BB 492 Midterm 1 Flashcards
1/163
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
164 Terms
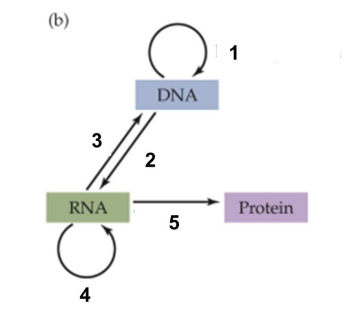
Central dogma
flow of biological information in these routes
replication
transcription
reverse transcription
RNA replication
translation
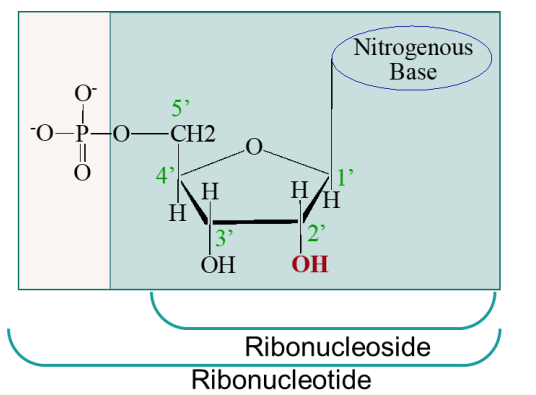
ribonucleotide vs ribonucleoside
ribonucleotide also includes the phosphate attached to 5’ C
what do nucleotides do?
roles in:
macromolecule metabolism
coenzymes
regulation
information
energy currency
which carbon is it oxygen/deoxygenated?
2’
pyrimidine
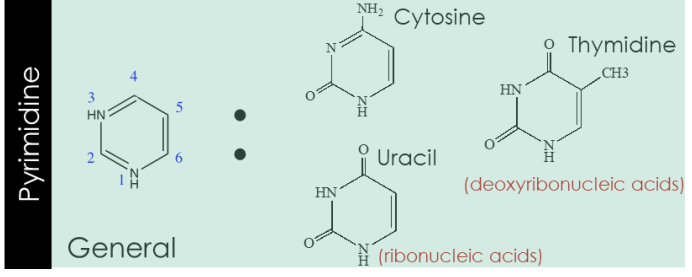
purine
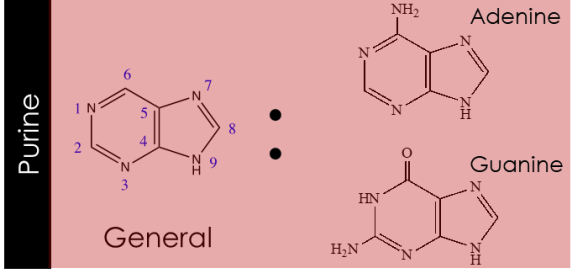
due to electron rich nature of purine and pyrimidines
they can tautomerize, but there will be a dominant form under physiological pH
in nucleobases what are the H donors and H acceptors
H donors: amino groups
H acceptors: nitrogen ring and oxygen
How many H bonds between the different nucleic acids?
A:T =2
G:C =3
key conventions with deoxyribonucleotides and ribonucleotides
all phosphodiester links have the same orientation → nucleic acid polarity
the 5’ end lack a nucleotide at the 5’ end, and the 3’ end lacks a nucleotide attacked to the 3’ of the ribose
5’ → 3’ refers to the ends of the strand orientation of individual nucleotides it doe NOT affect the orientation of particular phosphodiester bond
• How is base pairing of nucleotides determined?
complementarity: Only combinations of A w/ T and G w/ C allow for the alignment of "donors" (atoms with a hydrogen) and "acceptors" (electronegative atoms). If A tried to pair with C, their hydrogen bonding groups would clash or fail to align.
What is the purpose of DNA in the central dogma?
the purpose of DNA is to act as the primary, stable storage system for an organism's genetic information
why is uracil used in RNA and thymine in DNA?
DNA uses thymine because its chemical structure allows the cell to easily detect and repair mutations. RNA uses uracil because it is significantly cheaper to produce, and since RNA is short-lived
secondary structure of DNA
While the primary structure is simply the linear sequence of nucleotides (A, T, C, G),
the secondary structure = base-pairing interactions and stacking that give the molecule its three-dimensional shape.
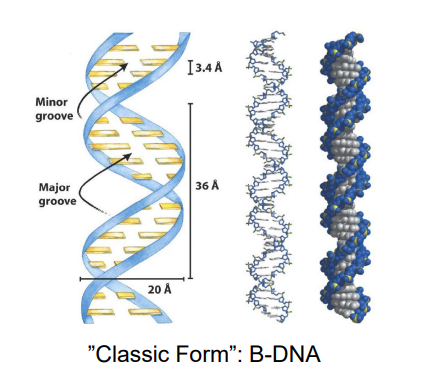
DNA’s Structure “Classical Form”: B-DNA
two antiparallel strands that wind in right handed manner around common axis
the bases occupy the CORE (of the helix), the sugar phosphate back bone makes up the outside and form the major and minor grooves
each BP has same width = symmetrical
~10BP/turn w/ a helical twist of 36 degrees per BP. They are stacked tightly on each other with a helix pitch (rise per turn of 34 A)
A-DNA
rare
can only find in dehydrated state
differs from B-Form by 20 degrees of rotation of helix perpendicular to the axis
shows a deep major groove and flat minor groove
Z-DNA
forms left handed helix
greater distance between BP’s vs B-DNA
may occur in limited segments in vivo
transient (only lasts a short time)
specific proteins for Z DNA
biological function is still unclear…
RNA will adopt A-from like confirmations!!
A-RNA (RNA-II)
•11 basepairs per turn
• Helical pitch of 30.9 Å
• Base inclination of 16.7°
DNA-RNA hybrids also have A-form-like conformations
• 10.9 basepairs per turn
• Helical pitch of 31.3 Å
• Base inclination of 13.9°
how does RNA take on multiple secondary structures?
do complementary intra-strand base pairing to achieve these structures
what’s a common secondary RNA structures?
hairpin!!
What three forces stabilize DNA’s strucutre?
