mcdb 6 midterm 2 (week 4-7)
1/84
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
85 Terms
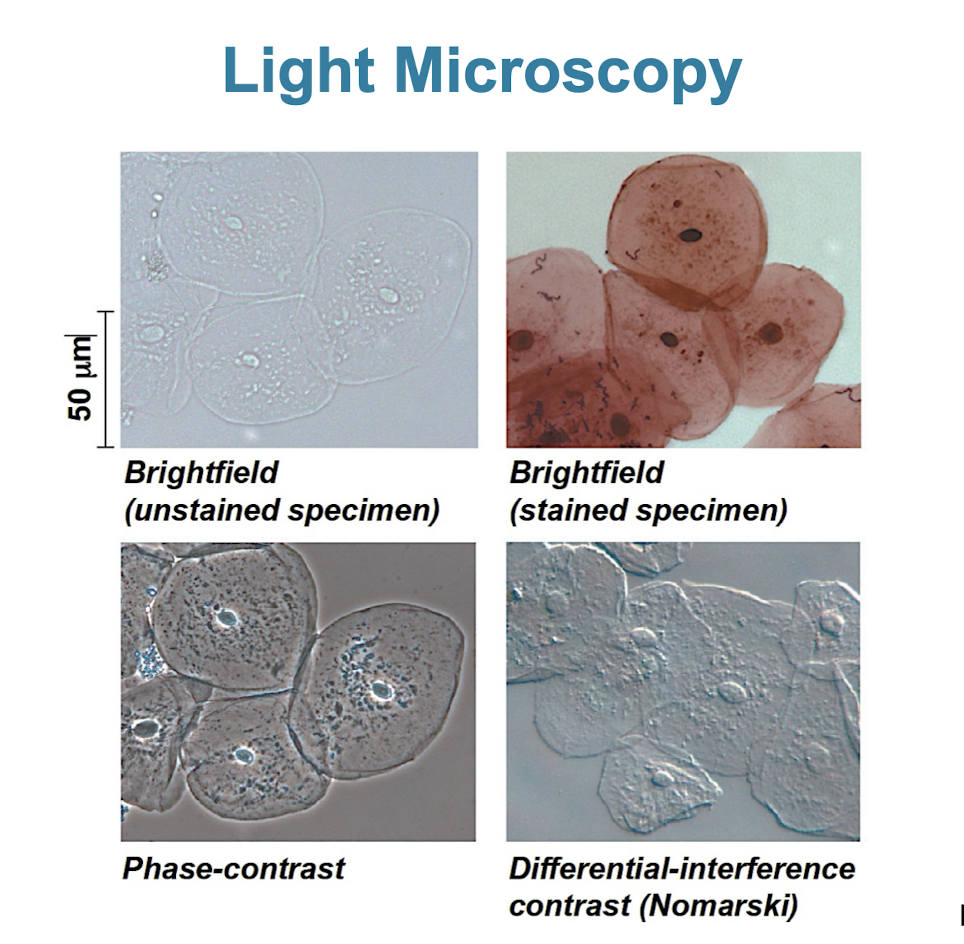
Light microscope (LM)
visible light is passed through a specimen and then through glass lenses
lenses refract/bend the light to magnify image
not able to see membrane organelles/sub-cellular structures (up to 200nm)
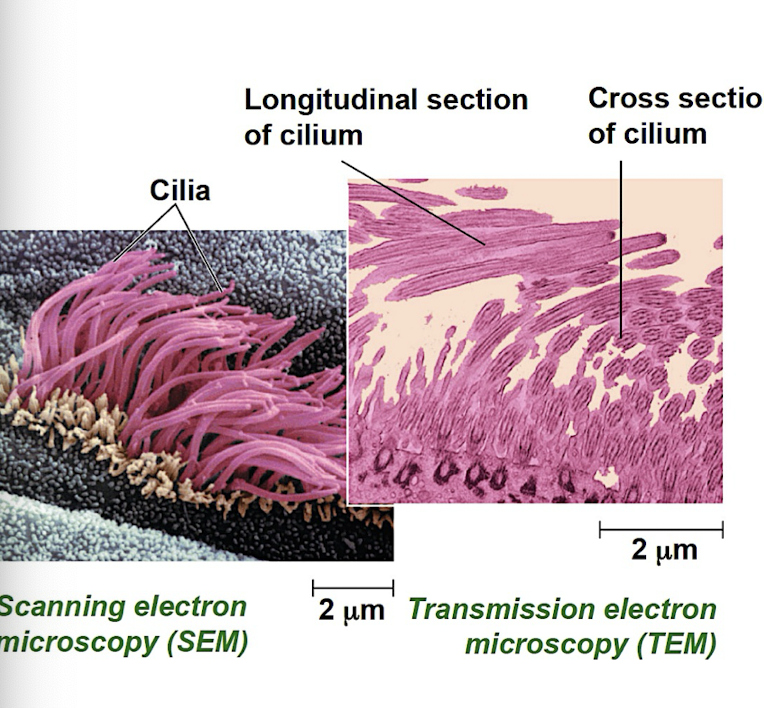
Electron Microscope (EM)
Scanning electron microscope (SEM): focus beam of electrons onto the surface of specimen producing 3D images
Transmission electron microscope (TEM): focus beam of electrons through a specimen to study internal structure of cells
size of plant and animal cells
10-100 μm
size of nuclei, bacteria, and mitochondria
1- 10 μm
3 important parameters of microscopy
Magnification (ratio of image size to real size)
Resolution (minimum distance between 2 distinguishable points)
Contrast (difference in brightness between light and dark areas of image)
advances in light microscopy
fluorescent markers label molecules to improve visualization of details
confocal microscopy w/ sharpened images of tissues/cells
