Ch17 DNA Replication, Repair, and Recombination
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
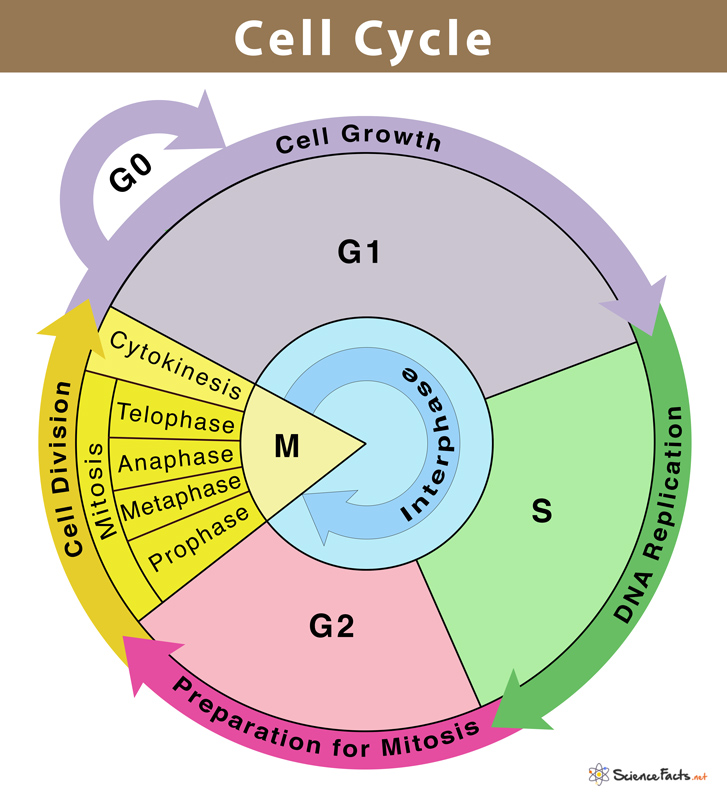
the cell cycle
G0 → G1 → S → G2 → M (PMAT)
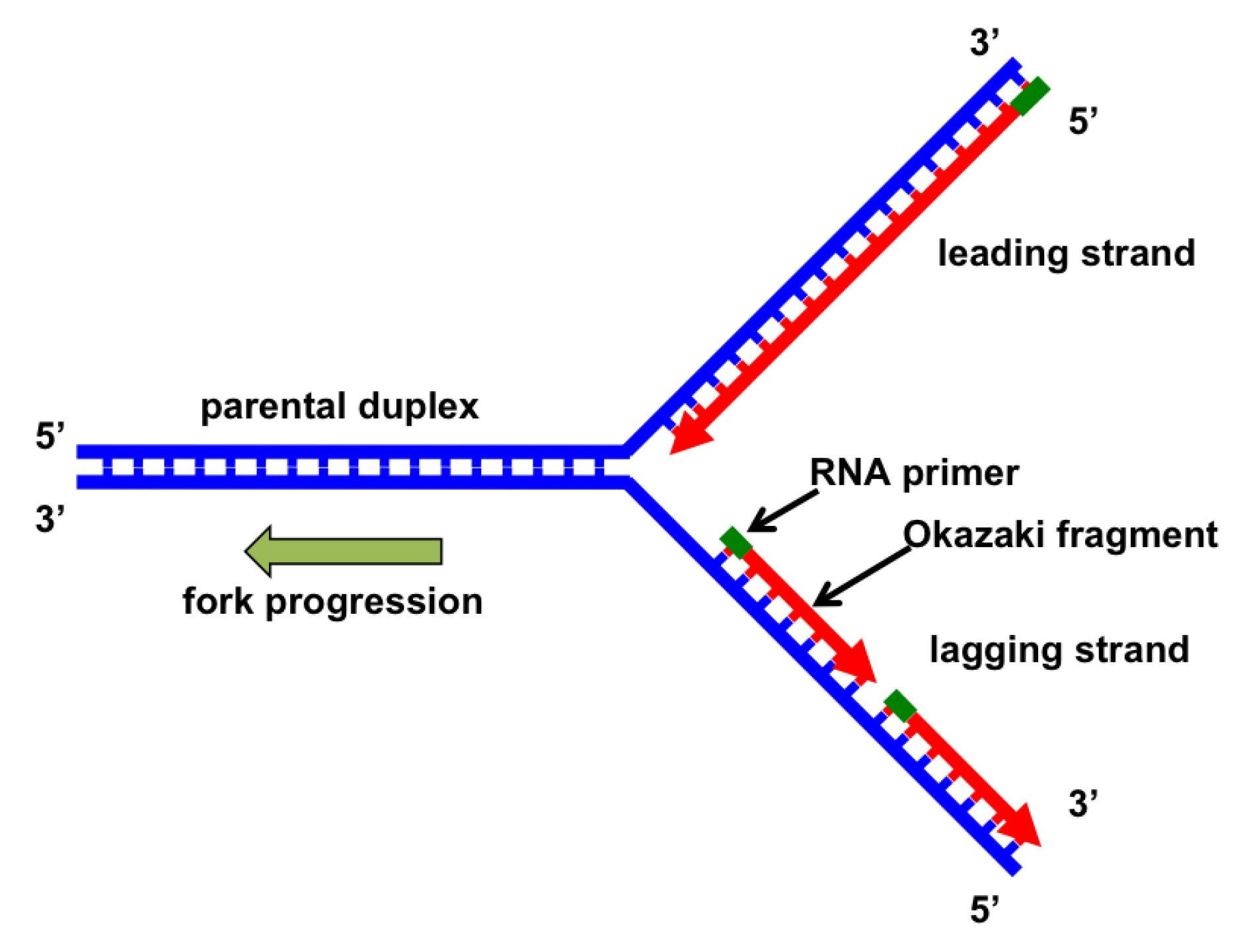
replication forks
formed where replication begins and proceeds bidirectionally, away from the origin
unwind DNA and copy both strands
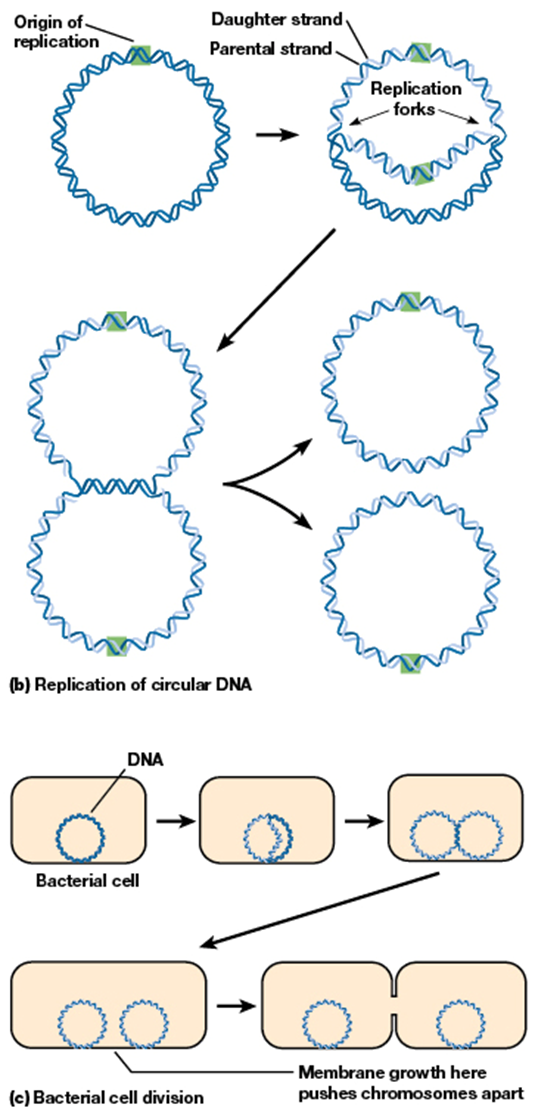
theta replication
mechanism of replicating circular DNA
uses replication fork
replicons
replication site or replication bubbles
multiple per linear chromosome
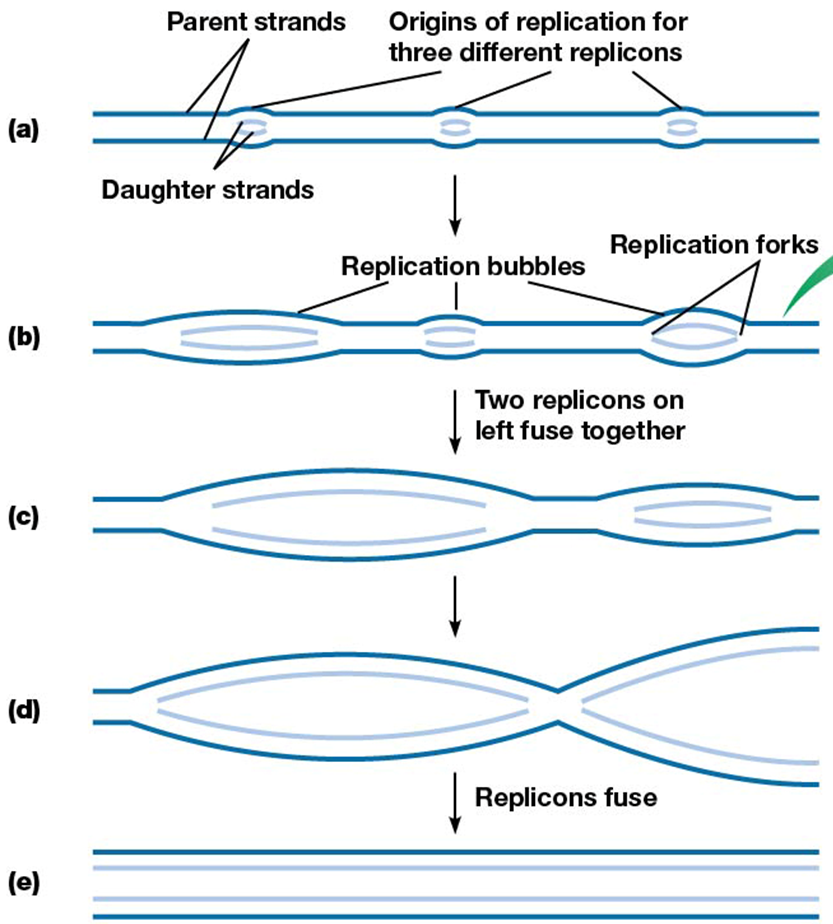
origin of replication (and E. coli)
site of DNA replication initiation
OriC of E. coli is AT rich and about 245 bp in length
