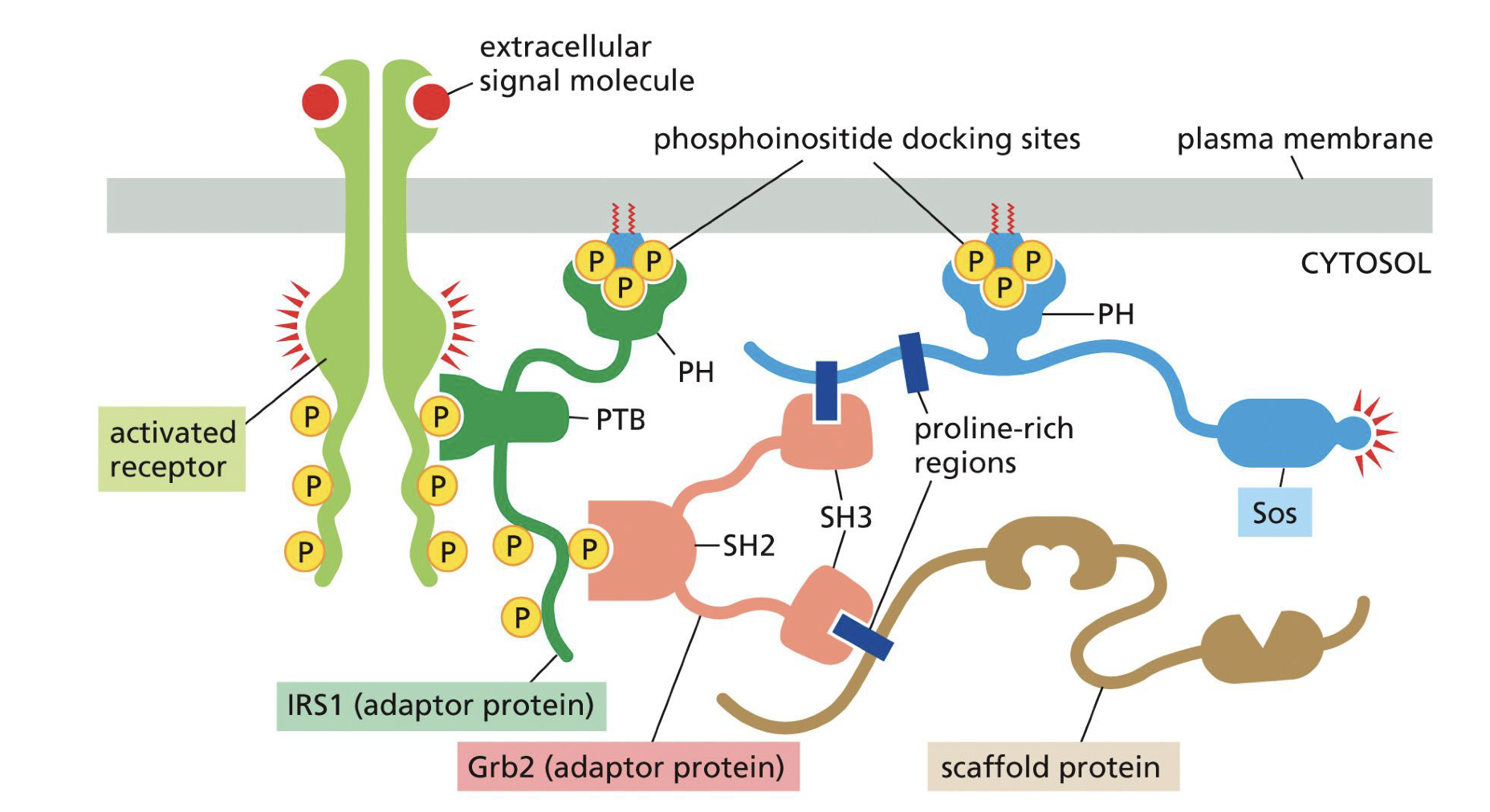
Protein domains specify connections (Proteins interact with each other by domain)
Define the following domains, function/role and to which domains belong on same protein:
PH
PTB
SH2
SH3
All 1 protein
PH = Plextrin Homology: phospholipid docking, localizes to membrane
PTB = PTB: PhosphoTyrosine Binding: links to peptide of RECPTOR with phosphorylated tyrosines (pTr).
All 1 proteinSH2 = Sarc Homology binds to phosphorylated tyrosines. (Can also bind to receptor)
SH3 = Sarc Homology binds to proline rich sequences, assemble signaling complex (like scavenging complexes, so to set up actual complex)
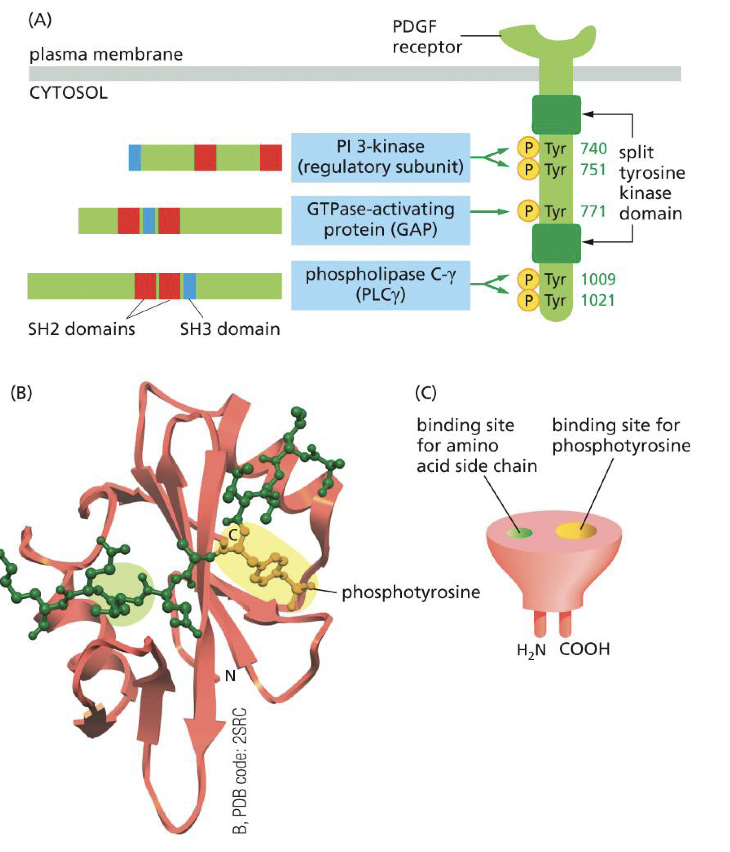
SH2 domain
To which receptor does it bind?
Does SH2 have high specificity?
Does binding location on SH2 matter
To activated RTKs
Tyrosine is ofc very important (SH2 binds to phosphorylated tyrosines), but yes is important but many mutations/types of the SH2 domain
Yes depending where on RTK leads to different cell signal.
So when certain ligand binds on RTK leads to certain phosphorylation which leads to certain cell response
G-proteins, give general mechanism plus what subunits (with function).
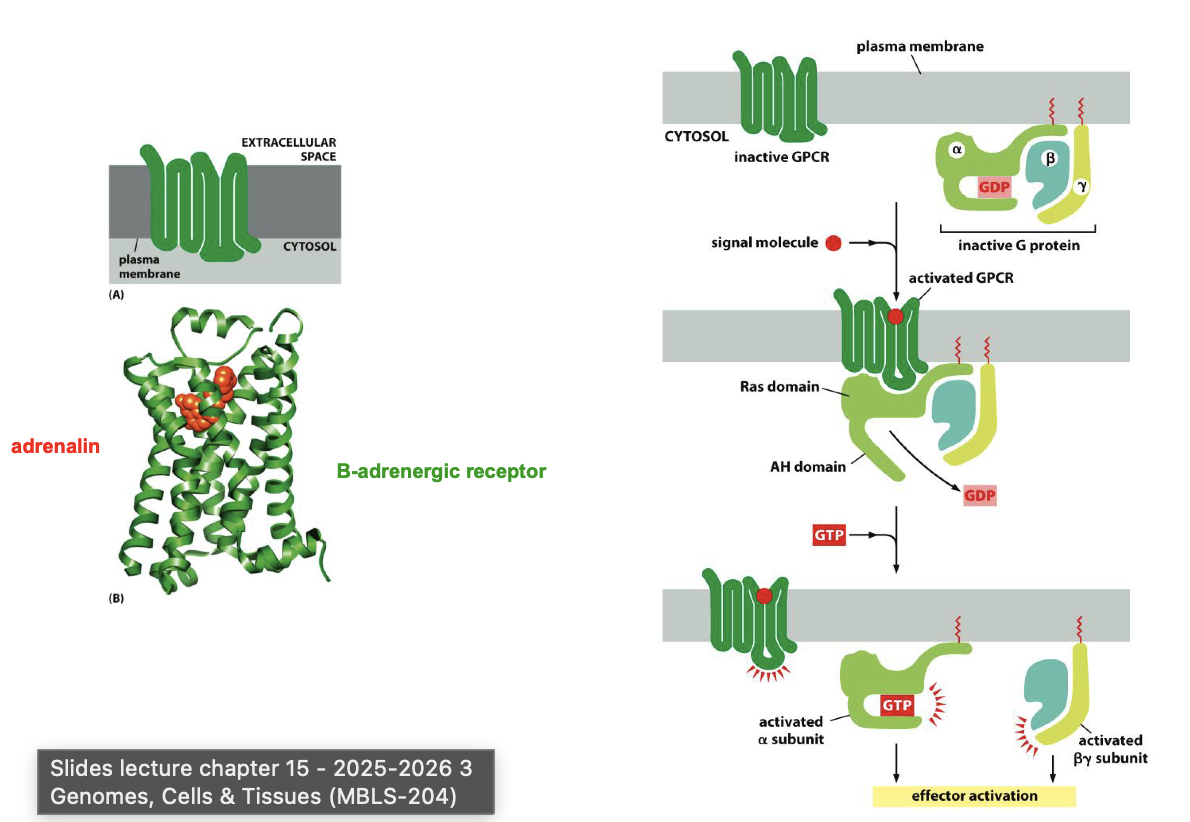
G proteins name effect of following types:
Gs
Gi
Gq
Gs = activate Ca2= channel
Gi = Inhibits adenyl cyclase (I = inhibit) + activates k+
Gq = activates phospolapse C-beta
G protein: PKA signaling:
Why/function
Relavent enzymes/cascade
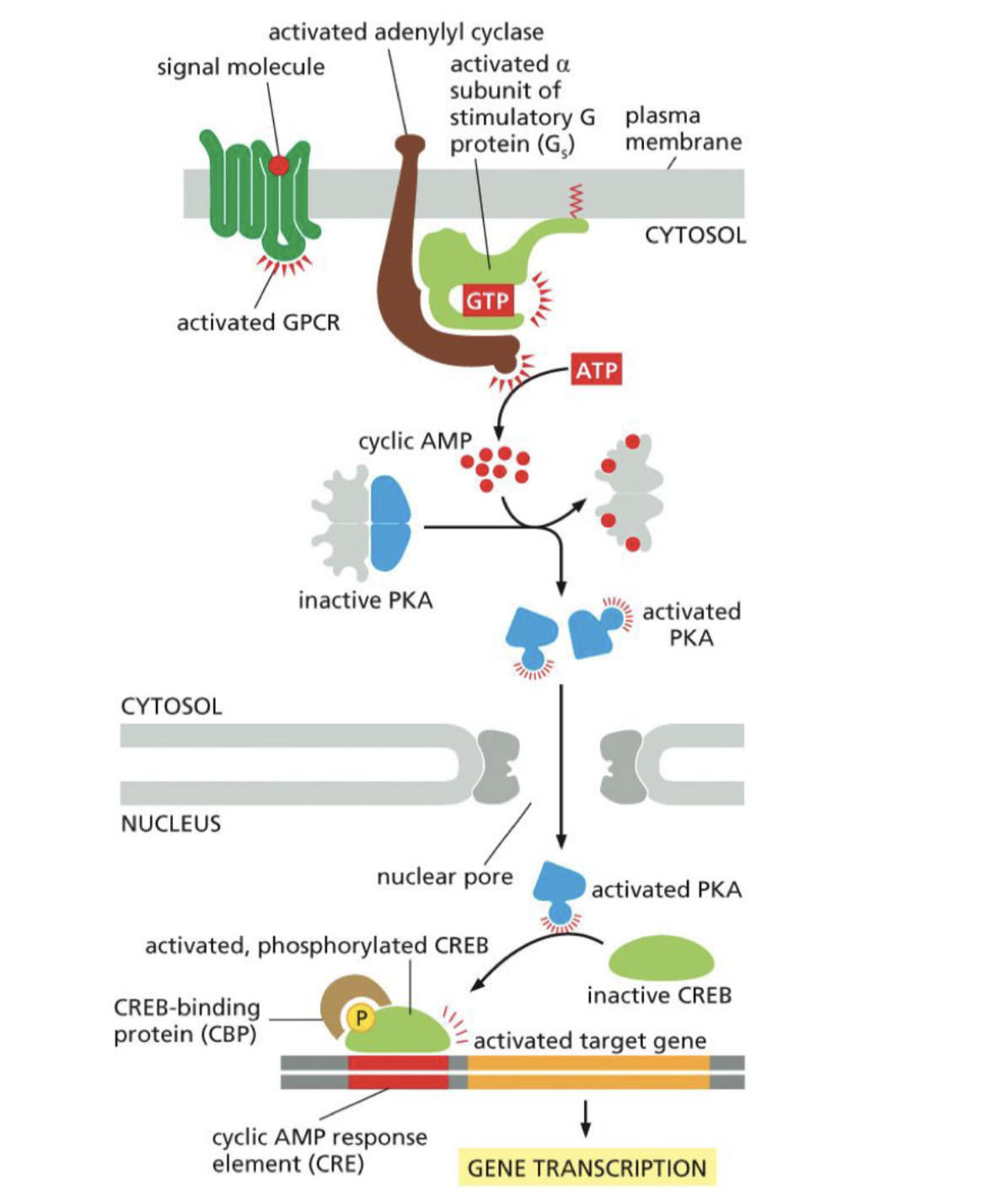
How gene transcription can be activated from outside signail:
Active G-protein
cyclic AMP inhibits by holding it PKA
Active G-protein with ATP inhibit cyclic AMP
PKA let go, activated PKA
PKA activates gene transcription
Arrestin
Function
Mechanism
Inactivates GPCR (g-protein coupled receptors)
Happens after phosphorylation of GPCR:
Binds to GPCR, so G-protein can not bind
Marks for decration
GTPases
Rho
Ras
Rab
Ran
Rho = relay signals from cell membrane to cytoskeleton.
Ras = relay signals from RTKs
Rab = Regulate vesicle transfer
Ran = Regulate mitotic spindle
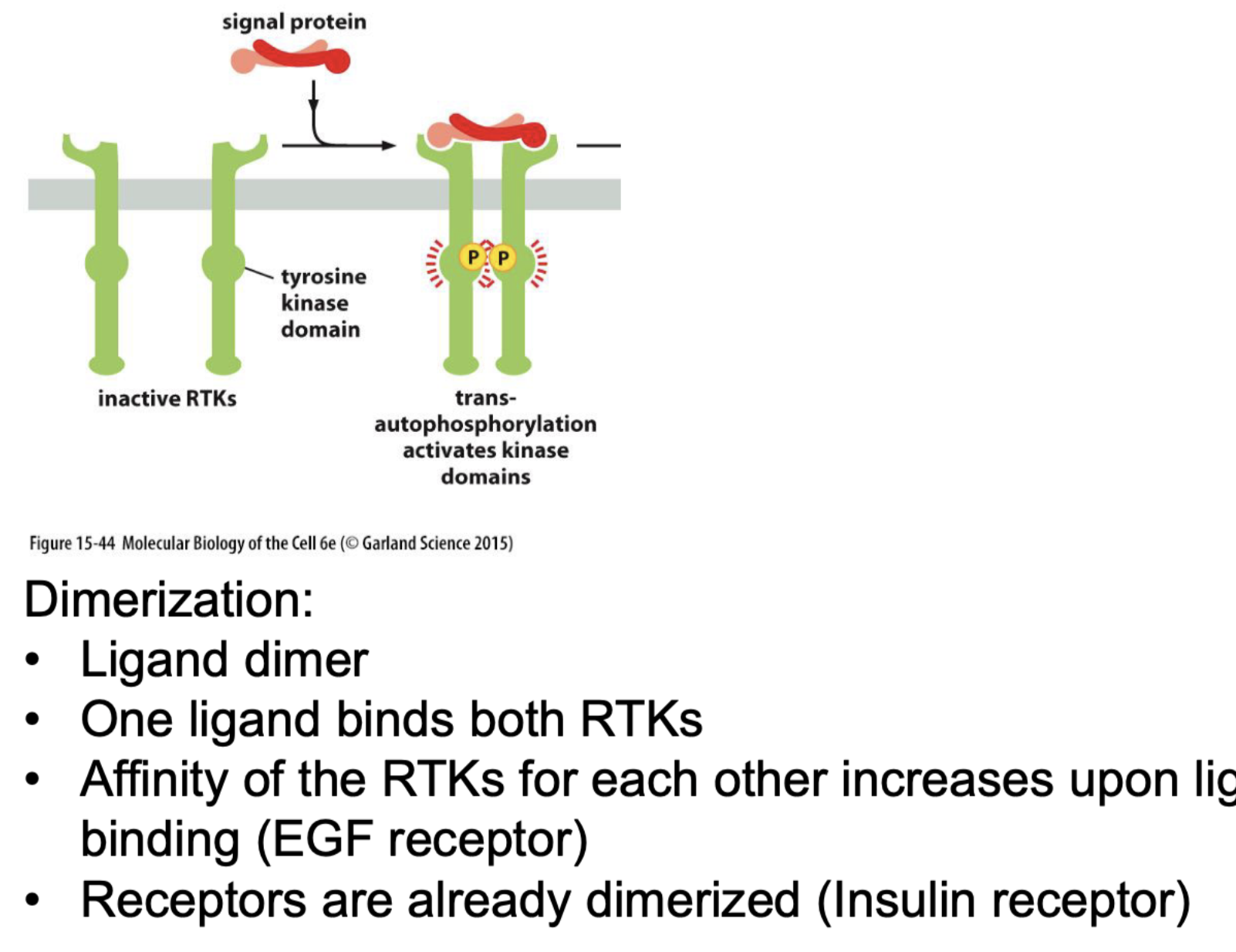
RTKs receptor = tyrosine kinase receptor
Do you have different types???
How does it dimirise?
What happens next?!?!?!?!
Yes many (know growth one EGF as described earlier)
Mechanism:
Ligand bind to both, dimers.
bringing them together, RTK affinity for each other increases when closer together.
Then they bind through a tyrosine kinase domain.
Als een relatie therapeut dan trouwen ze en activeren ze en krijgen kinderen.
Ones dimerized:
SH2 and PTB can bind to receptor, to create the intracellular signaling complexes, to activate protein relay signal.