improved resolution (as small as 10–20 nm)
Cell fractionation
breaks up the cells (sonication) and separates the components using centrifugation (based on size and weight)
help correlate cell function with structure
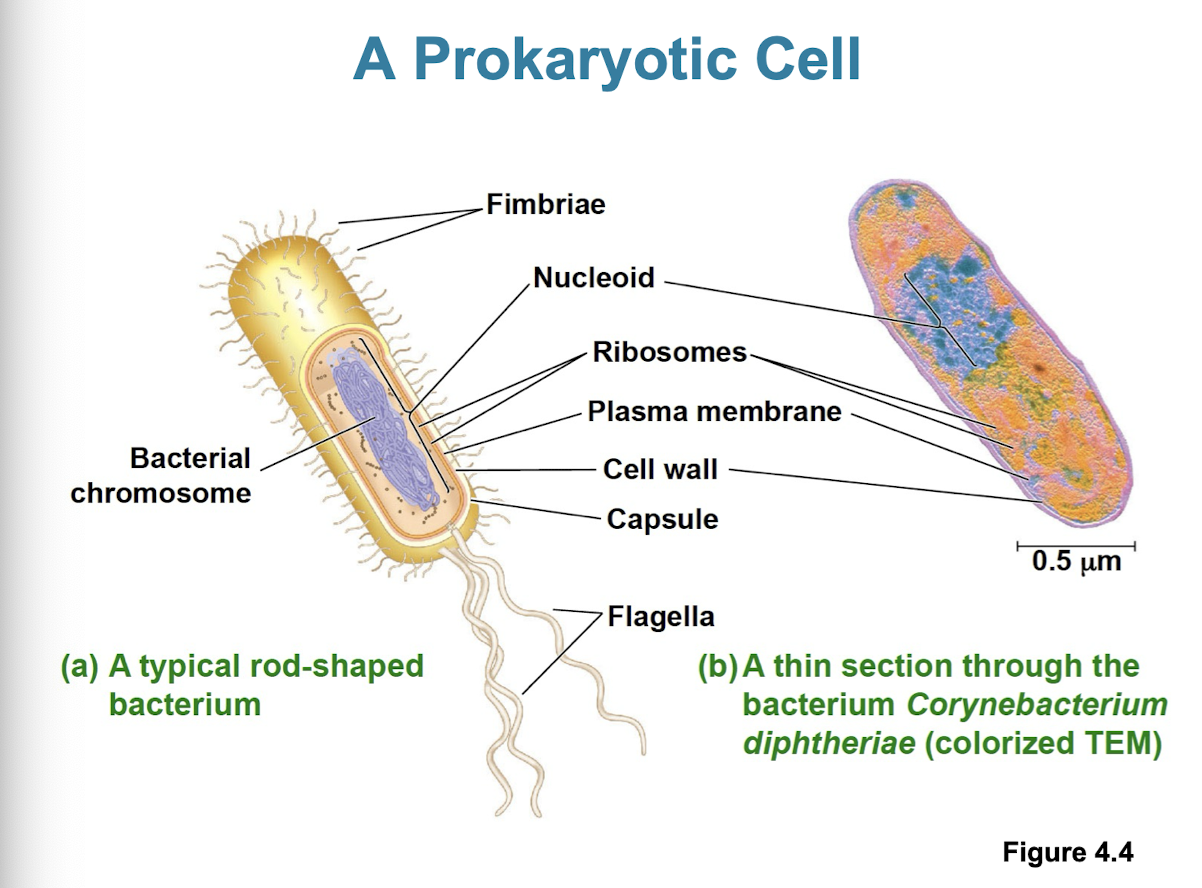
Prokaryotic cells (Archaea/Bacteria)
no nucleus/ membrane bound organelles
DNA is an unbound region called nucleiod
plasma membrane w/ cytoplasm inside and rigid cell wall on outside
1–5 μm
divides by binary fission
Eukaryotic cells (animal, plant, fungi, and protist cells)
DNA is in the nucleus (organelle bound by double membrane)
larger than prokaryotic cells
plasma membrane w/ cytoplasm inside
10–100 μm
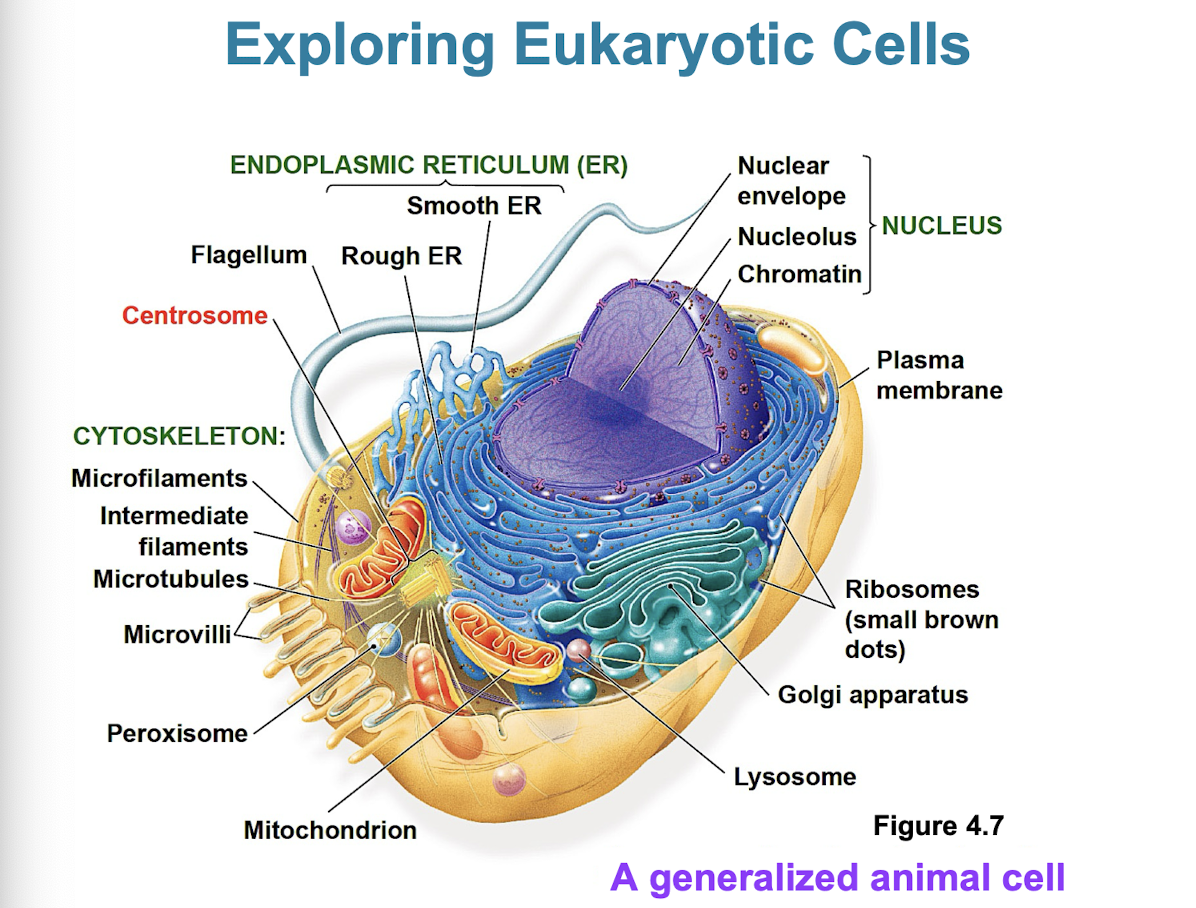
Basic features of all cells
plasma membrane
semifluid substance (cytosol)
chromosomes (carry genes)
ribosomes (make proteins)
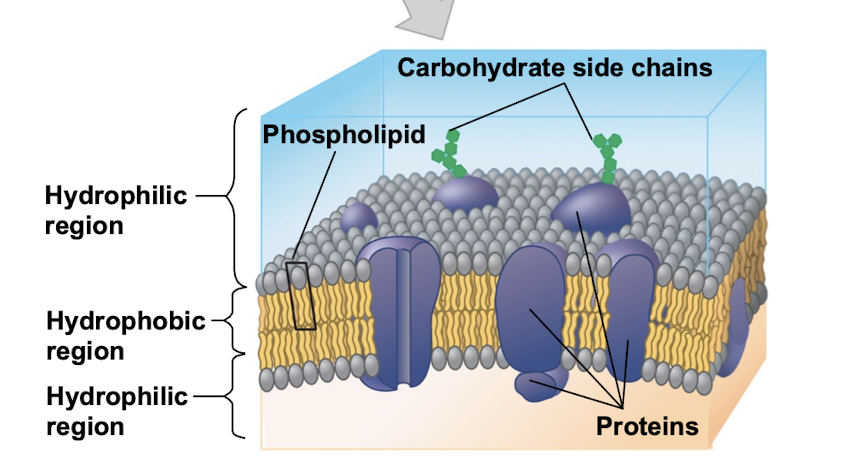
Plasma membrane
selective barrier that allows the passage of oxygen, nutrients, and waste to service the volume of every cell
double layer of phospholipids (bilayer)
proteins embedded inside
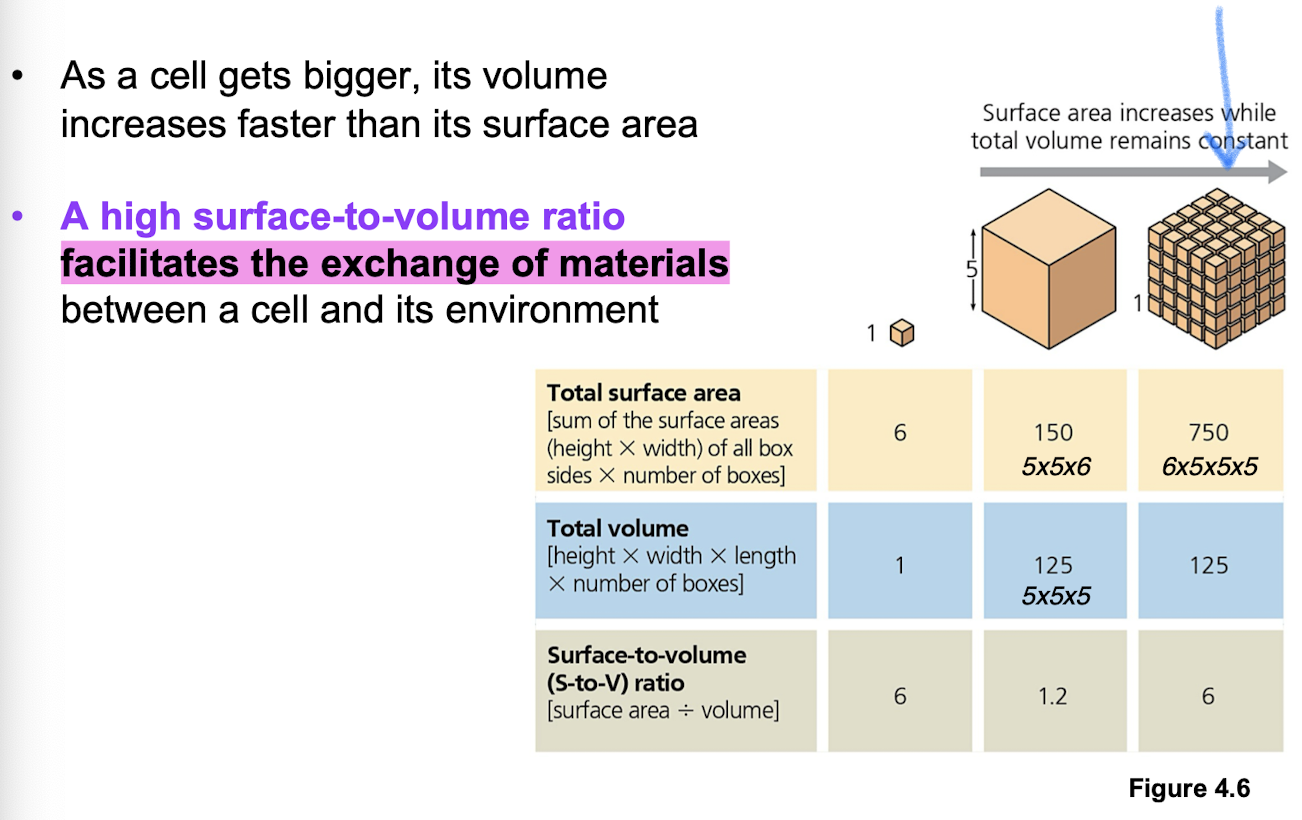
What limits cell size?
