MOL unit 4
1/61
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
62 Terms
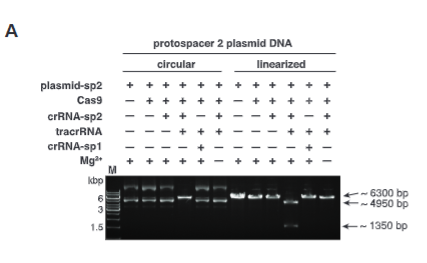
explain fig 1 A Jinik
an agarose gel with both linear and circular forms of a plasmid. The circular one shows that Cas9 alone does not cut it into a linear form, so they add more and more things until it shows cutting. They used the already linearized plasmid as a confirmation of the cutting abilities.
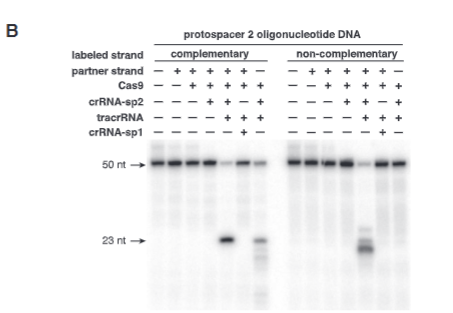
explain fig 1B jinik
this is a polyacrylamide gel of single-stranded DNA. They added radioactvitiy to the 5’ end of the noncomplimentary strand then assmebed the gel. They separated to run single stranged on the gel and then when they cut can onyl see the radioactive lable.
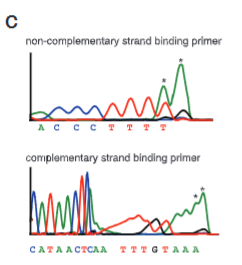
explain fig 1C jinik
sequencing reactions to see where Cas9 is cutting DNA, we see stopping in the graphs where cleavage takes place.
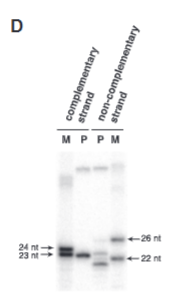
explain fig 1D jinik
they wanted to confirm the size of cutting using different DNA ladders
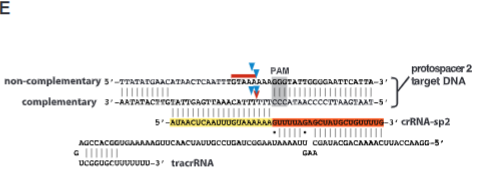
explain fig 1E jinik
this set up is showing how tracrRNA and protospacer and crRNA work together. schematic representation of tracrRNA, crRNA-sp2, and protospacer 2 DNA sequences. Regions of crRNA complementarity to tracrRNA (orange) and the protospacer DNA (yellow) are represented. The PAM sequence is shown in gray; cleavage sites mapped in (C) and (D) are represented by blue arrows (C), a red arrow [(D), complementary strand], and a red line [(D), noncomplementary strand].
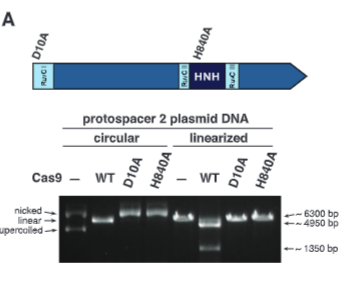
explain fig 2A jinik
at H840 amino acid there is a mutation in the HNH domain. At D10A there is a mutation in the RuvC1 domain. The mutated areas do not cut like the wild type Cas9, so these domains are domains important for cutting.
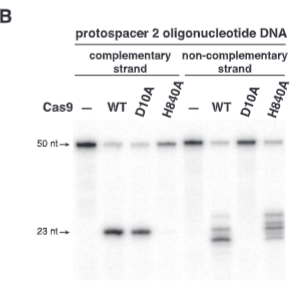
explain fig 2B jinik
HNH mutation affects the complementary strand (it does not cut), RuvC1 domain mutation affects the noncomplementary strand (it does not cut). HNH is responsible for cutting the complementary strand, and Ruvc1 is responsible for the noncomplementary strand.