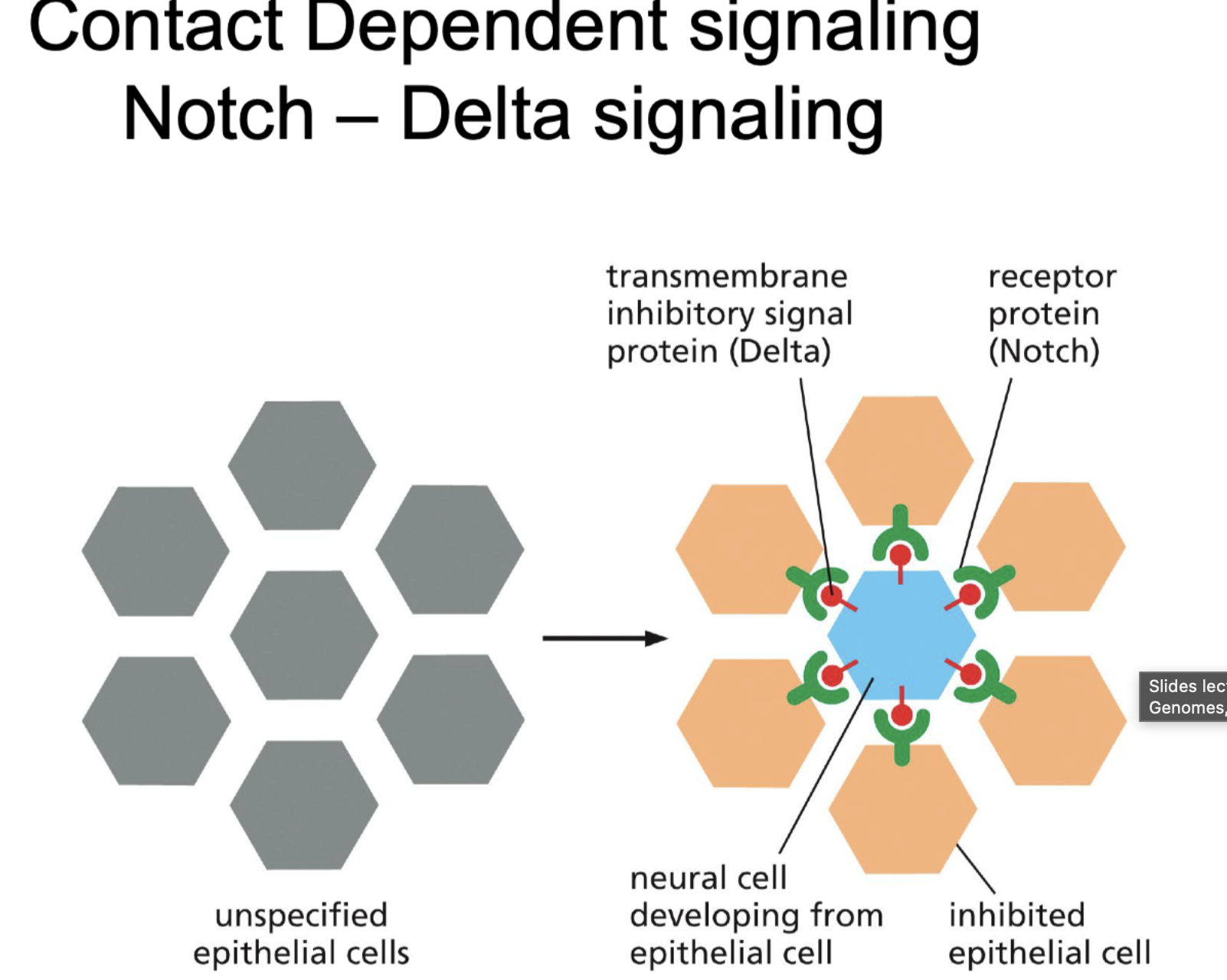
What is noch delta signialing?
Notch?
Delta?
Function
Notch workings
Its contact dependent signaling.
Notch = receptor protein (is heavily glycocilated)
Delta = Inhibery signal
So that only one cell turns into neuron and the rest can not because they are inhibited.
If notch does not bind, part of receptor breaks of and is used to activate gene expression
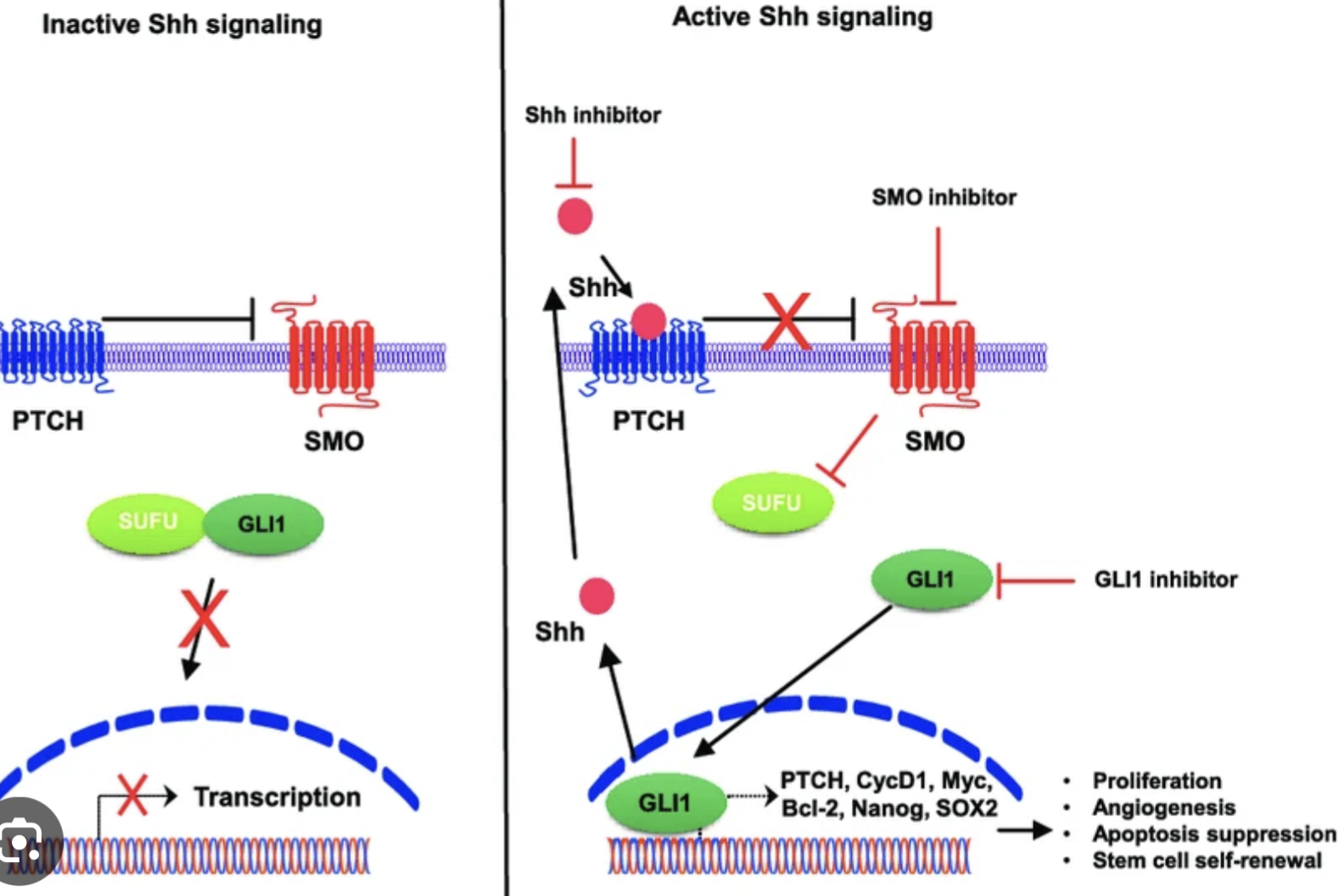
Hedgehog signaling:
Function?
Is it localized?
How does it work?
Forms neural tubes
Creates polilized segments (like stripes)
It is localized.
Hh ligand inhibits → Ptch, what stops the inhibition of Smo → Smo activates Gli2 protein → activates Hedgehog signals.
So controlled by HH
Name all cell cycles + basic functions
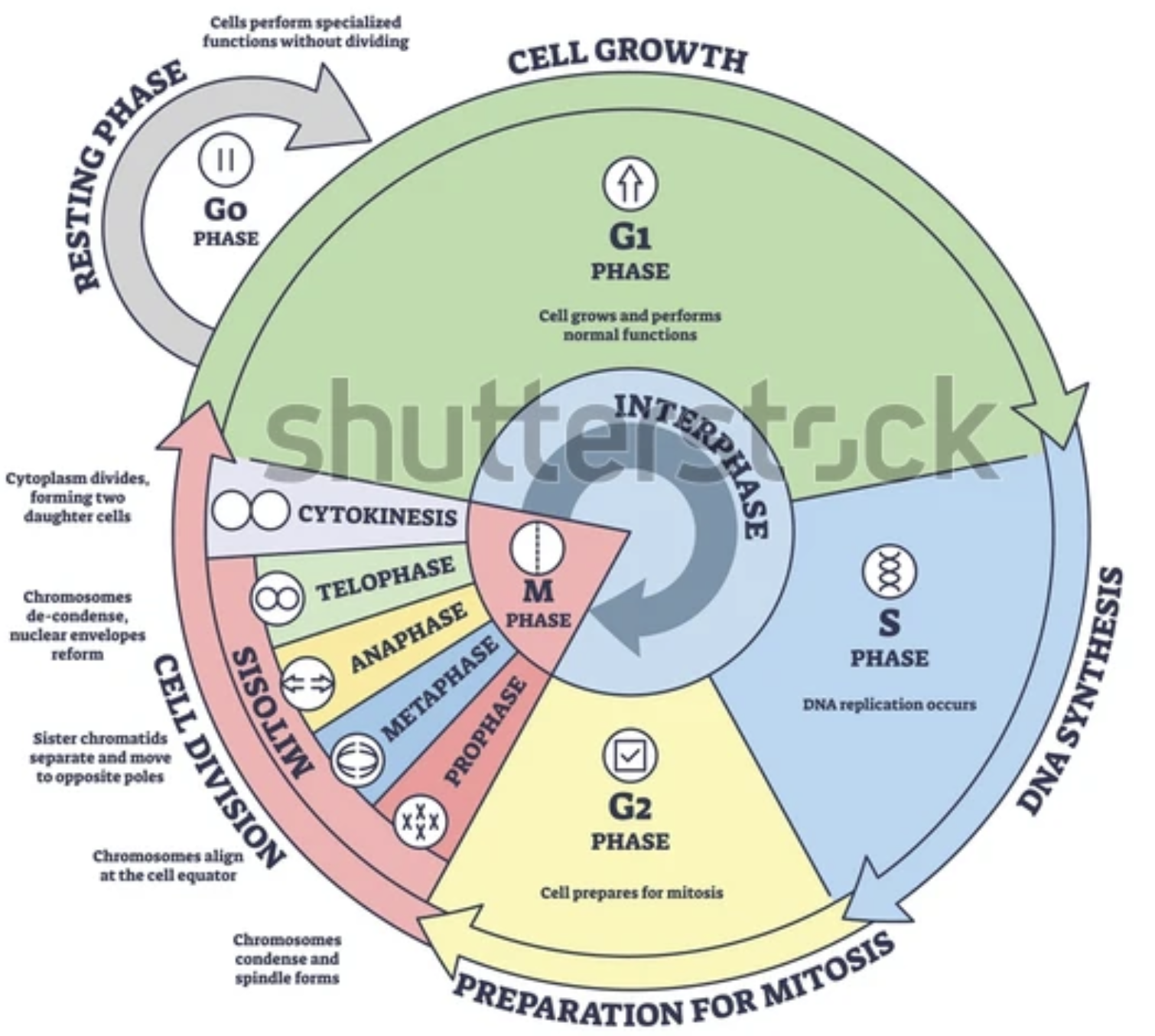
CDKs (Cyclin dependent kinases) and cyclins form complexes together
Are there different Cyclin CDK combination complexes?
For what CDK1?
Yes, depends on cell cycle
CDK1 = for M-fase and G2 fase depening on cyvlin
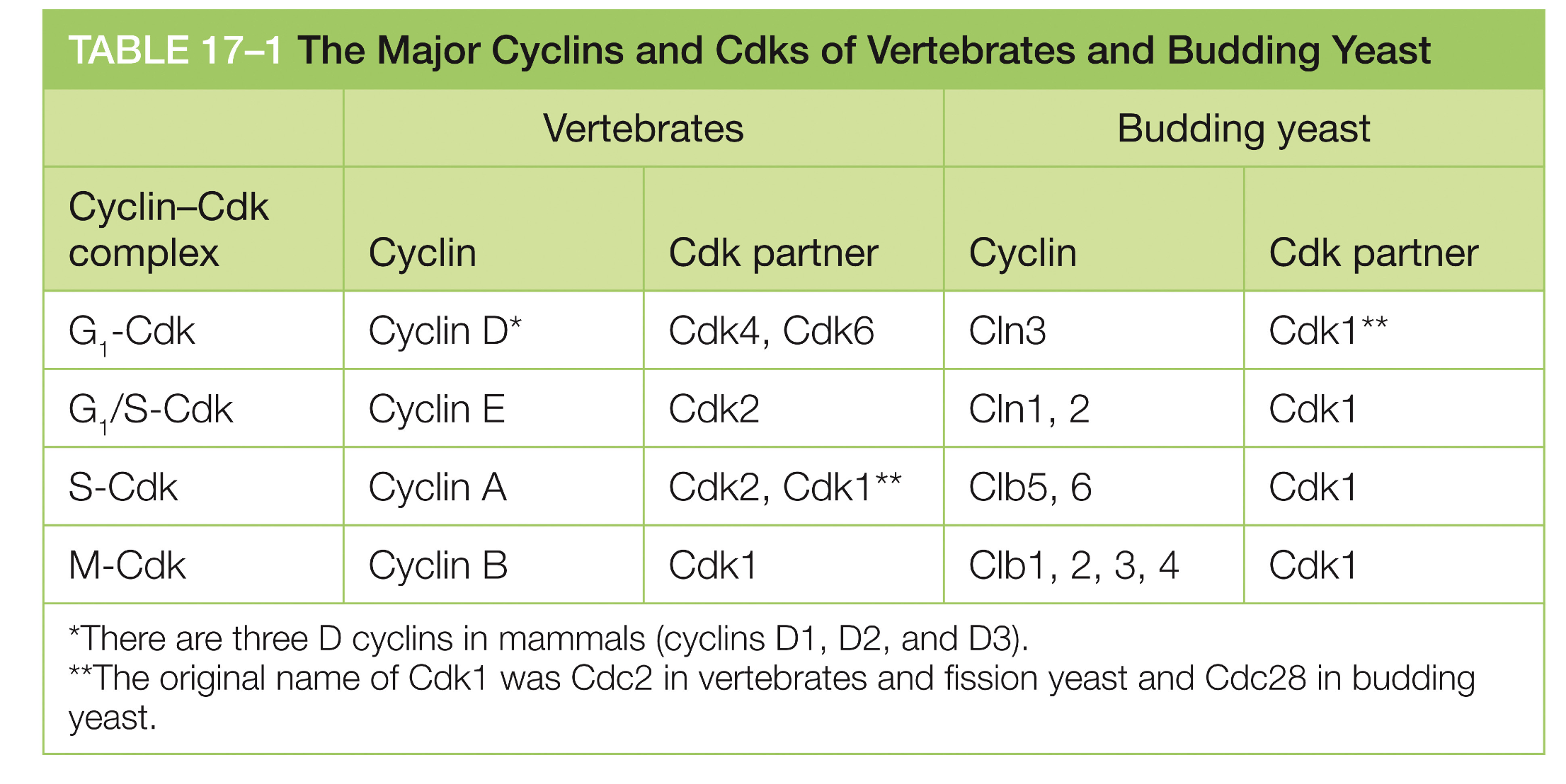
What are the levels of CDKs regulation:
Gene expression of cyclin type
Positive and negative CDK phosporylation
CAK and T-loop
Wee1
Cdc25
Transcription regulation of CDK?
Transcription of Cyclin in regulated (Not of CDK)
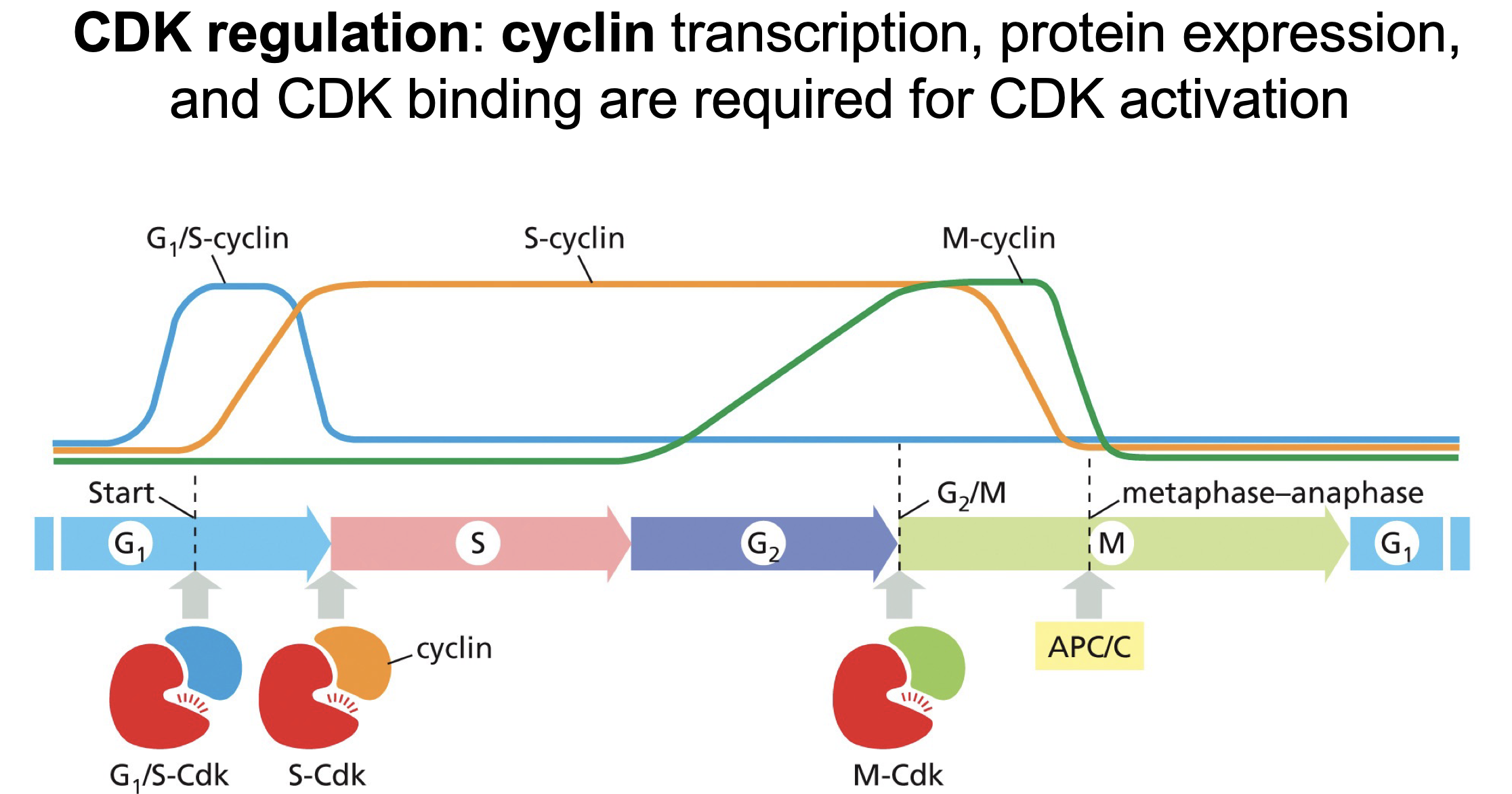
Now CDK and Cyclin form complex:
How does it form a complex?
Which enzyme regulates this.
Does it already work?