Diffusion + metabolism (upper limits) —> ratio of surface area to volume is critical to facilitate exchange of materials (n² sa and n³ volume)
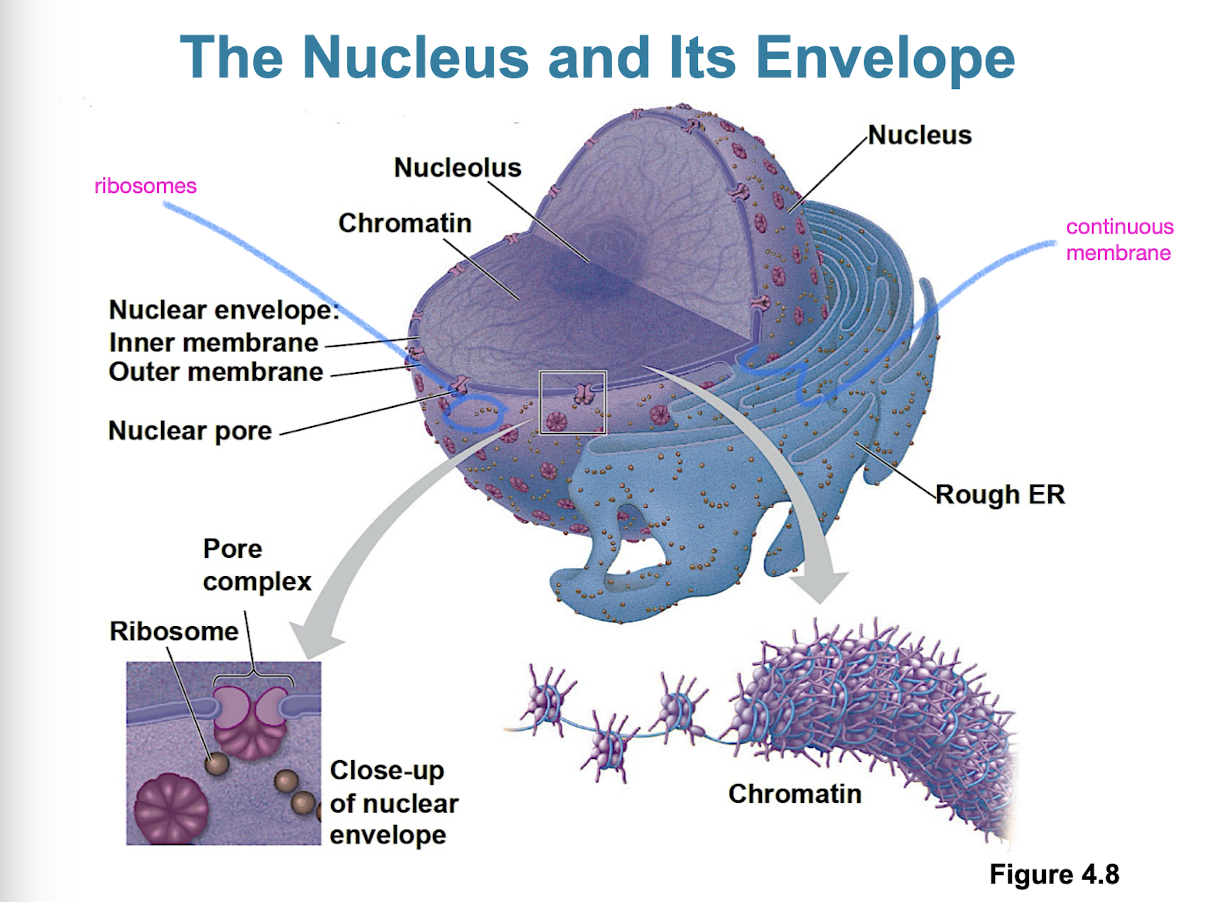
The nucleus
contains most of cell’s genes
nuclear envelope encloses nucleus
double membrane (2 lipid bilayers)
nuclear pores regulate entry/exit of molecules
nuclear lamina to provide mechanical support of nucleus
nucleolus on inside (site of rRNA synthesis)
Ribosomes
complexes (not organelles) of rRNA and protein
carry protein synthesis in cytosol (free) and outside of ER or in nuclear envelope (bound)
Endomembrane system components + function
nuclear envelope, ER, Golgi, lysosomes, vacuoles, and PM (continuous or connected through vesicles)
regulates protein traffic and performs metabolic functions
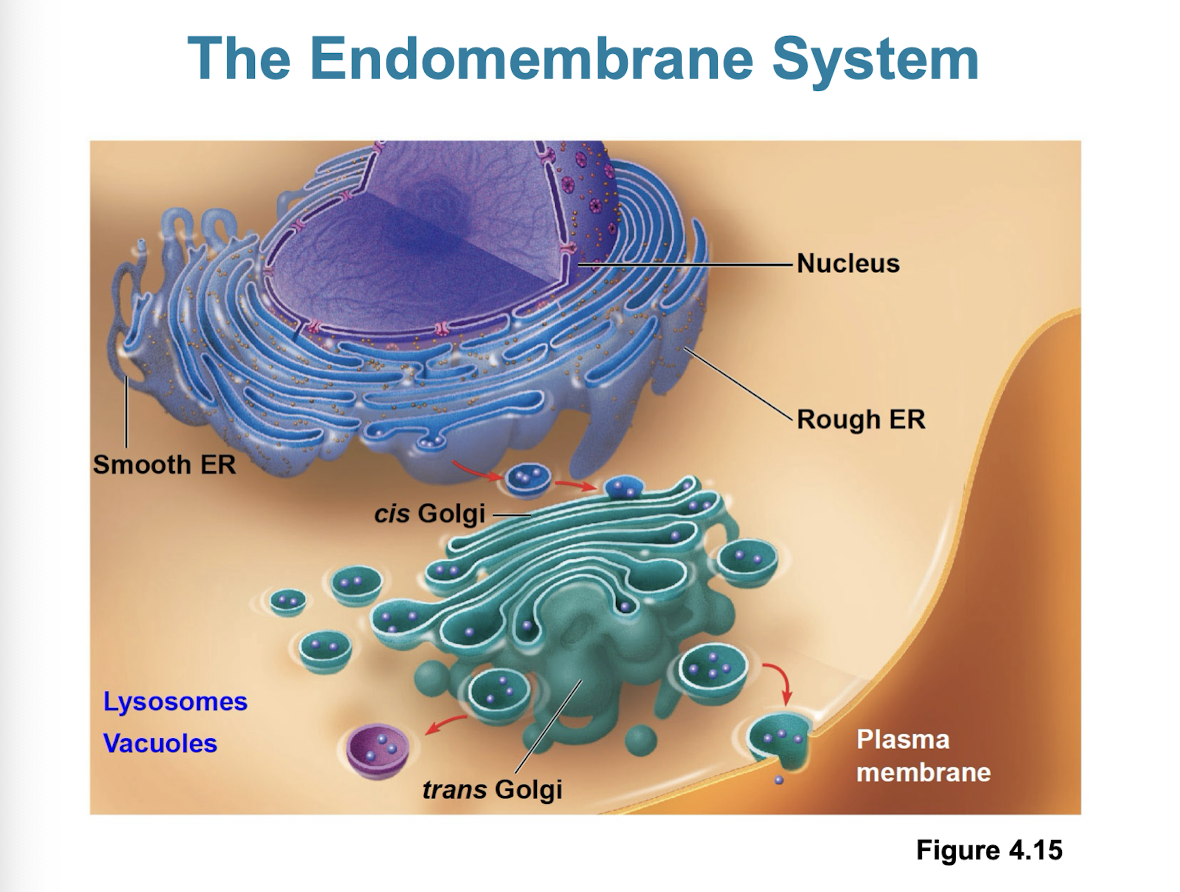
Endoplasmic Reticulum
can account for more than half the total membrane in eukaryotic cells
continuous with nuclear envelope
Smooth ER + Rough ER
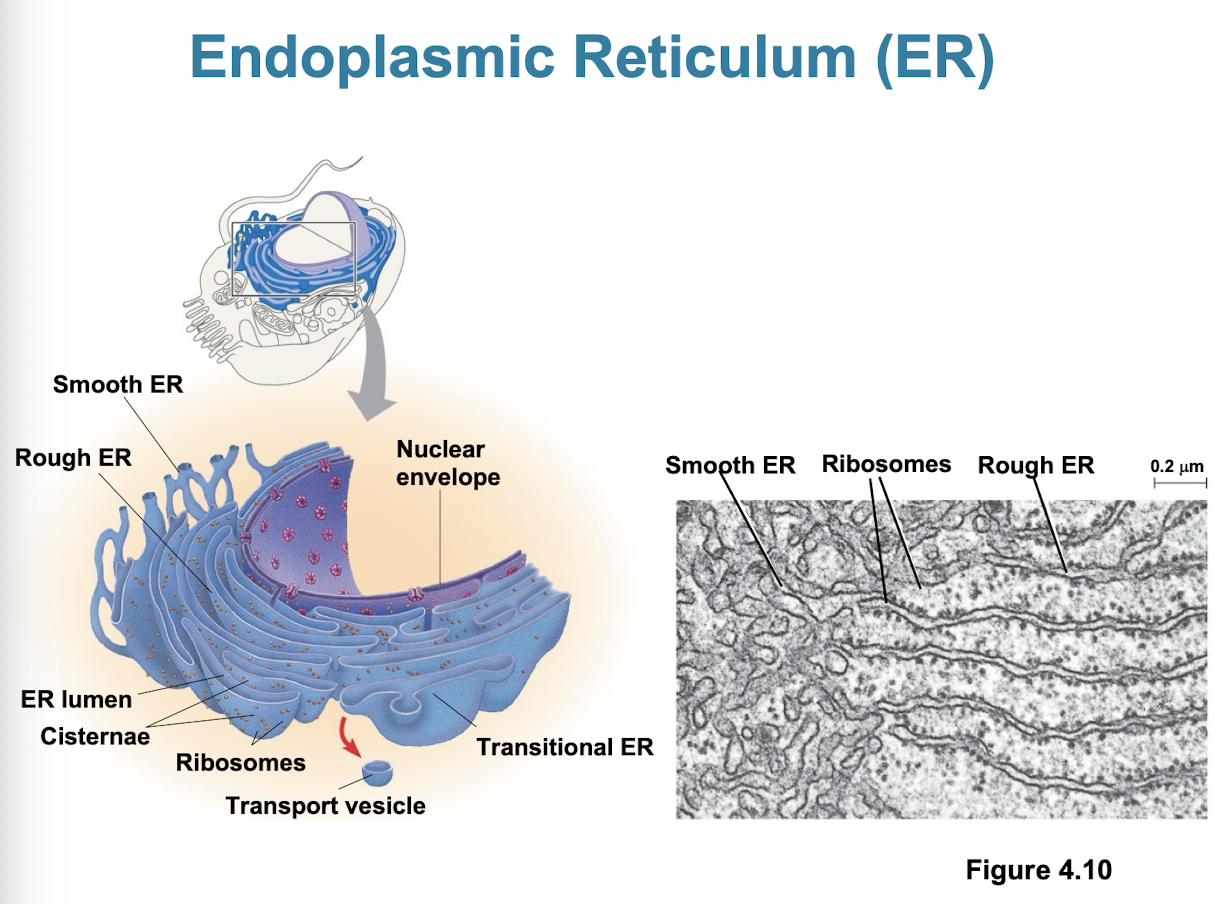
Smooth ER
synthesis of lipids (cholesterol, sex hormones)
metabolism of carbs (glycogen/cellulose)
detoxification of drugs/poisons
calcium ion storage
Rough ER
has bound ribosomes (secrete glycoproteins)
distributes transport vesicles
membrane factory for cell