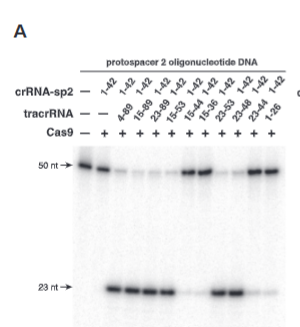
explain fig 3A jinik
tracrRNA nucelotide imporance shown in A
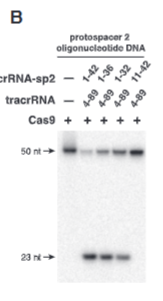
explain fig 2B jinik
crRNA nucelotide imporance shown in B. 1-10 are critical becasue without them it doesnt cut.
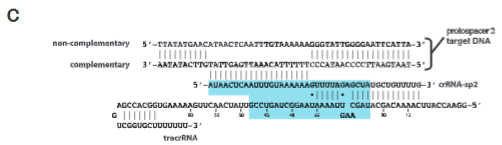
explain fig 2C jinik
Minimal regions of tracrRNA and crRNA capable of guiding Cas9-mediated DNA cleavage (blue shaded region).
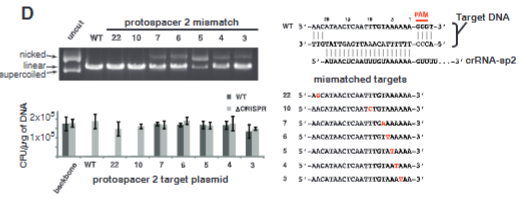
explain fig 3D jinik
More bacteria that die the more critical that sequence is, the closer to the PAM site the mroe critical protospacer matching to PAM becomes. Plasmids containing WT or mutant protospacer 2 sequences with indicated point mutations (right) were cleaved in vitro by programmed Cas9 as in Fig. 1A (top-left) and used for transformation assays of WT or pre-crRNA–deficient S. pyogenes (bottom-left). The transformation efficiency was calculated as colony-forming units (CFU) per microgram of plasmid DNA.
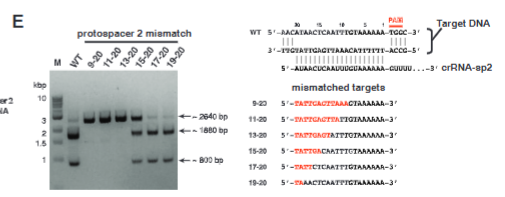
explain fig 2E jinik
The more change in nucleotides the more affected the cutting is
what is PAM
Protospacer Adjacent Motif- a short motif that dictates R-loop formation. PAM is found downsteream of crRNA binding sequence NGG following.
when can you find NGG seqeunce
maybe every 8 nucelotides
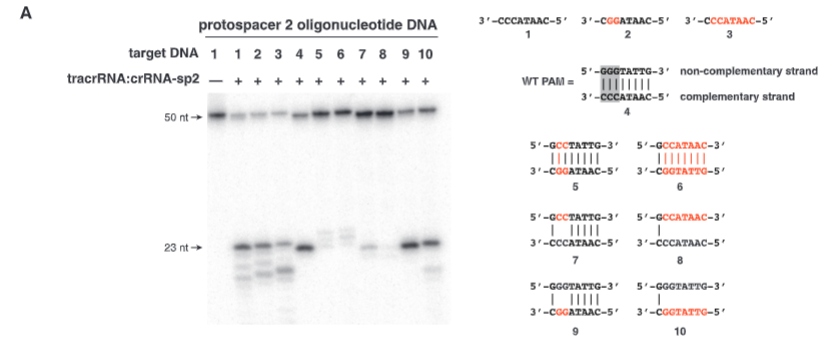
explain fig 4A jinik
Polyacrylamide gel, N-labeled, and tested the need for the NGG sequence. The results showed that GG being in the noncomplementary spot is important for cutting to occur.
