BIO150 Final
1/123
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
124 Terms
Protein
Monomer:
Covalent Bonds:
Non-Covalent Bonds:
Key Roles:
Amino Acids
Peptide Bonds
H-Bonds, ionic, hydrophobic, Van Der Waals, disulfide interactions
Enzymes, large structures, signaling, connecting cells
DNA/RNA
Monomer:
Covalent Bonds:
Non-Covalent Bonds:
Key Roles:
Nucleotides
Phosphodiester Bonds
H bonds (b/w bases) & Van Der Waals (Base Stacking)
Info storage, gene control, protien production
Carbohydrates
Monomer:
Covalent Bonds:
Non-Covalent Bonds:
Key Roles:
Monosaccarhides
Glycosidic Bonds with a/B linkages
H-bonds, Van der Waals
short-term energy (glycogen & starch), structural support (cellulose & chitin), cell recognition
Lipids
Monomer:
Covalent Bonds:
Non-Covalent Bonds:
Key Roles:
Glycerol and Fatty Acids
Ester Bonds w/ triglycerides & phospholipids
Hydrophobic interactions & Van der Waals
Energy Storage, Membranes, & Signaling
Condensation Reaction
Forming Macromolecules with water as a byproduct
Hydrolysis
Breaking down macromolecules by adding water
Central Dogma
DNA -(transcription)→ RNA -(translation)→ Protiens
Prokaryotic protein production
Speed is the priority; transcription and translation happen in the cytoplasm; transcribed genes are translated immediately
Eukaryotic protein production
Fine control is the priority; transcription in the nucleus; translation in the cytoplasm; transcription and translation are independently regulated
Transcription