site of protien synthesis: manufactures, packages, and transports proteins designated for cell membranes, other organelles, or secretion outside the cell
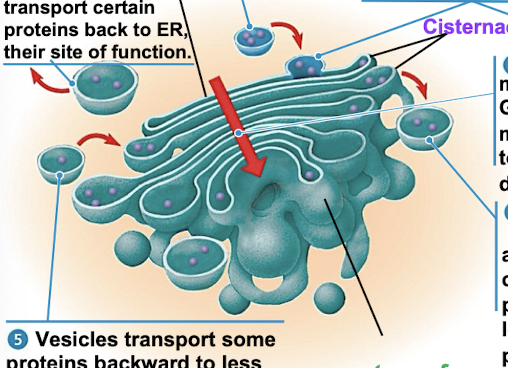
Golgi Apparatus
shipping + receiving center (amazon)
consists of flattened membranes sacs called cisternae (reservoir or pita bread)
modifies products of ER
manufactures macromolecules
sorts/packages materials into transport vesicles
cis + trans faces (receiving + shipping sides)
Lysosomes
single membrane bound organelle found in many animal cells
spherical (3D) vesicle containing hydrolytic enzymes (digest molecules)
enzymes work best in acidic environments
involved in secretion, PM repair, and energy metabolism (reusing materials)
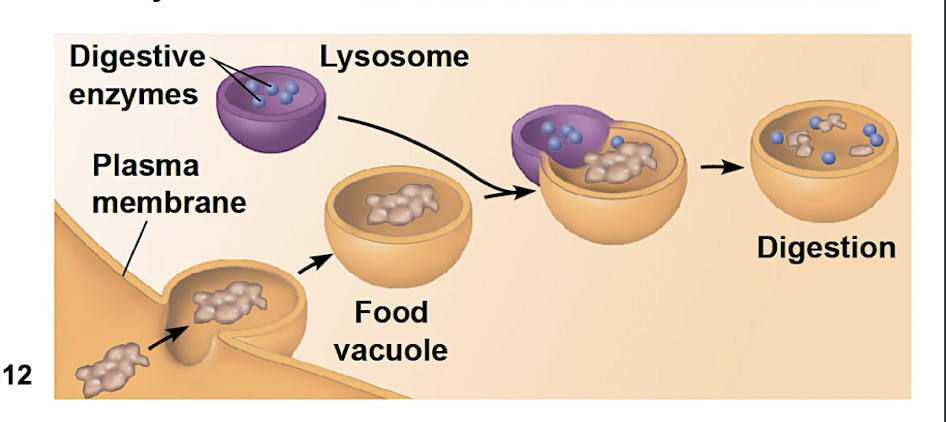
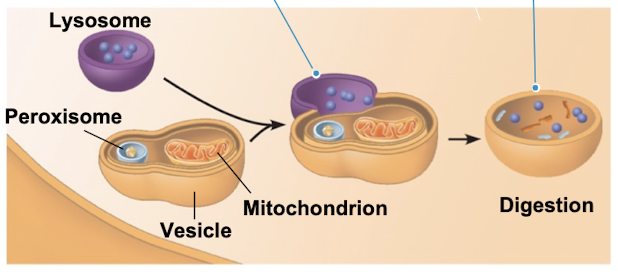
Phagocytosis
when cells engulf another cell to form food vacuole
lysosome fuses with the food vacuole + enzymes digest with molecules
Autophagy
recycling cells own organelles + macromolecules
mechanism used by the lysosome to recycle the cells own dead or damaged organelles and macromolecules
Tay-Sachs Condition
inherited disease where lysosomes are unable to breakdown certain membrane glycolipids due to enzyme defect (glycolipids accumulate in brain cells —> death by 3-4 years old)
Vacuoles
vesicles derived from ER + golgi w/ inside solution differing from cytosol