H-bonding (watson kirk base pairing)
stacking interactions (van der Waals Forces
cationic shielding
How does cationic shielding work
DNA, which is very negative because of the phosphate groups, is stabilized by Na+
Li+, K+, and Mg2+ can also help to stabilize it.
conditions favoring denaturation
high temperature
low salt concentration (remember this does shielding)
high pH (basic)
More of what bonds increase Tm (melting temperature)
G and C
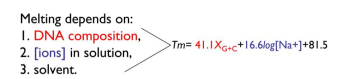
Melting depends on:
DNA composition (amount of G+C)
ions in solutions
solvent
Why is DNA flexibility important?
for the sequence-specific recognition of DNA by the proteins that process genetic infromation
Types of DNA flexibility
phoso-ribose segments have 6 degrees of freedom
note internal constrains that affect angles reducing rotational freedom
ribofuranose ring puckers to relive crowding of the ring substituents, this out of plane atom usually goes endo to the 5’C
nucleobase can rotate between syn and anti confirmation, but note this rotation is greatly hindered
Chromosomal DNA is often orders of magnitudelonger than the cells or viruses housing it!; how can cells package it?
DNA’s ability to coil about itself for compaction
what causes supercoiling?
underwinding (removal of some turns), DNA is kinda like fighting to return to its relaxed state;
you can also do overwinding, but less common
negative supercoiling
twisting against the helical conformation (twisting in a left-handed fashion)
which preferentially underwinds and "straightens" the helix at low twisting stress,
knots the DNA into negative supercoils at high twisting stress
positive supercoiling
twist the DNA even tighter, (in right handed fashion, until helix begins to distort and knot
Linking number (Lk)
Lk = the sum of all intersections made by 1 DNA strand across the surface carved by another DNA strand; think of like a chain, and number of links in the chain is the linking number.
If either strand of a closed circular DNA has a break, can it have a linking number?
No, if either strand of a closed circular DNA has a break (a "nick"), it does not have a linking number (Lk).
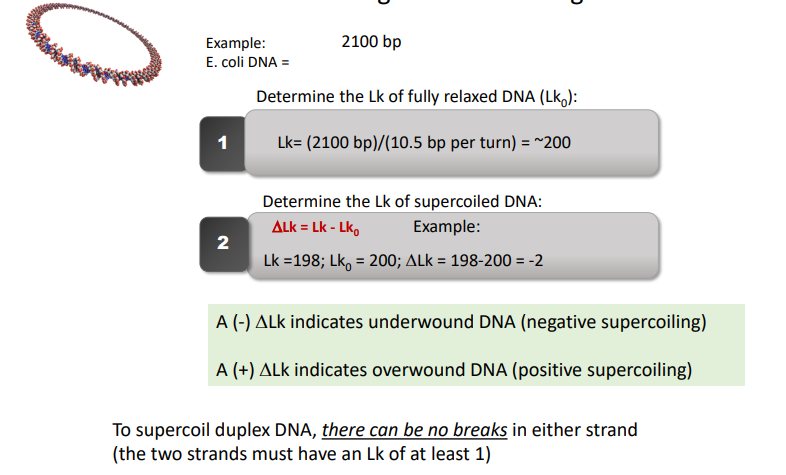
relaxed Lk equation
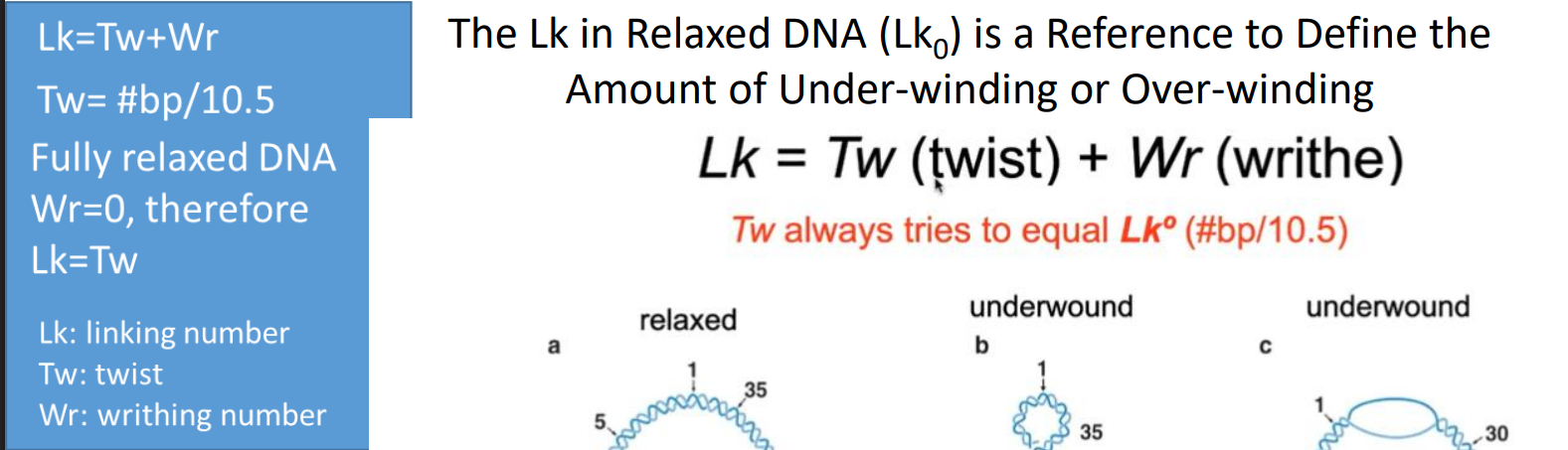
Calculate the approximate linking number of the following E. coli DNA (10,500 bp) in its fully relaxed state.
If the DNA in question 22 is under-wound by a type II topoisomerase called gyrase what is its
new linking number?
Why do we like underwinding (aka negative supercoiling so much)
Negative supercoiling has been selected for by evolution so that it is very similar for all organisms.
The resulting strained state of the DNA represents a form of stored energy.
The underwinding of DNA in vivo makes it easier to separate DNA strands and thereby gain access to the information they contain.
Facilitating strand separation is one important reason for maintaining DNA in an underwound state.