How RNA is made; DNA copied into RNA by complementary base pairing; forms raw RNA, which must be edited; many RNAs are never translated into proteins but proteins are the final working molecule
Complementary Pairing
Complementary bases prefer to H-bond with each other; A-T (U in RNA) has 2 H-bonds and G-C has 3 H-bonds
Primary Structure
Amino acid sequence, covalent bonds between peptide sequence
Secondary Structure
Alpha helix and beta pleated sheets. A-helices have right-handed (most common), coiled-coil, amphipathic, Beta sheets have parallel, antiparallel (most stable), mixed. Different amino acids prefer to bond with different structures. They bond by forming H-bonds between the backbone atoms
Tertiary Structure
R group interactions, non-covalent interactions with some covalent bonds, are affected by environmental conditions (pH, ions, temperature, chemical modifications)
Quaternary Structure
Multiple Subunits; folding driven by non-covalent interactions between tertiary subunits
Translation
Proteins fold as they are translated. Evolution favors fast folding; the final shape is thermodynamically favored but not guaranteed. Chaperones assist folding by preventing aggregation, providing a safe environment, and using ATP-driven cycles to give proteins multiple chances to reach their proper structure. The N-C-C backbone of a misfolded protein attracts chaperones
Directionality of protiens
-N-C-C- Chain in center, polarity, gives directionality to entire macromolecule
Disulfide Bonds in proteins
between cysteines, connect two different parts of a chain
Van der waals in proteins
complementary shapes bind together
H-bonds in proteins
