biopharmaceutical stability
1/103
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
What are monoclonal antibodies and what does it mean that they are quaternary glycoproteins?
Monoclonal antibodies are large protein molecules made by identical immune cells. Being quaternary glycoproteins means:
Quaternary structure: They are made of multiple protein chains (usually 2 heavy + 2 light chains) forming a complex 3D structure.
Glycoprotein: They have sugar groups attached, which are important for stability, recognition, and function.
Q2: How do monoclonal antibodies direct immune cells (ADCC, ADCP, CDC)?
ADCC (Antibody-Dependent Cell-Mediated Cytotoxicity): mAbs bind to target cells and recruit immune cells to destroy them.
ADCP (Antibody-Dependent Cellular Phagocytosis): mAbs coat target cells and trigger phagocytosis by macrophages.
CDC (Complement-Dependent Cytotoxicity): mAbs activate the complement system, causing cell lysis.
Q3: How do monoclonal antibodies act on soluble ligands or receptors?
They can bind soluble molecules, such as TNF-α, preventing them from interacting with their natural receptor and blocking inflammatory or disease processes.
Q4: How do monoclonal antibodies act as receptor antagonists?
They bind to receptors on target cells and block the natural ligand, preventing the receptor from triggering a response (e.g., growth signals in cancer).
Optional simple analogy:
ADCC/ADCP/CDC: Antibodies are like flags on enemy cells that call immune soldiers to attack.
Soluble ligand/receptor blockade: Antibodies are like sponges soaking up a signal so it doesn’t reach the target.
Receptor antagonism: Antibodies are like locks on a door, stopping the key (ligand) from entering.
What are monoclonal antibodies and what does it mean that they are quaternary glycoproteins?
Monoclonal antibodies are identical proteins made by immune cells that target a specific molecule.
Quaternary protein: made of several protein chains (2 heavy + 2 light chains) forming a complex structure.
Glycoprotein: has sugar molecules attached that help with stability and function.
How do monoclonal antibodies use the immune system to attack cells?
They can flag cells for destruction in different ways:
ADCC: antibodies mark target cells so immune cells kill them.
ADCP: antibodies coat target cells so macrophages eat them.
CDC: antibodies activate the complement system, which punches holes in the cell membrane.
How do monoclonal antibodies stop harmful molecules in the body?
Some mAbs bind to soluble molecules like TNF-α, blocking them from acting, which can reduce inflammation or disease activity.
How do monoclonal antibodies block receptors on cells?
They attach to a receptor on a cell so the normal signal (like a growth signal) cannot bind, stopping the cell from responding in a harmful way.
Extra simple analogies to understand:
ADCC/ADCP/CDC: antibodies are like flags marking enemy cells for destruction.
Soluble ligand blocking: antibodies are like sponges soaking up a signal before it reaches the target.
Receptor blocking: antibodies are like putting a cap on a door, so the key (signal) can’t open it.
Q2: What types of biopharmaceuticals exist?
They include:
Monoclonal antibodies (mAbs) – full antibodies
Antibody fragments (Fabs) – only part of an antibody
Give an example of an antibody fragment and how it relates to a full antibody.
Ranibizumab is a single Fab fragment derived from bevacizumab, which is a full monoclonal antibody. This fragment retains the ability to bind to the target, but is smaller and often better for certain clinical uses.
What is an ScFv in biopharmaceuticals?
An ScFv (single-chain variable fragment) is a small antibody fragment containing the variable regions of the heavy and light chains, designed to bind a specific target.
Give an example of an ScFv and its target.
Brocluzimab is an ScFv that binds VEGF-A, a molecule involved in angiogenesis. Its molecular weight is ~26 kDa.
Why are smaller antibody fragments like ScFvs advantageous in therapy?
smaller molecules:
Penetrate tissues more effectively
Can be delivered in smaller volumes
Achieve higher concentrations at the therapeutic site, e.g., in the eye
Q4: How does ScFv therapy benefit ocular treatments?