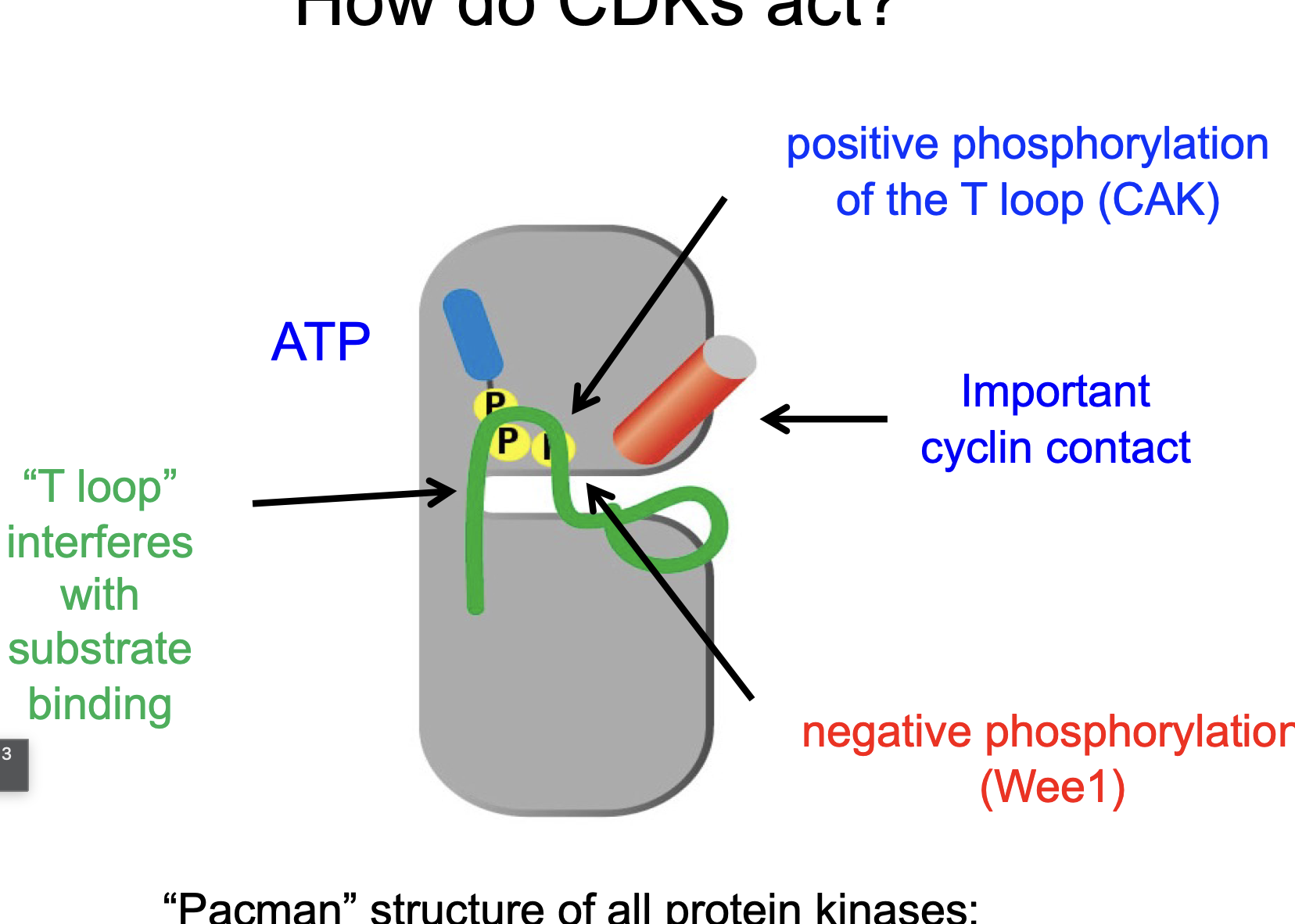
CAK phosphorylates the T-loop (an alpha helix) thereby unwinding the T-loop and allowing binding between the 2.
CAK of poep begins,
No
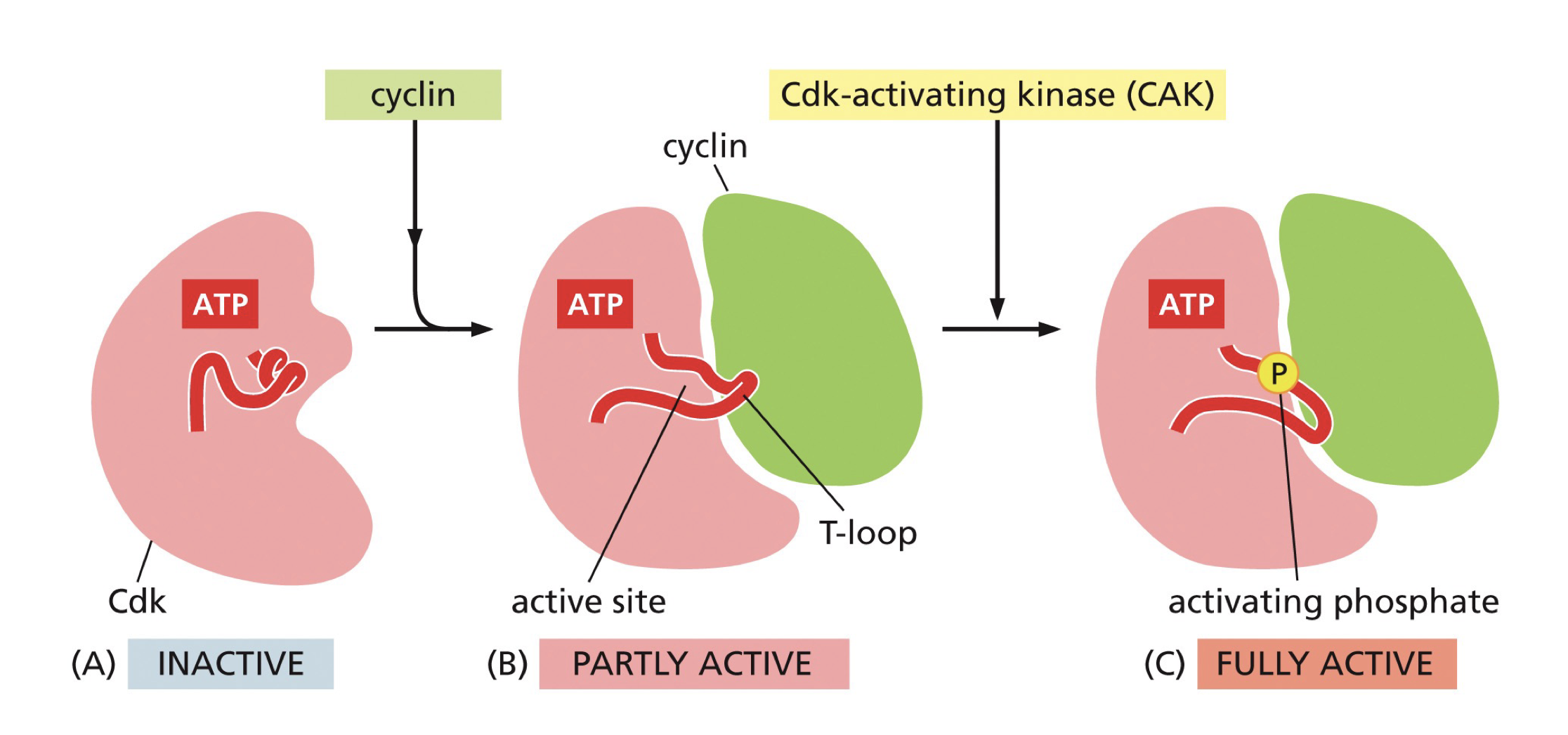
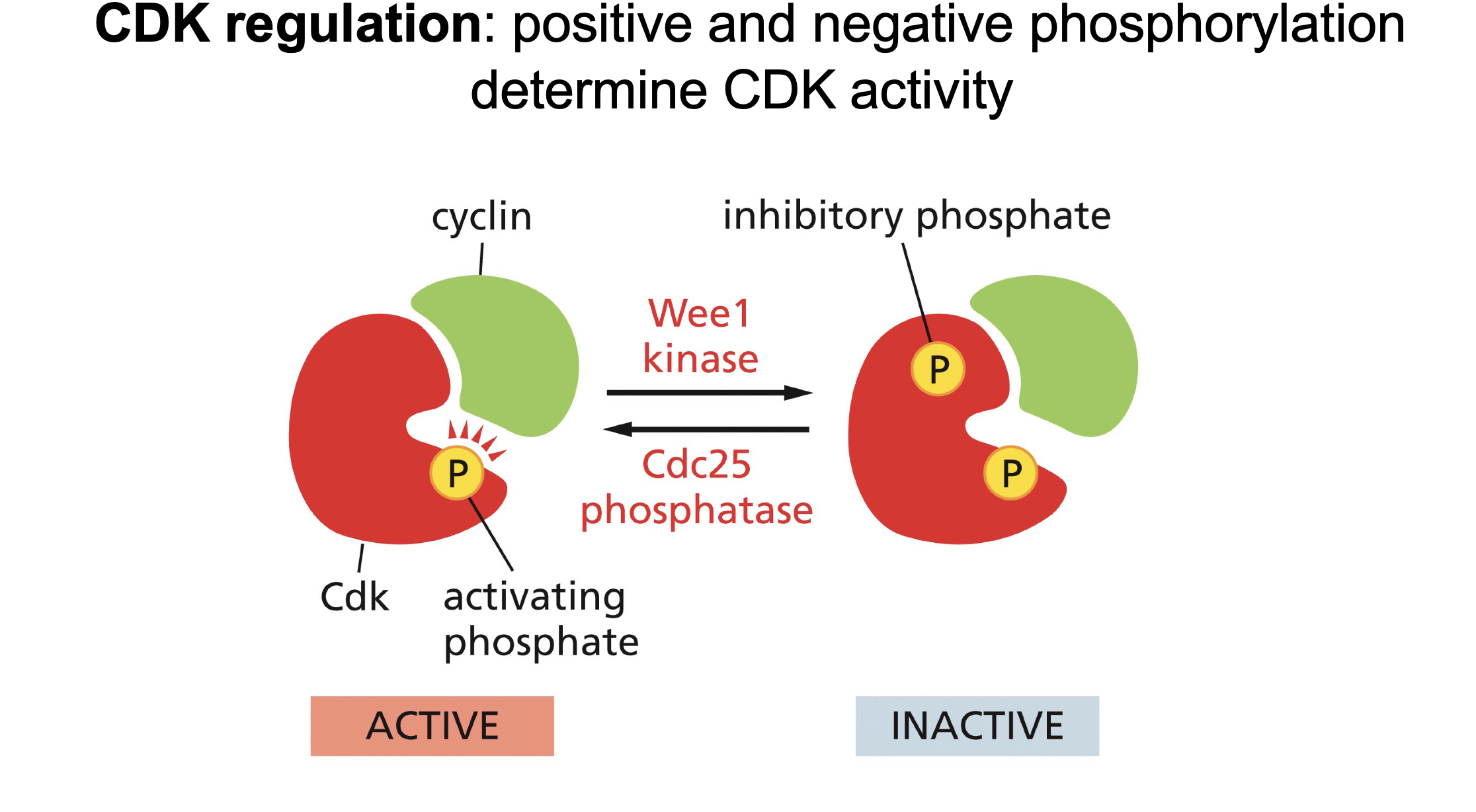
Phosphorylation regulation of CDK-cyclin complex:
How many phosphors for on, How many phosphors for off (plus points if you say location)?
Which enzymes?
Switches between 2 (off) and 1 (on) phosphate:
Begins with 2 phosphates, Cdc25 removes phosphate (activating)
Wee1 adds one phosphate inactiting it.
CAK?
CAK phosphorylates the T-loop (an alpha helix) thereby unwinding the T-loop and allowing binding between the 2.
CAK of poep begins,
Binding of Cyclin moves T-loop, so things can enter…
Wee1?
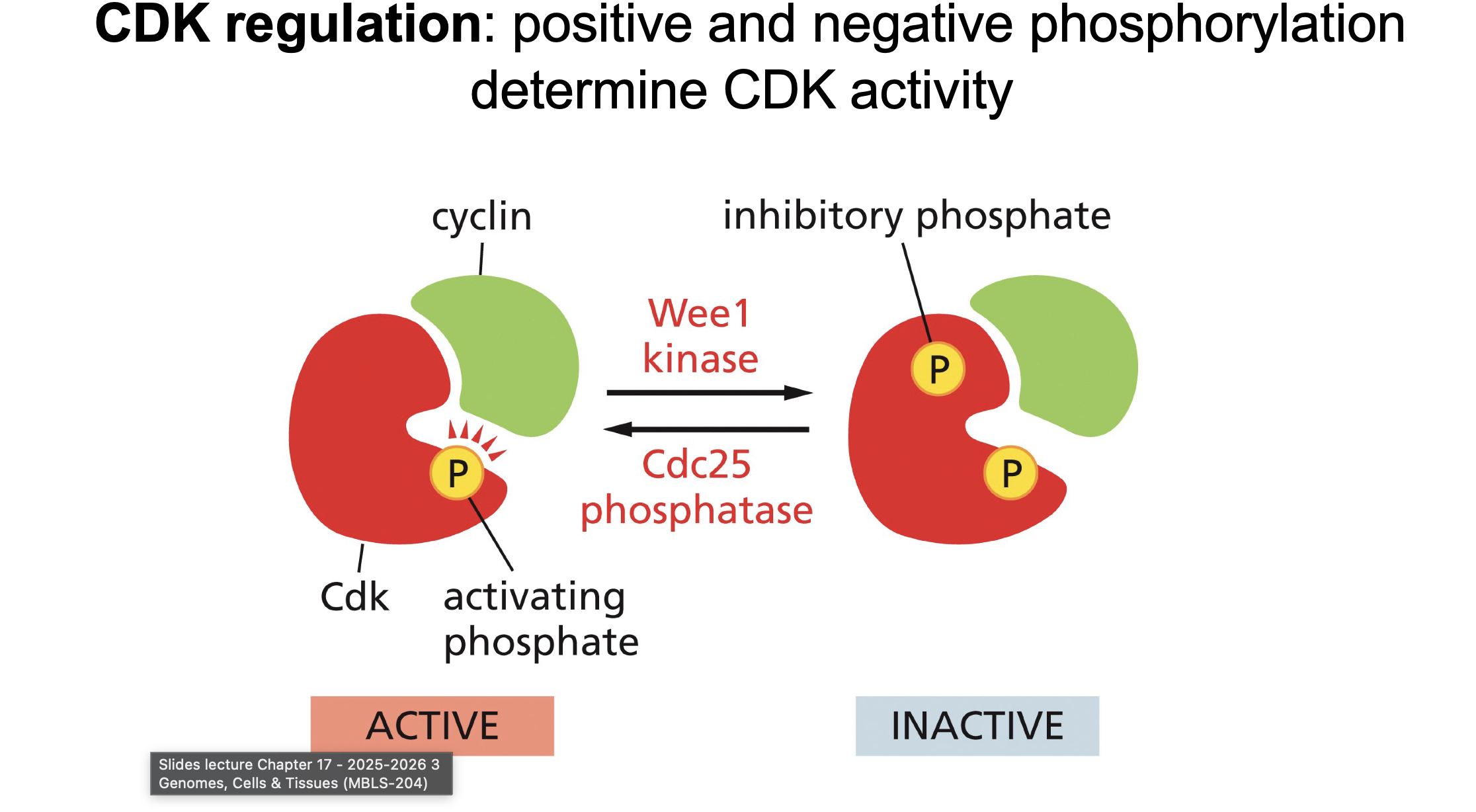
Cdc 25
When can ATP be hydrolyzed?
After CAK and Cdc25
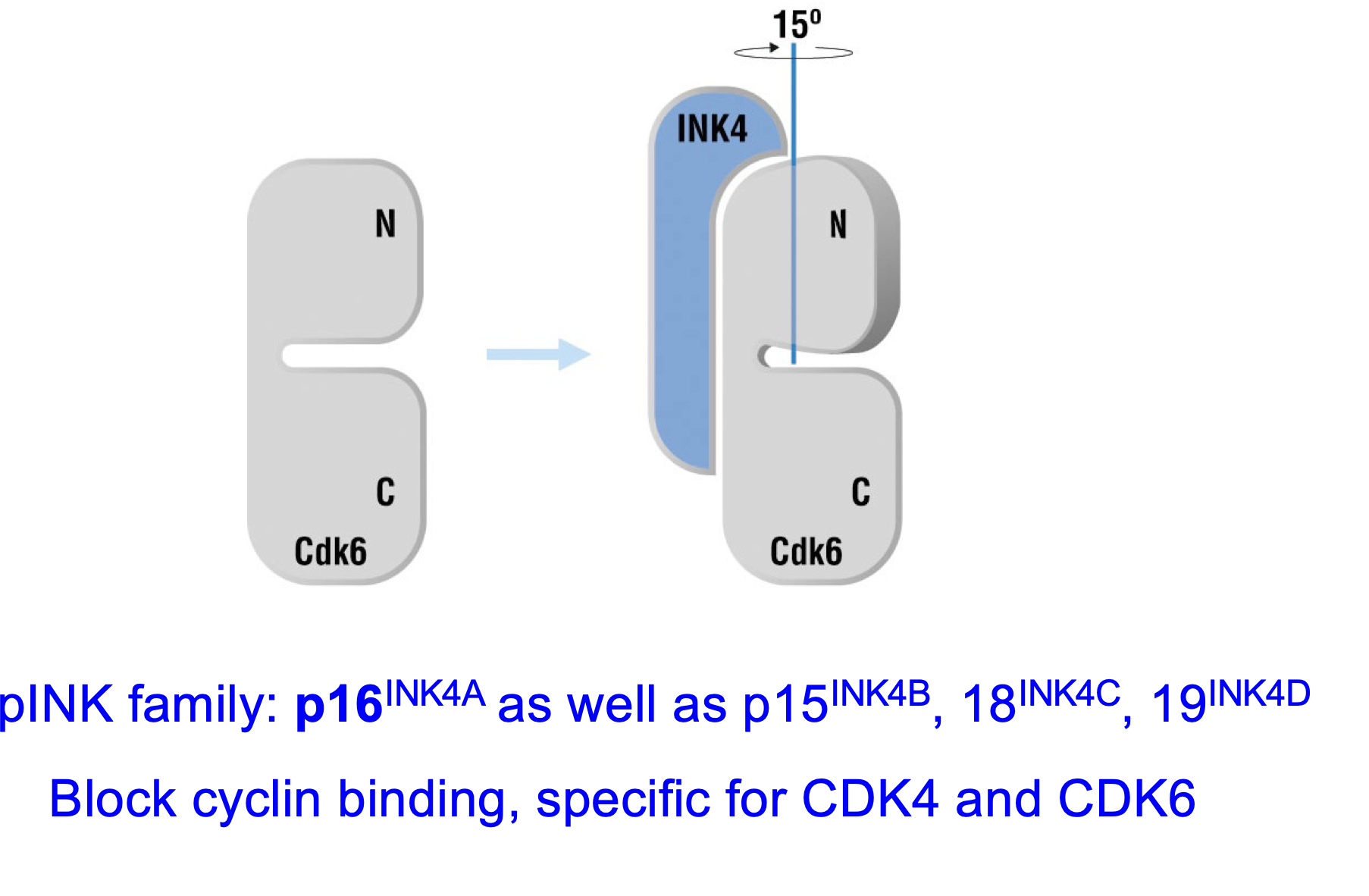
pINK family: p16INK4A etc: p16, p15 and p18
Block cyclin binding so formation of complex
Different members of pINK block different things:
p16INK4A blocks CDKA4 and CDK6
CIP/KIP families: like p21, p27, p57
Binds to full complex inhibits by preventing substrate from binding
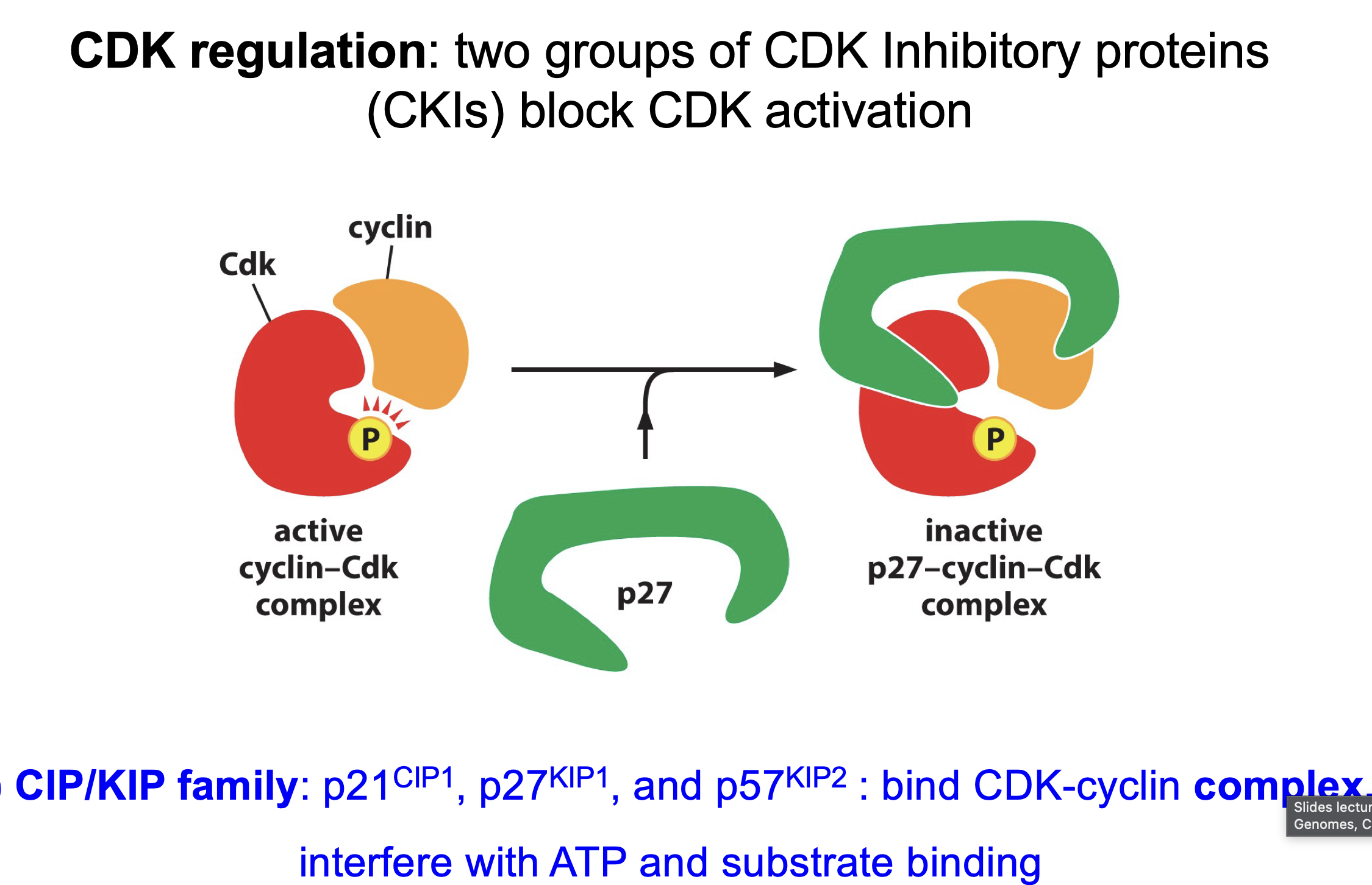
Why CDK so effective
Positive feedback
Can phosporolate a lot
APC/C + Cdc20
E3 ubiquitniastion enzyme, needs Cdc20 and phosphate to be active.