between side chain and backbone or between side chain and another side chain
Hydrophobic interactions in proteins
R groups avoid water and aggregate
Carbohydrate structure: monomers
monomers: carbon backbones with multiple OH groups and one double-bonded O group. Can spontaneously form a ring (-O-)
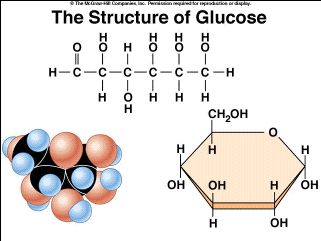
Carbohydrate Structure: Polymers
disaccharides and polysaccharides; glycosidic bonds link because of condensation reaction, they form between 2 -OH groups
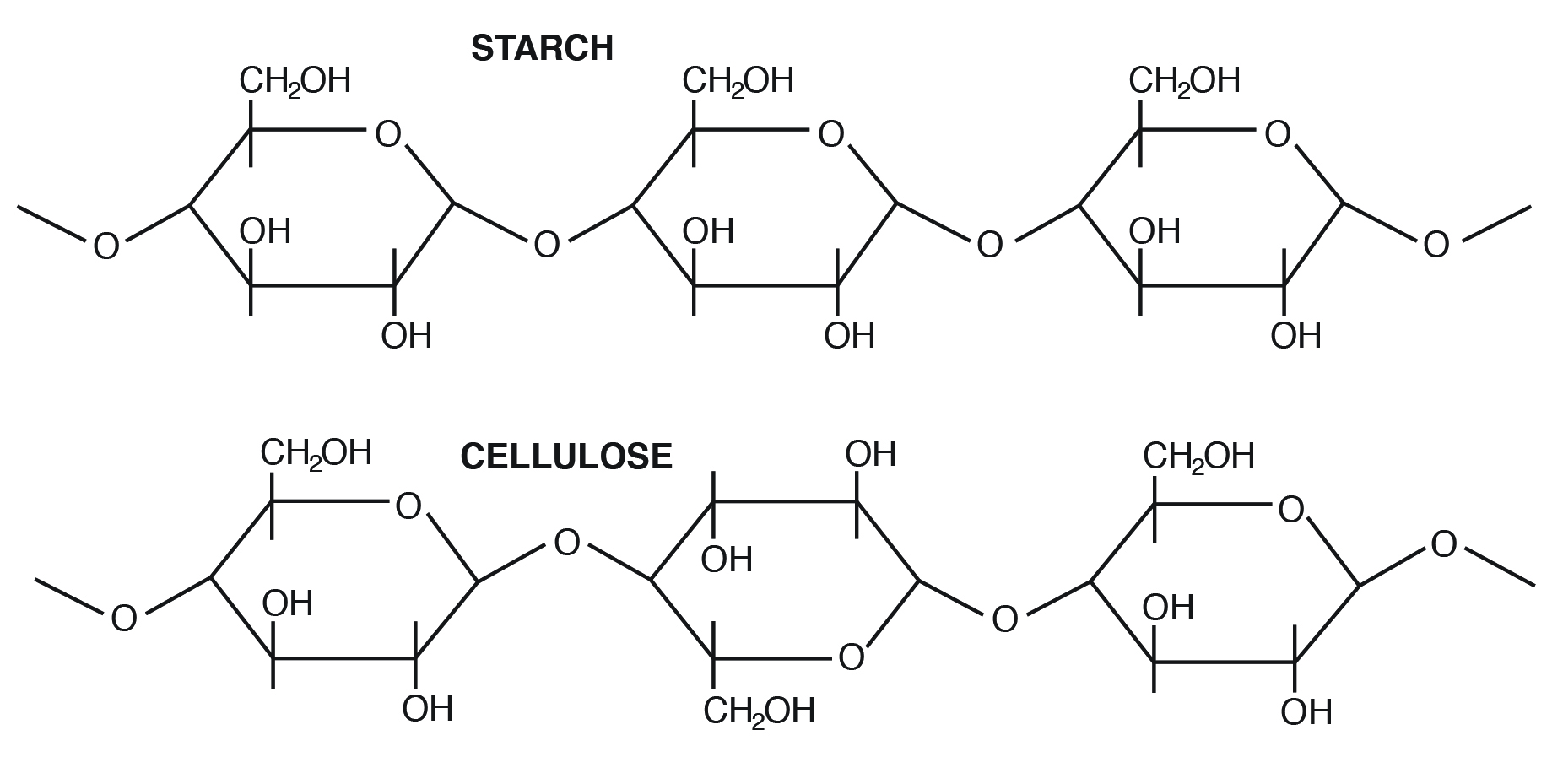
Small sugars are used for:
Bulk Transport because they are a source for rapid release energy (ex: glucose)
Polysaccharides
They form long fibers (ropes), strands have covalent and non-covalent interactions, used for storage and structural support (ex: cellulose)
Charged polysaccharides
attract water, form a cheap space filling mass (ex: pectin in plant cell wall, mesoglea in jellyfish)
Fatty Acids
Carboxyl head group (-COOH) → Charged; nonpolar (C:H) tails that differ in length and number of double bonds (saturation)
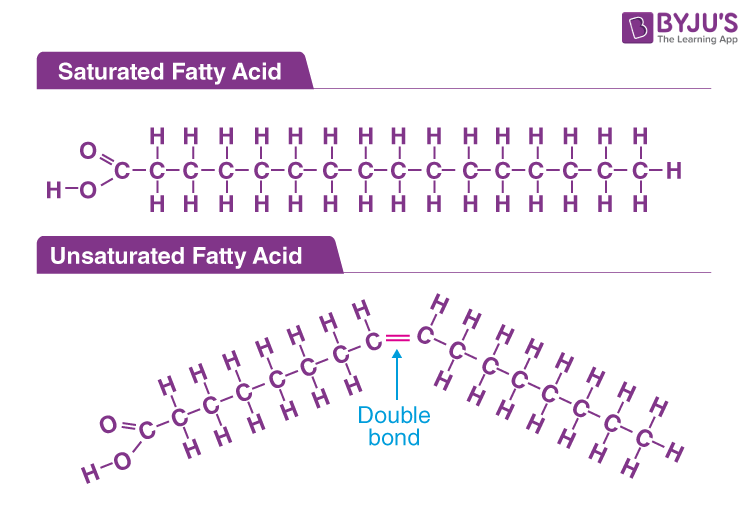
Triglycerides
Also called triglycerols (TAGs), 3 fatty acids linked to glycerol via ester linkage, hydrophobic
Functions: energy storage, physical padding, insulation
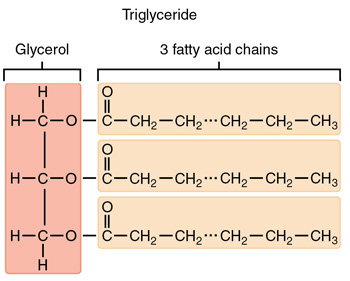
Phospholipids
2 fatty acid chains linked to glycerol, phosphate linked to 1 of 4 possible head groups, they are amphipathic
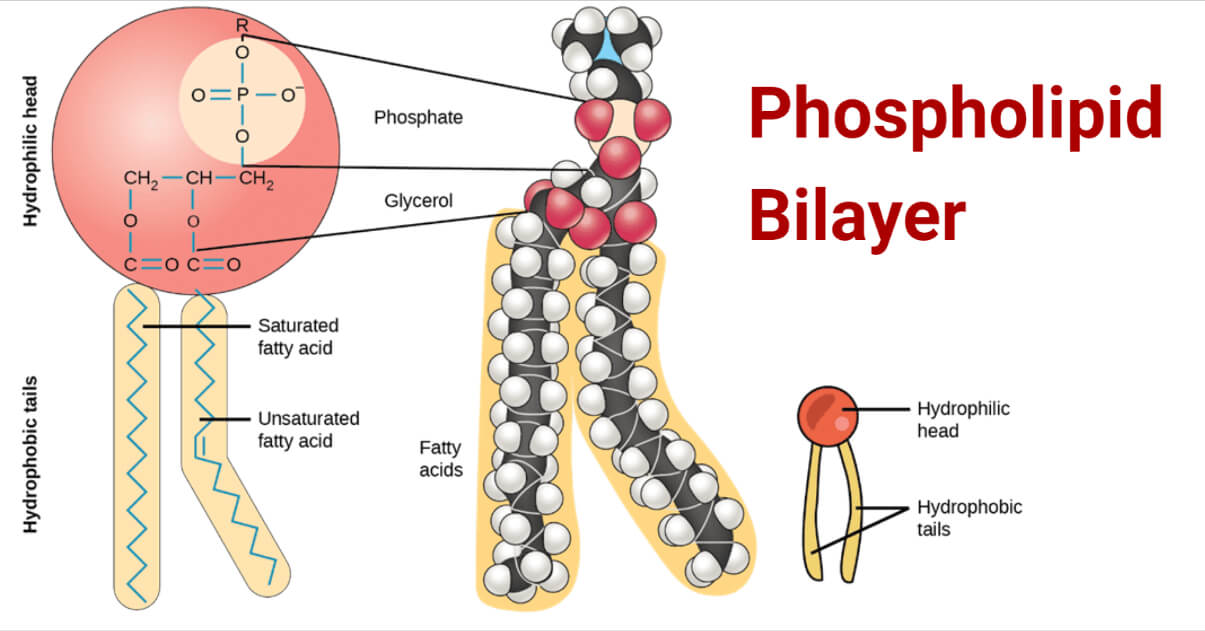
Steroids
3 6-Carbon rings and 1 5-Carbon ring, side chains, and Carbon Carbon double bonds that modify function.