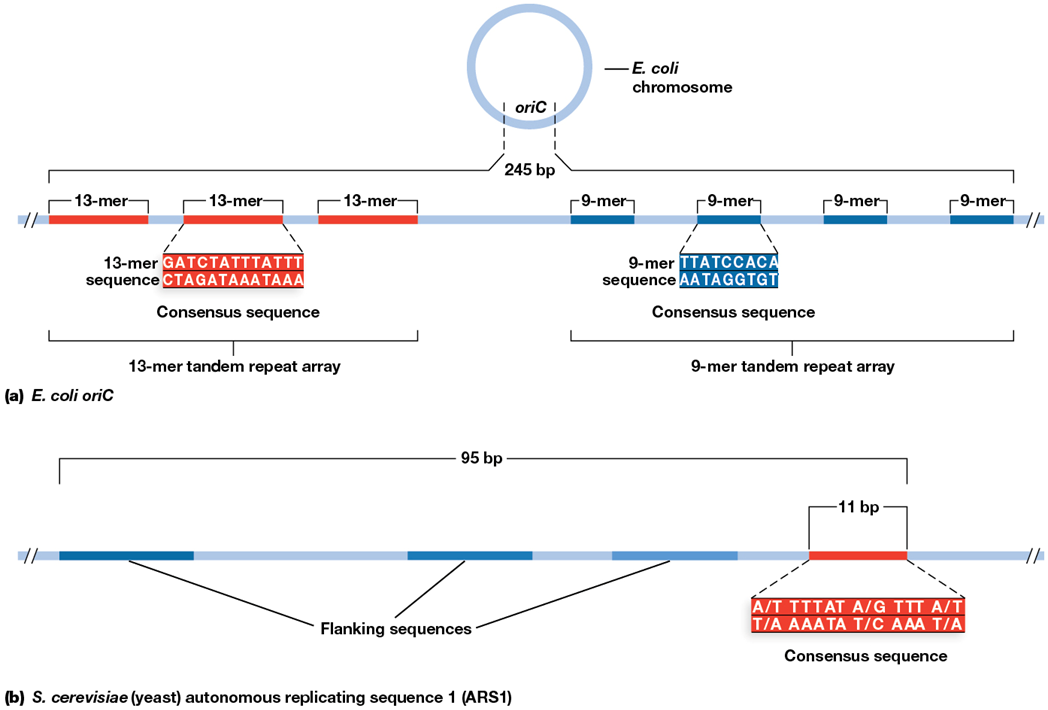
consensus sequences
common sequences conserved across various species
example is the AT rich oriC of E. coli
3 enzymes in E. coli that initiate replication
DnaA, DnaB, DnaC
each bind the oriC
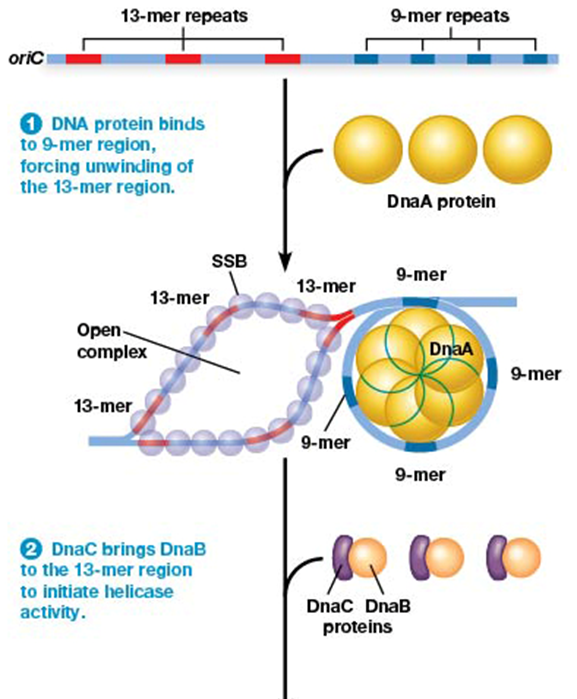
role of DnaA (E. coli) in initiating replication
binds to the conserved sequence 9-mer of oriC
result is unwinding of DNA at 13-mer sites
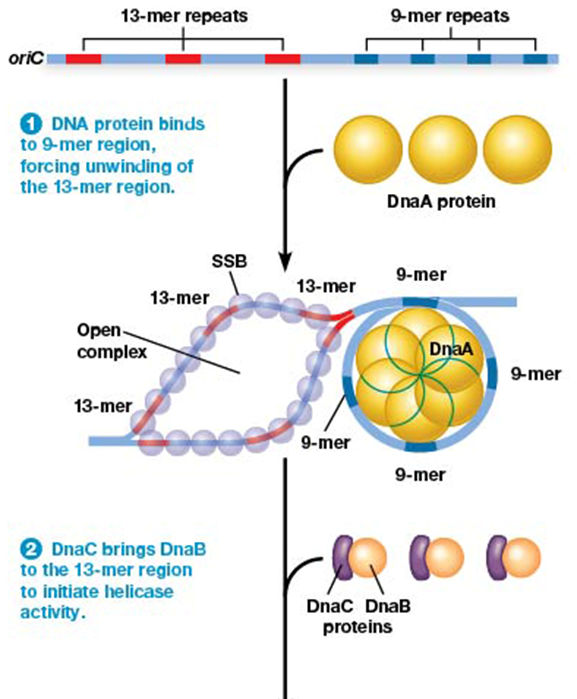
role of DnaB (E. coli) in replication
acts as DNA helicase to unwind DNA strands
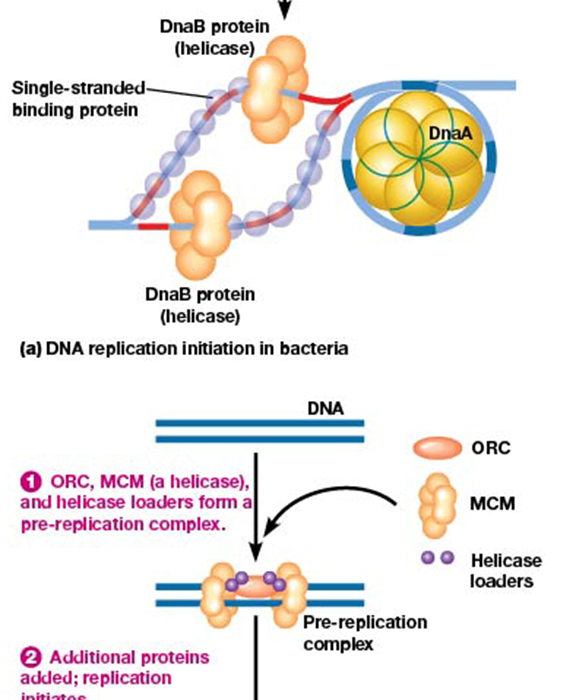
DNA replication initiation steps in E. coli
1) DnaA binds the 9-mer sequence
2) unwinding at 13-mer sequence
3) single stranded protein binds unwound regions