Because the fragments are small and highly concentrated, they can be administered in low injection volumes while still delivering effective doses to the target tissue, reducing side effects and improving efficacy.
Q1: What is a fusion protein in biopharmaceuticals?
A fusion protein is a biopharmaceutical made by combining parts of two or more proteins to enhance therapeutic properties, such as target binding and half-life.
Q2: Give an example of a fusion protein and its molecular weight.
Aflibercept is a fusion protein with a molecular weight of approximately 97 kDa.
How does aflibercept compare to other ocular therapies like bevacizumab and ranibizumab?
Studies show aflibercept can be favoured after a year of treatment, potentially due to:
Smaller size improving tissue penetration compared to bevacizumab
Conserved Fc domain extending half-life in the patient through FcRn interactions
Q4: Why is the Fc domain important in fusion proteins like aflibercept?
The Fc region interacts with the neonatal Fc receptor (FcRn), which protects the protein from degradation, increasing the drug’s half-life in the circulation
Q1: What is an antibody-drug conjugate (ADC)?
An ADC is a biopharmaceutical that combines a monoclonal antibody with a cytotoxic drug. The antibody targets specific cells, delivering the drug directly to the disease site.
Q2: How do ADCs work?
The antibody portion binds to a specific target (e.g., a cancer cell).
The cytotoxic drug is delivered directly to the target, minimizing damage to healthy cells.
Once inside the target cell, the drug is released, killing or inhibiting the c
Q3: What is the advantage of using an ADC over a free cytotoxic drug?
A3: ADCs increase specificity, delivering the drug to diseased cells while reducing systemic toxicity, improving safety and effectivenes
Q4: What are ADCs used for clinically?
They are mainly used in cancer therapy, where precise delivery of potent cytotoxic agents is critical.
Q1: What are multi-specific antibodies?
Multi-specific antibodies are biopharmaceuticals engineered to bind two or more different targets simultaneously, often combining multiple antibody fragments or domains.
Q2: What structural components can multi-specific antibodies include?
They can include combinations such as:
Fab fragments (antigen-binding regions)
Fc domains (constant region for half-life and immune recruitment)
Q3: Give an example of a multi-specific antibody and its use.
Mosunetuzumab is a multi-specific antibody used in cancer therapy, designed to engage both tumor cells and T cells to enhance immune-mediated killing.
What is the main advantage of multi-specific antibodies over traditional monoclonal antibodies?
They can target multiple pathways or cells simultaneously, increasing efficacy and potentially overcoming resistance mechanisms in diseases like cancer.
Q1: What is a biosimilar?
A biosimilar is a biologic medicine that is highly similar to an already licensed biologic, with no clinically meaningful differences in quality, safety, or efficacy. (Royal Pharmaceutical Society, 2018)
Why can a biosimilar never be an exact copy of the original biologic?
: Because biologics are produced in living systems, there is inherent heterogeneity due to:
Charge variants
Glycan profile differences
Minor structural variations
This makes an identical generic copy impossible.
Q3: What is the purpose of developing biosimilars?
They are specifically developed and licensed to treat the same disease(s) as the original innovator biologic, offering more treatment options and often lower cost.
Q4: How does the source of the antibody affect a biosimilar?
The host species and growth media can affect glycosylation patterns (addition of sugar chains), which influence:
Drug function
Half-life in the body
Immunogenicity (potential to trigger immune response)
Why is correct glycosylation important for monoclonal antibodies (mAbs)?
Even though oligosaccharides contribute only ~5% to the mAb size, they have a major impact on function, including:
Sialic acid removal → exposes galactose → faster excretion, decreases half-life (e.g., EPO from 2h → 10 min)
Loss of alpha-1,6 fucose → enhances ADCC activity for IgG
How do glycosylation differences influence biosimilar behaviour?
Glycan patterns affect:
Drug efficacy (e.g., ability to trigger immune cell killing)
Drug stability and half-life
Immunogenicity – different sugar patterns may provoke an immune response depending on the species used to produce the mAb
Extra simple summary:
Biosimilars are “highly similar but never identical” to the original biologic. Small changes in sugar chains can dramatically change how the drug works, lasts, and interacts with the immune system.