Superhelical Density Equation
σ = Wr/Tw
• Negative σ means DNA is negatively supercoiled
• Positive σ means DNA is positively supercoiled
• DNA from bacteria and eukaryotes, s = -0.06
ΔLk Equation
A (-) ΔLk indicates underwound DNA (negative supercoiling)
A (+) ΔLk indicates overwound DNA (positive supercoiling)
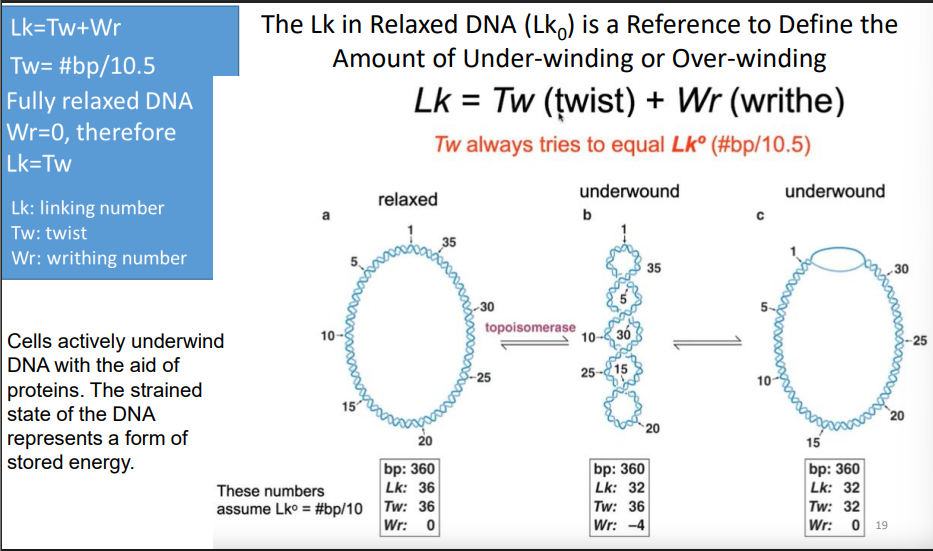
Lk equation
Lk=Tw(twist)+ Wr(writhe)
topoisomerases
enzymes that increase or decrease the extent of DNA supercoiling
Topoisomerase I (Topo I):
Changes the linking number by 1:
The enzyme transiently breaks one of the DNA strands and passes the unbroken strand through the break.
Topoisomerase II (Topo II):
Changes the linking number by 2
Transiently breaks both DNA strands and passes the unbroken portion through the break.
What catalyzes the relaxation of supercoiled DNA
Topo I does the actual cutting; other important factors at play:
Mg2+ -requiring monomeric proteins that have a hinge region, which can fold to create a 20 x 28 angstrom hole
Duplex DNA with negative supercoils are substrates The enzyme changes the Lk by 1
Critical Tyrosine residue that can attack the phosphate bond of the deoxynucleotide strand
Effect of a type 1 topoisomerase on supercoiling
one less supercoil
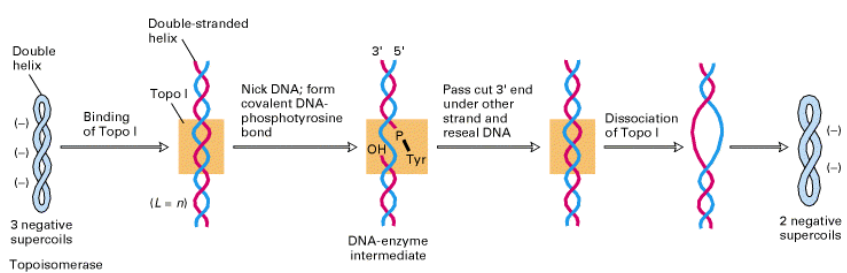
Type I A topoisomerases
function by a strand-passage mechanism. A transient phospho-tyrosine bond (to the 5’-terminal phosphoryl group) is formed. No additional energy is required to reseal the nick.
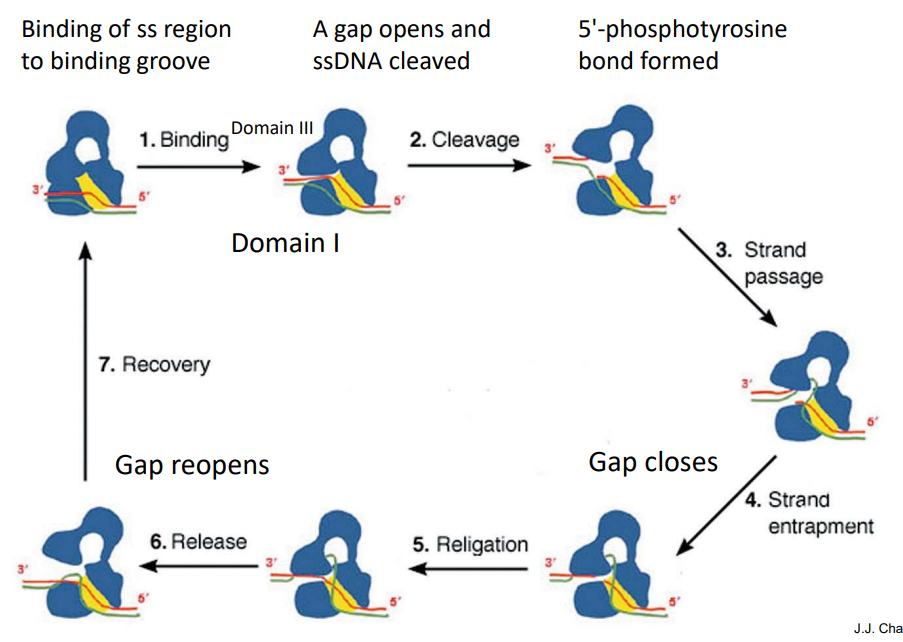
Type I B topoisomerases
Functions by controlled rotation mechanism. A transient phosphodiester bond with tyrosine (to the 3’-terminal phosphoryl group) is formed. The super-helical tension drives the rotation, so no additional energy is required.