Function: membrane fluidity (cholesterol) and signaling molecules (testosterone, estrogen)
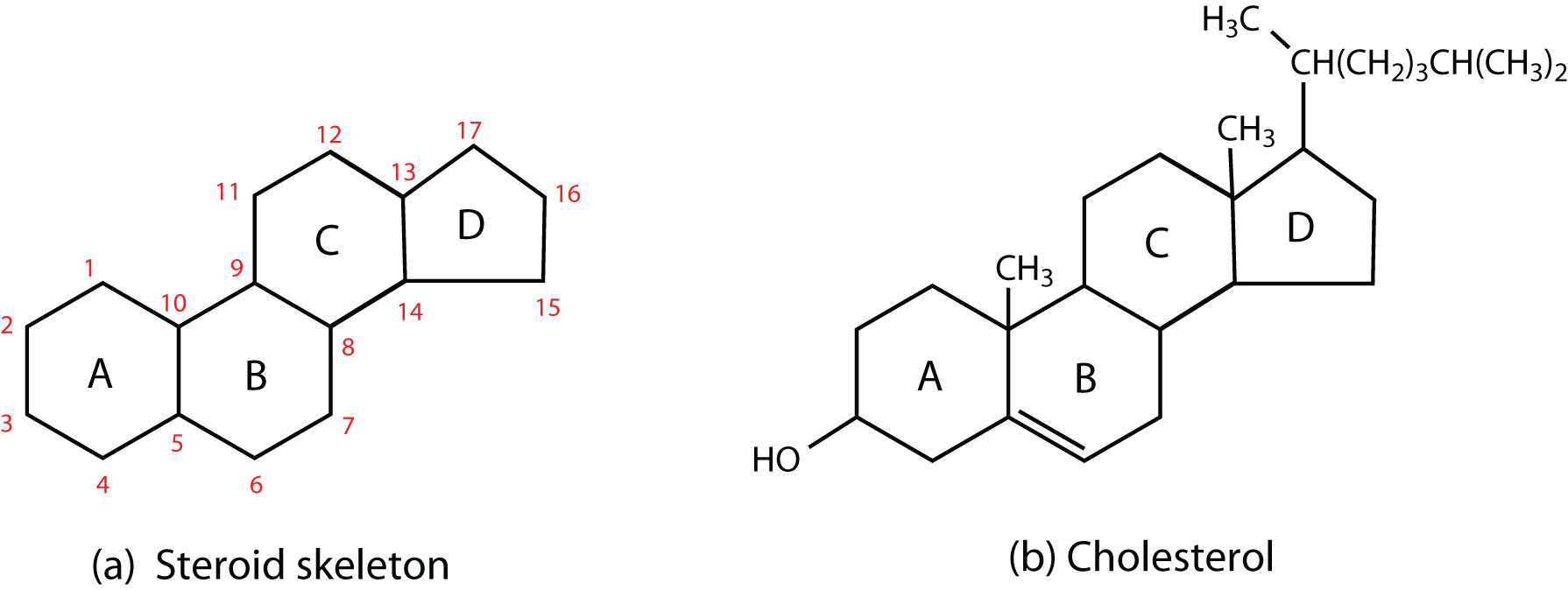
Phospholipid Bilayer: Fluid Mosaic Model
The lipid bilayer is the main barrier; integral proteins, outside surface has glycoproteins attached
Phospholipid bilayer structure
Hydrophilic heads on the outside (attract solutes and interact with H2O); Hydrophobic tails are on the inside (Hydrophobic effect causes them to cluster together so fewer of them are exposed to water; water pushes them together), they form a semi-permeable membrane
Fluidity Depends On:
Proportion, Saturation, Cholesterol/Steriods
Proportion of head groups and tails on packing
Bigger heads = less packing
shorter tails = less packing ( decrease in hydrophobic interactions)
Saturation of C C double bonds on packing
more C C double bonds = less packing
more C C double bonds = melt at a lower temperature
Cholesterol/Steroids on packing
High Temp: Reduce fluidity = reduces movement
Low Temp: Increase fluidity = prevents tight packing
Steroids act as a buffer, stabilizing membrane fluidity
How do bacteria maintain fluidity?
change length and saturation
How do eukaryotes maintain fluidity?