Cdc aperntly dont like CDKs
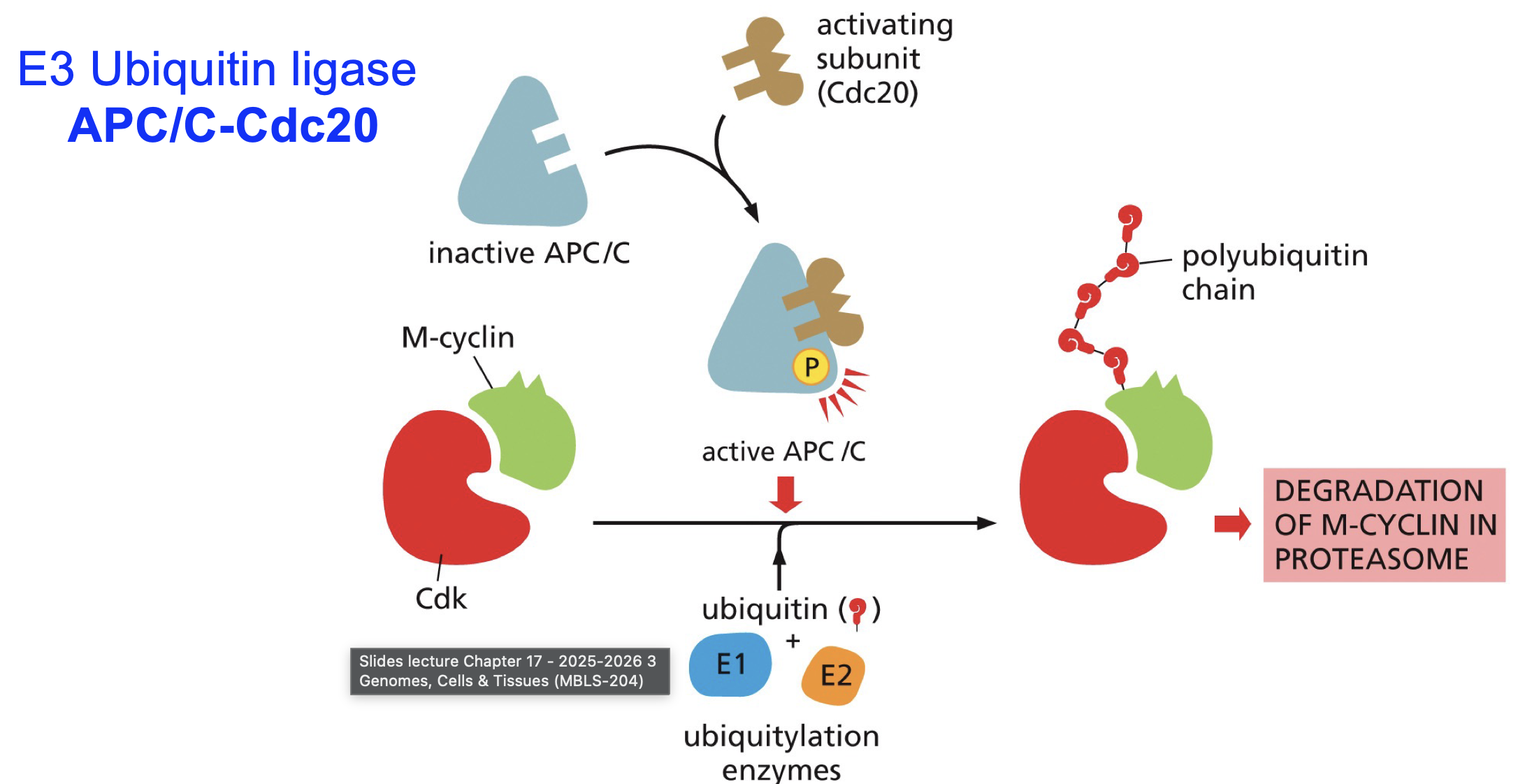
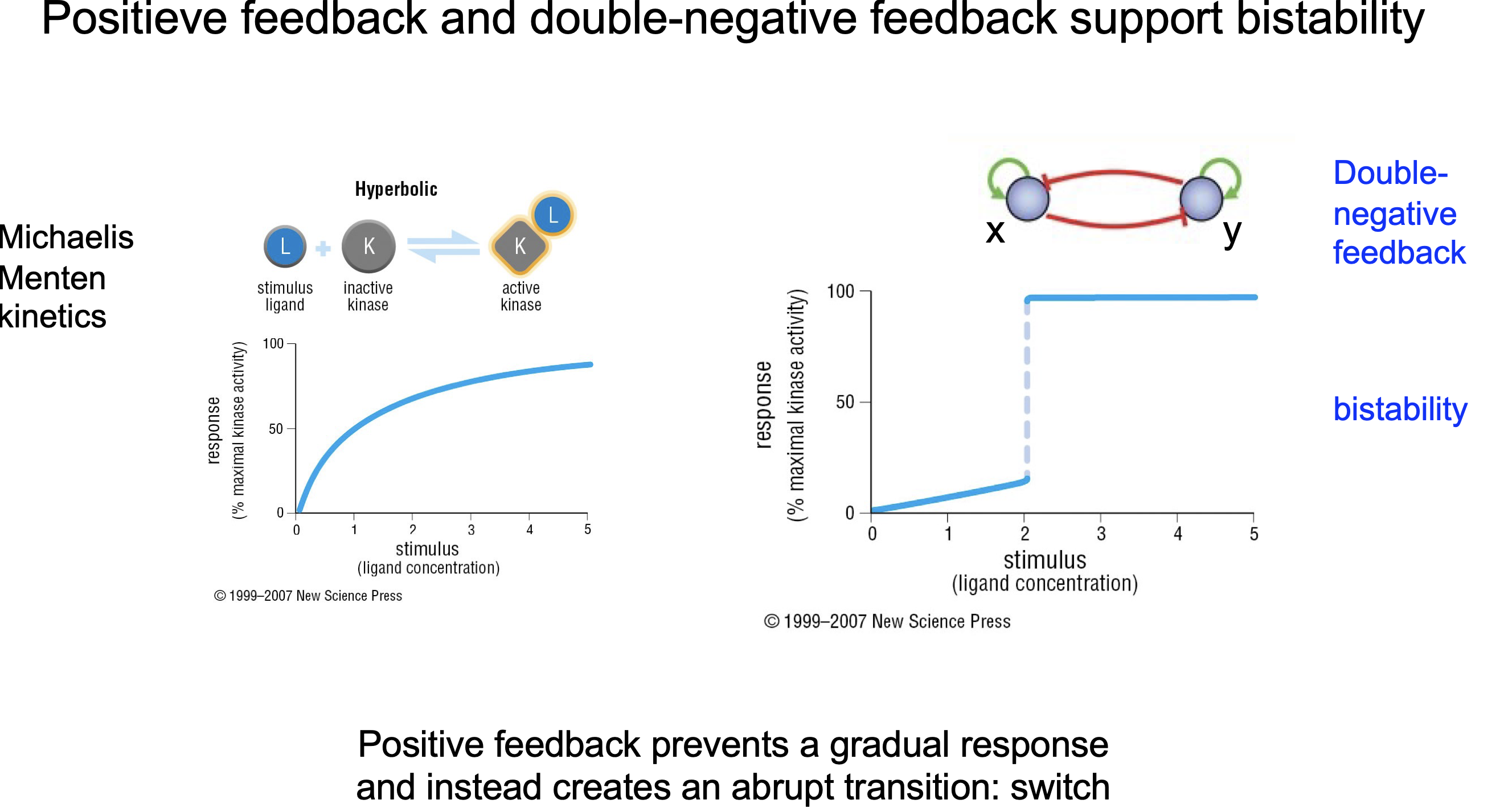
How do CDKs achieve “all-or-nothing decisions”?
What is Dubble negative feedback?
In bio mostly transition state unstable
Positive feedback
Negative feedback
Dubble negative feedback see picture
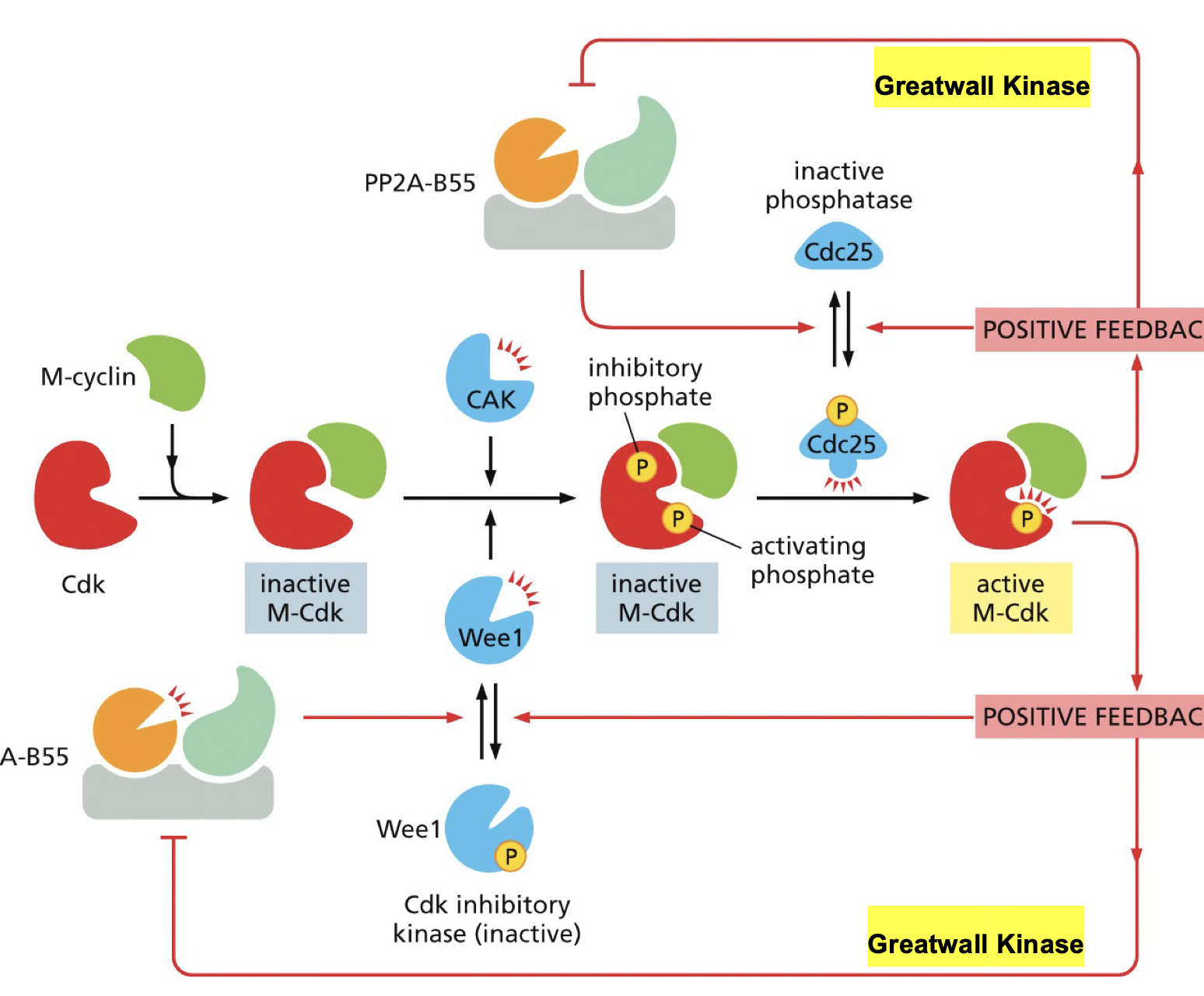
Greatwall kinase.
Removes phosphate from Cdc25 thereby inactivating cdc25 → leading to more active CDK what leads more great wall
Removes phosphate from wee1 thereby activating it→ leading to more active CDK what leads more great wall
Feedback loops in activation of CDK-cylin:
Snap afbeeld
As can see, if active it will be crazy active or it will. be of, the system is so that only on or off, no in between…
How can CIP/KIP be reugulated (p21, p27, 57)?
Is this positive feedback for CDK-cyclin?
Ubiquitin-dependent degradation
Yes ofc
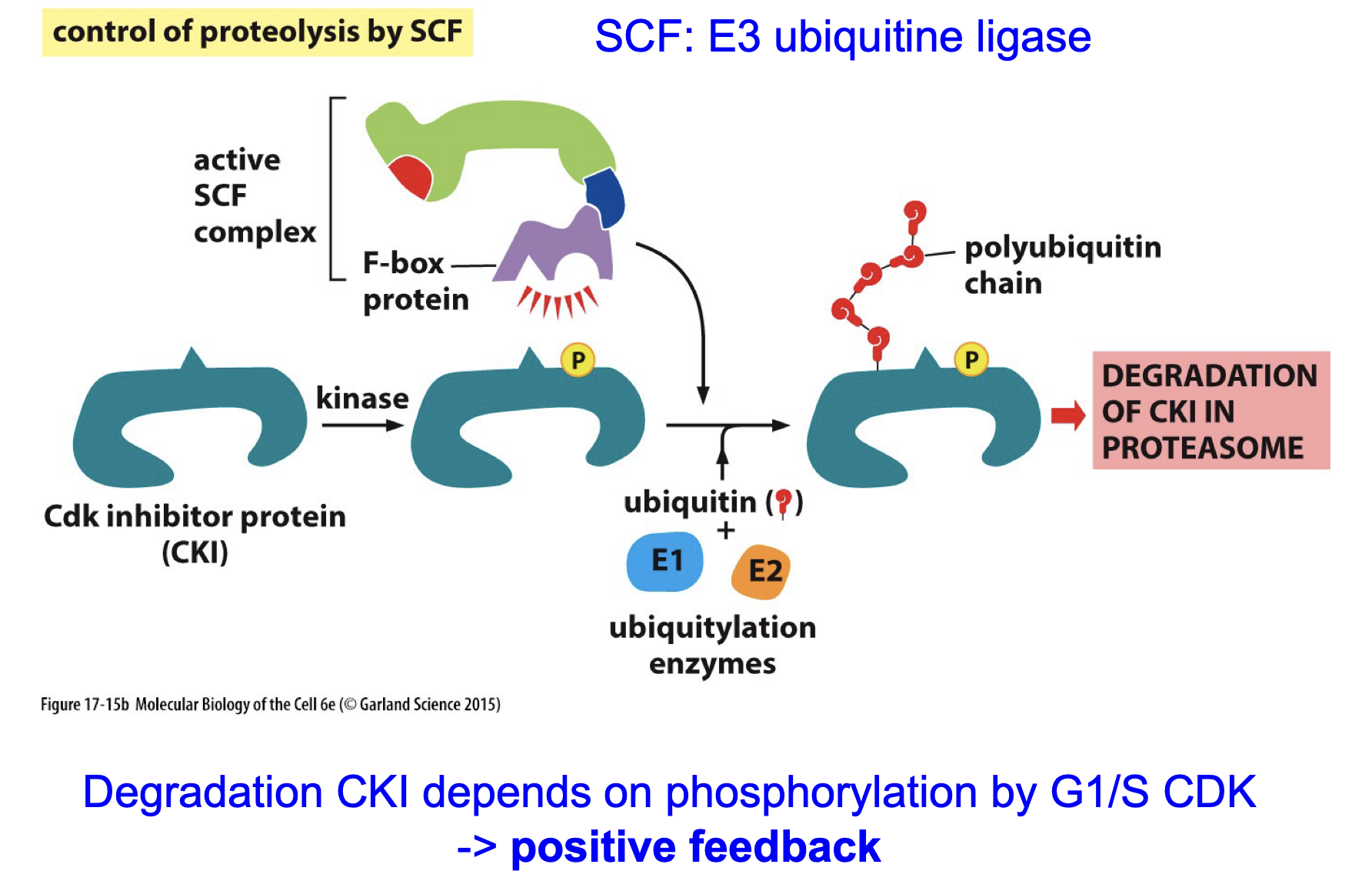
Cdh1?
Promoted by CDK-cyclin.
binds in the APC/C binding site blocking Cdc20 binding, thereby preventing the activation of the inhibitory complex.