Q1: What is the difference between primary and higher-order structures in drugs?
Primary structures (like aspirin or paclitaxel) are small molecules with strong covalent bonds, giving a simple and stable structure.
Higher-order structures (secondary, tertiary, quaternary) exist in large biologics, such as hemoglobin or monoclonal antibodies, and are held together by a mix of strong covalent and weaker non-covalent bonds (hydrogen bonds, van der Waals forces, ionic interactions).
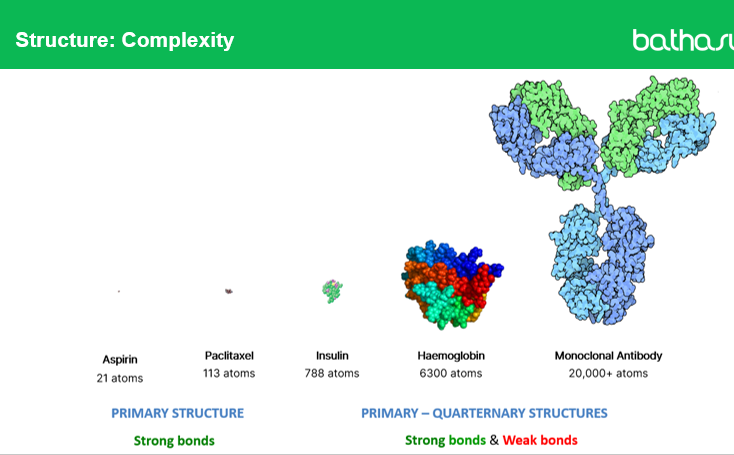
Q2: Why do quaternary structures like monoclonal antibodies have more complex stability considerations than small molecules?
Because quaternary structures:
Depend on weak bonds that can be disrupted by temperature, pH, agitation, or chemical environment
Are sensitive to denaturation, aggregation, or degradation, making handling and formulation more complex than small-molecule drugs
Q3: Give an example of a drug with a simple primary structure and one with a quaternary structure.
Primary structure example: Aspirin, paclitaxel
Quaternary structure example: Hemoglobin, monoclonal antibodies (mAbs)
Q4: How does the complexity of monoclonal antibodies affect their manufacturing and formulation?
A4: Complex structures mean:
Careful control of pH, temperature, and ionic conditions is needed
Formulations must preserve protein folding and glycosylation
Small changes can alter activity, half-life, or immunogenicit
Q1: What is protein aggregation and why does it occur?
: Protein aggregation is the abnormal association of proteins into larger structures, which tend to be insoluble. It occurs when native monomers unfold, exposing regions that stick together to form clusters, nuclei, oligomers, and aggregates.
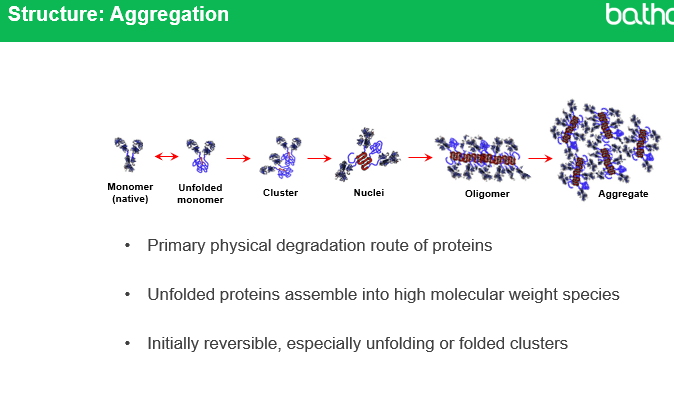
Q2: What is the typical size range for antibody monomers and aggregates?
mAb monomers: ~10 nm
Soluble aggregates: detectable with laser diffraction
Insoluble aggregates: up to ~50 µm detected with microscopy
Aggregates >50 µm: sometimes visible to the naked eye
Q3: Are all stages of aggregation reversible?