3) DnaB (helicase) unwinds DNA as sequence proceeds
4) DNA polymerase and more proteins are added
5) replication proceeds
DNA polymerase III vs DNA polymerase δ
III: replicative enzyme in E. coli
δ: replicative enzyme in eukaryotes
directionality of DNA replication
3’ (hydroxyl) → 5’ end
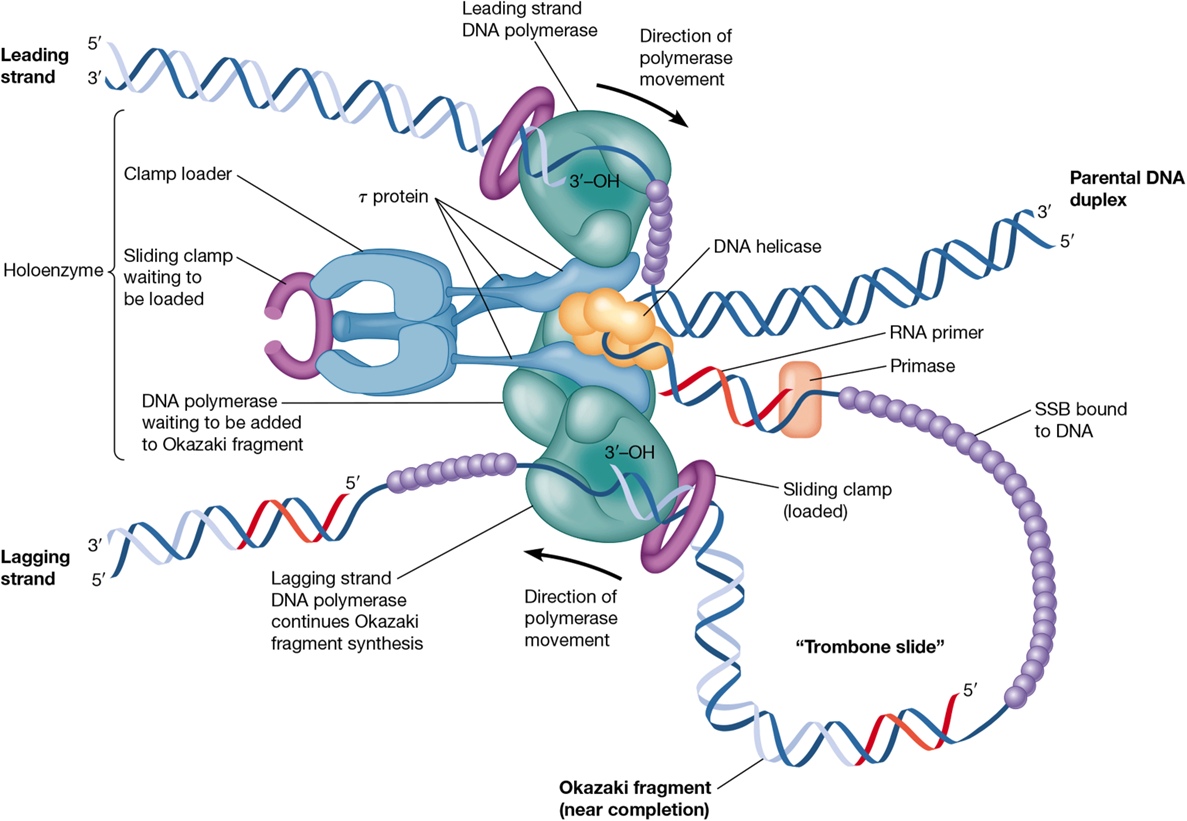
leading vs lagging strand
leading: synthesized continuously in 5’ → 3’ direction
laggind: synthesized in Okazaki fragments that make a 3’ → 5’ replicated strand
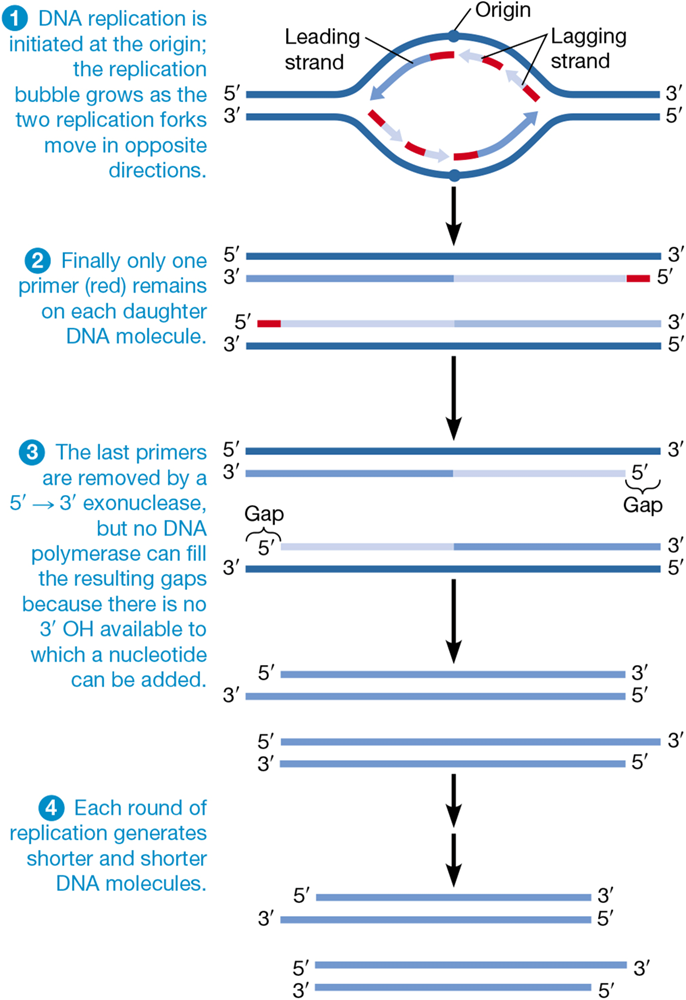
why do eukaryotes have telomeres
lagging strands are difficult to deal with because they need primers
each round of replication results in the loss of some nucleotides from the end of the sequence