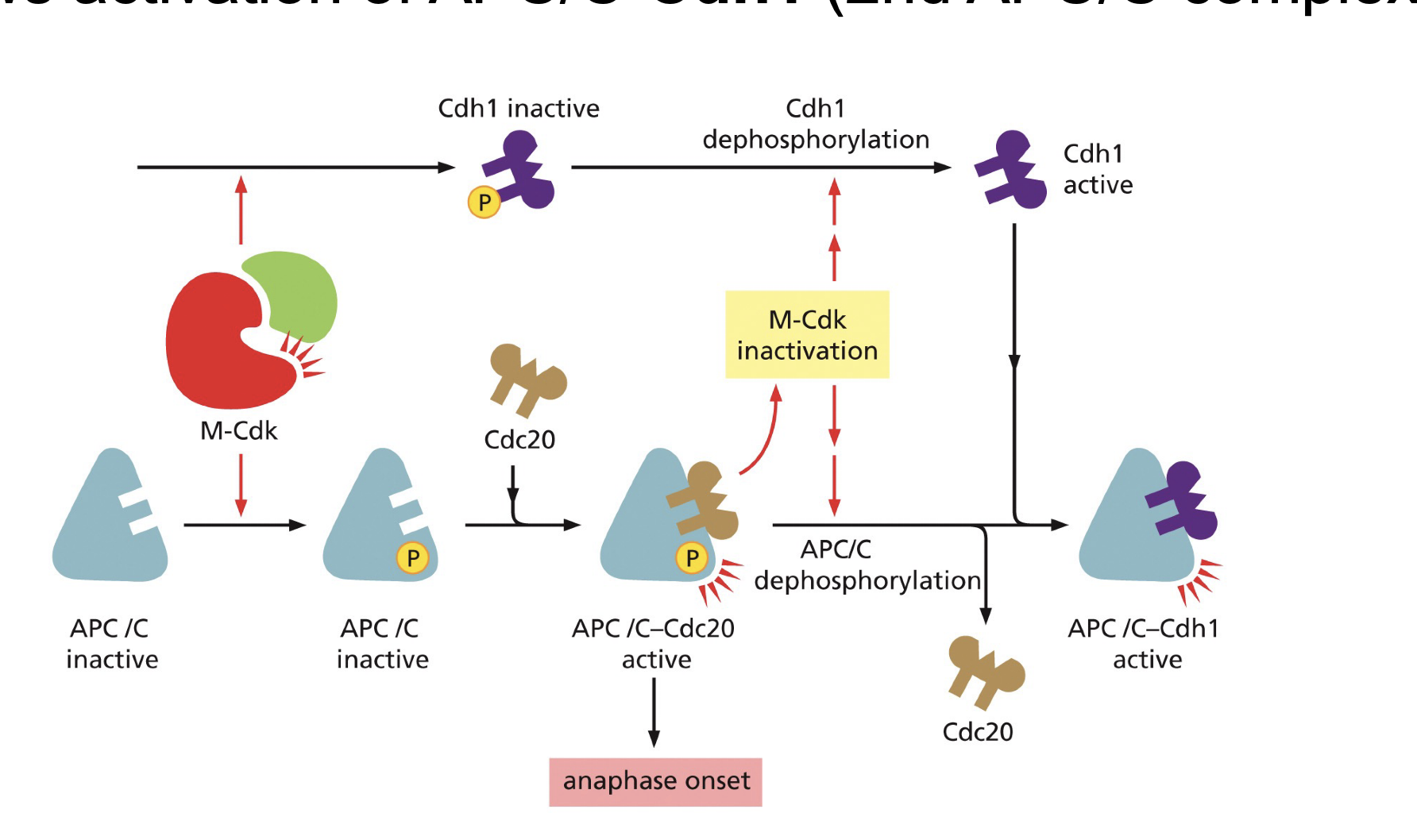
How does cell cycle start: G1→ S: starts the whole thing.
Groth factor binds
Was activates MAP kinase
Activates Genexpression
Activates D-type cyclins
What is first step towards cell cycle entry:
Transcription of D-type cyclins.
Trigers positive feedback loop
Forms CDK G1 plus D-type cyclins complex
E2F
Starts DNA synthesis.
Activated by G1-CDK-D-type cyclins, than activates it self, and creates a positive feedback loop to activate more CDK G1 plus D-type cyclins complex and CDK-S2 activation. What will lead to DNA synthesis
Switch confirmation dramatically after phosporlisation
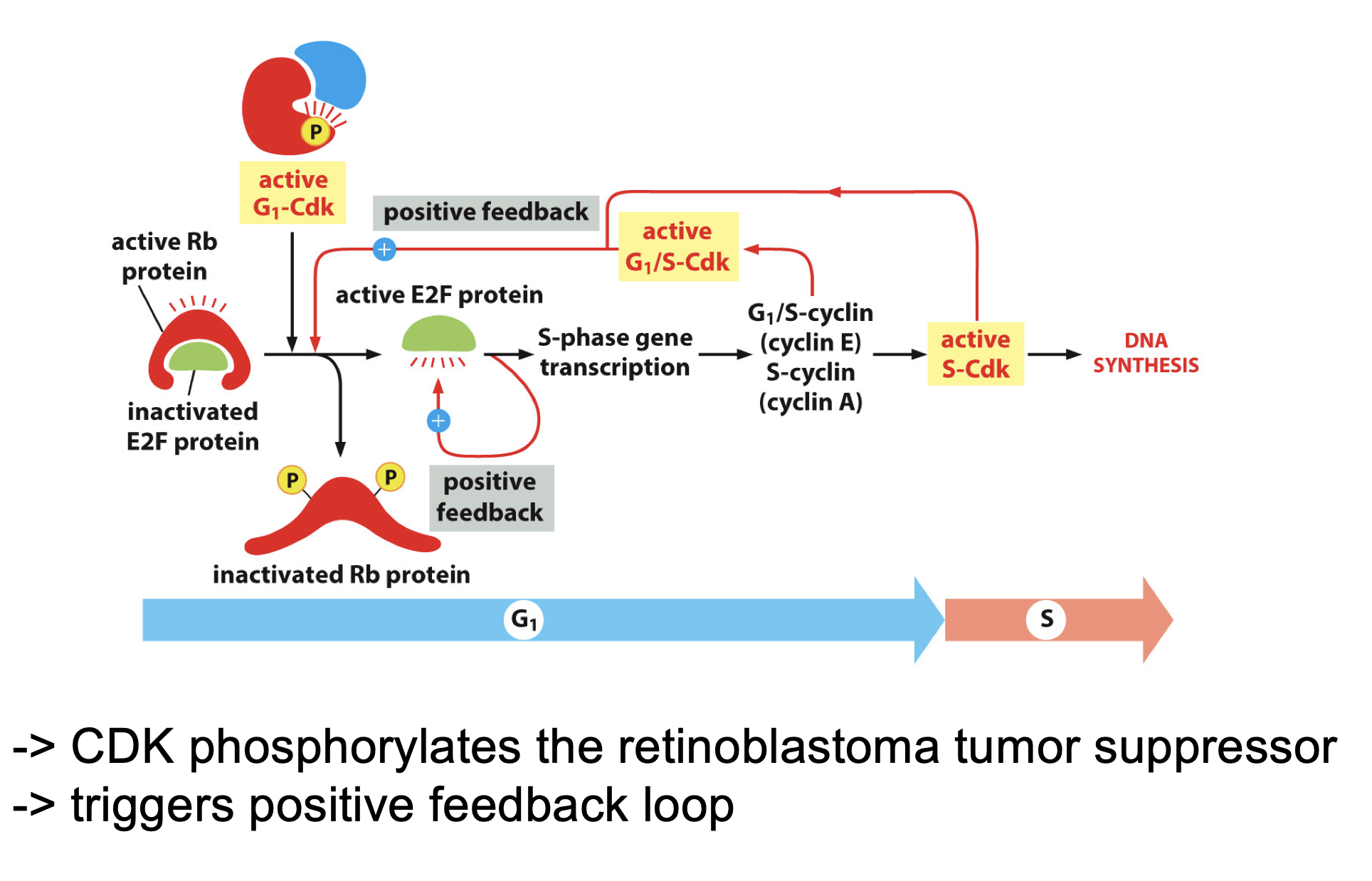
Is a balence in this activation:
But have this whole process mapped:
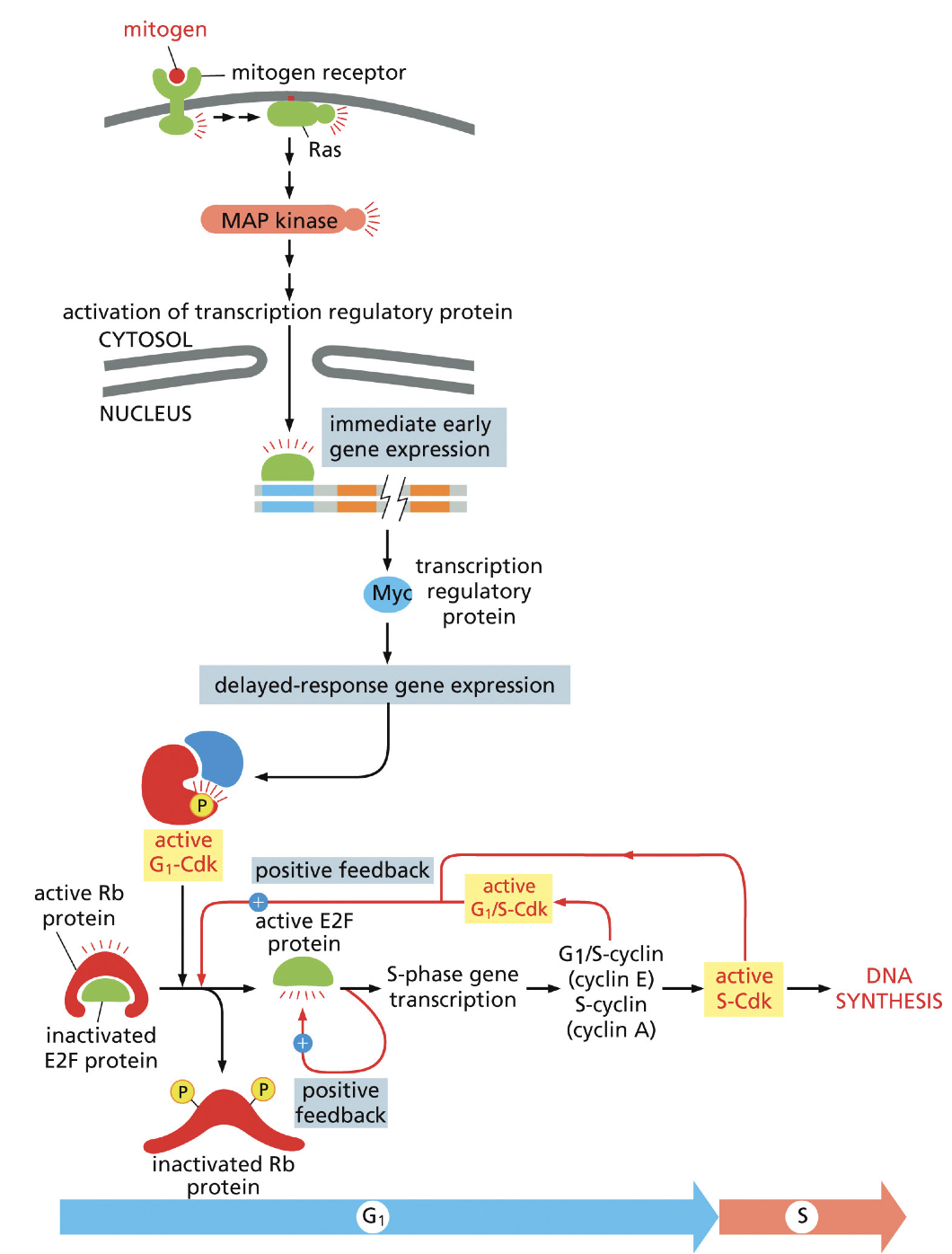
Rb protein
Inhibits Rb
Inactivated by G1-Cdk
Rb destroyd in almost all cancers..
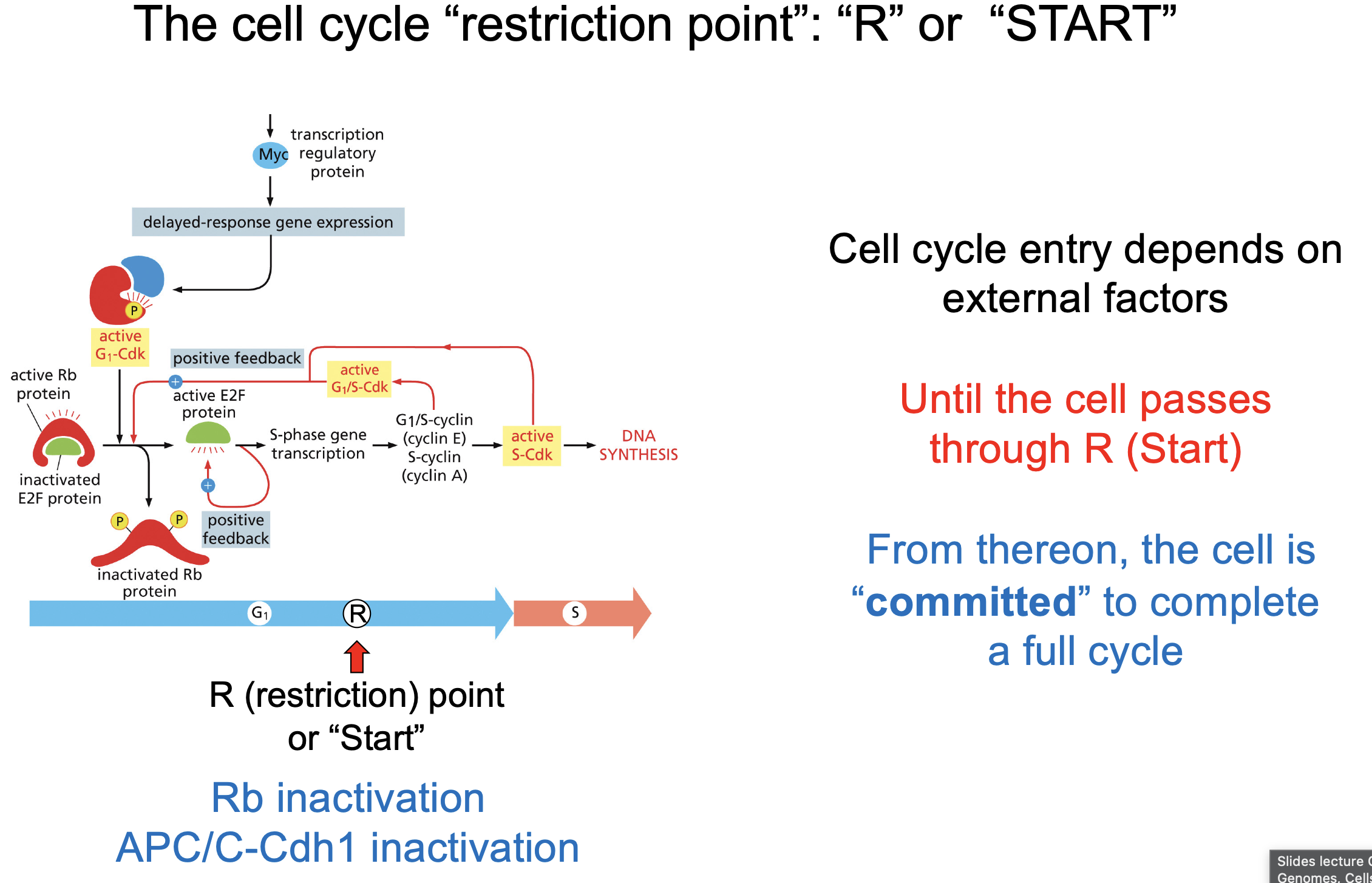
The cell cycle “restriction point”: “R” or “START”
After this point full cell cycle must be completed
enough Rb protein Inactivated.
But then it goes crazy very fast,
CDKs control DNA replication at multiple levels
Set up of replication complex →APC/C activation CDK inactivation (only when inactive complex can form)
Initiation → S-CDK
Chromosome seperation → M-CDK
Mitoses phases:
Prophase: Centromeres get in position + DNA condensation
Pro-metaphase: Mitotic spindle formed + Nuclear envelope vesiclased
Metaphase: Chromosomes alliant
Anaphase: Chromosomes pulled apart
Telophase: New nuclear envelopes formed + DNA decondensate + contracti starting to connect
Cytokinesis:: Cytoplasmic cleavage
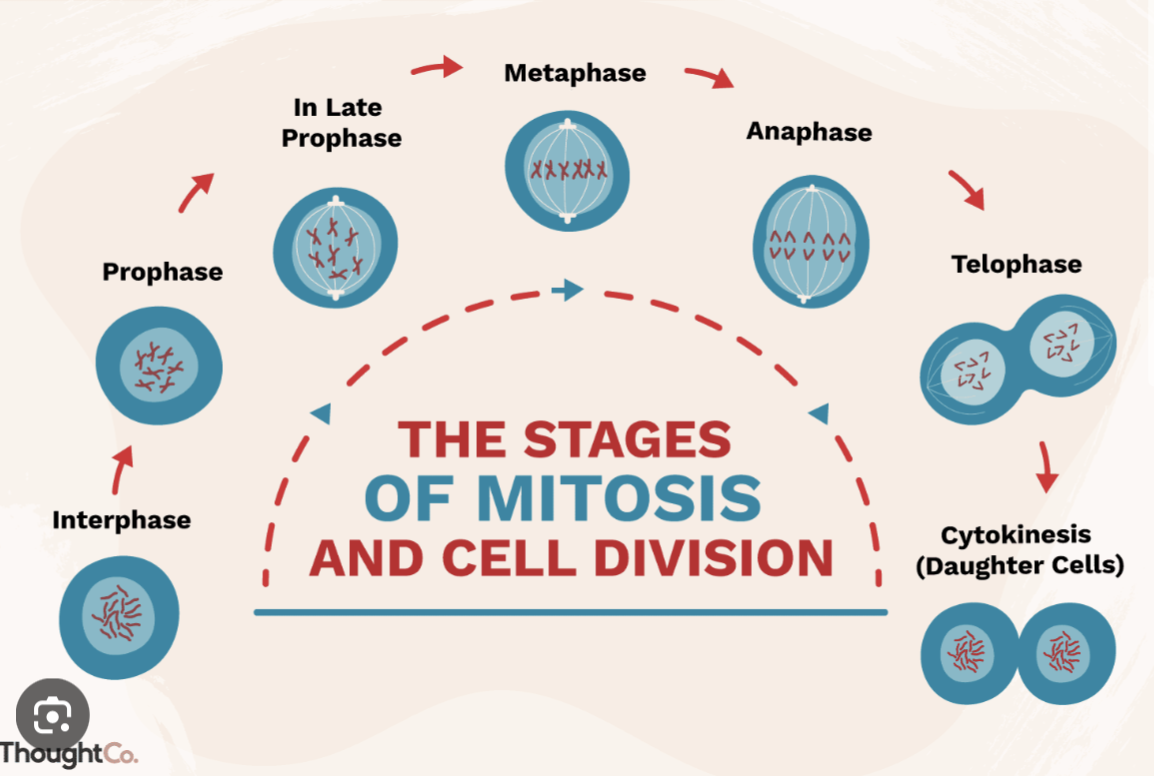
How CDK mitoses controls:
1. Chromosome condensation Histones, Condensin I
2. Reorganization microtubule cytoskeleton:
Formation mitotic spindle
Microtubule Associated Proteins (MAPs)
3. Degradation nuclear envelope Nuclear Lamins.
4. Initiation Anaphase Securin -> degradation cohesins
How CDK mitoses controls: Cytoskeleton .
CDK regulates:
Chromosome condensation (Histones, Condensin I).