Initially, some forms such as unfolded monomers or folded clusters can be reversible. Larger aggregates tend to be irreversible.
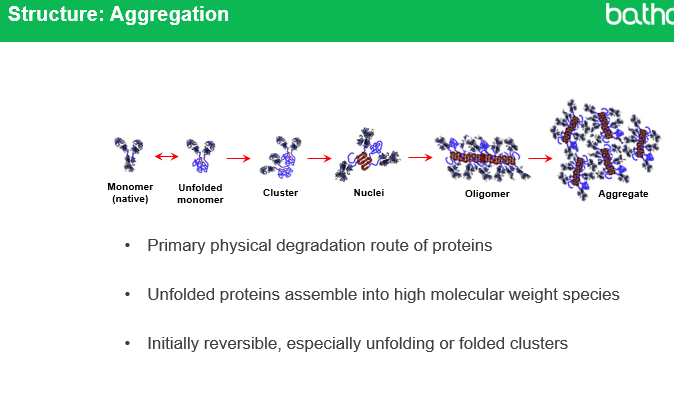
How does aggregation affect monoclonal antibody treatments?
Aggregation can:
Reduce effectiveness (loss of function)
Increase immunogenicity (aggregates are more likely to be taken up by antigen-presenting cells)
In severe cases, cause micro-vessel occlusion
Why is aggregation considered a primary physical degradation route for proteins?
Because unfolded proteins naturally assemble into high molecular weight species, making aggregation the main pathway by which protein therapeutics lose stability and function
means… Proteins have a specific 3D shape (fold) that they need to do their job. When they lose that shape (unfold), they become unstable and their insides get exposed — parts that were meant to be hidden away inside the protein are now on the outside.
Unfolded proteins are sticky → they clump together → clumping is the main reason protein drugs go bad and stop working
Extra note
Aggregation is a major concern in biopharmaceutical formulation, requiring controlled temperature, pH, and handling, as even small changes can trigger irreversible aggregation.
Q1: What is oxidation in the context of protein drugs?
Oxidation is the addition of oxygen to a molecule, which can alter the protein’s structure and function. In monoclonal antibodies,
methionine residues are most commonly oxidized,
but solvent-accessible tryptophan, cysteine, lysine, and histidine can also undergo oxidation.
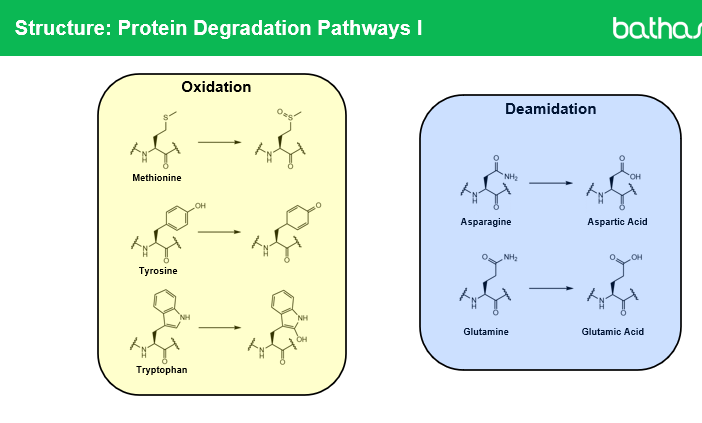
Q2: What is deamidation in proteins?
Deamidation is the conversion of asparagine residues into aspartic acid or iso-aspartic acid, which changes the protein’s charge and can affect stability and folding.
Q3: How does oxidation affect biopharmaceutical stability?
Oxidation can:
Alter protein structure
Reduce therapeutic activity
Promote aggregation if exposed residues cause abnormal interactions
Q4: How does deamidation affect biopharmaceutical stability?