modify melting properties with steriods
Channels/Pores
fast, let the solute diffuse down the gradient
can be open continuously but most are gated
filter by size/charge → lets selected ions and small molecules move in
uses simple diffusion
less specefic
Transporters
Carriers and Pumps
bind, transport specific solutes
Carriers
Type of transporter
Use passive, facilitated diffusion
move solutes through membranes by making conformational changes
Bind and transport to specific solutes
Rate plateaus (like enzymes)
Ex: GLUT1
Pumps
Active transport, Endergonic (movement against the gradient) coupled with exergonic reaction (ATP → ADP)
Primary Active Transport
exergonic process is ATP breakdown; creates own gradient
Secondary Active Transport
Indirectly, energy is from a concentration gradient; it uses the existing gradient created by primary transport, used by prokaryotes and eukaryotes
Ex: SGLT, glucose is transported used Na+ gradient
More efficient because if glucose pimp was used, you would need 1 ATP for each glucose. This amplifies the energy and is easier to control/regulate
Types of Membrane Protiens
Integral and Peripheral
Integral proteins
Single or Multi-pass
always a-helices, have side groups that bind to lipid tails, can form pores and channels when paired
Peripheral
On either side of the membrane, anchored by a covalent bond to lipid/protein or by a non-covalent interaction
Endocrine
Long distance
Paracrine
Short Distance
Gases and Non-polar molecules as a stimulus
diffuse through the bilayer to a macromolecule ( ex: enzyme) that causes a response or an intracellular receptor in the cytoplasm, signal is not amplified, slower response
Small polar molecule as a stimulus
Can’t diffuse through bilayers, uses surface receptors, signal is amplified so faster response
Surface Receptor Types
Ion channels, G Protein, and Kinase
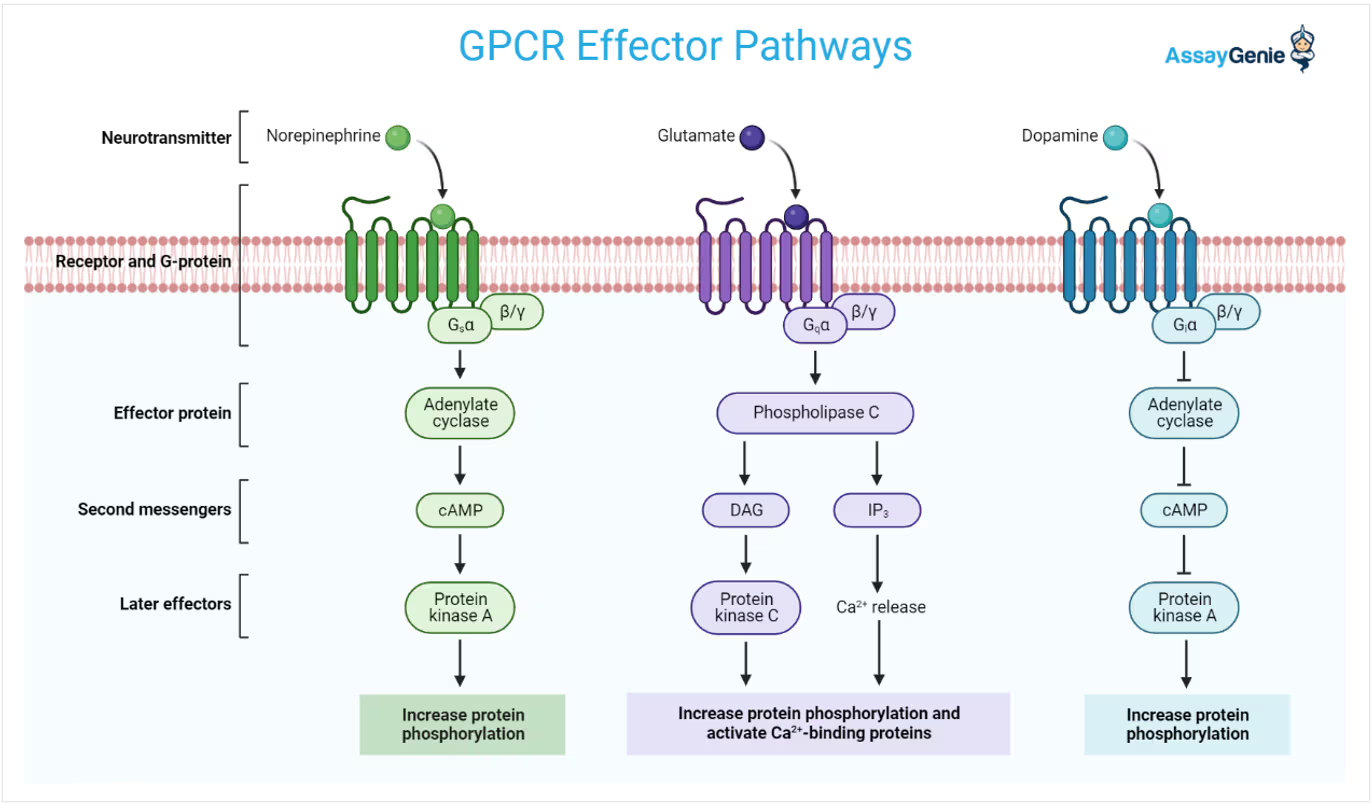
G-Protein coupled receptors (GCPRs) Steps
1) ligand changes the receptor’s shape
2) The new shape attracts the G-protein, which turns on the alpha subunit by GTP exchange
3) a- subunit leaves and turns on the enzyme generating secondary messengers
4) Secondary messengers activate protein kinase A
5) Protein kinase a activates new enzymes to control downstream events and/or a change in transcription factors that control DNA
G-Protein coupled receptors (GCPRs) Deactivation
1) release/destroy ligand
2) turn off the receptor
3) switch G protein off (GTP→GDP); done by G-protein coupled receptor kinases
4) destroy secondary messengers
5) turn off kinases, turn on phosphatases
What molecules are GCPRs used for
small soluble molecules
Kinase Receptors
1) ligand (small proteins) binds to receptors
2) receptors dimerize, trans-phosphorylate (they phosphorylate each other)
3) Adaptor proteins bind, and Ras (G-Protein) is activated
4) Ras activates phosphorylation cascade of MAP kinases
5) final kinase turns on transcription factors and/or increases expression of cell growth genes
Deactivating Kinase Receptors
Takes longer than GCPRs
Phosphatases remove phosphates from kinases and transcription factors; gene products must be degraded
ECM
A network of proteins that surrounds the cell