repeated sequences at the ends of chromosomes solve this
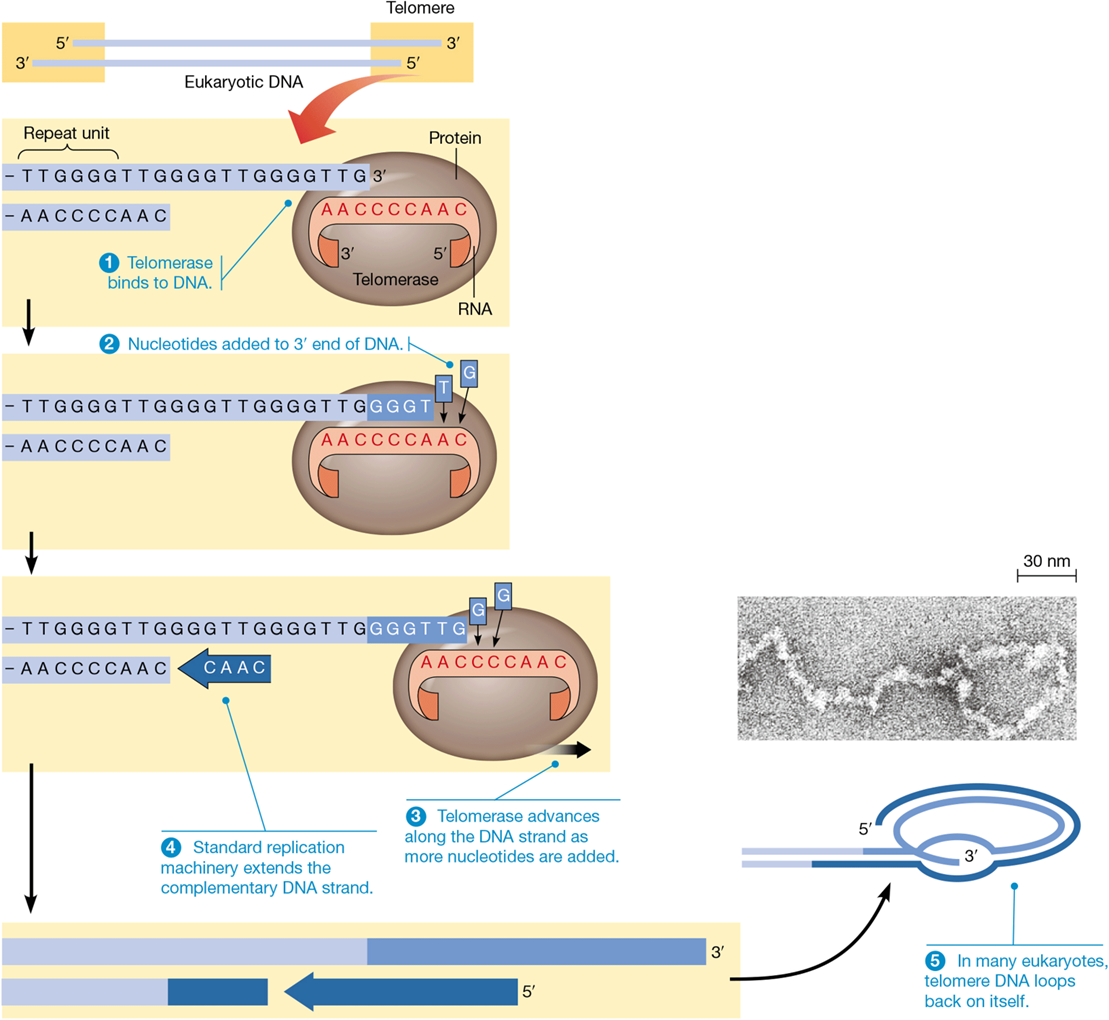
composition of telomeres
110 - 1500 copies of TTAGGG
noncoding
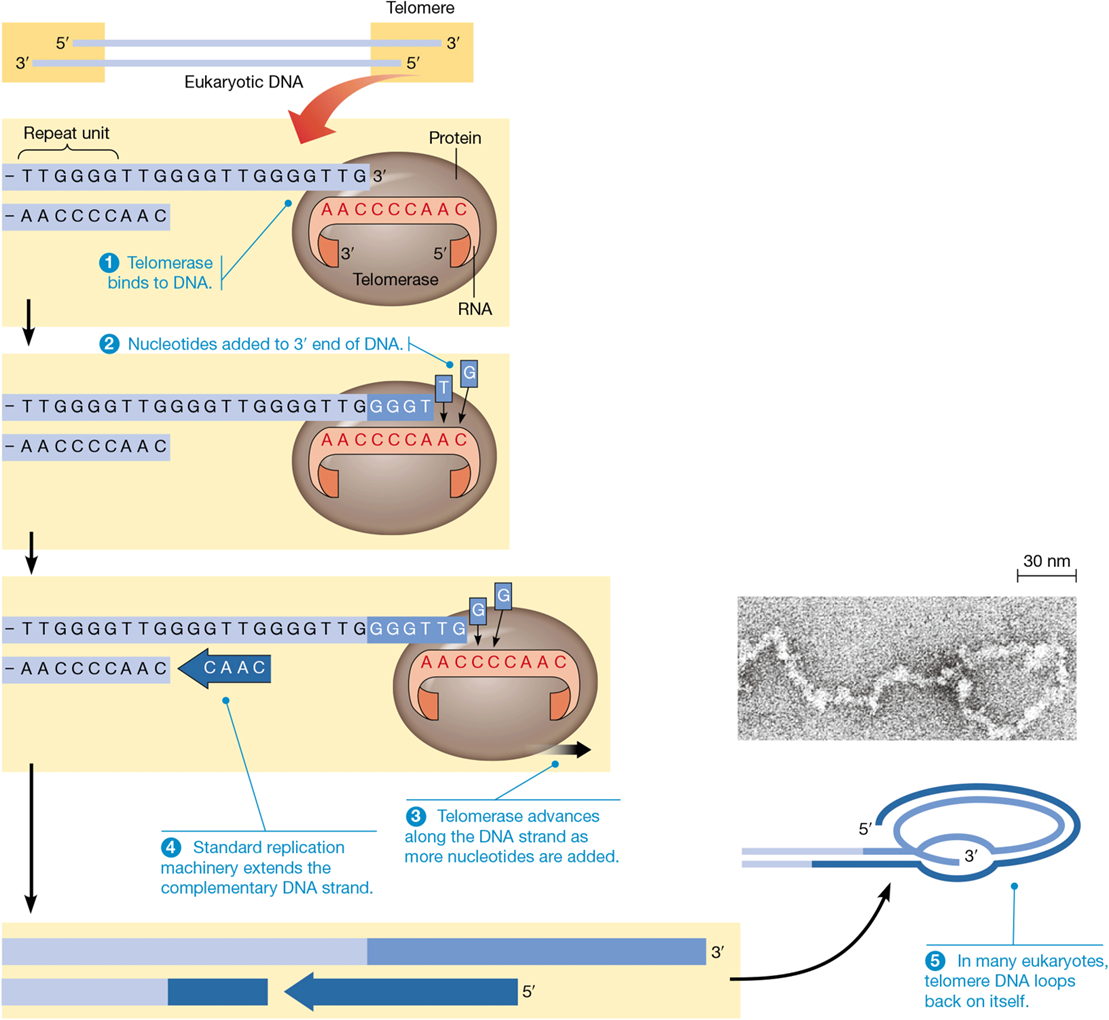
telomerase
catalyzes addition of telomeres to chromosome ends
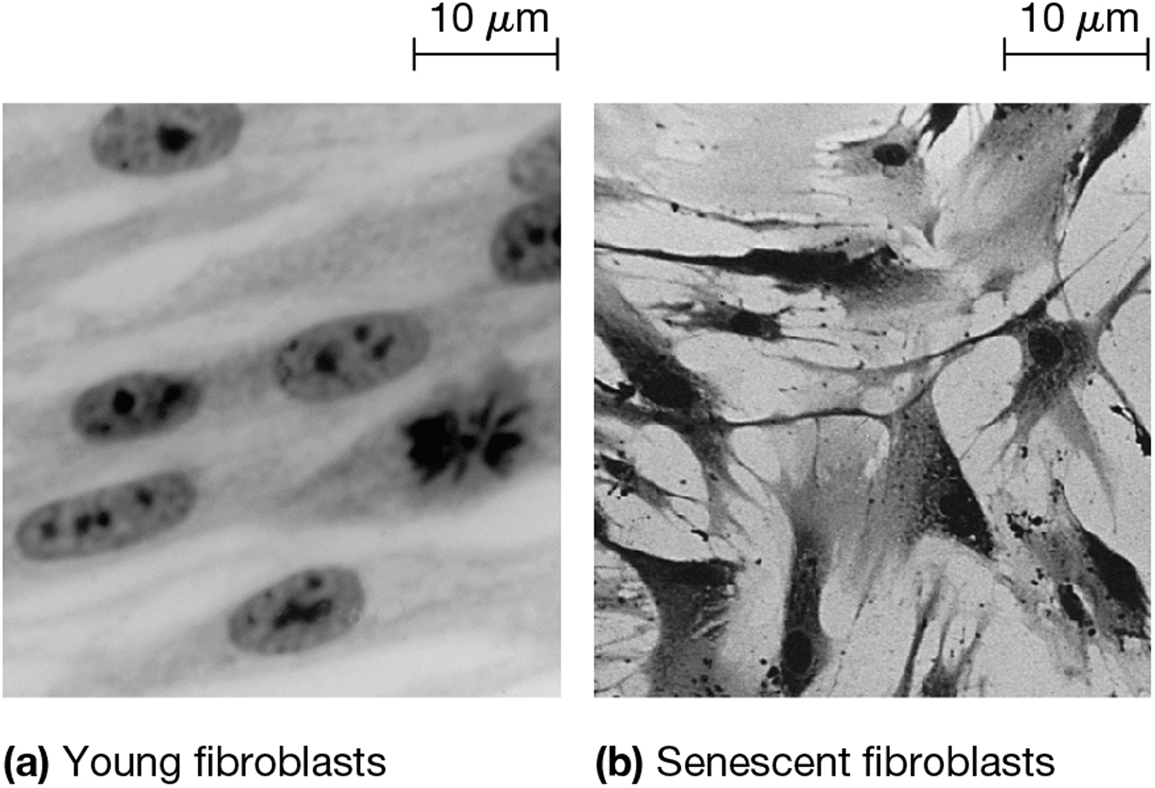
hayflick limit
amount of times a cell population can divide before reaching senesence (permenant cellular arrest) or apoptosis

how do cells circumvent the Hayflick limit
producing telomerase
link between telomerase and cancer
almost all cancer types have telomerases
telomere capping proteins
form T loop to protect linear ends of single stranded DNA
werner syndrome
patients lack a telomere cap protein WRN