: Deamidation:
Converts asparagine to aspartic or iso-aspartic acid
Alters charge state, which can reduce protein stability
Can promote aggregation due to charge and structural changes
Q5: How are oxidation and deamidation detected in proteins?
Oxidation is when a protein's amino acids (particularly methionine and tryptophan) react with oxygen, altering their chemical structure and potentially damaging the protein's function.
Deamidation is when certain amino acids (particularly asparagine and glutamine) lose an amine group, changing their chemical identity and altering the protein's charge and shape.
They can be detected using:
Mass spectrometry → detects changes in mass of the protein
Ion exchange chromatography → detects changes in charge due to chemical modification
Extra note:
Oxidation and deamidation are considered primary chemical degradation routes in proteins and must be monitored during formulation, storage, and handling to ensure safety and efficacy.
Q1: What is reduction in proteins?
Reduction is a chemical reaction that breaks disulfide bonds (-S-S-) between cysteine residues in a protein. This can:
Alter the protein’s 3D structure
Reduce stability
Promote aggregation if exposed thiols interact abnormally
Disulfide bonds broken → protein loses structural support → unfolds → aggregates
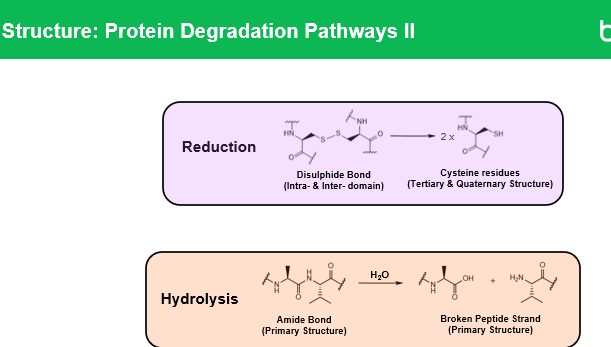
Q2: What is hydrolysis in proteins?
Hydrolysis is the breaking of peptide bonds in the protein backbone by water, which produces smaller fragments. It can be acid-catalyzed, base-catalyzed, or enzymatic, and can lead to:
Loss of monomeric active protein
Formation of smaller fragments that may aggregate
Changes in charge or solubility
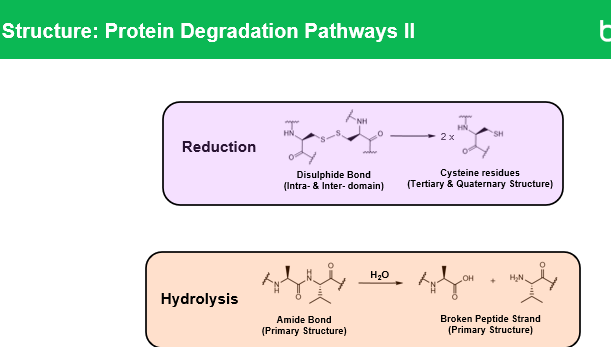
Q3: What happens during acid hydrolysis of antibodies?
Occurs at pH ~6
Produces protein fragments
Increases aggregation
Reduces monomer API (active drug)
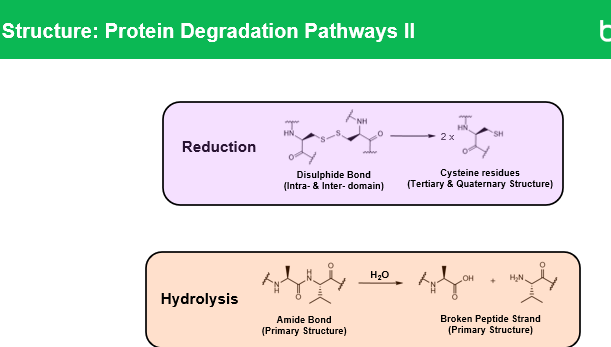
Q4: How is pH related to hydrolysis rates?
Acid hydrolysis slows at pH 8 and above
Most antibody formulations are between pH 5–7, where acid hydrolysis can still occur
Q5: What is isomerization in proteins?
