Evolution of Quantitative Traits
1/23
Earn XP
Description and Tags
Evo Bio Exam 3
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
Quantitative Genetics
the study of continuous traits
Ex:
height
color
speed
growth rate
fitness
the assumption is that these traits are polygenic ( controlled by many loci)
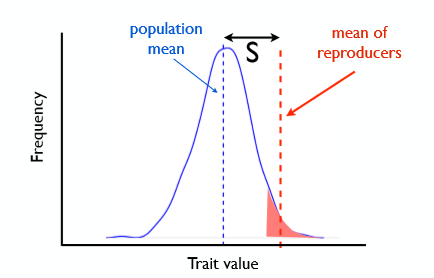
Selection differential (S)
mean trait value of the total population - mean trait value of the reproducers
describes the strength of selection in phenotype units
gives the difference in phenotypic mean of breeders and non-breeders
standardize fitness data (divide all fitnesses by the highest fitness)
steeper the slope = stronger selection or fitness gradient
Response to selection (R)
trait mean before selection - trait mean in the next generation
For a given S, what determines R?
Darwins postulates
trait exhibits variation
trait variation affects fitness
trait variation is heritable
for evolution by natural selection one needs traits that exhibit heritable variation for fitness
Breeder’s equation
R = h²S
the evolutionary response to selection (R) is a product of the heritability of a trait (h²) ( in that population at that time) and the selection differential (S)
Heritability
fraction of the total phenotypic variance in a population due to the genetic variance that causes resemblance between parents and offspring
range = 0-1.0
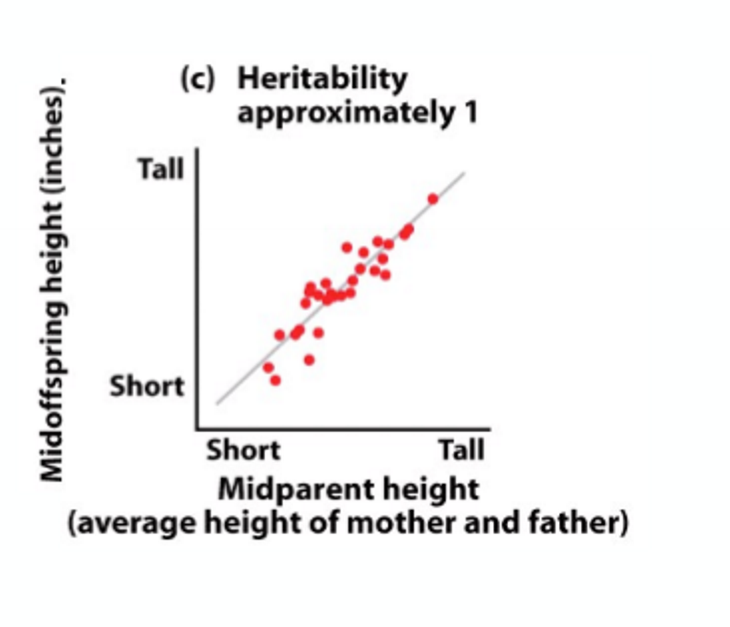
High heritability
high resemblance between parents and offspring
can predict offspring phenotype from parental phenotype
For a given selection differential, the higher the heritability in a population…
the larger the response to selection
2 Sources of phenotypic variation
Genetic (G)
Environmental (E)
Vp = Vg + Ve
phenotypic variance = genetic variation + environmental variances
always sums up to 1.0
Broad sense heritability (H)
Vg/Vp = Vg/ Vg + Ve
Deviations from the regression line may be due to:
Ve (environmental variance)
Non-additive genetic variance
dominance
epistasis
these obscure the relationship between allelic and phenotypic variation, decrease the resemblance between parents and offspring, and thereby impede the response to selection
Three components of genetic variation
V= Va + Vd + Vl
Va = additive variance
Vd = dominance variance
Vl = epistasis variance
Va : Additive variance
alleles are codominant - have additive effects
effects are independent of genetic background
this produces resemblance between parents and offspring and thus contributes to the response to selection
allelic variation consistently associated w/ phenotypic variation
Vd : Dominance variance
allele are dominant/ recessive - nonadditive at the same locus
ex: Aa has same trait value (same phenotype) as AA
V: Epistatic variance (interactive variance)
genetic variance resulting from epistatic (non-additive) interactions between alleles at different loci
effects are dependent on genetic background
allelic variation not CONSISTENTLY associated w/ phenotypic variation
What does epistasis do to the resemblance between parent & offspring?
reduces resemblance by masking or modifying phenotypic expression of genes
How does epistasis affect heritability?
can hide genetic variance?
Heritability complications
environmental effects
non-additive effects
Vp= Va + Vd + Vi + Ve
Narrow-sense heritability
h² = Va/Vp
produces the resemblance between parents and offspring and thus contributes to the response to selection
Additivity
because alleles have independent effects, offspring resemble parents
consistent relationship between alleles and phenotype means there will be a good response to selection
strong environmental effects can cloud the resemblance between parents and offspring (lower Va/Vp), meaning that the relationship between allelic and phenotypic variation is less consistent- lowering heritability and reducing the response to selection
Deviations from the regression line may be due to
Ve (environmental variance)
Non-additive genetic variance
Epistasis
the effect of an allele at one locus depends on the alleles at one or more other loci
add effects of alleles at each locus
multiply effects between loci
Epistasis: Little resemblance between parents and most offspring
Poor correlation caused by:
recombination
independent assortment
non-additivity
Recombination
eliminates LD, decreasing resemblance between parents and offspring by changing the allelic combinations at interacting loci
b/c loci 1 and 2 interact non-additively and the allele combinations at these loci change each generation, offspring do not resemble parents and these cannot contribute to the response to selection