premature aging
types of mutations that occur during DNA replication
spontaneous mispairing due to transient formation of tautomers (resonance structures of nitrogenous bases)
slippage (example is trinucleotide repeats)
spontaneous damage to individual bases (depurination and deamination)
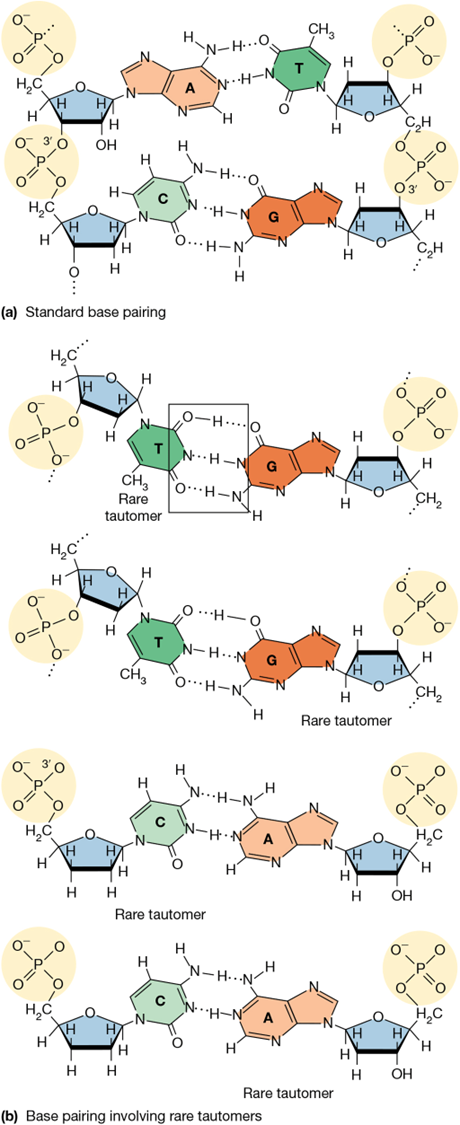
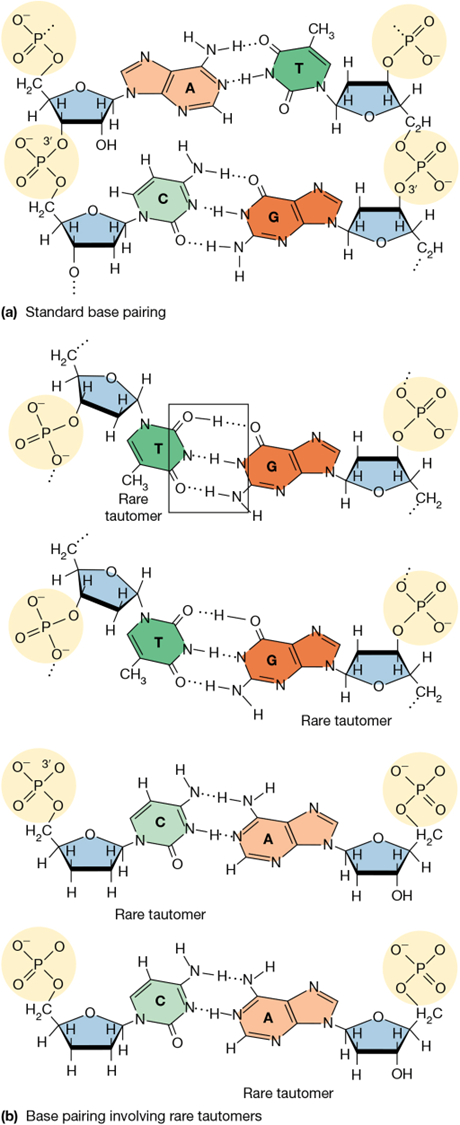
most common mutation in DNA replication
mispairing due to tautomers
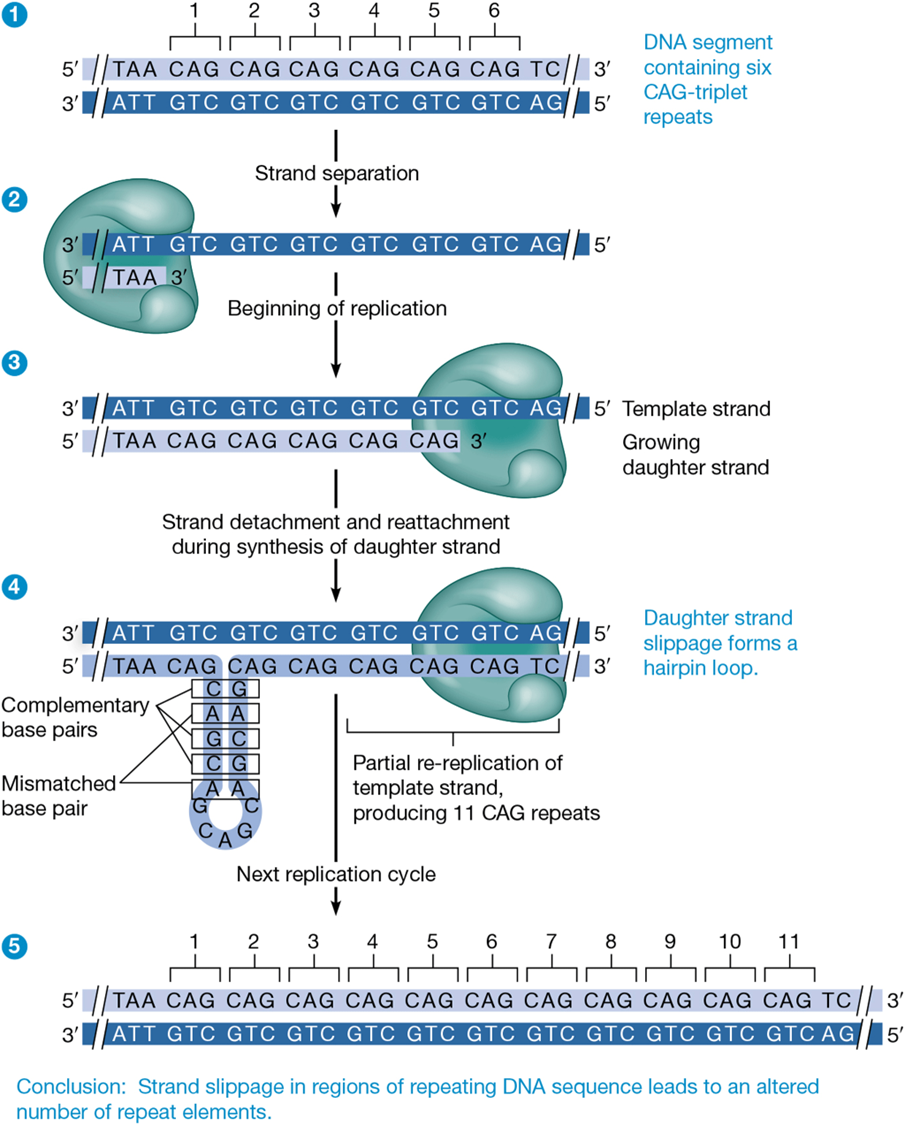
trinucleotide repeats
spontaneous replication error that occurs in a region with repetitive DNA
causes slippage errors
depurination DNA damage
loss of a purine
human cell can have 1000s/day
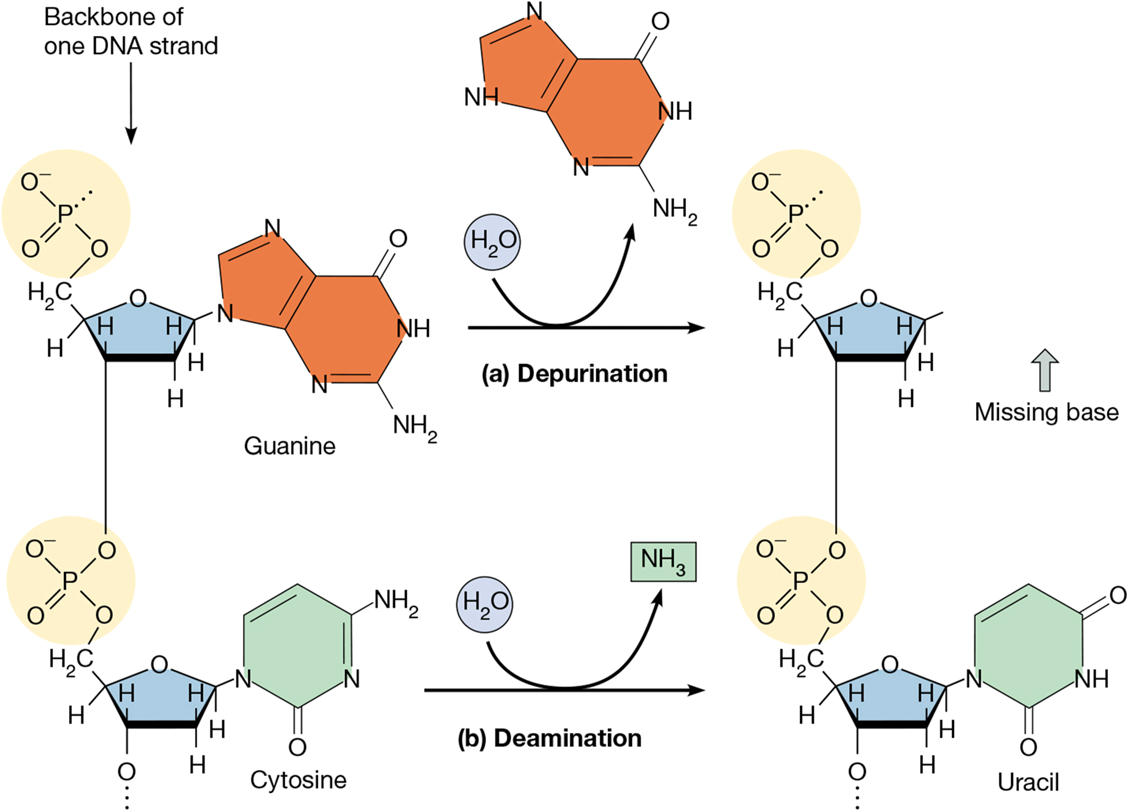