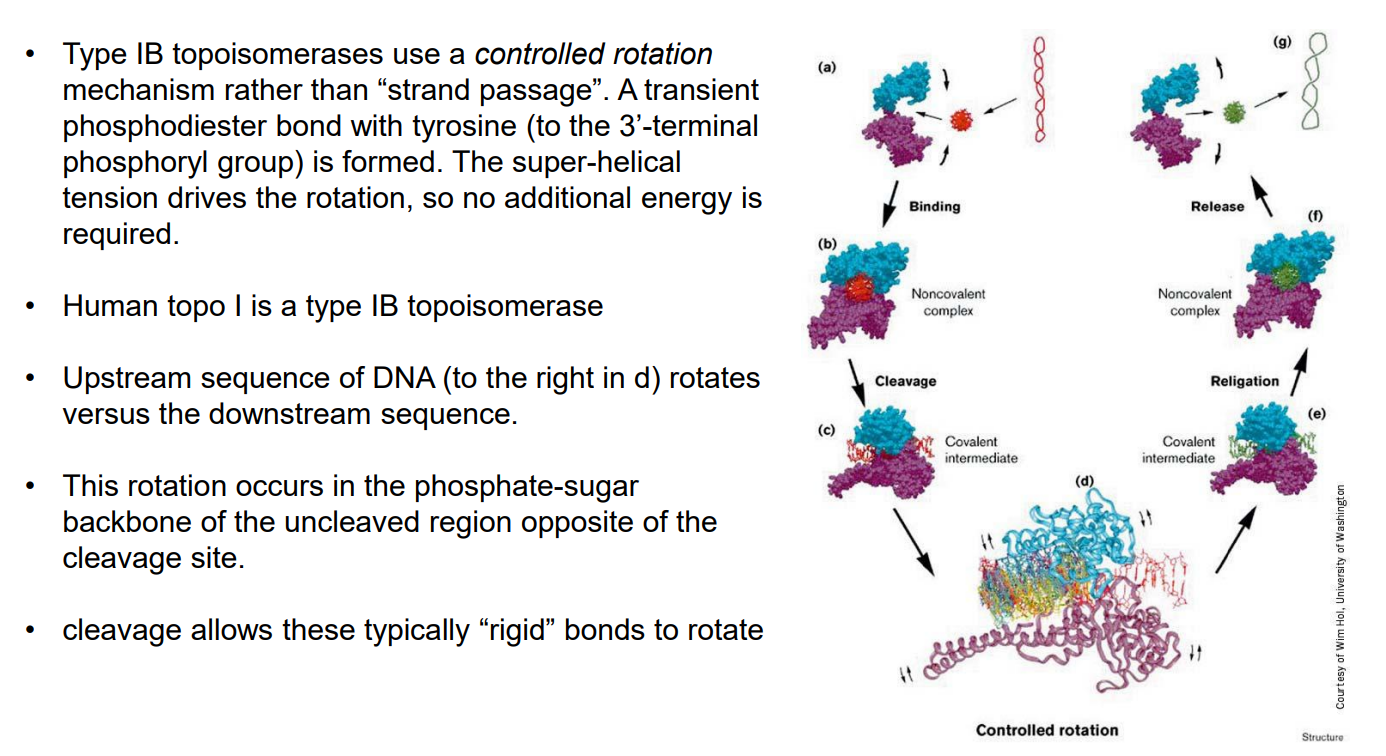
Topoisomerase II (Topo II)
Multimeric (multi-polypeptide) proteins
Requires ATP hydrolysis
Requires Mg2+
Changes the Lk by 2
Can relax (-) and (+) supercoils (eukaryotes)
Which topoisomearase introduces negative supercoils?
DNA gyrase (a Type IIA prokaryotic topoisomerase) is the primary enzyme that introduces negative supercoils into DNA. It requires ATP to do so. All other type II topoisomerases will still hydrolyze ATP, but can only relax supercoils
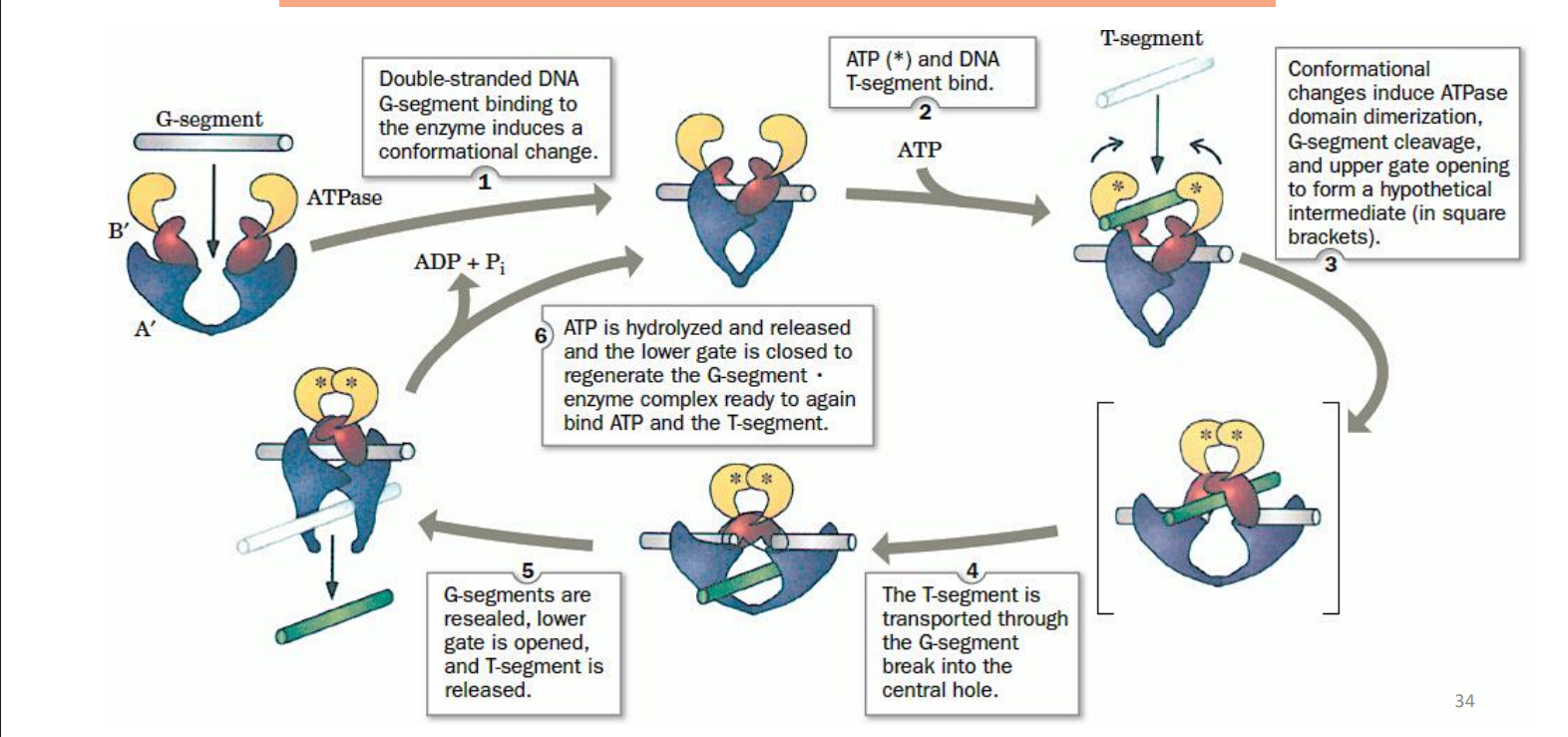
Mechanism of TOP II
doubled stranded DNA and the G-segment binding to the enzyme induces a conformational change
ATP and DNA T segment bind
confirmational changes induce ATPase domain dimerization, G-segment cleavage, and upper gate opening to form “intermediate”
the T-segment goes through the G-segments break into the central hole
G-segments are resealed, lower gate is opened, and T-segment is released
ATP is hydrolyzed and released and the lower gate gets closed to regenerate the G segment x enzyme complex ready to again bind ATP and T-segment
Why are topoisomerase inhibitors effective anti-cancer drugs?
DNA cannot be replicated once it has been nicked!
example: Camptothecin (quinoline alkaloid) and its derivatives inhibit type IB topoisomerases by stabilizing the topoisomerase I-DNA complex
We can also kill the cell!
function by changing the rate at which type II topoisomerase cleaves dsDNA and/or at which the enzyme reseals the breaks. Breaks become permanent, thus killing the cell.
examples include: Ciprofloxacin, doxorubicin, and etoposide
What are two important functions of supercoiling DNA?
compacting the DNA to fit into the genome and regulating gene expression
Chromatin
helps with stability in supercoiling of DNA and further packs it as well
allows DNA to be condensed but in a way that allows access to genetic information
predominant protein in chromatin