food vacuoles
contractile vacuoles (found in freshwater protists to pump excess water out of
cells)
central vacuoles (found in mature plant cells to hold organic compounds and
water)
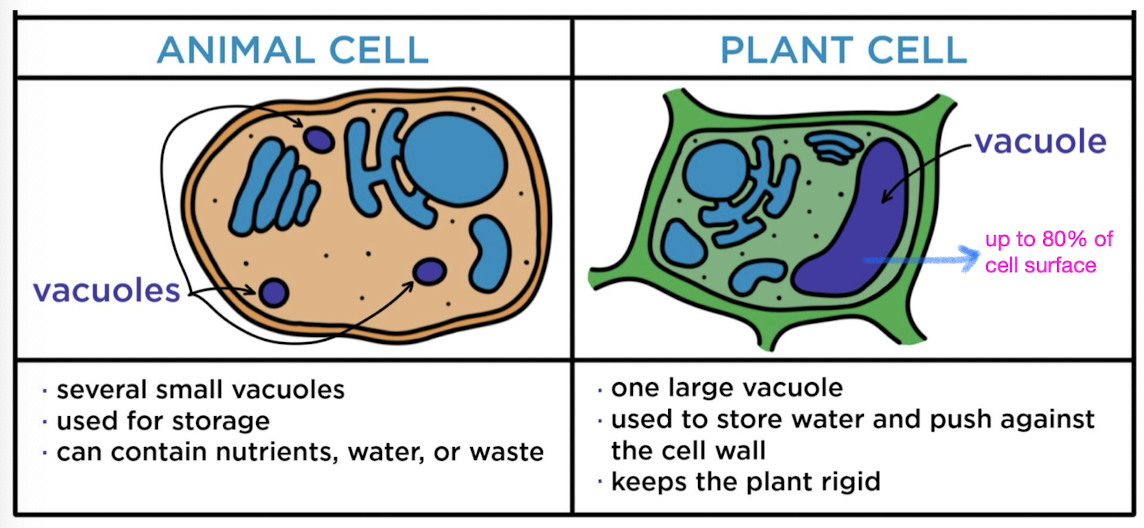
Peroxisomes
single membrane bound organelles that lack own genetic material
produce hydrogen peroxide then convert it to water (detoxifying + oxidizing molecules)
Cellular respiration
A metabolic pathway breaking down glucose (C6H12O6) with oxygen to produce energy (ATP), water, and carbon dioxide
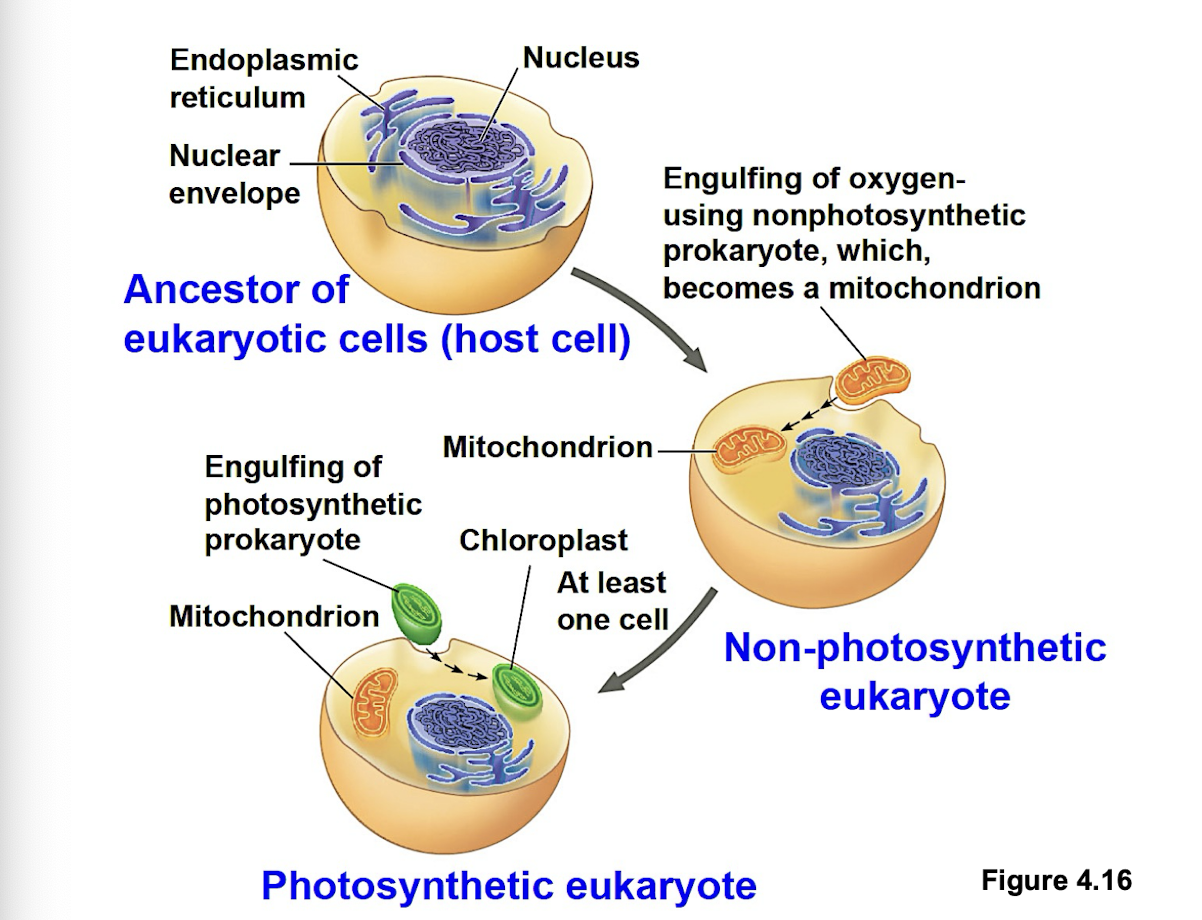
Endosymbiont theory
early ancestor of eukaryotic cells engulfed oxygen-using prokaryotic cell (bacteria) which formed endosymbiont relationship with the host. Host cell and bacteria merged into single eukaryotic cell with a mitochondrion. One of these cells might have later taken up photosynthetic prokaryote becoming the ancestor of cells that contain chloroplasts.
Sites of cellular respiration
mitochondria
Similarities between mitochondria/chloroplasts and bacteria
enveloped by double membrane
contain ribosomes and multiple circular DNA molecules (plasmids)
grow and reproduce somewhat independently in cells
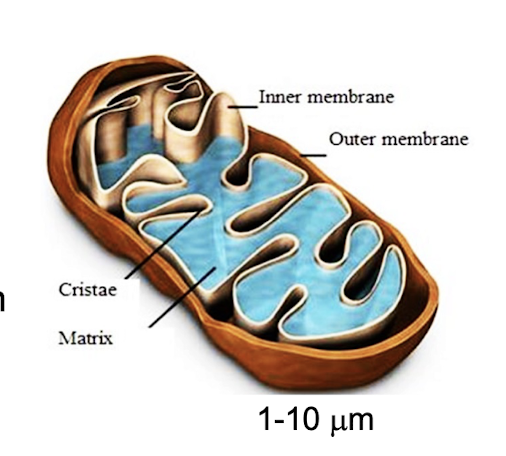
Mitochondria
present in nearly all eukaryotic cells