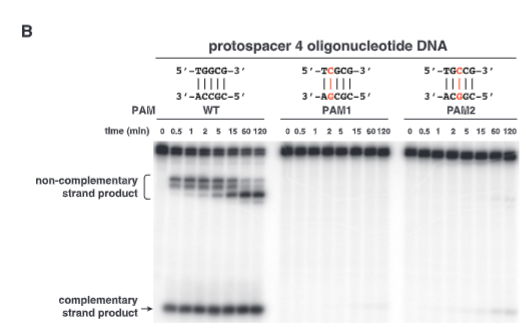
explain fig 4B jinik
3 different PAM sites, changing either of the G sites show dramatic decrease in cutting
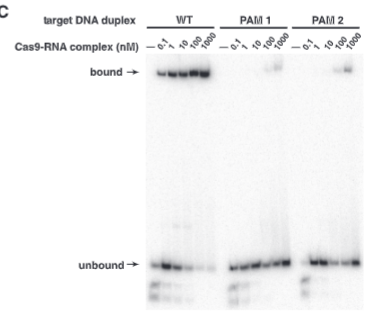
explain fig 4Cjinik
3 PAM sites with mutations with different amounts of Cas9-rna complex added, shows that no matter how much Cas9 compelx there is the PAM site is very imporant for binding
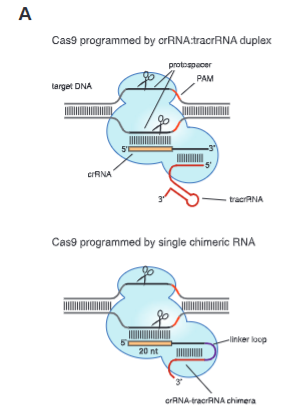
explain fig 5A jinik
Top) In type II CRISPR/Cas sys- tems, Cas9 is guided by a two-RNA structure formed by activating tracrRNA and targeting crRNA to cleave site-specifically–targeted dsDNA (see fig. S1). (Bottom) A chimeric RNA generated by fusing the 3′ end of crRNA to the 5′ end of tracrRNA.
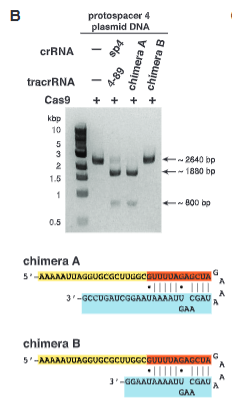
explain fig 5B jinik
Chimera A cuts, but Chimera B does not, this shows that the sequence is important, because Chimera B lacks some of the tracrRNA sequence and does not work
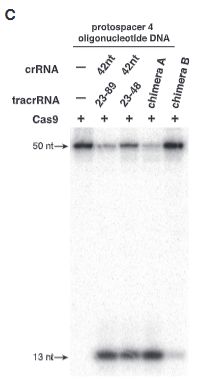
explain fig 5C jinik
shows Chimera B not cutting as in fig 5B
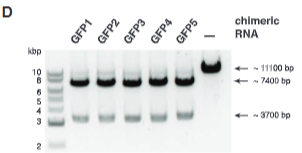
explain fig 5D jinik
Created chimeric RNA around PAM sites, and everything they target gets cut. Five chimeric RNAs designed to target the GFP gene were used to program Cas9 to cleave a GFP gene–containing plasmid. Plasmid cleavage reactions were per-formed as in Fig. 3E. In all five cases, Cas9 programmed with these chimeric RNAs effi-ciently cleaved the plasmid at the correct target site (Fig. 5D and fig. S15D), indicating that ra-tional design of chimeric RNAs is robust and could, in principle, enable targeting of any DNA sequence of interest with few constraints beyond the presence of a GG dinucleotide adjacent to the targeted sequence
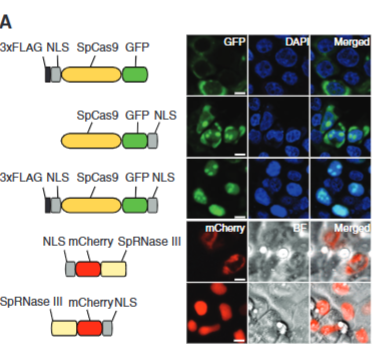
explain fig 1A cong
Transfected into cell and wanted to see if protien is made (to see if Cas9 gets into the nucelaus) Satined DNA w/ Dap1 to see the nucelus and labled Cas9 with GFP (green), the first line across shows Cas9 not in the nucelus, so they moved the NLS site to the Carboxyl Terminus and saw some staining in the nucleus. They then tagged both ends of the Cas9 with NLS and say clear staining in the nucelus. The red one shows RNase 3 labled with red dye and used RNase 3 because it is needed with Cas9 fro processing, so they needed it in the nucelus as well. Results showed that NLS is needed on the carboxyl terminus.