Isomerization is the conversion of one amino acid form to another (e.g., asparagine → iso-aspartate) without changing the overall sequence. It can:
Alter charge distribution
Promote aggregation
Reduce protein activity
Q6: How does hydrolysis differ from acid hydrolysis specifically?
Hydrolysis is the general breaking of peptide bonds by water.
Acid hydrolysis occurs under slightly acidic conditions (pH ~6) and can produce fragments and aggregates more rapidly than neutral or basic conditions.
Extra summary:
Primary chemical degradation routes: oxidation, deamidation, hydrolysis, isomerization, and reduction
Implications: Loss of monomer, formation of aggregates, reduced activity, increased immunogenicity
Q1: What is meant by protein fragmentation in biopharmaceuticals?
Protein fragmentation is the breaking of a protein into smaller pieces, known as fragments. This can happen due to chemical or physical degradation and can reduce the therapeutic activity of the protein and increase the risk of immunogenicity.
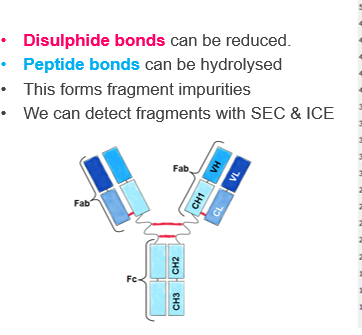
Q2: How can disulfide bonds cause fragmentation in proteins?
: Disulfide bonds (-S-S- bonds between cysteine residues) hold parts of the protein together. If these bonds are reduced or broken, the protein can separate into smaller domains, forming fragments. In antibody-drug conjugates (ADCs), some disulfide bonds are intentionally reduced to attach the cytotoxic drug, which is a controlled form of fragmentation.
Q3: How does peptide bond hydrolysis contribute to protein fragmentation?
Peptide bonds are the links between amino acids in a protein backbone. Hydrolysis, which is the reaction of water with these bonds, can break them, generating smaller protein fragments. This can happen spontaneously under certain pH or temperature conditions, producing impurities that reduce drug effectiveness.
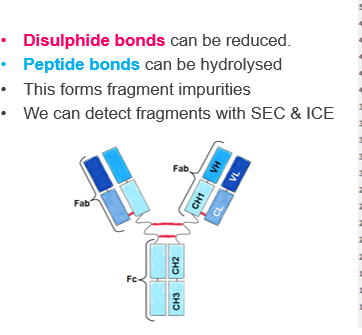
Q4: How are protein fragments detected in biopharmaceuticals?
Fragment impurities can be detected using analytical techniques such as:
Size Exclusion Chromatography (SEC): separates fragments based on size
Ion-Exchange Chromatography (ICE): separates fragments based on charge differences
These methods help ensure the protein remains intact and functional in the final product.
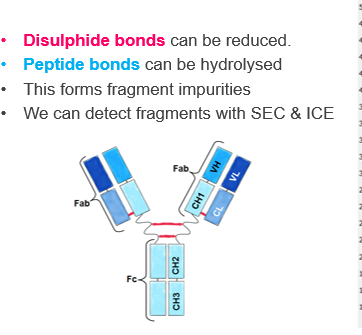
Q5: Why are Fab regions particularly susceptible to fragmentation?
The Fab regions of antibodies are more exposed and flexible compared to the Fc region. This makes them more prone to disulfide bond reduction or peptide bond hydrolysis, leading to the formation of fragments that can affect activity and stability.
Q1: What is a pH-sensitive linker and give an example?
A pH-sensitive linker breaks in acidic environments, like inside a cell’s lysosome, to release the drug.
Example: Gemtuzumab ozogamicin
Simplified: Think of it like a candy wrapper that melts in heat — the acidic environment “melts” the linker and frees the drug.
Q2: What is an enzymatically cleavable linker and give an example?
An enzymatically cleavable linker is cut by specific enzymes inside the target cell, releasing the drug.
Example: Brentuximab vedotin
Simplified: It’s like a string that only a certain pair of scissors (enzyme) can cut to release the payload.