primary function to generate large quantities of energy in form of ATP
contains smooth double membrane (outer membrane and inner membrane folded into cristae (large surface area for enzymes))
inner membrane made out of inter membrane space and mitochondrial matrix
Chloroplasts
site of photosynthesis (production of sugar + O2 while converting sunlight into ATP)
contain green pigment chlorophyll and other enzymes for photosynthesis
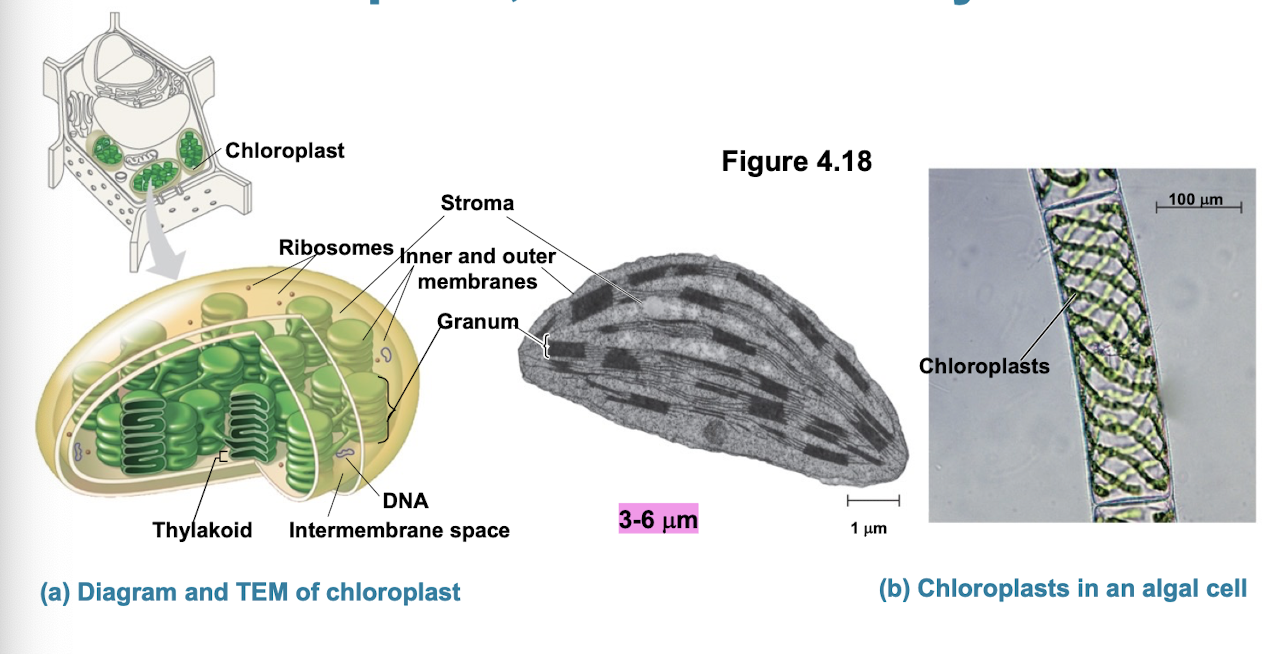
The cytoskeleton
network of fibers extending through cytoplasm
organizes cells structures + activities
helps support cell + maintain shape
anchorage for organelles + molecules (very dynamic)
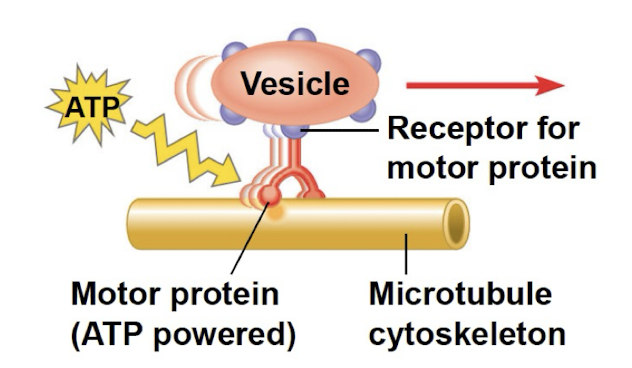
interacts with motor proteins to produce motility
vesicles/organelles use motor protein feet to walk along tracks of cytoskeleton
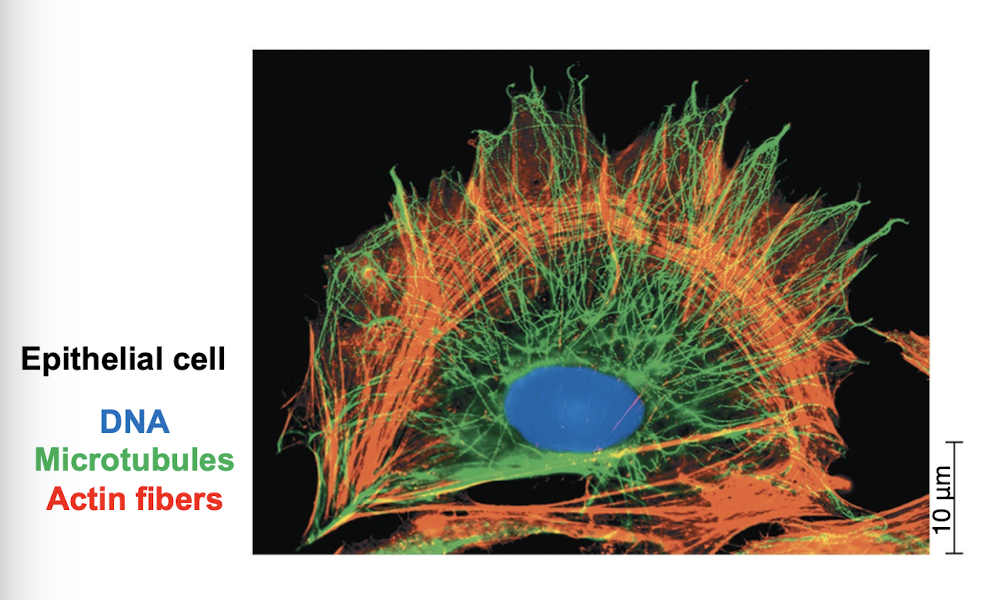
Three main fibers of cytoskeleton
microtubules: thickest
intermediate filaments: diameters in middle range
microfilaments/actin filaments: thinnest
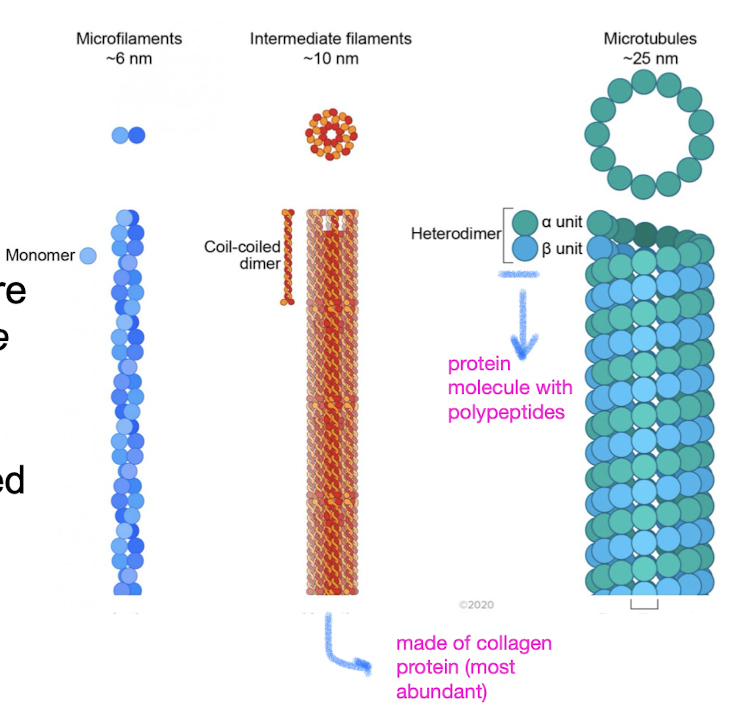
Microtubules
hollow dynamic rods constructed from tubulin, globular protein dimers
shape + support of cell
guide movement of organelles
separate chromosomes during cell division
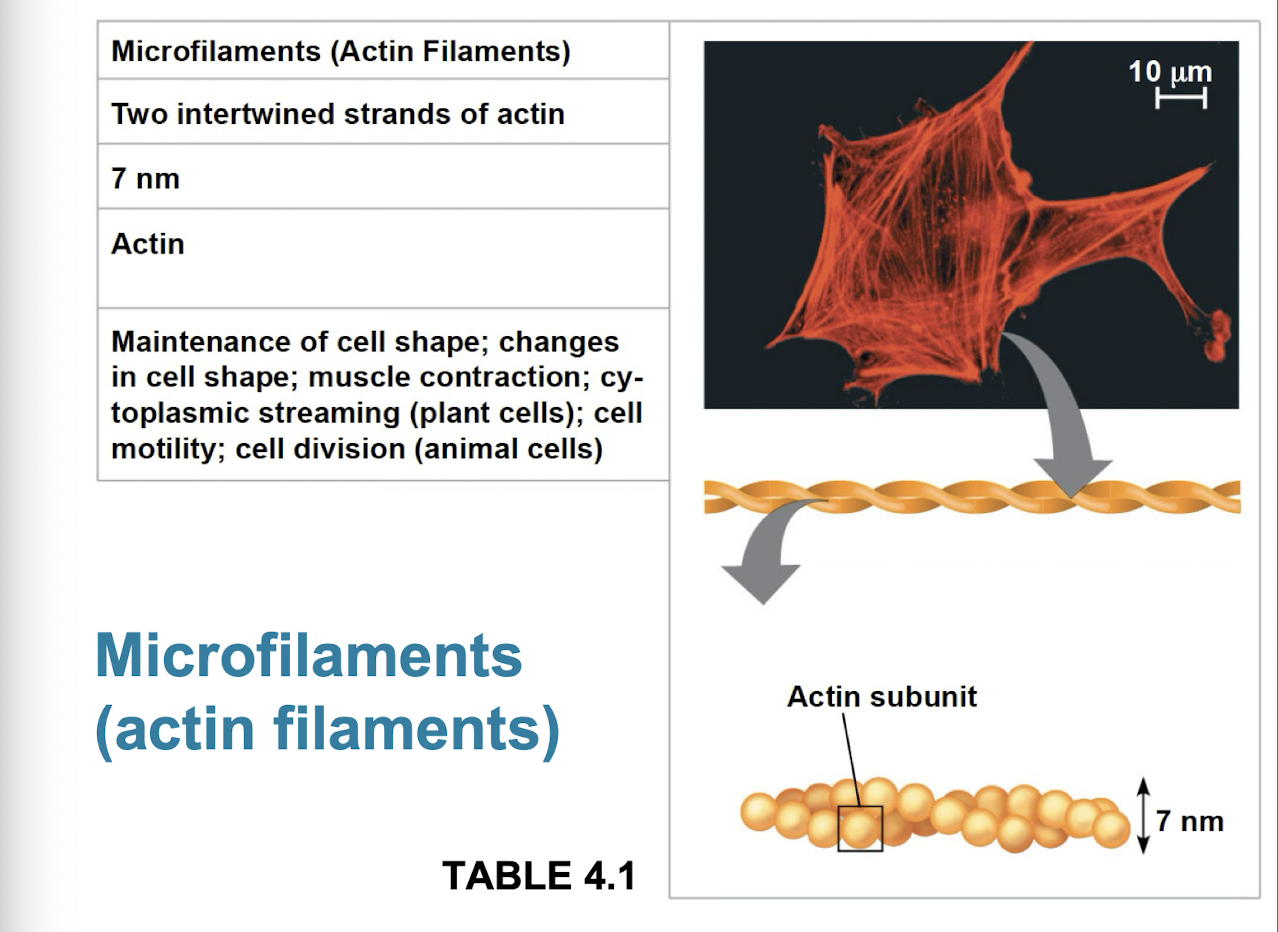
Microfilaments