what is NLS
Nuclear localization signal, it is a tag that tell the cell machinery to take it to the nucleus
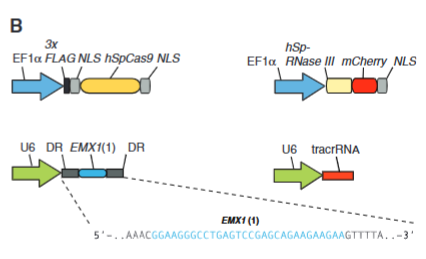
explain fig 1B cong
This tries to answer how the A seqeunces get transcribed. Promoter EF1 alpha gets recognized by the RNA polymerase 3 for trasncription. Use EF1 alpha as promoter for Cas9 and RNase 3. Used EMX 1 as it has a PAM site, and this is the DNA they wanted to target to cut. They also had to make tracrRNA. Mamma-lian expression of human codon–optimized SpCas9 (hSpCas9) and SpRNase III (hSpRNase III) genes were driven by the elonga-tion factor 1a (EF1a) pro- moter, whereas tracrRNA and pre-crRNA array (DR- Spacer-DR) were driven by the U6 promoter. A protospacer (blue highlight) from the human EMX1 locus with PAM was used as template for the spacer in the pre-crRNA array. EF1A was used as it efficiently recruits RNA polymerase II. U6 was used as it recruits RNA Polymerase III.
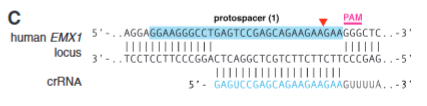
explain fig 1C cong
Shows protospacer w/ guide RNA and EMX1 set up that has a PAM site
what does DR stand for
direct repeats (protospacer-repeat-protospacer-repeat)
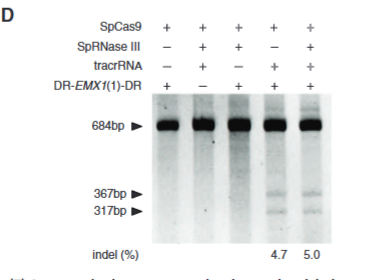
explain fig1D cong
Shows what is needed for cutting, RNase 3 not super important for guiding the sequence for where cutting is needed.
stage 1 of CRISPR-associated Cas Response
the bacteria aquires the pathogens DNA and inserts it into a Locus on the 5’ end. Upstream of this locus (crispr) is a promoter and upstream of that are Cas genes. CRISPR locus gets trasncribed to RNA this is called precrRNA. There are regulary repeated palindromes in the pre-crRNA that allow formation of stem loop structure
stage 2 of CRISPR-associated Cas Response
CRISPR RNA gets processed and clipped (durign this Cas and trasncribed to protiens and these protiens then associate w/ individual clipped RNA from CRISPR)
stage 3 of CRISPR-associated Cas Response
RNA guided targeting of the viral element, now bacteria has specific seqeunce (guided missle) to fight the virus.
the type 2 CRISPR system
uses RNase protien to do the processing of pre-crRNA and requires base-pairing structure between crRNA and tracrRNA to be able to allow Cas9 (specific to type 2) to cleave phage DNA
explanation given by Cong authors as to why RNaseIII is not super imporant for adding when cutting cells?