Histones:
H1
• H2A
• H2B
• H3
• H4
Nucleosome
Histone +DNA wrapped around it
characteristic of histone proteins tells us that they are very important
HIGHLY conserved across different organism… haven’t changed much throughout evolution.
histones are modified by
phosphorylation
methylation
acetylation
Nucleosome structure
histones form an octamer (2 each of H2A, H2B, H3 & H4)
H1 is peripheral relative to core
histone is highly positive because rich in lys and arg
this allows tight binding of phophoribosal backbone of DNA (very negative)
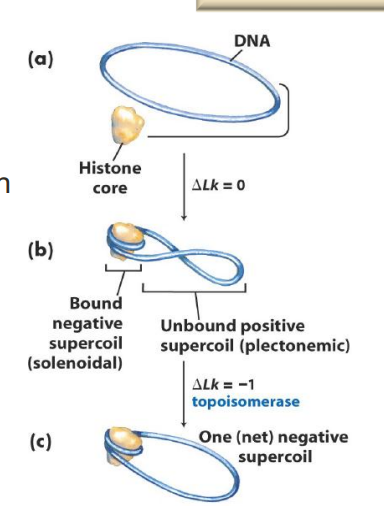
How do topoisomerases facilitate wrapping of DNA around histones?
Negative supercoiling facilitates DNA association with histones
BUT this causes more strain (+ supercoiling) in the remainder of the DNA
Topoisomerases come in and cause net negative supercoil… relieving the strain
Heterochromatin
densely packed and inactive
histone modification can alter chromatin in this way
Euchromatin
less densely packed and active
histone modification can alter chromatin in this way
Describe the different levels of DNA compaction in eukaryotic cells. What is the approximate packaging ratio?
Levels:
nuceleosome
chromatin fiber irregular zig zag
interdigitated polymer melt packaging
compact chromatin domain
interphase chromosome
territory in the nucleus
Compaction maintains DNA organization; Packaging ratio: >8000-fold.
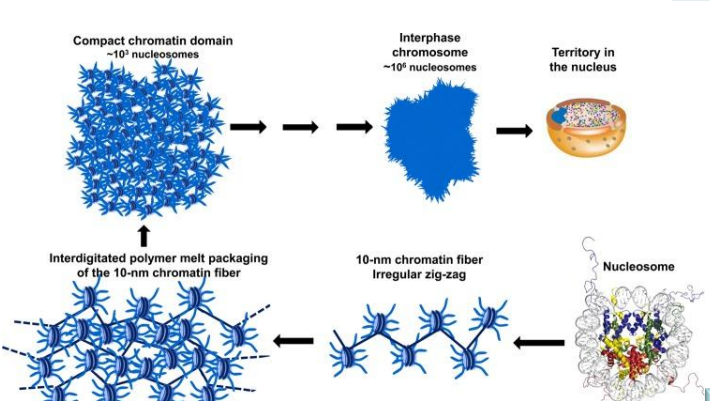
What is the new model for the folding and packing of DNA in eukaryotes?
It is hypothesized that the hierarchical globular organization of an interphase chromosome is fundamentally established by the self-interacting properties of a 10-nm zig-zag array of nucleosomes, while histone post-translational modifications, histone variants, and chromatin-associated proteins serve to mold generic chromatin domains into specific structural and functional entities.