thin solid robs built from dynamic polymers (protein molecules of globular actin subunits)
bears tension + resits pulling forces within cell
function in cellular motility + interact with motor protein myosin
actin + myosin interact to cause muscle contraction, amoeboid movement of white blood cells, and cytoplasmic streaming in plant cells
bundles of microfilaments make up core of microvilli of intestinal cells that increase cells surface area
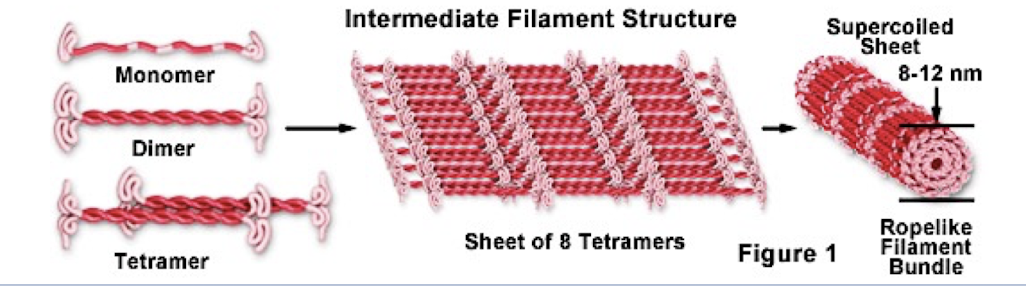
Intermediate filaments
only found in cells of some animals (like vertebrates)
reinforce cell shape and fix organelles in place (anchor nucleus)
formation of nuclear lamina
more permanent cytoskeleton elements that other 2
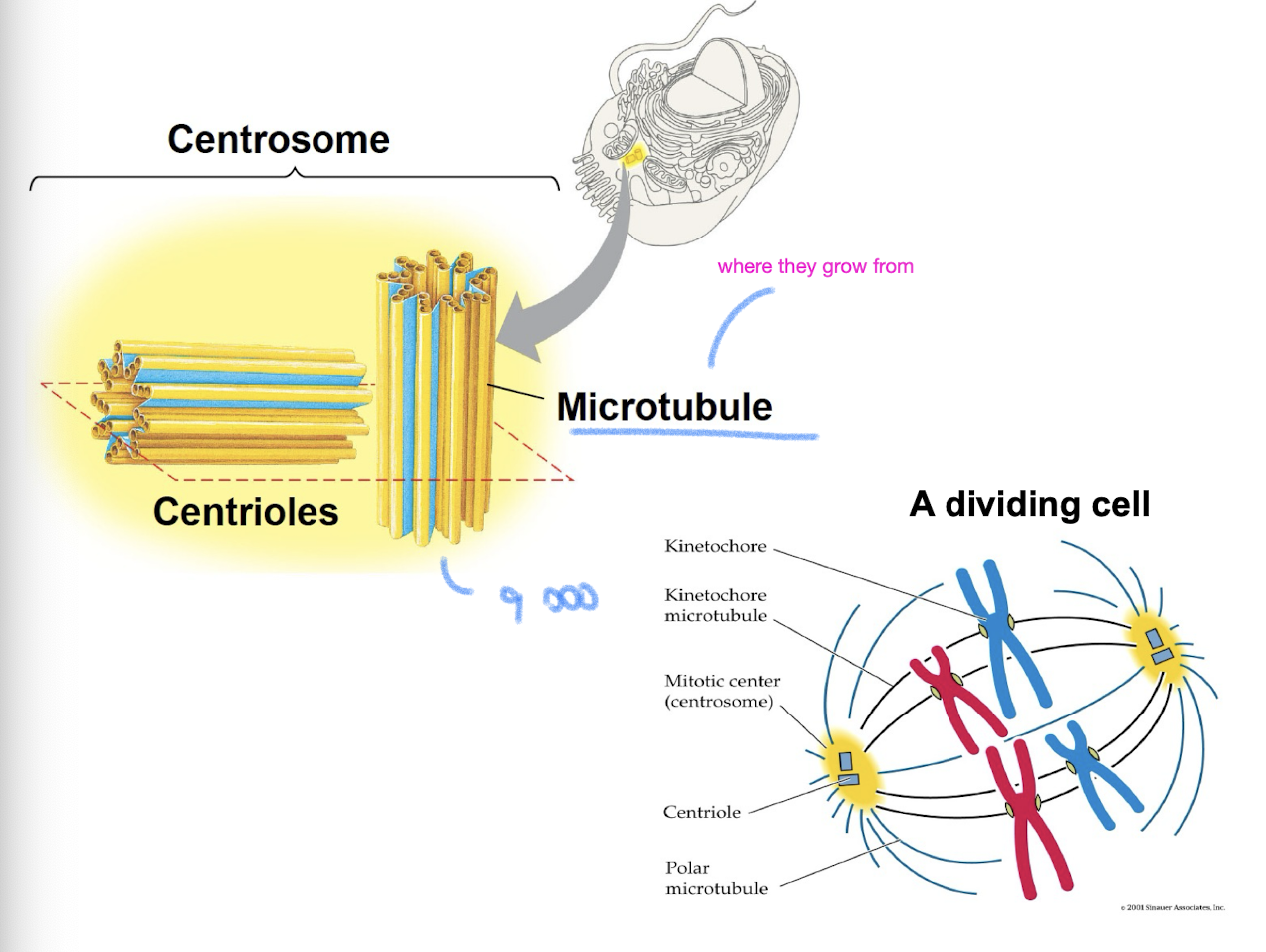
Centrosomes
in animal cells, microtubules grow out from a centrosome near the nucleus
the centrosome is the “microtubule organizing center”
centrosome has a pair of centrioles each with 9 triplets of microtubules arranged in a ring
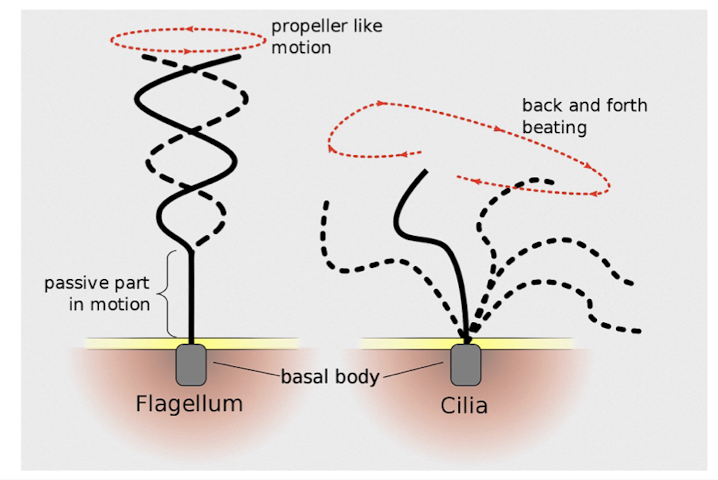
Cilia and Flagella
microtubules control the beating of cilia/flagella, which are microtubule-containing extensions projecting from some cells
Flagella are limited to one or few per cell
Cilia occur in large numbers on cell surfaces
cilia and flagella differ in beating pattern