Since cells already have RNase, it might be doing the processing naturally without having to add additional RNase to the cell.
what is an indel
insertion or deletion as a result of an NHEJ
what is an NHEJ
Non homologous end joining- DNA repair mechanism that allows cells to quickly repair double-strand breaks (DSBs) without the need for a template. Howver, this leaves mutations as it is not always the best.
Surveryor assay
runs through the DNA samples to find indels, the more indel the more cutting since it shows more need for NHEJ to repair.
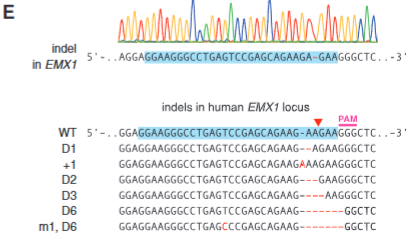
explain fig 1E cong
indel in EMX1 shown, found a variety of indels showing that indels do happen
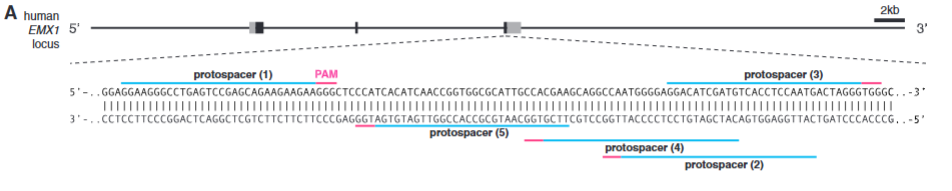
explain fig 2A Cong
Schematic of the human EMX1 locus showing the location of five protospacers indicated by blue lines with corresponding PAM in magenta.
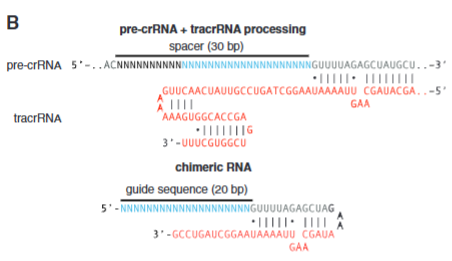
explain fig 2B Cong
Model that shows that you can make chimeric RNA to only use 1 RNA instead of 2
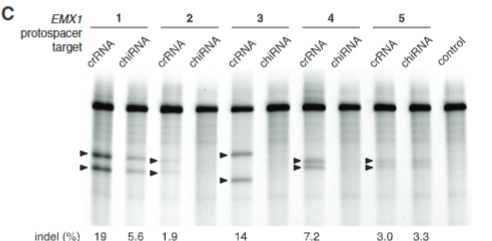
explain fig 2C Cong
compares how well chimeric or crRNA cut results show that crRNA cuts better, but chimeric still cuts so it still may be woth it
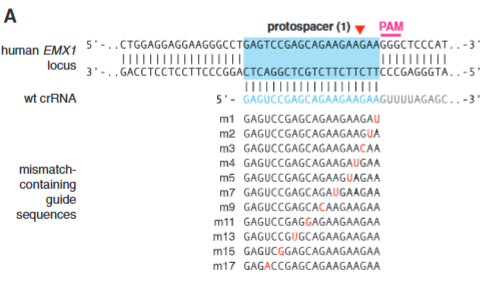
explain fig 3A Cong
Shows mismatched crRNA at differnt location close adn far from the PAM sites
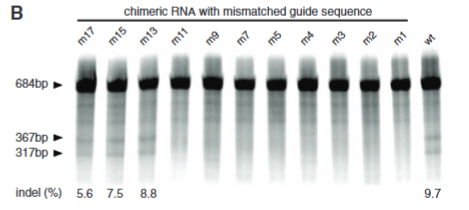
explain fig 3B Cong
this tests the mismatch from fig A and shows the closer to the PAM site the mismatch the less effetive Cas9 is at cuting

Explain fig 3C Cong
model showing TALEN gene cutting
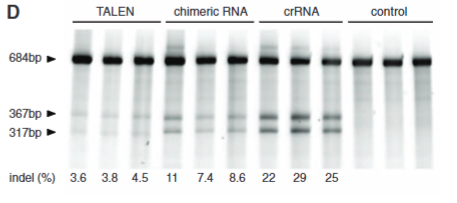
Explain fig 3D Cong
Tests the effecieny of TALEN, Chimeric, and crRNA shows that crRNA works best
explain the overall of Cong
The researchers engineered the Streptococcus pyogenes Cas9 system so it could function inside mammalian cells. They demonstrated that Cas9, guided by short RNA molecules, could target specific DNA sequences and create double-stranded breaks at chosen genomic locations. These breaks were then repaired by the cell, leading to targeted genome modifications.; One of the important findings in Multiplex Genome Engineering Using CRISPR/Cas Systems was that the researchers engineered Cas9 into a “nickase” version to improve homology-directed repair (HDR). Normally, wild-type Cas9 creates a double-stranded break in DNA. In this paper, the authors mutated the RuvC domain of Cas9 (D10A mutation), producing a Cas9 nickase (SpCas9n) that cuts only one DNA strand instead of both.