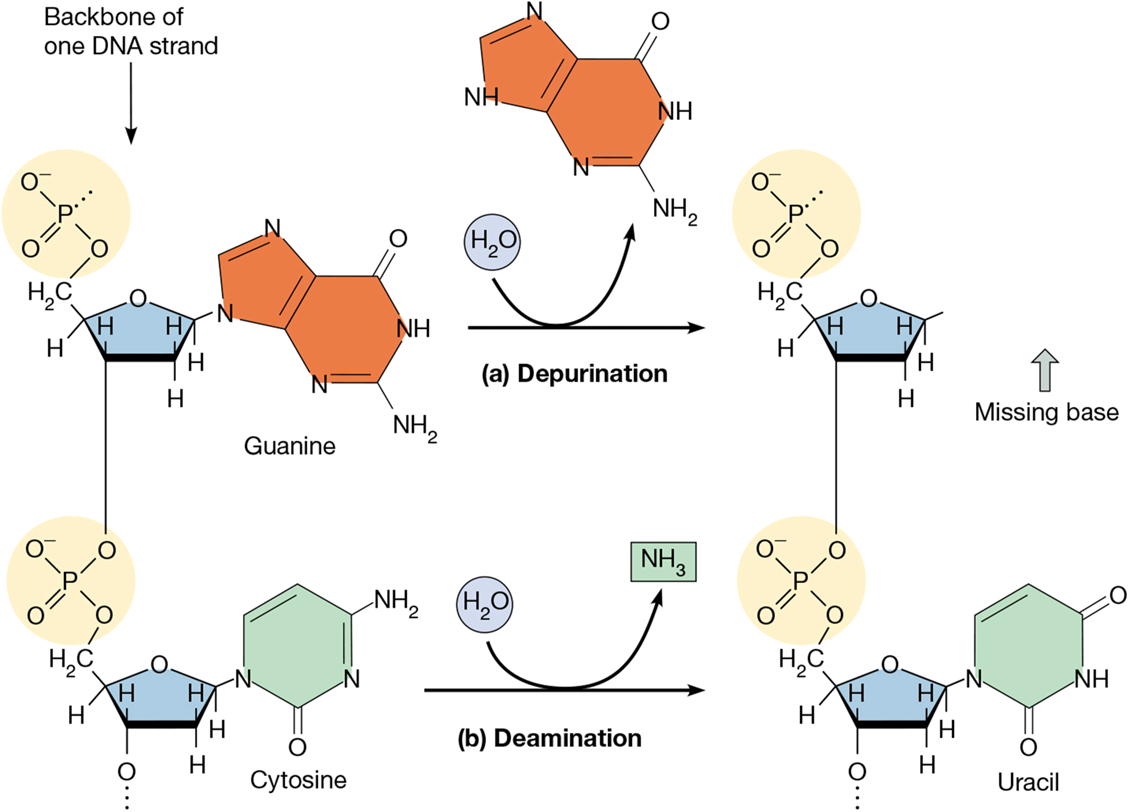
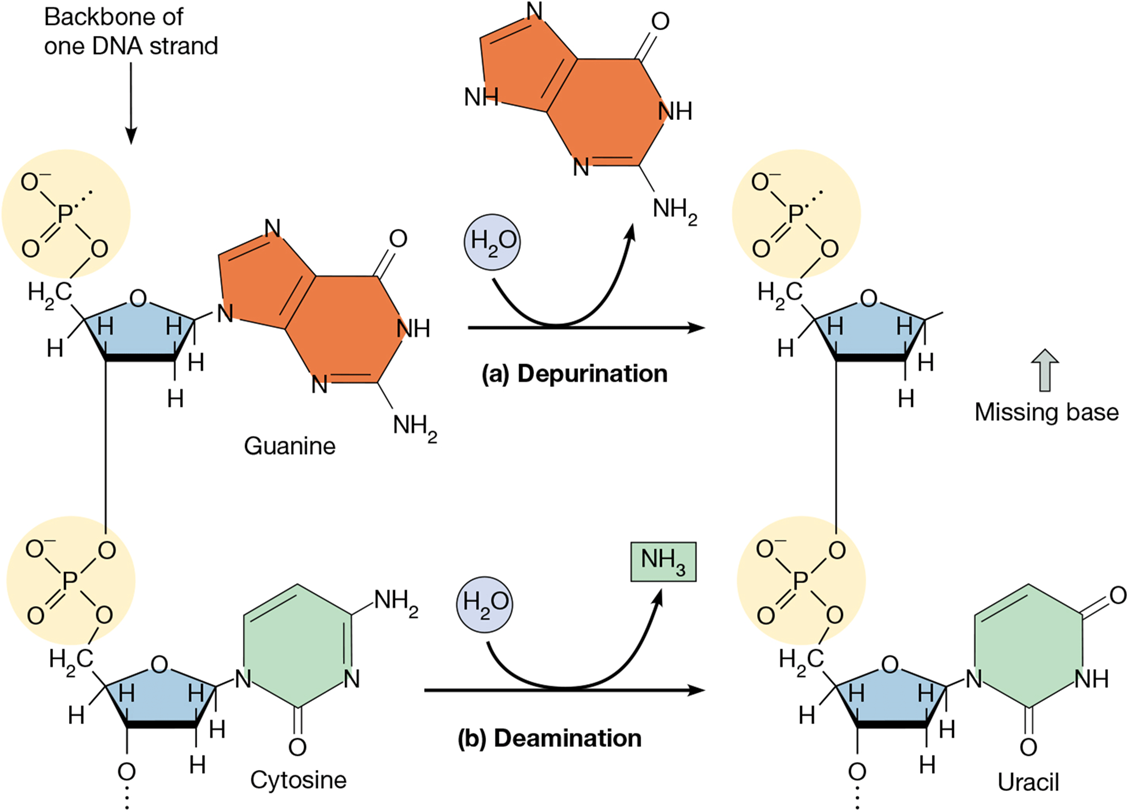
deaminations
loss of a bases amino group
human cell can have 100/day
Ethyl methansulfonate (EMS)
chemically alters base so it will mispair in next replication
adds an ethyl group to bases
nitroguanidine
chemically alters base so it will mispair in next replication
adds methyl groups
nitrous acid (HNO2)
chemically alters base so it will mispair in next replication
increases likilihood of deamination
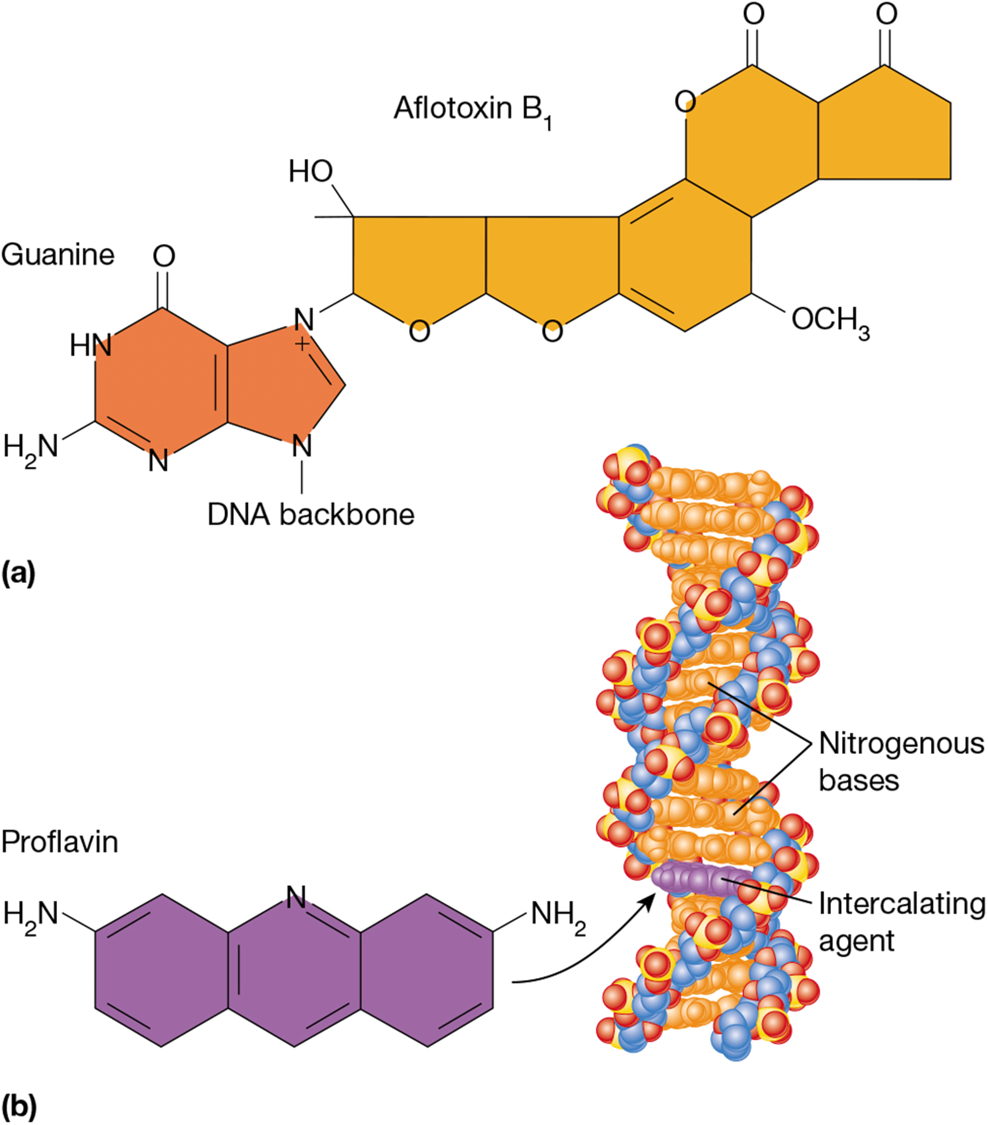
aflatoxin B1
add bulky DNA adducts to DNA
attaches to guanine and causes depurination
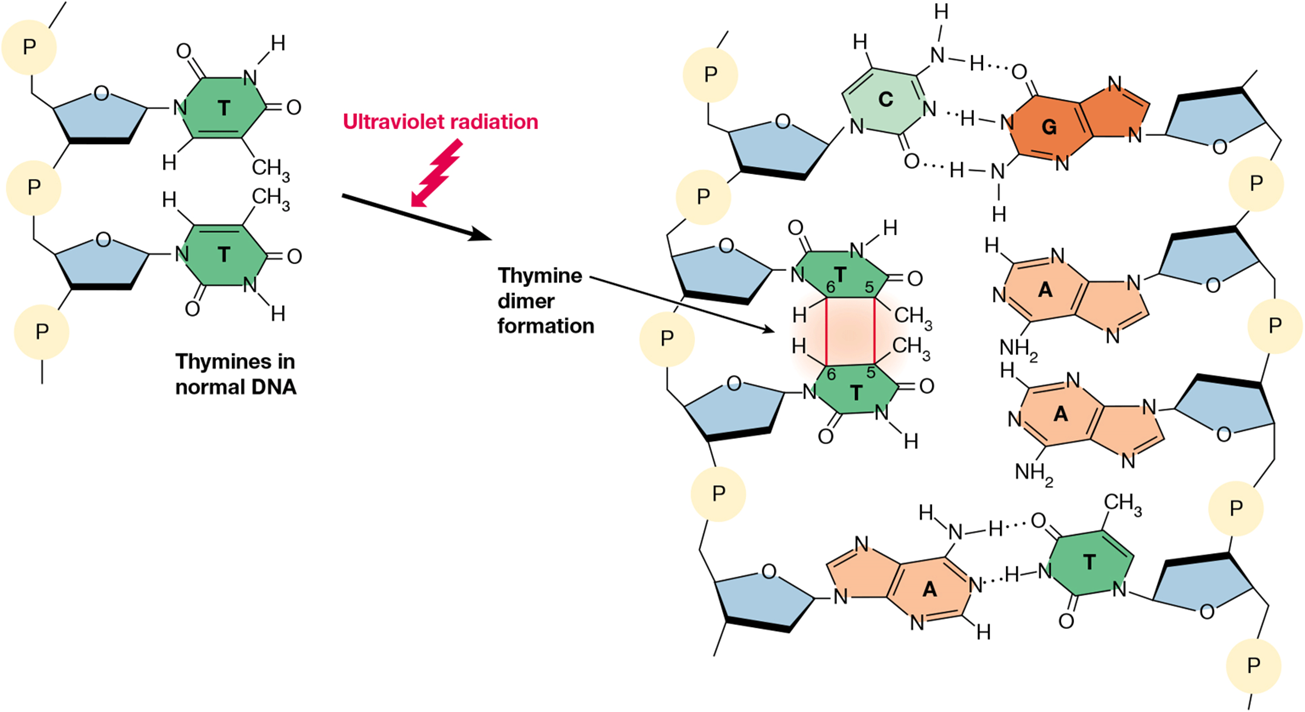
pyrimidine dimer
triggered by UV radiation
covalent bond forms between adjacent pyrimidines