Strongly enhanced microtubule dynamics (for centromeres).
Activity microtubule motor proteins boosted (dynein and kinesins) (again for centromeres).
Cohesin
Keep sister chromatids together,
For Anaphase needs to be broken down, so cell can pull chromosomes apart.
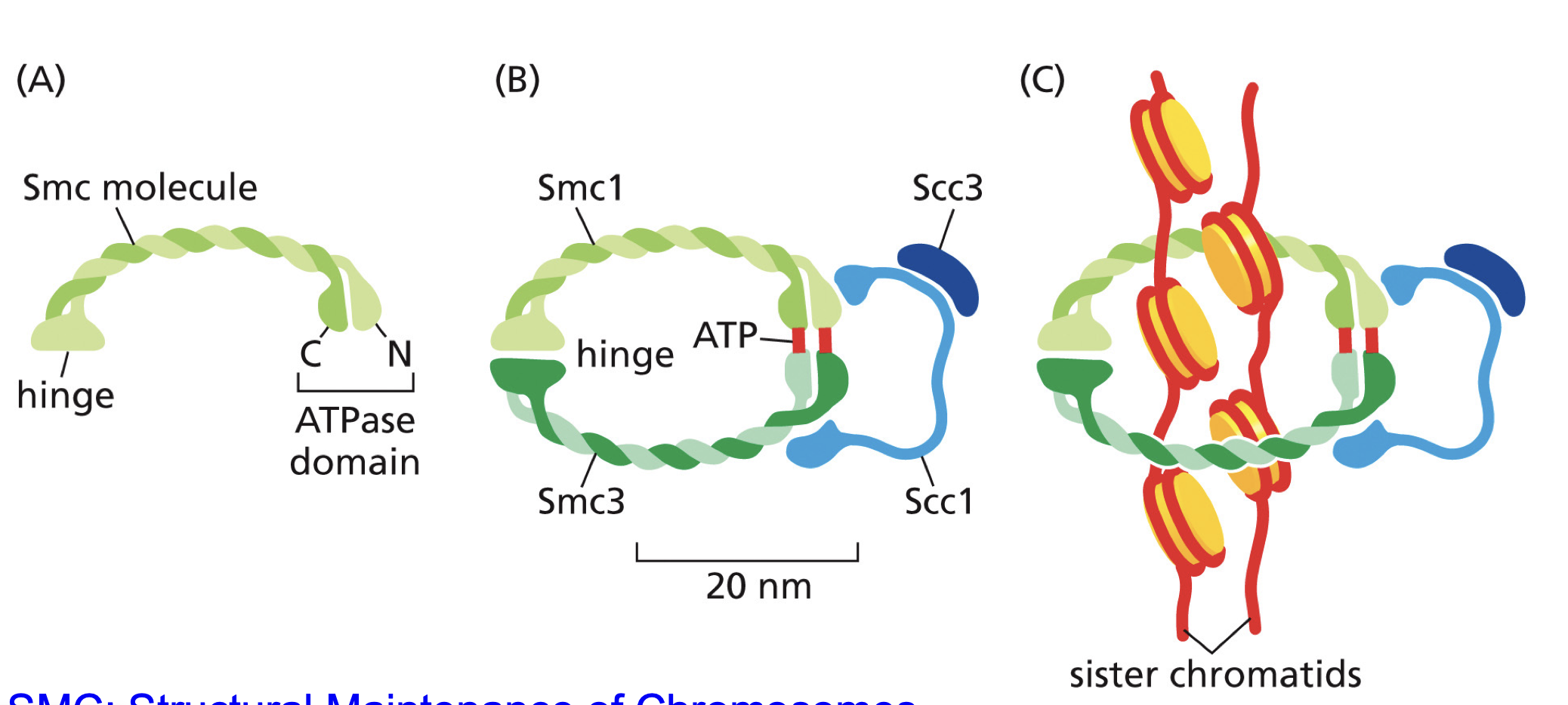
M-CDK-Cyclin B? (decreasing of Cohesin) what’s the pathway???
M-CDK phosporlyates APC/C.
APC/C binds Cdc20
Degrades securin (secure inhibits separase).
Active separase sapareses chromatids.
Anaphase…
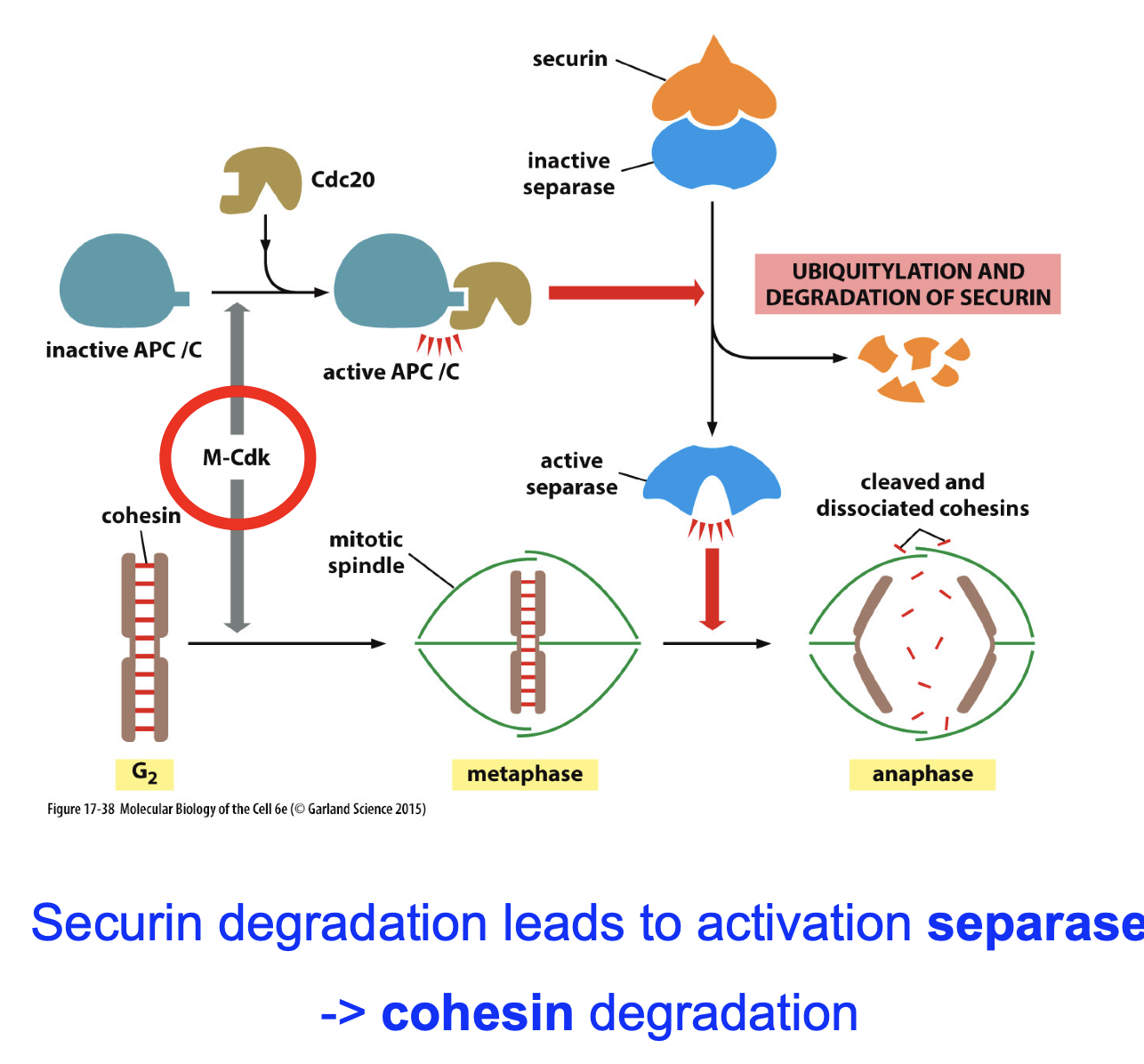
Checkpoints
all steps must be completed and did right…
Stops cell cycle
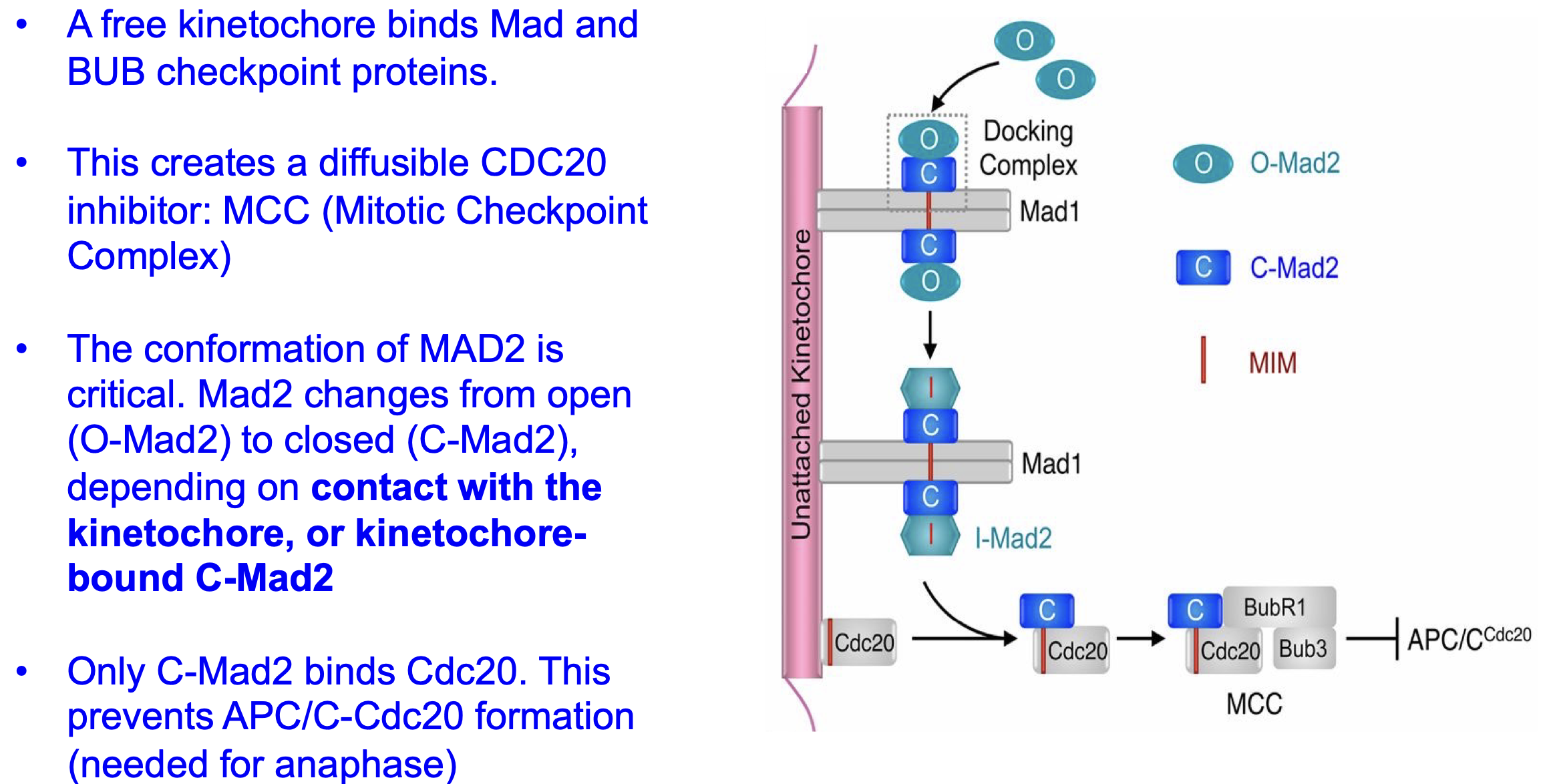
Mad2
Checkpoint protein:
Prevents APC/CCdc20 activation and thus separase to activate (so that anaphase does not start yet) till all microtubes are bind in the same right way.
MAD1 and MAD2 workings
If microtonal not bound = free kinetochore.
Mad 1 binds to free kinetochore.
O-Mad 2 binds to Mad 1.
Now O-Mad2 → C-Mad2.
C-Mad2 will bind and inhibit cdc20
and thus APC/CCdc20 activation and thus inhibitseparase
Can be fixed ones all kinechore are bound.
Microtubeles need to be connected to the kinechore in a stable manner. So that the centromere has the right one. What enzyme checks this?
Aura B, inhibits anaphase. Is removed from chromosome by the right tension thereby losing it ability to inhibit. what allows anaphase to start.
Hij farmed well Aura B Spiderman stijl
Are way more checkpoints sooo cool
apoptosis vs necrosis
Apoptosis:
• Regulated
• Caspase dependent
• Membrane “intact”
• Blebbing of the membrane
Necrosis:
• Not regulated
• Membrane rupture
• Spillout of cytotoxic proteins
What does Propidium iodide staining?
Propidium iodide (rood) DNA staining
For Necrosis
Cells that are more sensitive for apoptotic signals:
Developing organisms
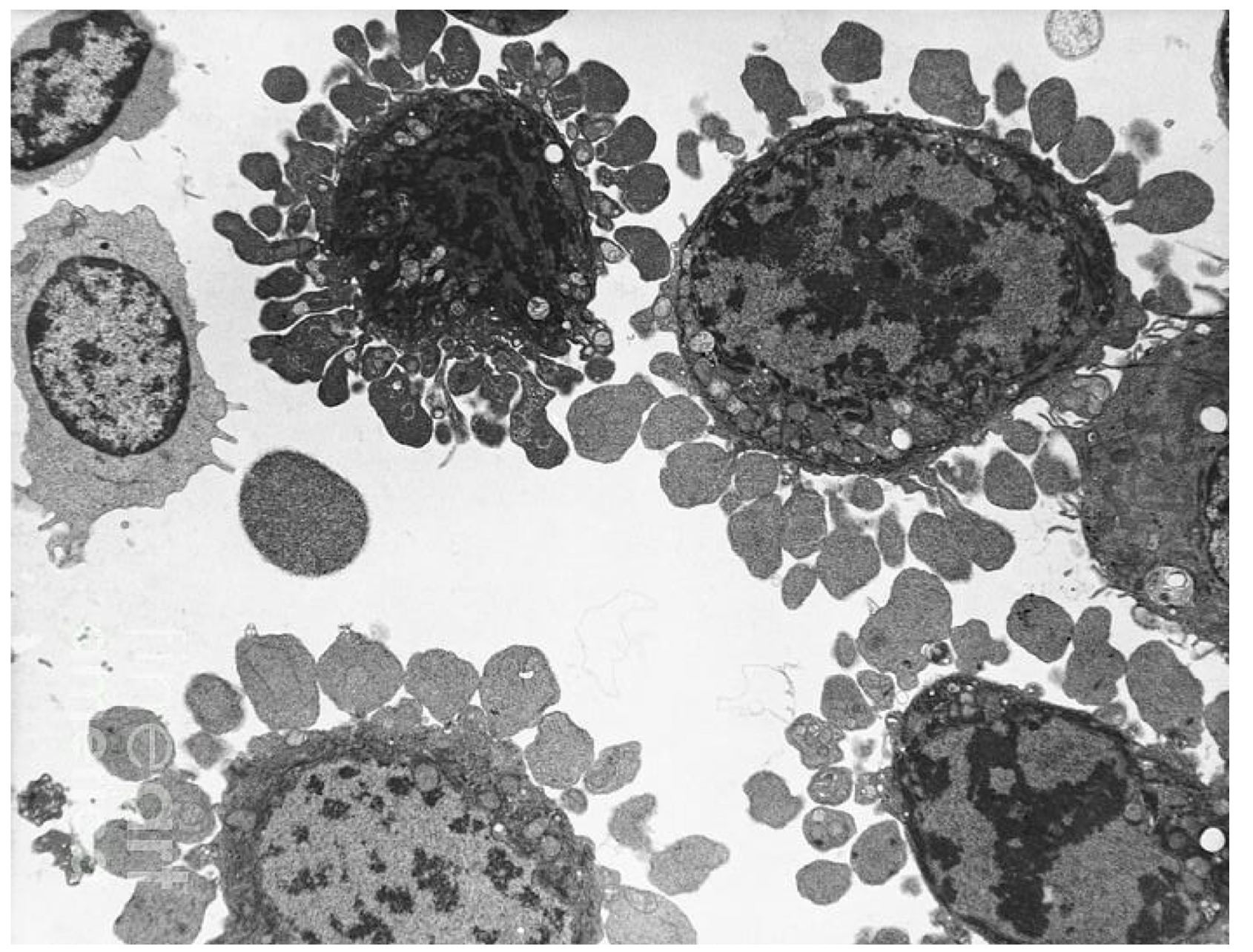
Aptosis steps:
1) Cell shrinkage
2) Activation of the caspase cascade
3) DNA fragmentation
4) Disassembly of the nuclear lamins & nuclear
envelope (dark patches)
5) Formation of large vacuolar structures (light
structures)
6) Collapse of the cytoskeleton
7) Blebbing (not present yet in the picture)
8) Removal by macrophages
How caspase is activated? anf formed
Apotatic signal binds to 2 caspase monomers dimerizing them.
Top part of dimer cleaved off and removed
Part of the active domain cleaved of to link the dimmers
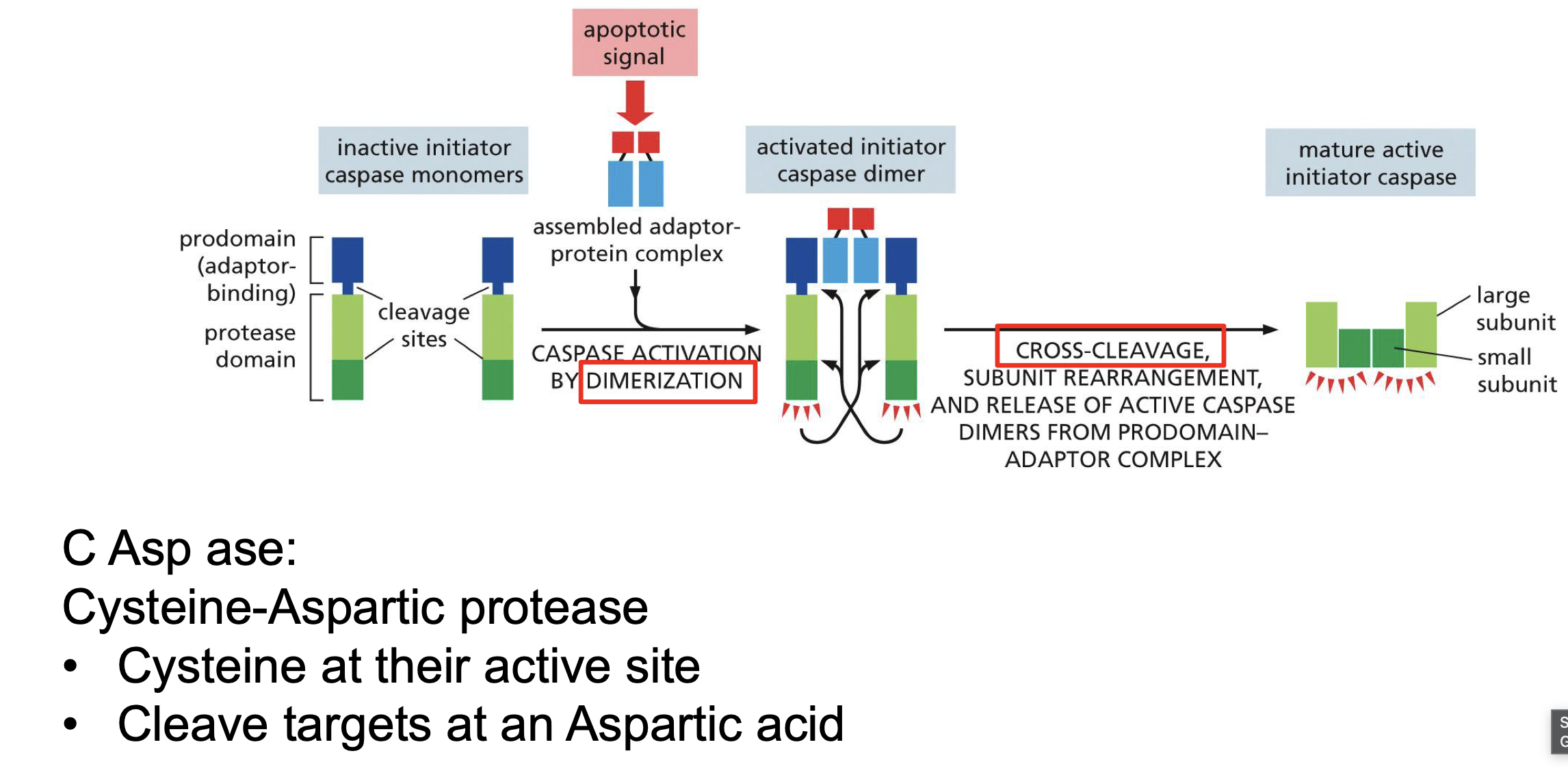
Initiator caspases?
activates Executioner caspases..