what did Cong want to look at
they wanted to look at how to edit the genome at more than one site, they specifically used mamallian cells not just a plasmid.
differences between type 1 2 and 3 crispr/cas systems
Type I and Type III systems use multi-protein complexes to recognize and cleave targets.
Type II systems use a single large protein, Cas9, guided by RNA to perform both target recognition and DNA cleavage.
Type II systems became the preferred genome-editing tool because changing the target only requires changing the guide RNA, making the system easier to engineer and use in mammalian cells.
process of crispr/cas9
The process works like this:
Scientists design a guide RNA that matches the gene they want to target.
The guide RNA binds to Cas9.
The guide RNA leads Cas9 to the matching DNA sequence next to a PAM site.
Cas9 cuts the DNA.
The cell repairs the cut:
Non-homologous end joining (NHEJ) may create small mutations or knock out a gene.
Homology-directed repair (HDR) can insert or replace DNA if a repair template is provided.
two things needed for crispr/cas9 system
The system has two main parts:
Guide RNA (gRNA)
A short RNA sequence designed to match a specific DNA target.
It acts like a GPS or address label that tells Cas9 where to go.
Cas9 protein
An enzyme that cuts DNA.
Once guided to the correct sequence, Cas9 makes a double-stranded break in the DNA.
explain the importance of use of tracrRNA
The main job of tracrRNA (trans-activating CRISPR RNA) is to help Cas9 recognize and use the crRNA to cut target DNA.
In Type II CRISPR/Cas9 systems, tracrRNA:
binds to the crRNA through complementary base pairing,
helps process immature pre-crRNA into mature crRNA,
recruits and stabilizes the Cas9 protein,
and forms the active RNA-Cas9 complex needed for DNA targeting.
Without tracrRNA, Cas9 cannot properly recognize the guide RNA or efficiently cut DNA.
explain the importance of crRNA
The crRNA (CRISPR RNA) is important because it determines the DNA sequence that Cas9 will target and cut.
Its main role is to provide specificity to the CRISPR/Cas system. The crRNA contains a short guide sequence that is complementary to a target DNA sequence. This allows the CRISPR complex to recognize a precise location in the genome.