Q3: What is a glutathione-sensitive linker and give an example?
A glutathione-sensitive linker breaks in the presence of glutathione, which is abundant inside cells.
Example: Gemtuzumab ozogamicin
Simplified: Imagine a lock that only opens when it senses glutathione, releasing the drug inside the cell.
Q4: What is a non-cleavable linker and give an example?
A non-cleavable linker does not break on its own; the drug is released only when the whole antibody is degraded inside the cell.
Example: Trastuzumab emtansine (thioether linker)
Simplified: Think of it as a strong box attached to the antibody — it only opens when the cell completely breaks down the antibody.
Key concept simplified:
The linker type controls when and where the drug is released.
pH-sensitive, enzymatic, and glutathione-sensitive linkers → release quickly inside target cells.
Non-cleavable linkers → release slowly, only after antibody degradation.
Q1: What is DAR in antibody-drug conjugates (ADCs)?
DAR stands for Drug-to-Antibody Ratio, which is the average number of drug molecules attached to each antibody.
Q2: Where can drugs be attached on an antibody in an ADC?
: Drugs are typically attached to lysine residues or cysteine residues on the antibody.
Q3: How does a high DAR affect an ADC?
Reduces half-life → the antibody is cleared from the body faster
Increases toxicity → more drug is delivered systemically, which can harm healthy tissue
Increases aggregation → higher risk of protein clumping, reducing stability
Simplified explanation:
Q1: What is photodegradation in biopharmaceuticals and ADCs?
1: Photodegradation is the breakdown of drugs caused by light exposure.
In ADCs, light-sensitive warheads can be damaged by light.
Light exposure can also accelerate oxidation of biopharmaceuticals, especially if Polysorbate surfactants are present.
Simplified: Light acts like a “trigger” that damages the drug and makes it less stable.
Q2: What is oxidative stress in biopharmaceuticals?
Oxidative stress occurs when proteins or ADCs are chemically oxidized, which:
Reduces functional activity (drug works less effectively)
Increases aggregation, forming clumps that reduce stability and may increase immunogenicity
Q3: How does light exposure increase oxidation?
Light can generate reactive species that attack sensitive residues (e.g., methionine, tryptophan), especially in the presence of surfactants like Polysorbate, accelerating chemical degradation.
Why is controlling light exposure important for ADCs and biopharmaceuticals?
To prevent loss of activity, reduce aggregation, and maintain stability during preparation, storage, and administration.
Q1: How does high temperature affect protein drugs?
High temperatures accelerate chemical interactions and degradation, increasing the chance of misfolding and aggregation.
Antibodies typically unfold around 65 °C.
Temperature cycling (repeated heating and cooling) increases the risk of misfolding.
Simplified: Heat is like shaking a fragile origami — it can make it unfold and break.
Q2: What happens to proteins during freeze-thaw cycles?
Freeze-thaw cycles can cause:
Excipient crystallisation, which may damage the protein environment
Protein-ice interfaces, where the protein can unfold or aggregate at the ice boundary
Simplified: Freezing and thawing is like putting a sponge in ice — the edges get damaged and parts stick together.
Q3: Why is controlling temperature important for biopharmaceuticals?
To maintain protein structure, stability, and function, and to minimise aggregation and degradation during storage and handling.
Q1: How are protein-based medicines like antibodies usually formulated?
They are usually formulated as freeze-dried powders (lyophilisates), which are stable for storage but not ready to administer directly to patients.
What happens when biopharmaceuticals are compounded into a ready-to-use medicine?
Compounding drastically changes the drug environment:
Reconstitution exposes proteins to aqueous diluents
Shaking or shear forces can make hydrophobic portions of the protein unfold, leading to aggregation
Why must pharmacists be careful when handling compounded biopharmaceuticals?
Pharmacists must remember that:
Compounding involves changes from the marketed, stable form
Stability data may come from the manufacturer, but the hospital pharmacist is responsible for safe use
Improper handling can reduce drug effectiveness or increase immunogenicity
Q4: How can materials used in administration contribute to aggregation?