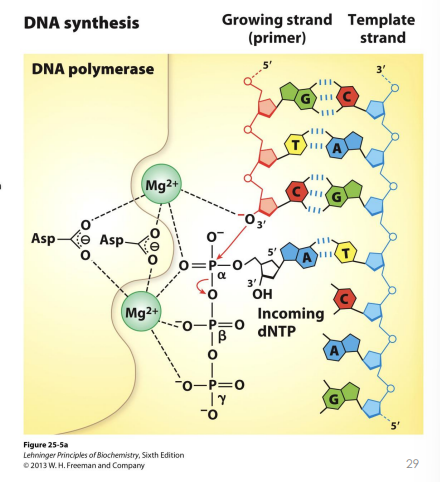
General Catalytic Mechanism of DNA Polymerases
A single stranded DNA template acts as a substrate along with an incoming dNTP
Mg2+ ions coordinate the phosphate groups and Asp residues
Direction of DNA Polymerase
synthesize DNA in the 5’ → 3’ direction but then proofreads/does a backspace in 3’→5’ activity (this is our exonuclease activity in case there is mis-incorporation
Polarity of the Template
The template strand has a 3’ → 5’ polarity relative to the direction of the replication fork's movement. Because DNA is antiparallel, the polymerase must "read" the template in the 3’ → 5’ direction so that it can "write" the new strand in the required 5’ → 3’ direction
What sort of reaction occurs on the growing chain?
The free
3’-hydroxyl (-OH
) group of the growing chain attacks the alpha-phosphate of the incoming deoxynucleoside triphosphate (dNTP).
This reaction releases a molecule of pyrophosphate (PPi
).
The subsequent breakdown of that pyrophosphate into two inorganic phosphates provides the energy (enthalpy) that makes the polymerization reaction irreversible.
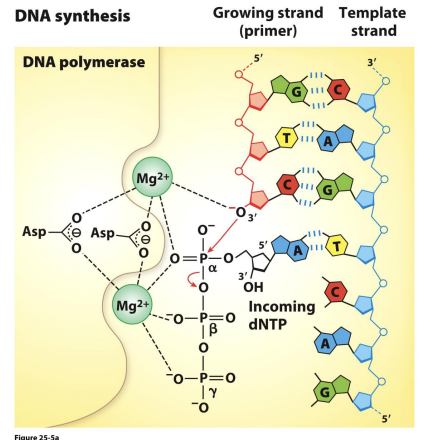
General Characteristics of DNA Synthesis
• Each original DNA strand acts as a template for DNA replication.
• DNA is always synthesized in the 5’→ 3’ direction.
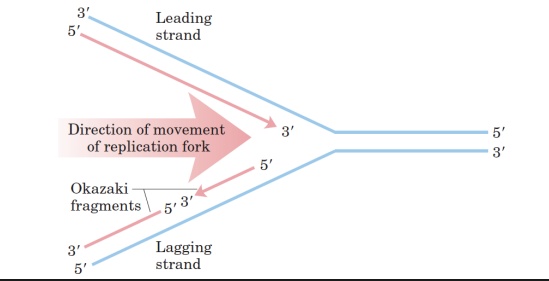
• One strand (leading strand) is continuously synthesized and proceeds (5’→3’) in the direction of the replication fork.
• The other strand (lagging strand) is synthesized in pieces, called Okazaki Fragments and then ligated together. Therefore, the two strands are synthesized semi-discontinuously!
• Lagging strand synthesis occurs in the opposite direction of the replication fork.
• The Okazaki fragments are later covalently joined by DNA ligase.
Stages for DNA replication
initiation
elongation and proofreading
termination
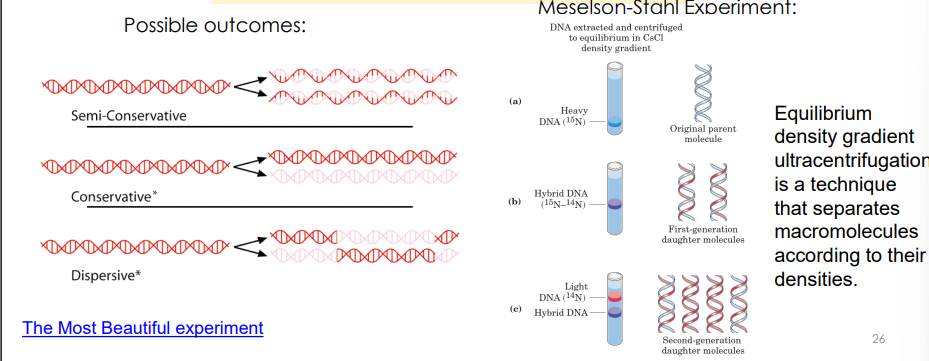
How did Meselson and Stahl demonstrate that replication of DNA was semi-conservative?
semi-conservative by using nitrogen isotopes to track parent and daughter DNA strands.
DnaA protein
recognizes the ori sequence; opens duplex at specific sites in origin
DnaB protein
unwinds DNA
DnaC protein
required for DnaB bindsing at origin
51. For prokaryotic DNA replication, what did the presence of theta structures tell researchers?
showed bidirectionality of replication
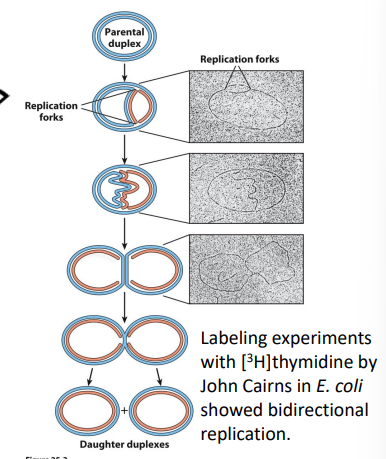
All polymerases exhibit:
5’ → 3’ Polymerization (Synthesis) and 3’ → 5’ Exonuclease (Proofreading);
DNA Pol I is special because it also has: 5’ → 3’ Exonuclease (Primer Removal)
DNA Polymerase III
The Main Replicase
DNA Pol III is the primary enzyme responsible for synthesizing the bulk of the new DNA strands.
Continuous Synthesis: It extends the leading strand continuously toward the replication fork.
Discontinuous Synthesis: It extends the lagging strand in short segments called Okazaki fragments.
High Processivity: It remains attached to the DNA for long periods, adding thousands of nucleotides without falling off. This is facilitated by a sliding clamp protein.
Proofreading: It features 3’ →5’
exonuclease activity, allowing it to "backspace" and remove mispaired nucleotides.