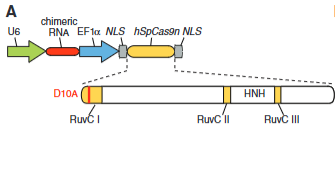
explain fig 4A cong
A model showing a mutation to RuvC in the 10th amino acid to ake RuvC inactive, making HNH domain in Cas9 a nicking enzyme. also shows addition of RNA and Cas9 in the same plasmid to simplify the system, instead of having two seperate ones
what is a nicking enzyme
nickase- when it only cuts one strand, not both
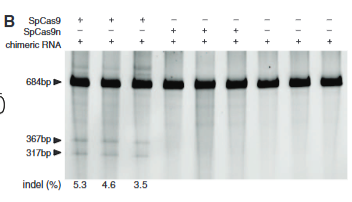
explain fig 4 B cong
survaor assay done to test the nicking Cas9 vs the regular Cas9 shows that nicking does not cut enough-or at least it doesn’t show up here as a survayer assay
explain fig 4C cong
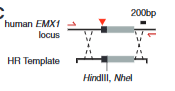
model of HDR- shows that bad DNA is cut out and new fixed seqeunce is instered with a recognition seqeunce that is recognized by enzymes that can cut it if desired to
what is HDR
Homology driven repair (fixes everything leaving no mutations unlike NHEJ and can edit in or out DNA)
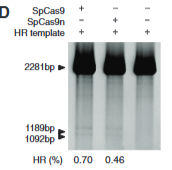
explain fig 4D cong
looks at Homologour recombination %- how often did swapping take place. this assay shows that nickase Cas9 does actually cut the single stranded-unlike B said- it was just not recognized by the indel assay
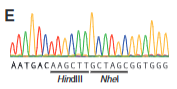
explain fig 4E cong
when they sequcned clones from HDR they saw successful recombination because the HindII/Nhel where not originally in it but they are seen now
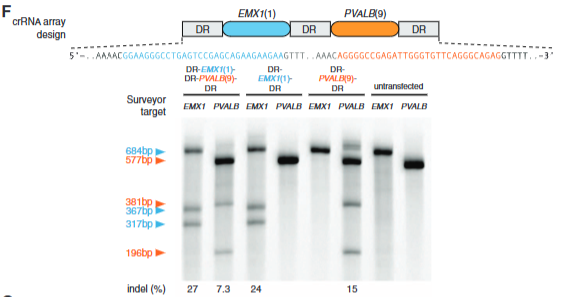
explain fig 4F cong
shows that both protospacers result in indels, cutting in both- at the same time, and shows that they can specifically target either or depending on the protospacer they use
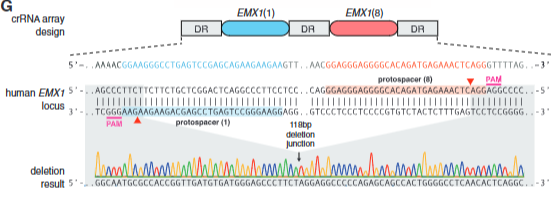
Explain fig. 4 G cong
construct w/ 2 protospacer targets in the EMX gene itself and show that they can cut out a specific segment precisely and then have recombination.
what does NHEJ target
double-stranded breaks
what did we learn from jinek?
Bacteria and archaea immune systems can remember virus infections
• Insert pieces of viral DNA (protospacers) into the CRISPR locus of their genome
• Transcribing this DNA yields crRNA used to guide an endonuclease (Cas9) to a new infection
• A second trans-activating RNA links Cas9 to the crRNA
• crRNA and tracrRNA can be made into a single, functional chimeric RNA
• A PAM site on the protospacer DNA completes the digestion mechanism
what did we learn from cong
By adding a nuclear localization signal (NLS), Cas9 can reach chromosomes in eukaryotic cells
• Using an RNA Pol III promoter results in an RNA molecule (e.g. it does not get translated)
− crRNA
− tracrRNA
• crRNA can guide Cas9 to digest endogenous DNA