ionizing radiation
removes electrons from molecules to generate damaging reactive intermediates
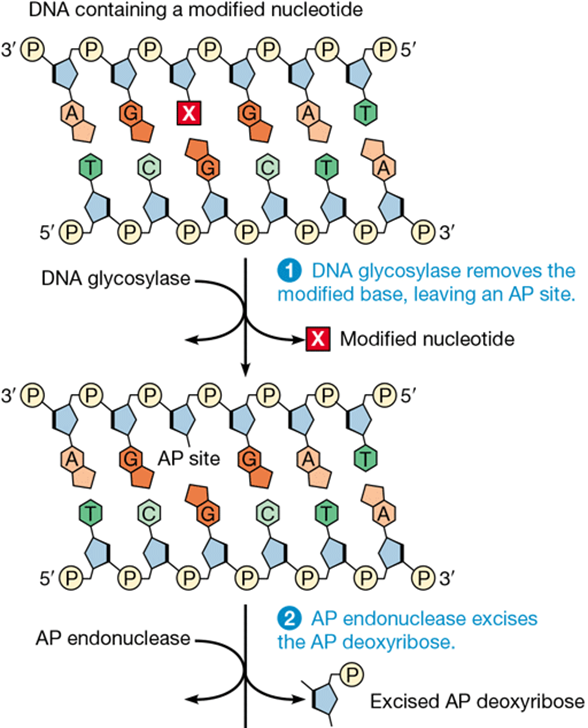
base excision repair and example
corrects single damaged bases
DNA glycosylase detects damage and cleaves base
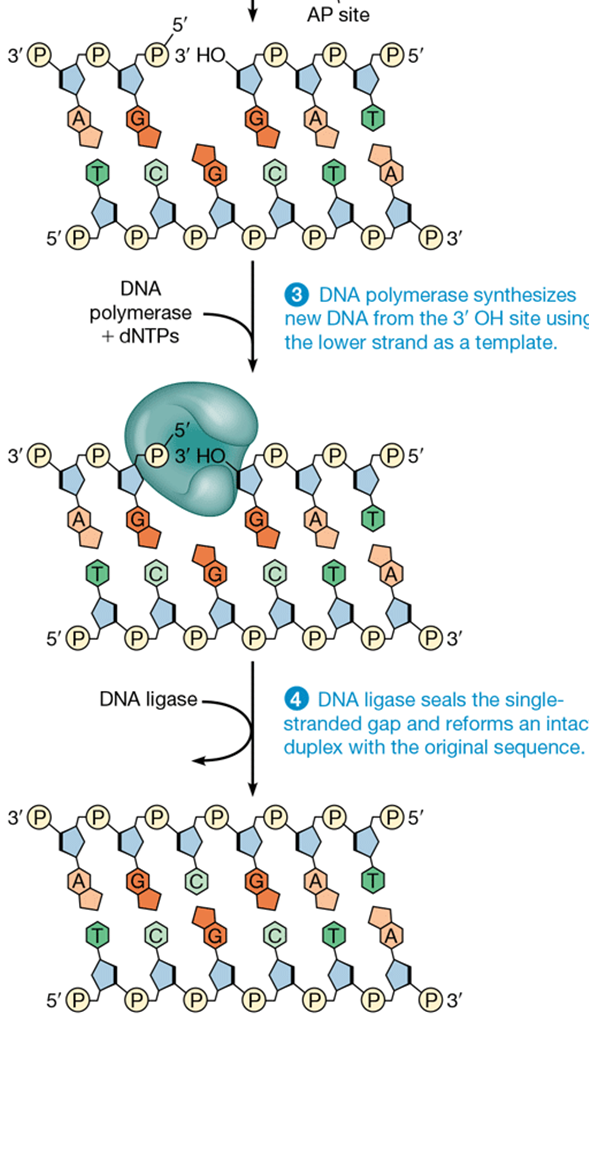
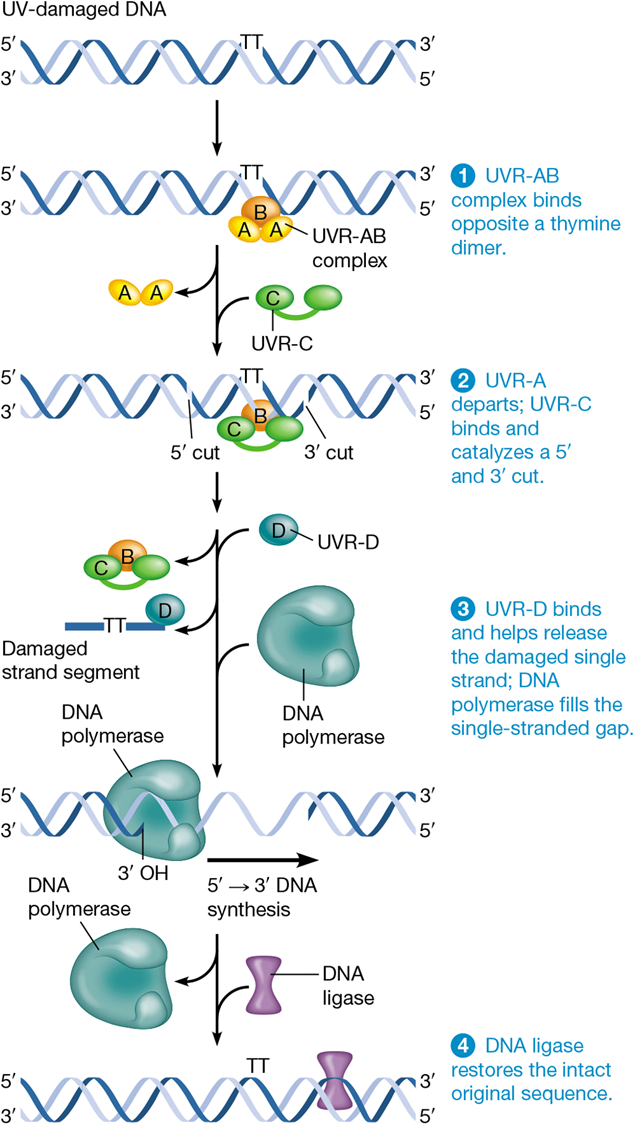
steps of nucleotide excision repair
1) proteins detect distortions in DNA helix
2) recruits NER endonuclease to cut DNA on both sides of lesion
3) helicase unwinds DNA between nicks (incisions) and frees distorted sequence from DNA
4) polymerase and ligase finish repair
steps of mismatch repair
repairs abnormal nucleotides after DNA replication
1) MutS detects mismatch