DNA Polymerase I
The Cleanup Crew
DNA Pol I acts as a supporting enzyme that handles "cleanup" tasks at the replication fork.
Primer Removal: It uses a unique 5’ → 3’ exonuclease activity to remove the RNA primers used to start synthesis.
Gap Filling: Once the RNA is removed, its polymerase activity fills the resulting gaps with the correct DNA nucleotides.
DNA Repair: It is also heavily involved in various DNA repair pathways, such as nucleotide excision repair
DNA Polymerase II
The Backup/Repair Specialist
DNA Pol II is primarily a repair enzyme rather than a main replicase.
Replication Restart: Its most critical role is restarting DNA replication if DNA Pol III stalls due to damage (like UV-induced lesions) on the template strand
High Fidelity Repair: It has 3’→ 5’
exonuclease activity for proofreading and is highly accurate, often acting as a backup for Pol III to maintain genome stability.
SOS Response: Its levels in the cell increase significantly during the SOS response (the cell's emergency repair mode).
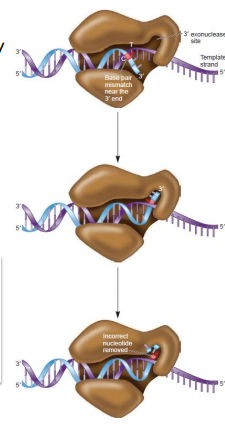
How do DNA polymerases with proofreading activity correct deoxyribonucleotide misincorporation?
Insertion of a wrong base by DNA Pol doesn’t allow normal Watson-Crick base pairing, causing the enzyme to pause
The 3’ end of the growing strand is shunted to an exonuclease site
The strand is hydrolyzed in the 3’ →5’ direction until the wrong base is removed
The strand moves back to the polymerization site
Primase (DnaG protein)
synthesizes RNA primers
single stranded binding proteins (SSB)
binds single stranded DNA
DNA gyrase (DNA topoisomerase II)
relieves torsional strain generated by DNA unwinding
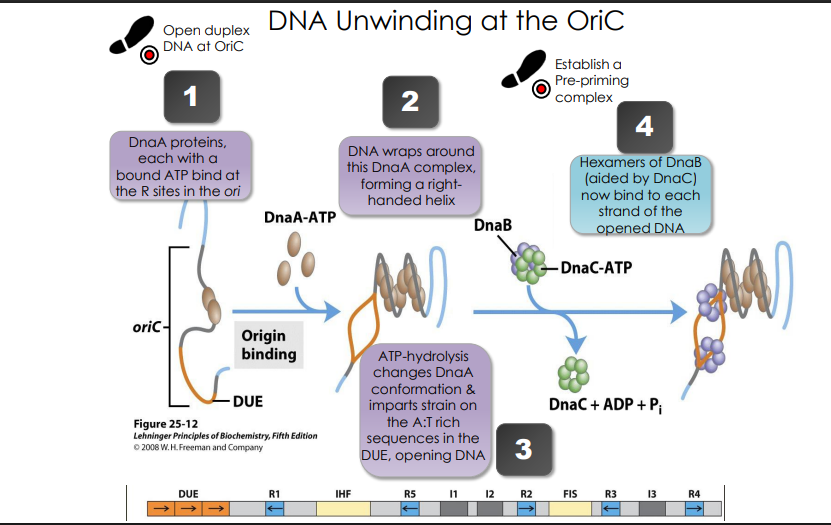
DNA Unwinding at the OriC
DnaA proteins, each with a bound ATP bind at the R sites in the ori
DNA wraps around this DnaA complex, forming a righthanded helix
ATP-hydrolysis changes DnaA conformation & imparts strain on the A:T rich sequences in the DUE, opening DNA
Hexamers of DnaB (aided by DnaC) now bind to each strand of the opened DNA
Mechanism of DnaB Hexamer
Hexamers of DnaB create a helicase. Mechanical movement supplied by ATP hydrolysis moves the helicase into the replication fork, forcing the two DNA strands apart in the process.
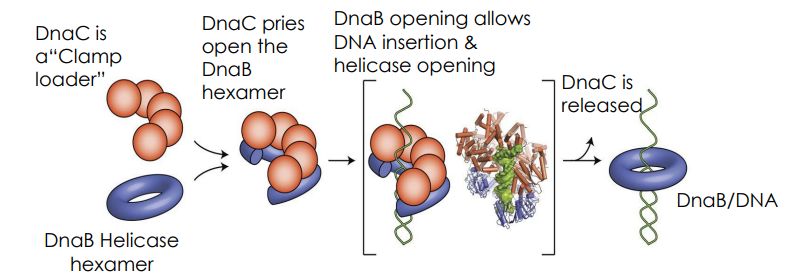
Once the duplex is open, what steps occur to generate a pre-primosome
DnaA opens up the duplex
DnaC pries open DnaB
DnaB hexamers attach to each strand of open DNA
DnaG associated with DnaB on both separated strands (+ five other proteins), creating a primosome
the primosome works with primase to make RNA primer
once the primer has been created, DNA Pol II can then add dNTPs to the free 3’ OH of the primer
Semidiscontinuous replication in lagging strand
occurs because RNA primer formation is more complicated here
The lagging strand template loops out to accommodate the primase laying down primers!
Primosome is propelled in 5’ → 3’ direction of the lagging strand (toward fork)
this is the OPPOSITE of the template reading direction
so primose reverses migration for a moment so primase can make RNA primer in 5’ →3’ direction
Note: The primosome is required to initiate each Okazaki fragment. It lays down 11 nt primers approximately every 1000 nts
Replisome
In prokaryotes, the Replisome is the total collection of all the proteins working together at the replication fork. This includes:
DNA Pol III Holoenzyme (usually two or three of them working together).
Helicase (DnaB).
Primase (DnaG).
SSBs (Single-strand binding proteins).
The Clamp Loader.
The DNA components are as follows:
Template Strands: The original "parent" DNA strands being copied.
RNA Primers: The short nucleic acid starters needed for Pol III to begin.
Okazaki Fragments: The short pieces of DNA built on the lagging strand.
Leading and Lagging Strands: The two newly synthesized strands.