Metals used in needles, such as tungsten, can interact with proteins and induce aggregation.
Simplified: The metal acts like a rough surface that makes proteins stick together instead of staying properly folded.
What are the main steps involved when reconstituting and diluting monoclonal antibodies (mAbs)?
Start with a lyophilised mAb (freeze-dried powder)
Reconstitute to make a concentrated solution
Dilute further to prepare a patient-ready dose (can be diluted 2–40×)
Q2: What chemical stresses can be introduced during reconstitution and dilution?
Chemical species that can catalyse protein degradation include:
Water
Metal ions (e.g., tungsten from needles)
Silicone (from stoppers or tubing)
Oxygen
Rubber bung materials
Simplified: These are like “pollutants” that can make the protein unfold, oxidize, or stick together.
Q3: How can adsorption affect biopharmaceuticals during manipulation?
Proteins can stick to container surfaces, leading to loss of active drug
This reduces the effectiveness of the medication
Stabilising excipients are used to prevent adsorption, but improper handling can override this protection
What equipment/materials can contribute to protein aggregation or loss during reconstitution?
Tungsten needles or polyolefin spikes can trigger aggregation
Surfaces in vials, syringes, and tubing can cause adsorption
Q1: How does agitation affect protein drugs?
Agitation physically increases interactions between protein molecules, which can lead to unfolding and aggregation.
Q2: How does agitation affect oxygen levels in a solution?
Agitation can dissolve more oxygen into the solution, which may accelerate oxidation of sensitive amino acids (e.g., methionine, cysteine).
Simplified analogy:
Shaking a bottle of protein solution is like stirring a fragile soup — it increases collisions and can let in extra air that “spoils” the protein.
Why agitation is harmful for mAbs
Increases protein-protein interactions
Shaking or stirring increases collisions between antibody molecules.
This can expose hydrophobic regions, leading to unfolding and aggregation.
Dissolves oxygen into the solution
More oxygen means oxidation of sensitive residues (like methionine or cysteine), which can reduce activity and increase immunogenicity.
Reduces stability
Agitation can accelerate chemical and physical degradation, shortening the shelf-life of the protein.
Q1: What materials are commonly used for syringes?
Syringes: Mostly polypropylene (PP) barrels, but can also include polycarbonate or polyisoprene, with rubber/latex plungers.
Q1: What materials are commonly used for infusion bags?
Infusion bags: PVC, polyethylene (PE), polypropylene (PP), ethylene vinyl acetate (EVA), and polyamide (PA).
Q2: How does container design affect protein stability?
Surface area to volume ratio: Higher ratio → more protein adsorption to the container.
Headspace (air above liquid): More headspace → more liquid-air interface, oxygen exposure, and potential physical agitation.
Q3: How does protein concentration affect aggregation?
High concentration increases protein-protein interactions, leading to more aggregation.
BUT, higher concentration can allow better excipient stabilisation, which reduces aggregation if formulated correctly.
How does the choice of diluent affect protein stability?
WFI (Water for Injection): Low tonicity → increases intermolecular attraction, promoting aggregation.
5% Glucose: Can cause glycation, not suitable for acidic proteins.
0.9% NaCl: Reduces intermolecular attraction, stabilising proteins.
Q5: How does viscosity affect protein stability?
High viscosity protects proteins from physical stress during handling and manufacture.
Q6: What excipients are used to prevent aggregation?
Detergents: Polysorbate 20 / 80 (TWEEN)
Polymers: Poloxamer-188
Cationic amino acids: Arginine, Lysine
Q7: What antioxidants are commonly used in biopharmaceuticals?
Amino acids: Methionine, Histidine
Non-reducing sugars: Sucrose, Trehalose
Q8: What metal chelators are used to stabilise proteins?
DTPA (Diethylenetriaminepentaacetic acid)
EDTA (Ethylenediaminetetraacetic acid)
These bind metal ions that could catalyse oxidation or degradation.