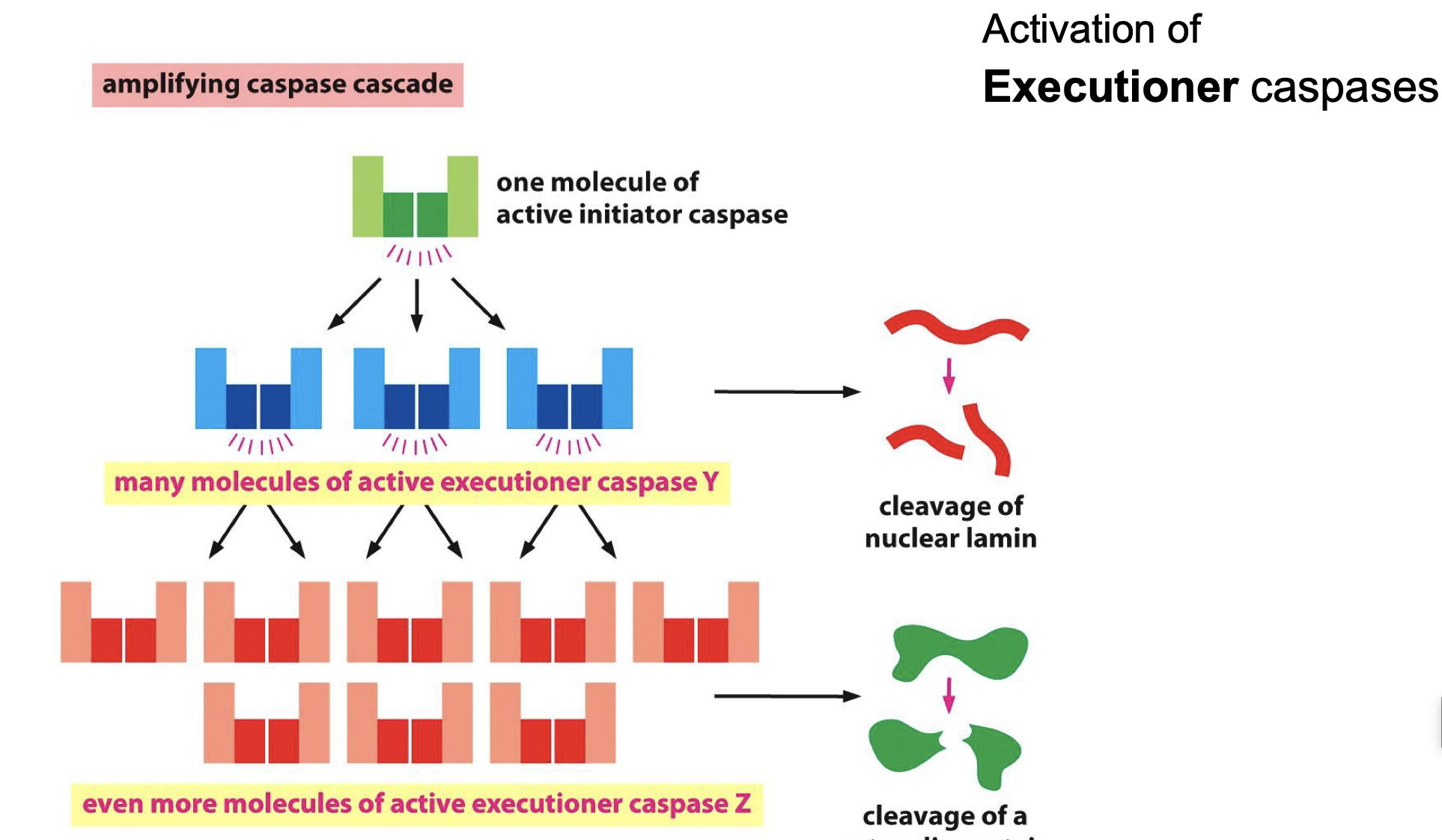
How DNA fragmentation?
Cad with iCad bound (iCad = inhibitor Cad)
iCad Cleaved by caspase
Cad cuts in between histones. (So euchromatin les cut up?)
Its special in aptotic cells can detect them with this
This how caspase works cleavage
TUNEL staining
Labels cut DNA
By binding to 3’OH
Mictophage and PhosphatidylSerine relation
How activated
PhosphatidylSerine on lumin side
PhosphatidylSerine is a eat me signal
Flipase turned into scrambalece
PS is transferred to the outer leaflet by two mechanisms:
1. Phospholipid flippases are inactivated.
2. A “scramblase” that transfers phospholipids nonspecifically in both directions betwee
Blebbing
Characteristic for apoptosis
stain: YFP-BAX
green, cytosolic
Mito-Cherry: stain
mitochondria
Hoechst 33342:
blue, nuclear stain
Phosphatidyl Serine
magenta
Activation of Apoptosis the 2 pathways:
Intrinsic pathway
Extrinsic pathway
Extrinsic pathway
Fas ligand Binds to Fas death receptor (Can come from cells or just ligands)
Activates DISC assembly
Activates by cross cleaves and dimerization caspase 8
Wich is an Initiator caspase and activation caspase 3 and 7
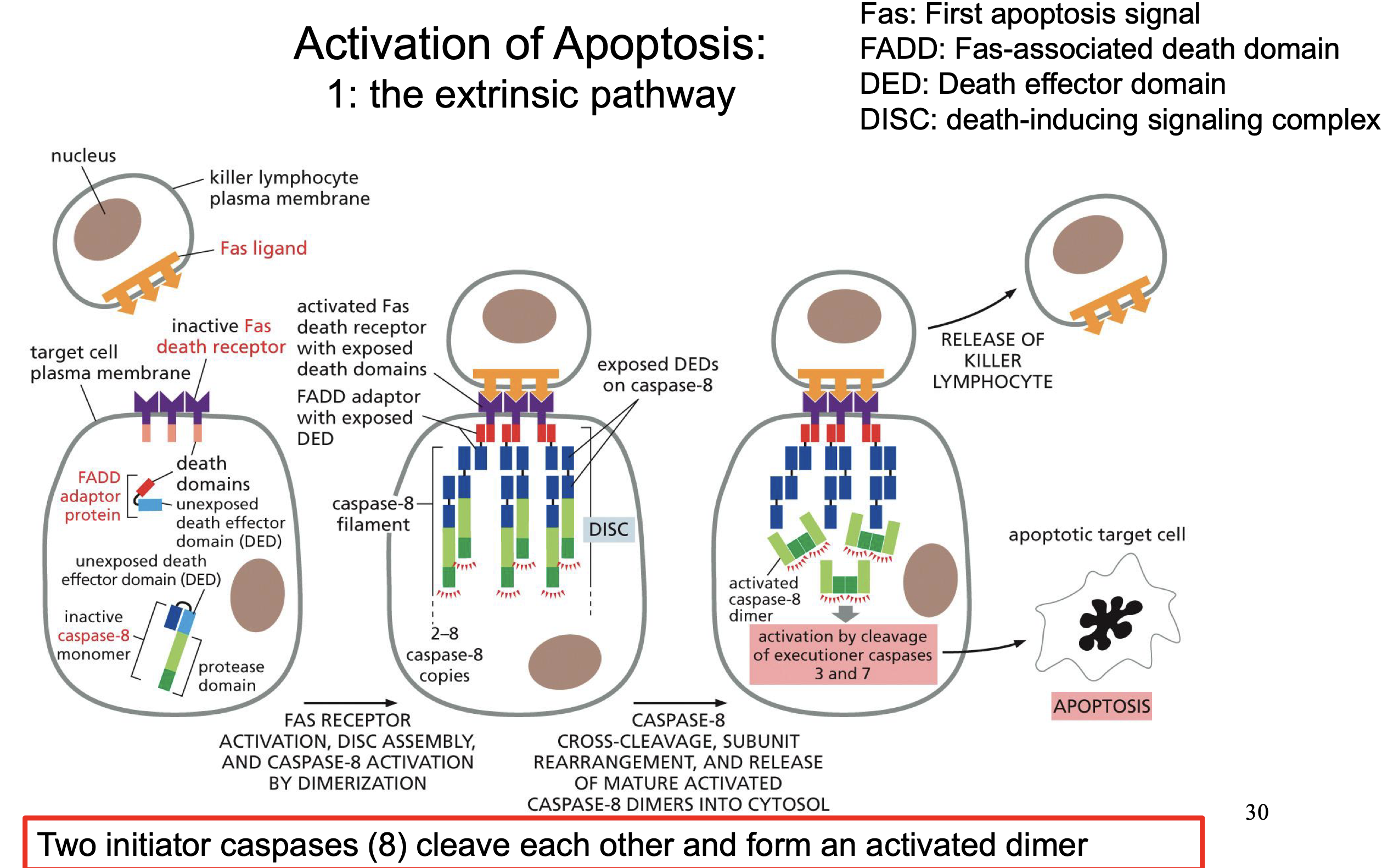
Fas ligand
Initation of Extrinsic pathway
(Can come from cells or just ligands)
Fas death receptor
Activates DISC assembly
DISC assembly
Activates by cross cleaves and dimerization caspase 8
Caspase 8
is an Initiator caspase
activates caspase 3 and 7
What induces the intrnincic pathway for aptosis
• UV irradiation -> DNA damage
• Hyperproliferative signals
• Malfunctioning protein kinases
• Mitochondrial dysfunction
p53
Can lead to:
Cell-cycle arrest
Senecence
Apotosis
Can be activated by:
UV irradiation -> DNA damage
• Hyperproliferative signals
• Malfunctioning protein kinases
• Mitochondrial dysfunction
SO if one on these situations occurs it leads to apoptosis
Withdrawal of trophic factors, growth factors, mitogens
Need survival factor to prevent apoptosis. If not than Bad not decreased so no Bcl2
KEY REMBER PICTURE
Tyrosine kinase
PDK1 and. AKT
AKT destroys Bad so active Bcl2
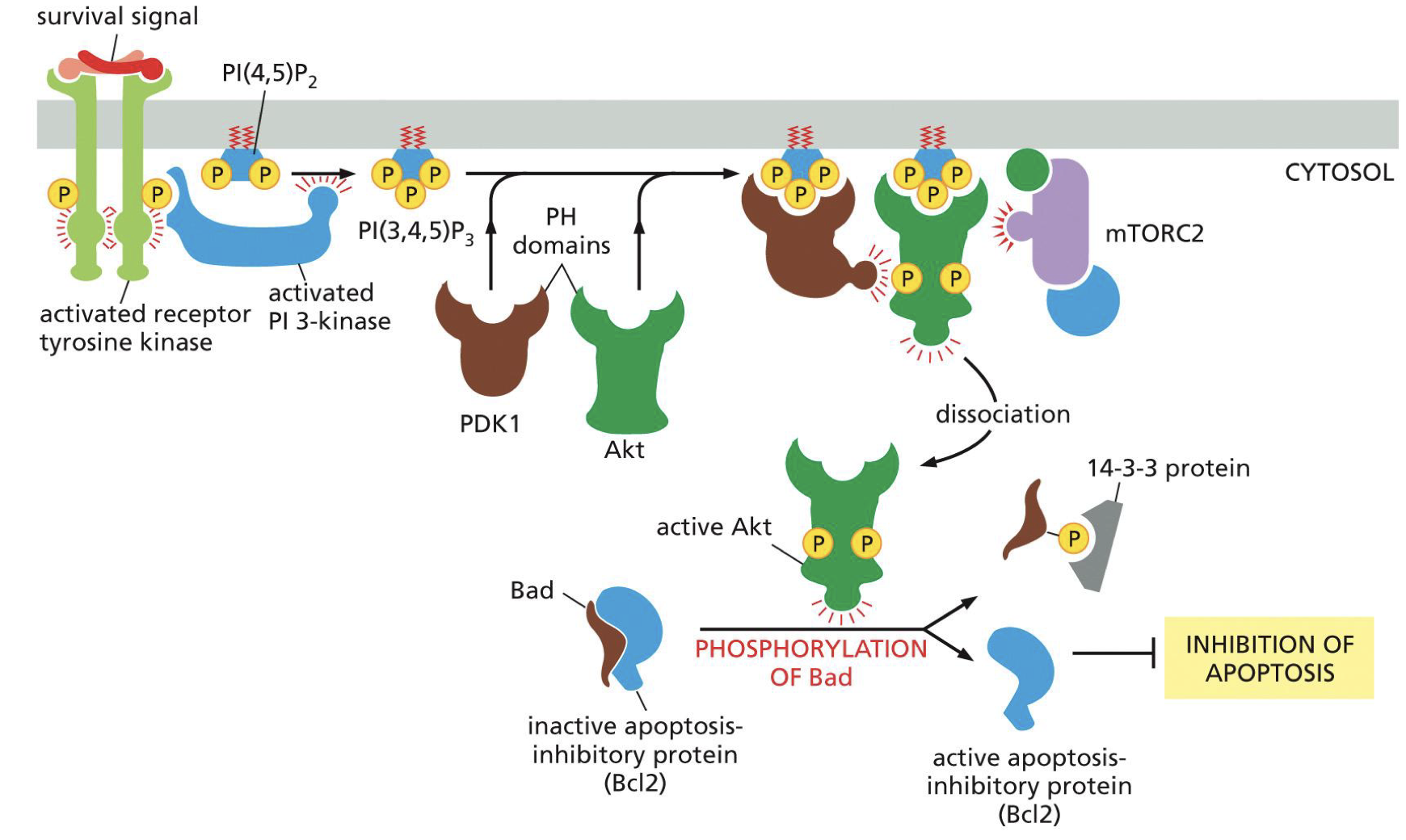
Cytochrome-C
Normal role in mitochondria.
But if it gets out of there it will Start some CRAZY SHIT
Activation of Caspase signaling by cytochrome C
Cytochrome C released in cytoplasm
Apaf1 binds cytochrome C which activates it
Active Apaf1 oligomerise (7subunits) into a ring
Recruitment of caspase 9 monomers
Activates caspase 9 monomers by cross cleavege and forming dimers
Caspase 9 is an initation caspase
Apoptosis
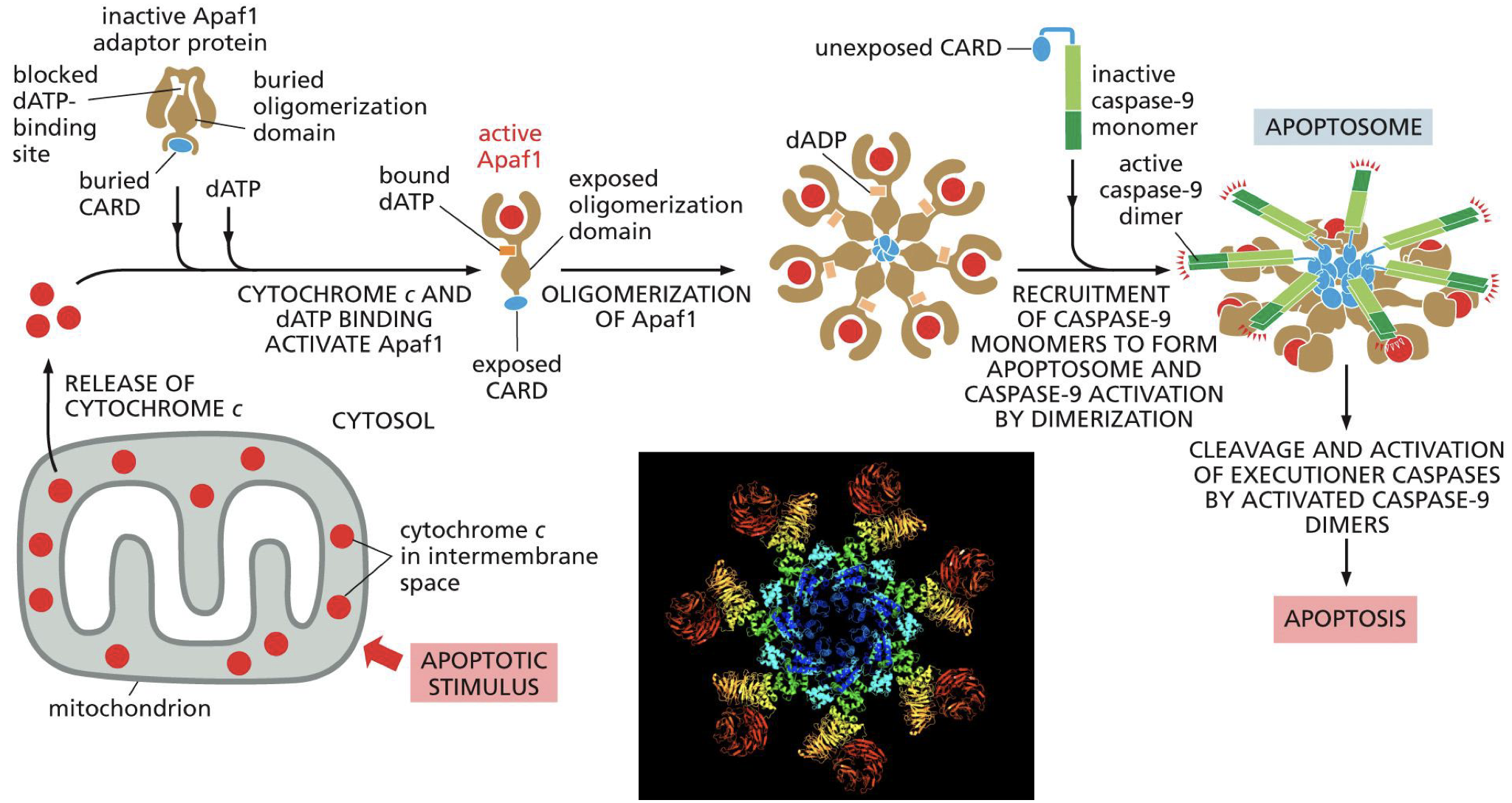
Apaf1
Cytochrome C released in cytoplasm
Apaf1 binds cytochrome C which activates it
Active Apaf1 oligomerise (7subunits) into a ring
Recruitment of caspase 9 monomers
Activates caspase 9 monomers by cross cleavege and forming dimers
Caspase 9 is an initation caspase
Apoptosis proteins (BCL-2)
Are all simelair in structure:
All use BH-3 domain
Stops Apoptosis
Bcl2
BclxL
All BH-3 only proteins stop apoptosis
Starts Apoptosis
Cytochrome-C
Bad
Bak
Bax
Bim
(Cool that Apotosis needs to be stop it is not activated)
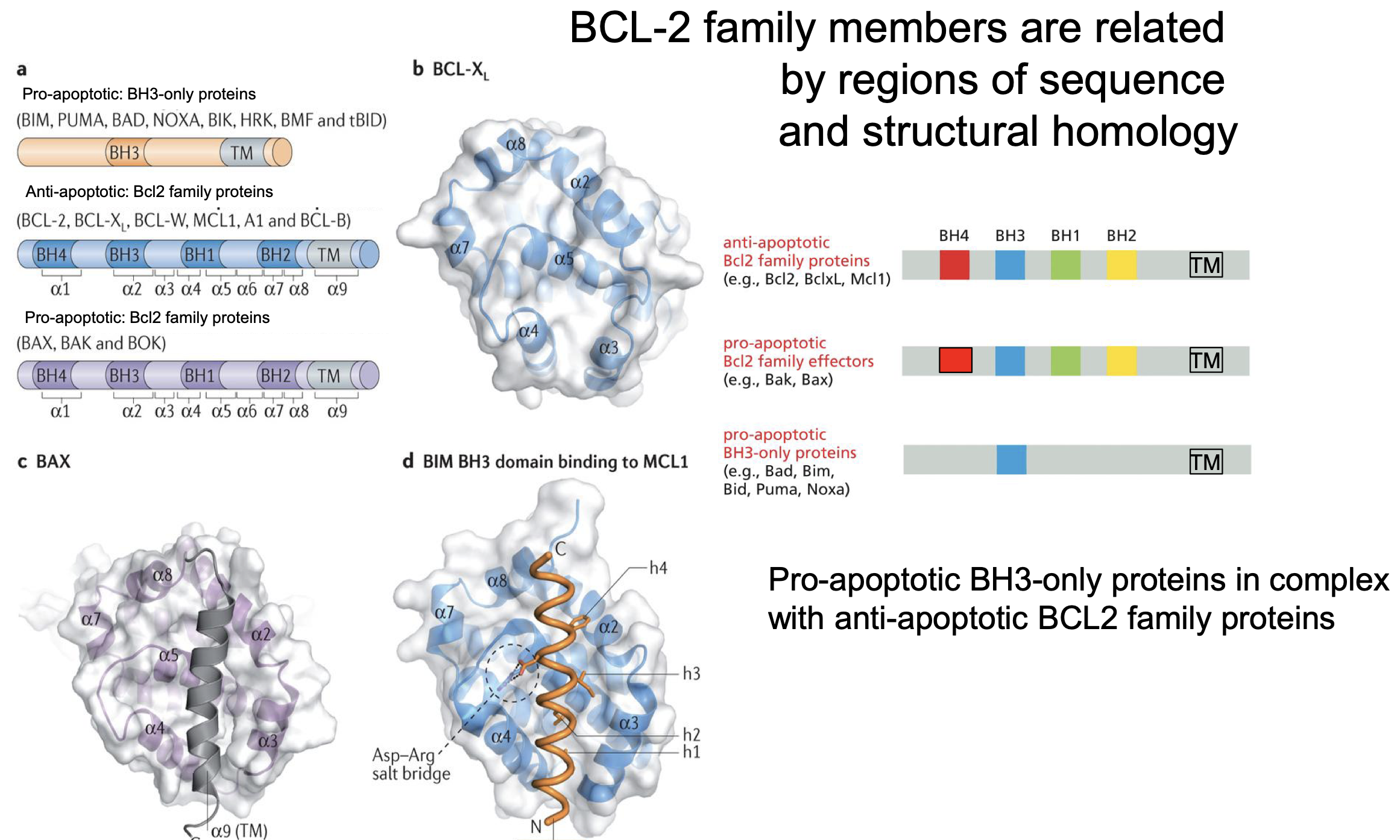
Pore forming proteins
Bak and Bax
When Bak or Max oligomerization to form a prores in mitochondria to let Cytochrome C out.