Made of:
Water
Jelly-like ground substance
fibers
secreted ions/molecules
adhesion molecules
Plant ECM
Cell Wall
Made of:
Pectin polysaccharides (space filling/jelly/adhesive)
Cellulose/ hemi-cellulose (strength)
Lignin (harden walls), silica
Stable, Stiff, not flexible
Fungal ECM
Cell Wall
Made of:
Glucans (space filling polysaccharide)
Chitin (Strength)
Glycoproteins (Cell-Cell Adhesion)
Septal pores: let adjacent cell membranes communicate
Properties of Animal ECM
Mechanical strength, protection, organization, specialization
Ground Substances
GAGS (glycosaminoglycans): chains of disaccharides, they attract Water and Na+ ions
Proteoglycans (GAGs + Protein anchor): The size of proteins and the number of GAGs control how much water it binds to (stiffness), resists compression forces, and is a cheap, adjustable filler
Fibers
Collagen, Elastin, and Fibrillin
Collagen
Many Types:
Fiber/Cable like → Type 1 (Cable), Type 3 (Fibers)
Net/Mesh like → Type 3 and 4
Fibers built from fibrils
Fibrils are 3 protein triple helix
High Tensile Strength: Resist stretch and length pulling
How collagen is produced
1) mRNAs translated into the ER Lumen
2) Golgi modifies some proteins to hydroxyproline, adds O-linked sugar
3) assemble procollagen triple helix
4) secreted with loose ends via vesicles
5) ends are trimmed
6) self-assemble into fibrils, then fibers
Stabilized by H-Bonds and cross-links
can bend but will not stretch in length
Elastin and Filbrillin
Elastin → Stretch
Fibrillin → limit
Found in all tissues that stretch, produced and secreted like collagen but elastin coils/links randomly; elastin will stretch until break without fibrillin
Adhesion Molecules
Fibronectin and Laminin
secreted out of the cell into the ECM, connect other parts of ECM to each other, help attach and create anchor points for cell
Fibronectin
Large protein in the ECM, V-shaped dimer, 2 covalently bonded chains, containing binding sites for integrins, collagen, and other ECM components
Laminin
T-shaped trimer, arms join to form flat sheet in the ground substance of the basal lamina → creates a selective barrier with epithelial tissue, separates cells from connective tissue; Tails form a triple helix which binds to cells
Secreted Materials
Minerals: stiffness and strength
Ca2+ and PO43-
Signal Factors
Tell cell what type of ECM
History
Growth Factors
Released for repair
Inactive when healthy
Metabolism
chemical reactions that sustain life
catabolism
breaking down molecules, releases energy
anabolism
building up molecules, uses energy
Endergonic Reaction
absorbs energy, Delta G is less than 0
Exergonic Reaction
Releases energy, Delta G is greater than 0
Endergonic Reactions are driven by:
coupling an unfavorable (endergonic) reaction with a favorable (exergonic) one
Steady State
Cells maintain a steady state, not equilibrium because they need ATP levels to be x10 higher than ADP
Enzymes
Catalyze Reactions:
lower activation energy, alters the rate of spontaneous reactions (causes spontaneous reactions to not be spontaneous anymore). Does not change delta G or make an endergonic reaction exergonic
How do enzymes catalyze reactions?