Pol III Holoenzyme
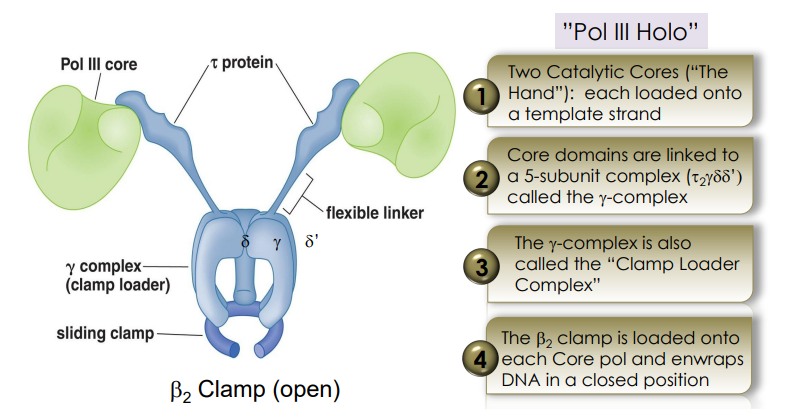
beta-clamp
part of Pol III Holoenzyme
Holds Pol III onto the DNA to keep it from falling off.
gamma complex
loads the beta-clamp onto the DNA
Which strand requires frequent loading of a new beta-clamp during replication?
lagging!!
Klenow fragment
a large protein fragment produced when DNA Polymerase I is cleaved by a protease.
It contains the 5’ → 3’ polymerase and the 3’ → 5’ exonuclease (proofreading) activities, but loses the 5’ → 3’ exonuclease activity
How are nicks in DNA repaired?
Nicks are sealed by the enzyme DNA Ligase, which creates a phosphodiester bond between the 3’ -OH of one fragment and the 5’ -phosphate of the next.
What molecule provides free energy for the ligase reaction in prokaryotes?
NAD+
What molecule provides free energy for the ligase reaction in eukaryotes?
ATP
The beta clamp isn’t perfect. how do we fix this?
on leading strand it will fall off after 500,000 nucleotides
on the lagging strand when it encounters the next primer pol III must let go so it can hop back to the next newly synthesized primer further up the fork (called polymerase cycling)
solution: the Clamp Loader uses ATP to open a new beta-clamp ring, slap it onto the DNA at the primer site, and recruit the Pol III core back to work.
DNA Replication Mechanism
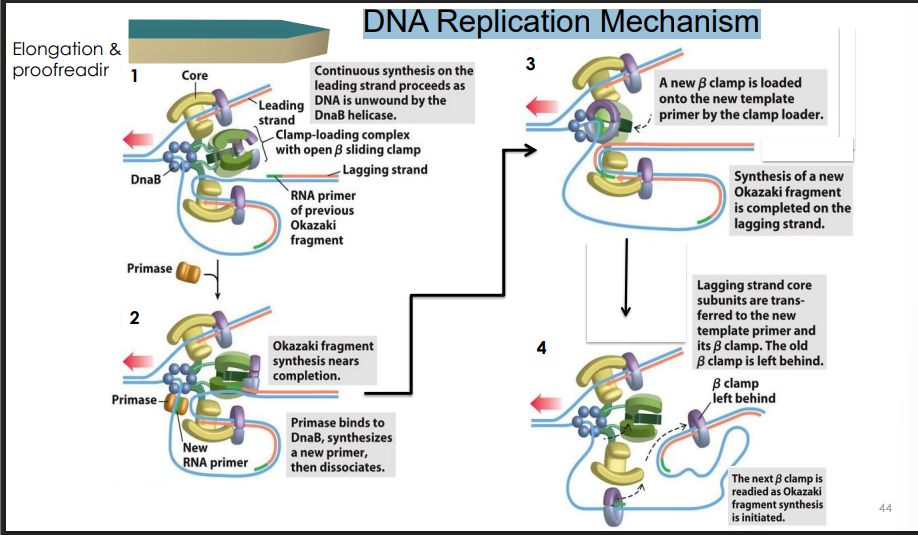
Nick translation overview
a nick is the gap between the end of a newly built DNA fragment and the start of the RNA primer of the fragment in front of it. There is a break in the "backbone" of the DNA strand here.
The Process: "Nick Translation"
Think of DNA Pol I like a snowplow that paves the road at the same time:
The Plow (5’ → 3’
Exonuclease): Pol I lands at the nick and chews up the RNA primer in front of it, one nucleotide at a time.
The Paver (5’ → 3’
Polymerase): As it removes an RNA nucleotide, it immediately replaces it with a DNA nucleotide, attaching it to the
end of the fragment it's sitting on.
The "Translation": Because Pol I is removing bits from the front and adding bits to the back, the "gap" (the nick) physically moves (translates toward the DNA’s 3’ end) down the strand until all the RNA is gone.
Once Pol I finishes replacing the RNA with DNA, it falls off. However, the backbone is still not connected—there is still a tiny break (a nick) between the new DNA and the old DNA. DNA Ligase comes in last to "glue" that final bond shut.
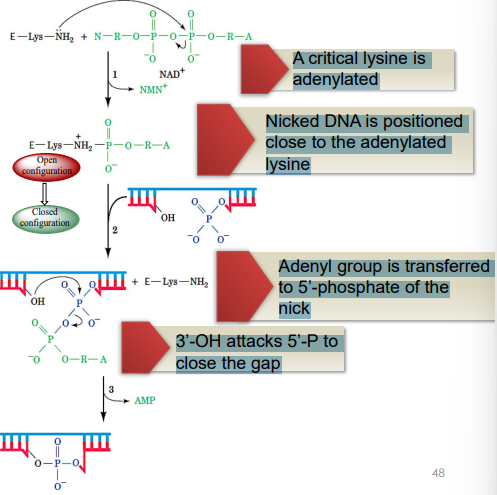
Process of DNA Ligase in Filling Nicks
has a conserved hydrophobic pocket for NAD or ATP to bind (depending on the species) & a cleft for DNA binding
A critical lysine is adenylated
Nicked DNA is positioned close to the adenylated lysine
Adenyl group is transferred to 5’-phosphate of the nick
3’-OH attacks 5’-P to close the gap