2) endonuclease MutH introduces a nick in unmethylated strand
3) exonuclease removes incorrect nucleotides from nicked strand, and these are replaced with correct sequence
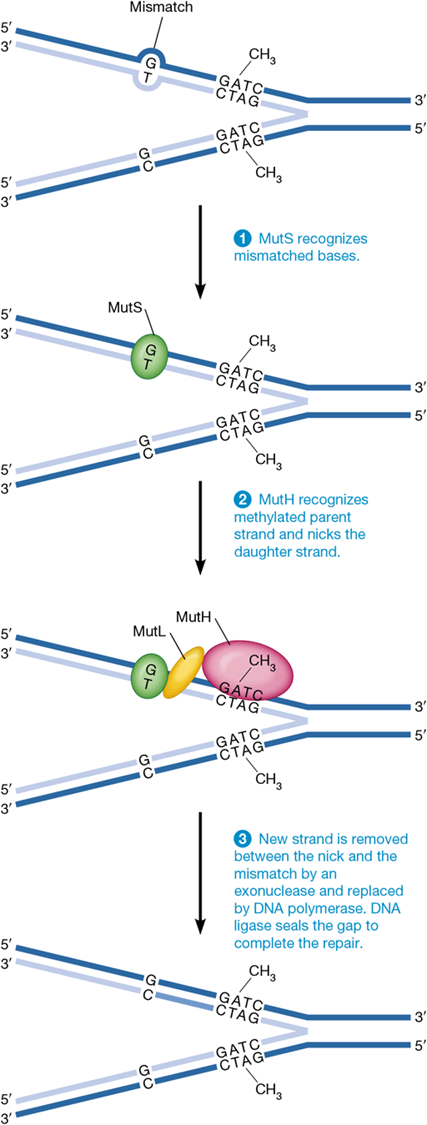
significance of methylation to mismatch repair systems
DNA methylation is not immediate after DNA replication allowing distinction between old and new strands
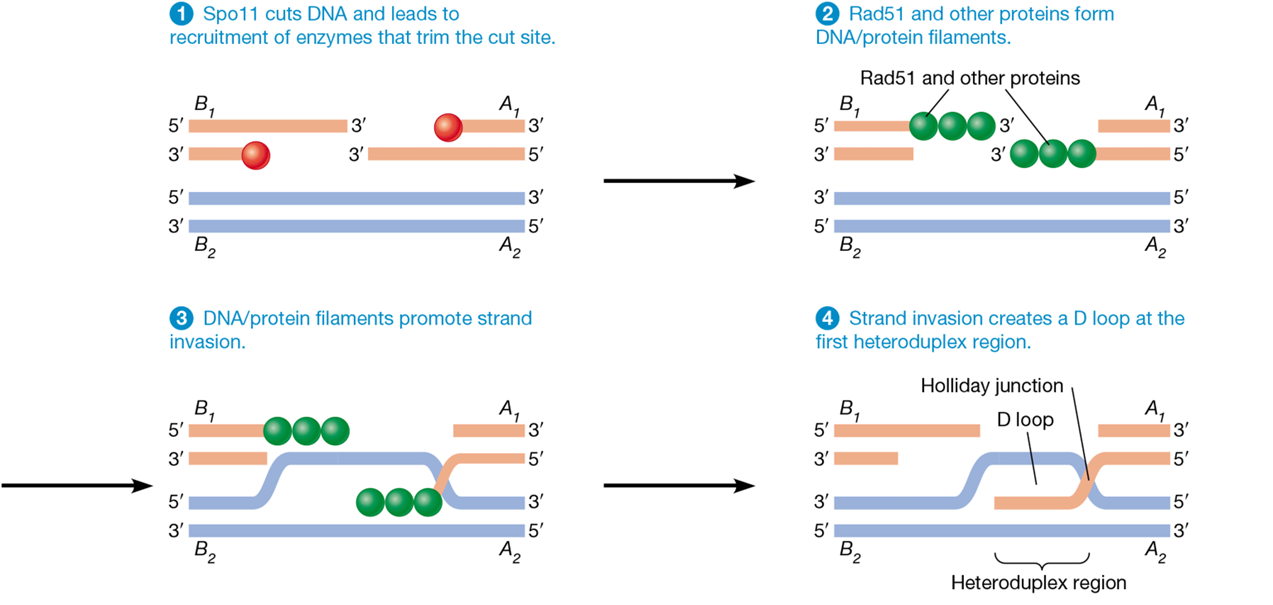
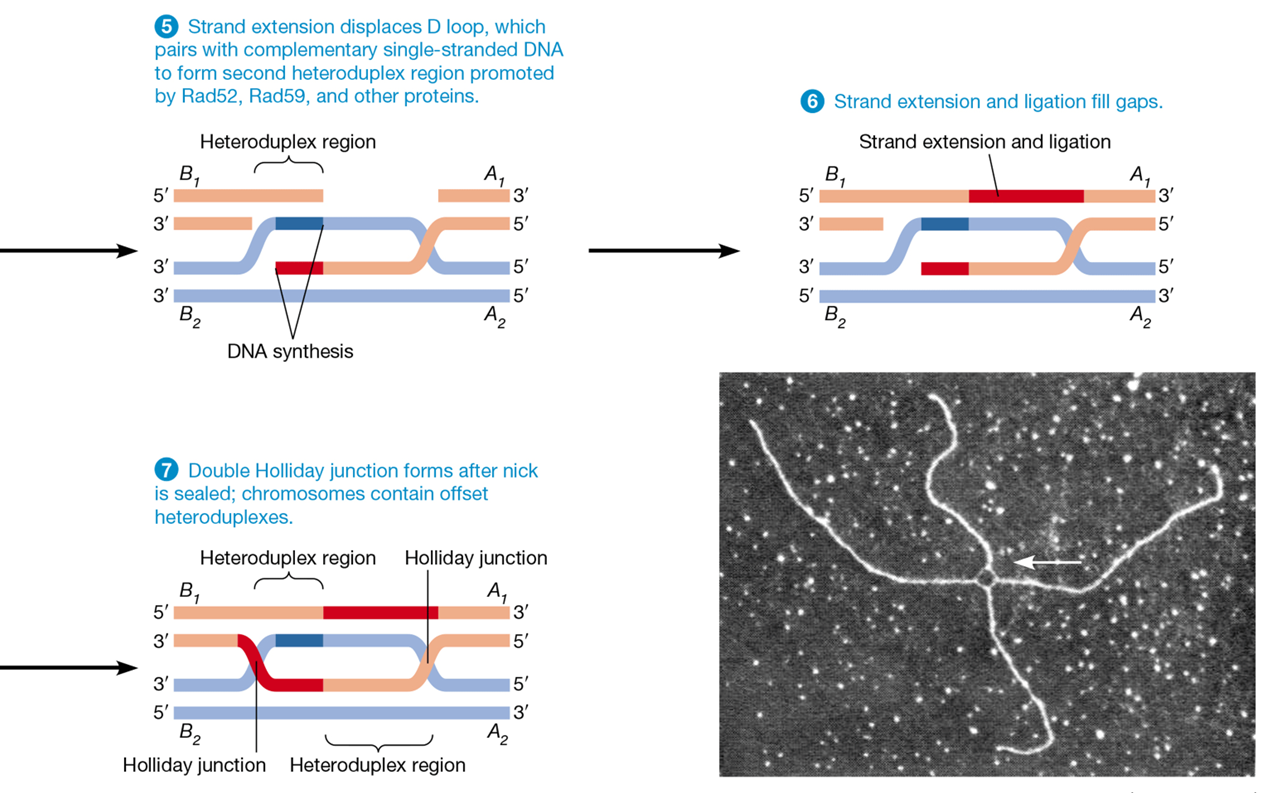
homologous recombination
uses the process of crossing over
homologue acts as template for accurate repair due to sequence similarity
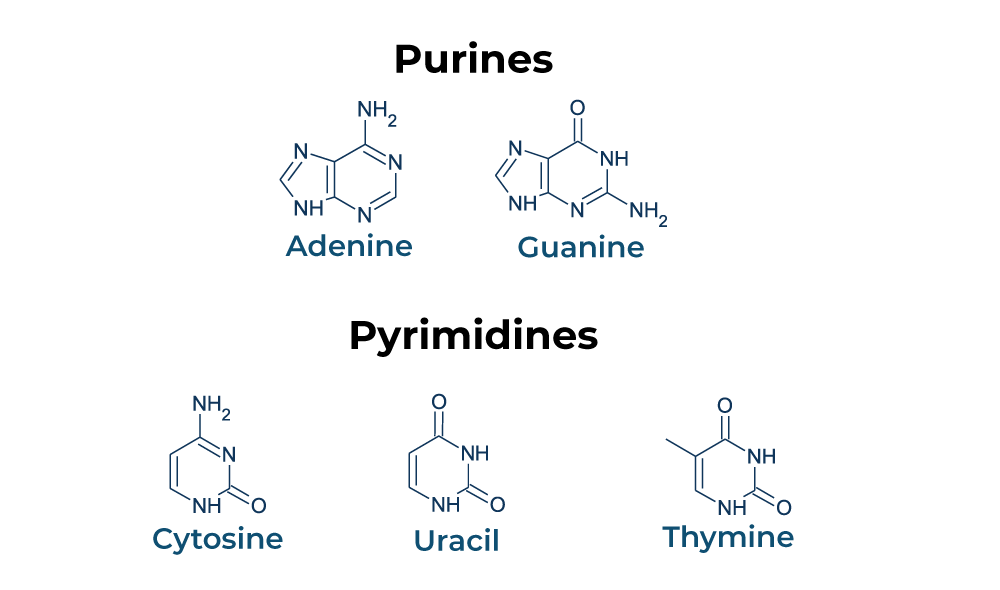
purines vs pyrimidines
Purines: AG, two rings (AG is silver! you want more!)
Pyrimidines: CUT, one ring