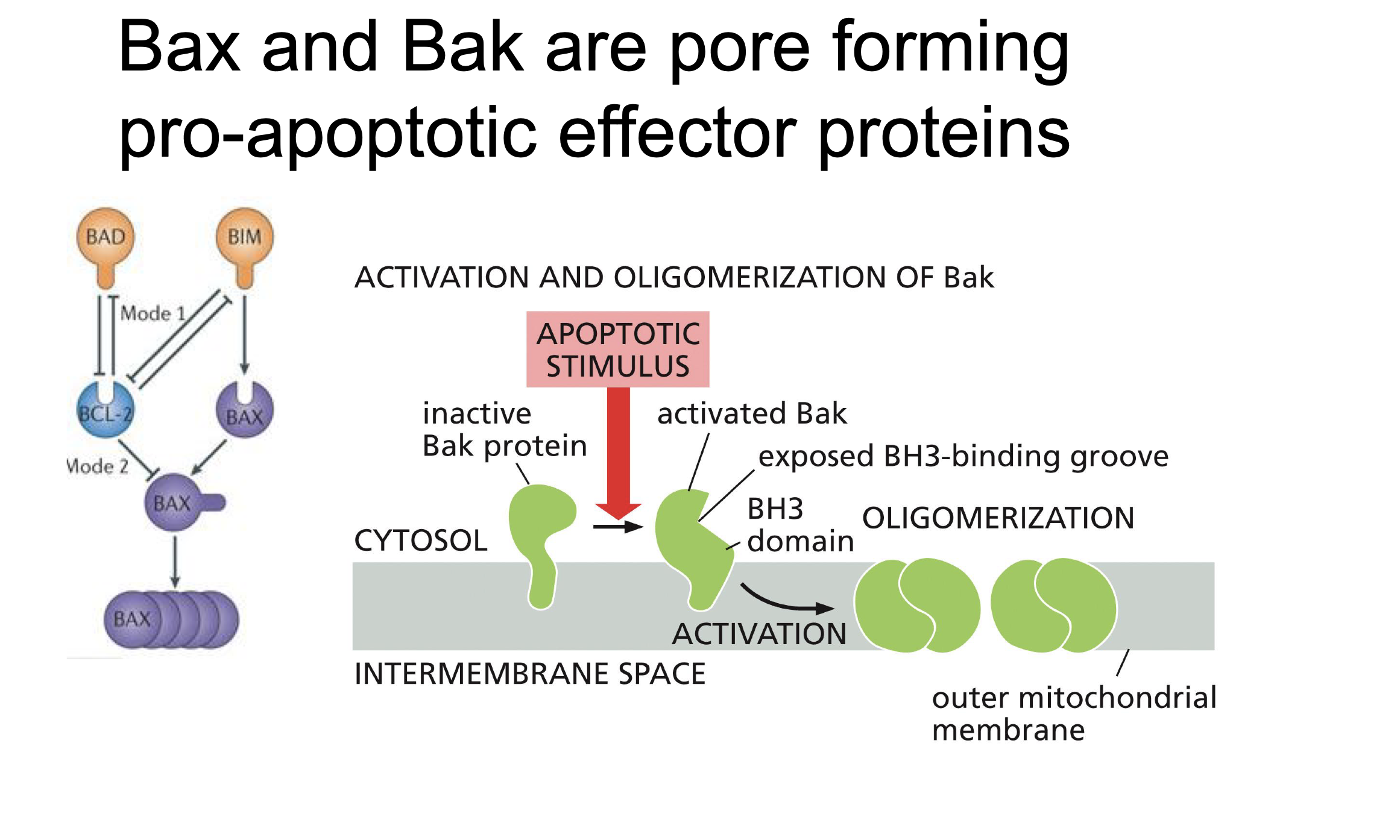
Bclxl
Binds to Bak and BAX so that it can not oligomerise
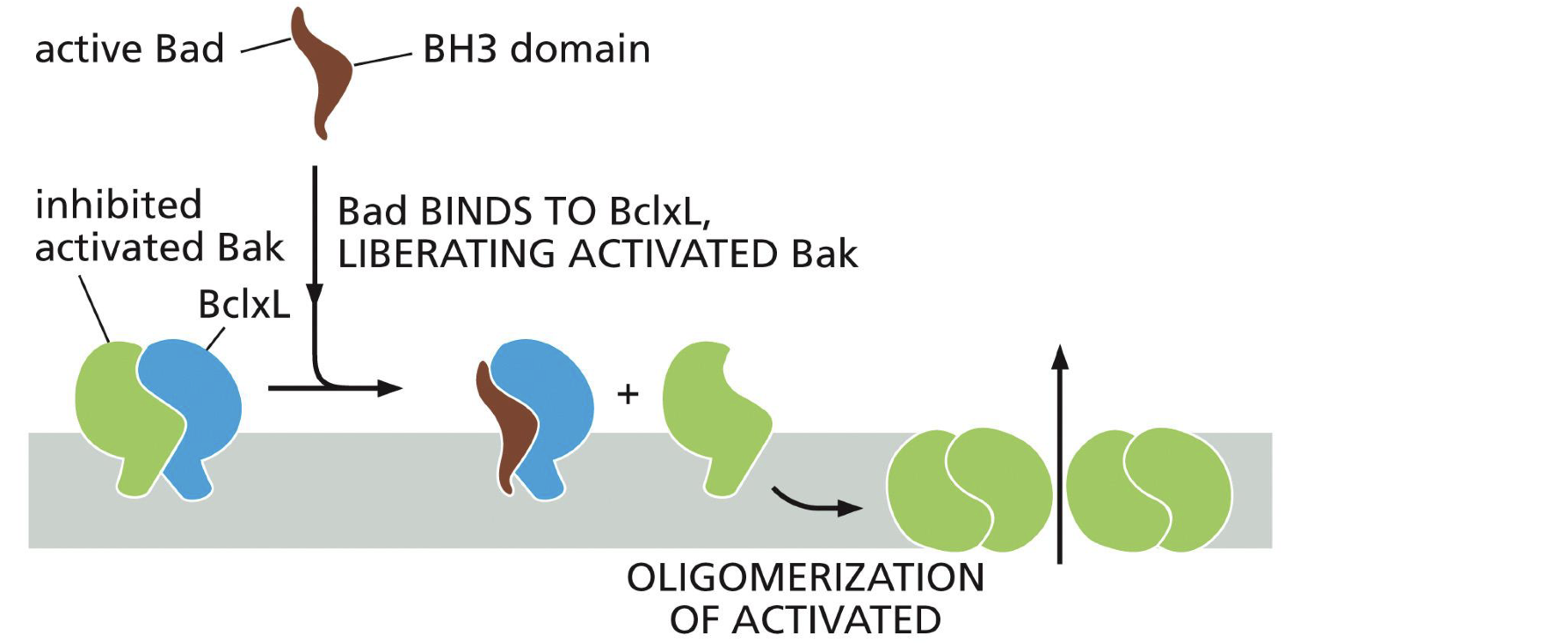
Bad
Binds to Bclxl
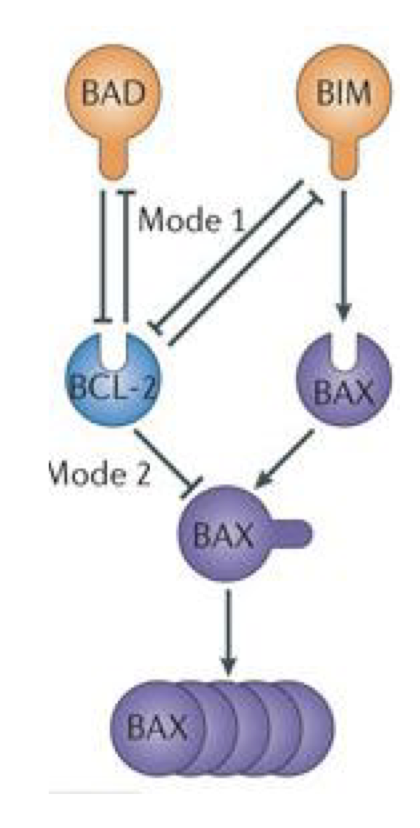
BAX reguation (BID)
In active form promotes it self
Inhibited by Bcl2 which is inhibited by BAD and by BIM
BIM promotes BAX
Bax cluster to form pores
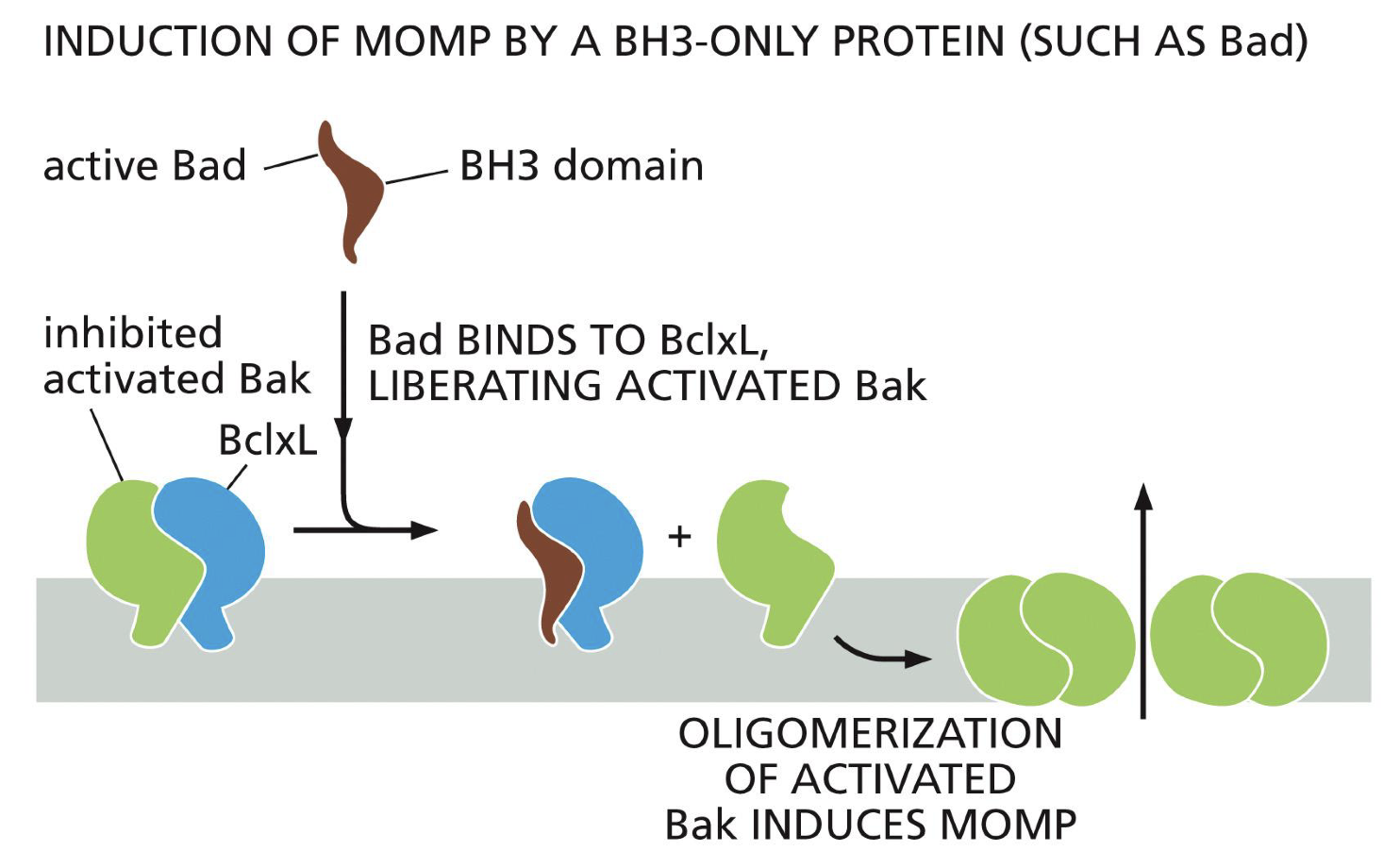
All BCL-2 proteins use BH-3, but what do proteins do with only a BH-3 domain
All BH-3 only proteins stop apoptosis
How is the extrinsic pathway is linked to the intrinsic pathway
By tBID
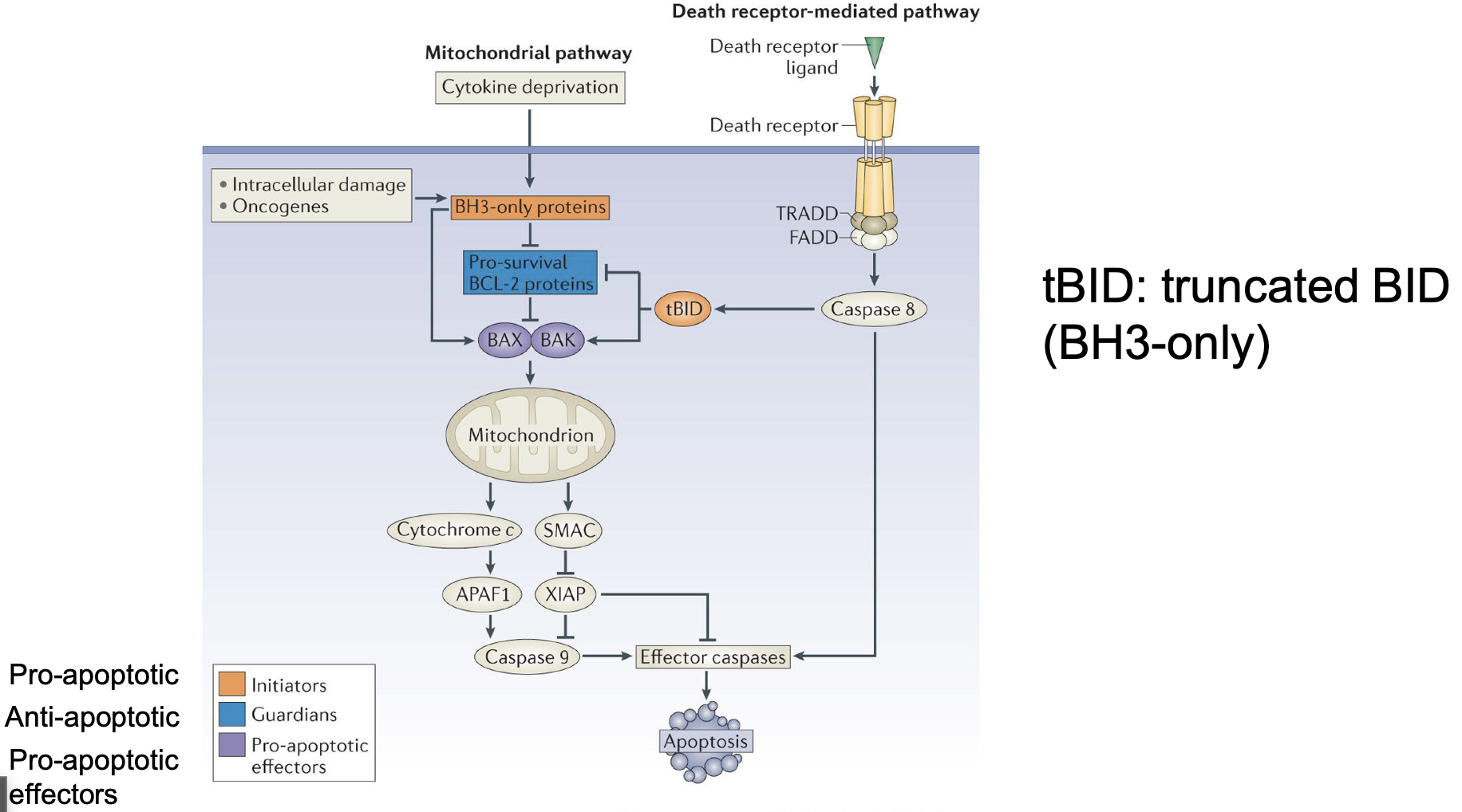
tBID
Represes pro survival BCL-2
Activate BAX and BAK (directly and through anti survival BCL-2)