1) align substrates so productive collisions are more likely
2) rearrange electrons so the substrate will spend more time in the transition state
3) distort bonds/shape of substrate so the binding site puts stress under bound substrate
How are enzymes regulated?
Chemical Modification: kinases and phosphatases
Activators: Small ions/molecules that can be required partners
Inhibitors:
Bind reversibly/irreversibly by a covalent bond
competitive
non-competitive inhibition
Allosteric Regulation
Competitive Inhibiton
inhibitor molecule binds to the active site, competing w/ substrate
Noncompetitive Inhibition
inhibitor molecule binds outside the active site and changes the enzyme shape so the substrate cannot bind to the active site
Allosteric Regulation
a downstream product directly inhibits an enzyme near the start of a series, this links products to production levels
Endomembrane system
ER → Golgi → vesicles
Rough ER
protein synthesis and initial processing
begins glycosylation (folding and modification)
proteins made to be secreted are made by the ribosomes attached to the rough ER
Smooth ER
Lipid synthesis, detoxification, and Ca2+ storage
makes phospholipids and steroids
detoxifies drugs (liver cells)
Stores Ca2+ for muscle contraction
Golgi
Modifies, sorts, and ships proteins
receives proteins and lipids from the ER
modifies (glycosylation)
sorts based on signals
Lysosomes
Breakdown and digestion
contains digestive enzymes
breaks down macromolecules, old organelles, and pathogens
Nuclear Membrane
A double-membraned structure that separates DNA from the cytoplasm
the outer membrane is continuous with the RER
Nuclear Pores
large protein complexes that control what enters/leaves
highly regulated transport
regulate: transcription factors, other proteins get in to control DNA expression, and ribosomes get out
Endosymbiosis
theory of how plastids (mitochondria and chloroplasts) were created. Cyanobacteria → chloroplasts and purple aerobic bacteria → mitochondria
Glycolysis
Glucose → 2 ATP + 2 NADH
Preparatory phase: consumption of 2 ATP
Glucose-6-phosphate is formed and trapped in cell because it is negatively charged
Fructose 6-phosphate is the irreversible step, dedicated to energy extraction after this step
Cleavage and rearrangement: 6C → 2 3C
Payoff
Reaction Releases energy and stores it in ATP via substrate phosphorylation
Takes place in the cytoplasm
Pyruvate Oxidation
3C pyruvate → 2C Acetyl +CO2
Acetyl + Coenzyme A → Acetyl CoA
Acetyl CoA is locked into the mitochondria
Citric Acid Cycle
Energy from oxidation reactions is captured and used elsewhere
Products: 10 NADH, 2 FADH2, 4 ATP, 2 CO2
Oxidative Phosphorylation
Regenerate FADH and NAD+
ATP Synthase
Uses H+ gradient to generate ATP
F1 (ATP Synthase)
uses rotational energy to catalyze ATP synthesis
F0
forms a channel that rotates as protons pass through
Degrading Fats
Beta oxidation
Degrading Amino Acids
Remove N-Group and rearrange backbone to be a more burnable molecule