common structure of group of microtubules sheathed by the PM and a Basal body that anchors cilium or flagellum
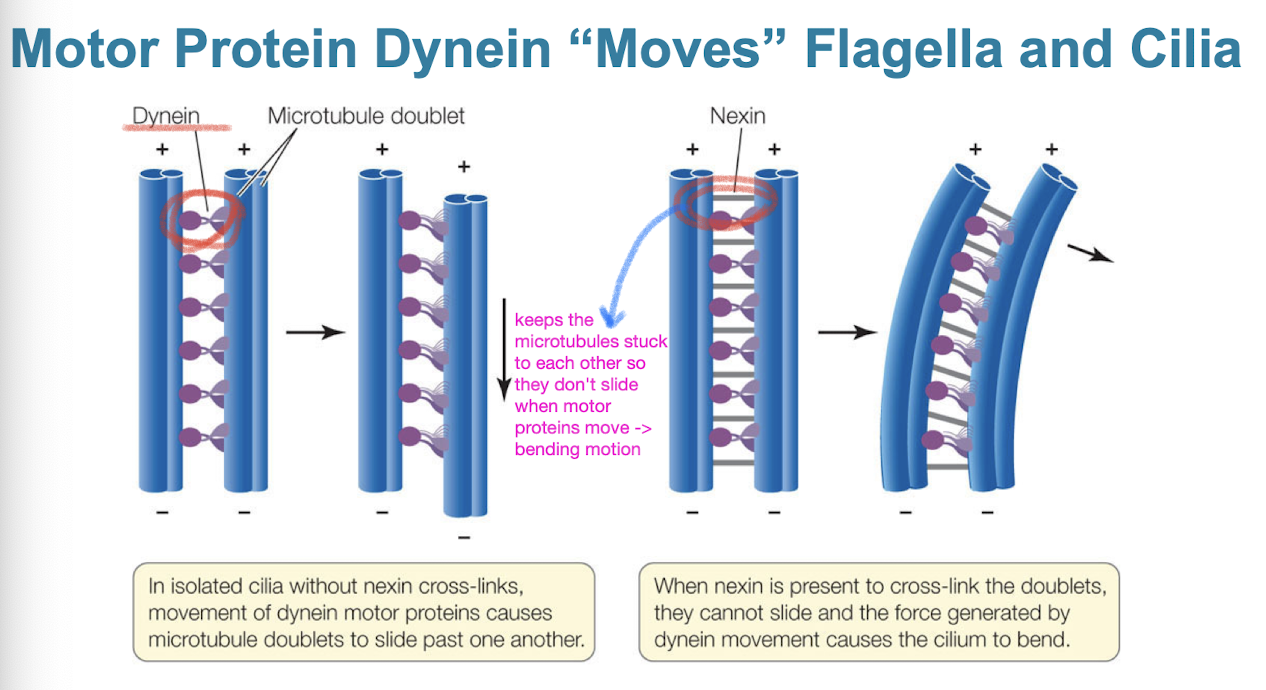
Dynein
motor protein that “moves” flagella and cilia
dynein arms alternately contact, move, and release the outer microtubules
movements of the doublet arms cause cilium/flagellum to bend
nexin keeps doublets from sliding
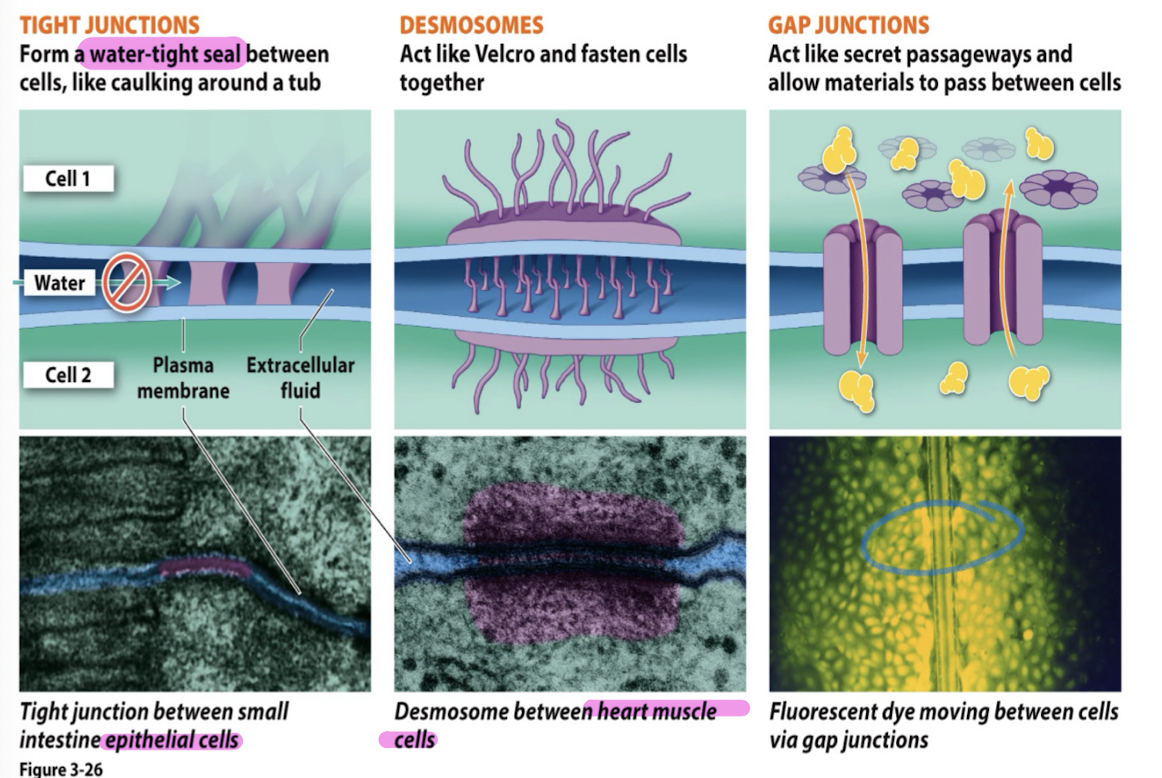
Cell junctions
plasmodesmata (plants); water, proteins, RNA can pass from cell to cell
tight junctions (animals): form water tight seal between cells (caulking around a tub) (intestine/skin cells)
desmosomes (animals): act like velcro + fasten cells together (heart muscle cells)
gap junctions (animals): act like secret passageways + allow materials to pass between cells
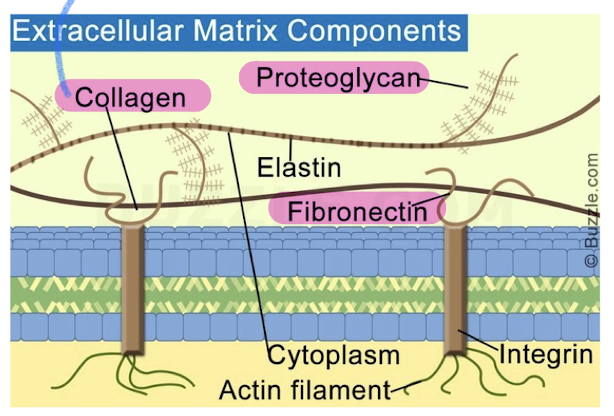
Extracellular Matrix (ECM) of animal cells
animal cells lack cell walls but are covered by elaborate ECM
made up of glycoproteins (collagen, proteoglycans, fibronectin)
ECM proteins bind to cell-surface receptor proteins in PM called integrins (membrane proteins with two subunits)
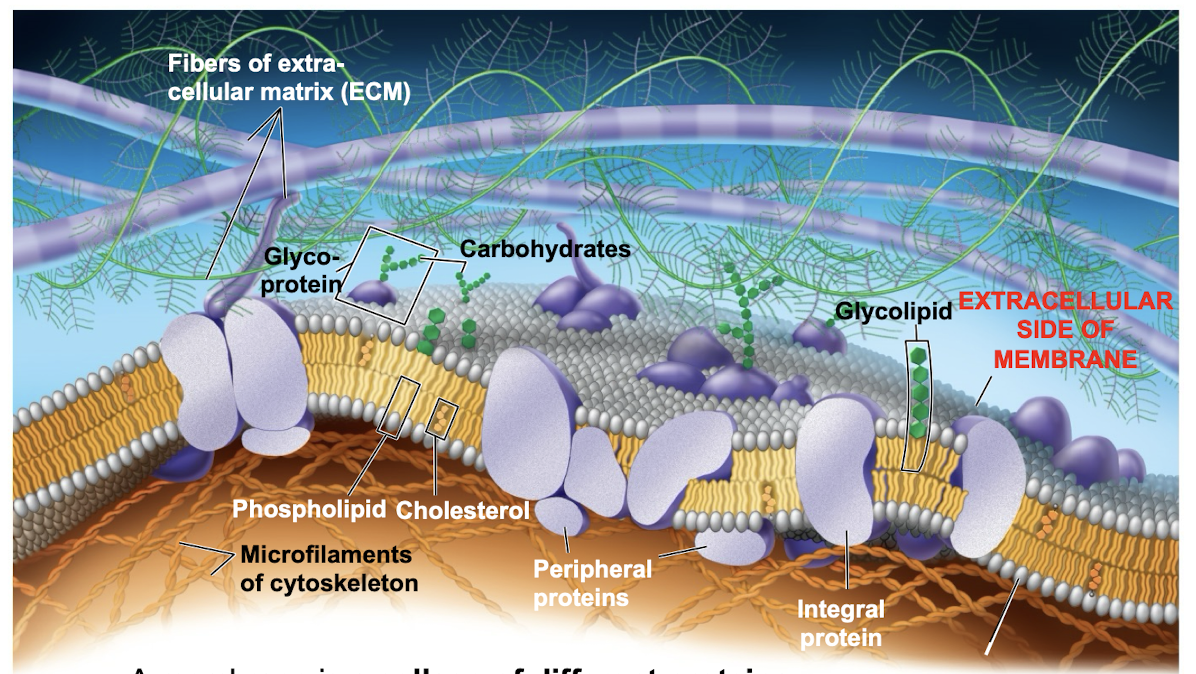
Role of membrane carbohydrates in cell cell recognition
cells recognize each other by binding to surface molecules (often containing carbs) on extracellular surface of PM
membrane carbs may be covalently bonded to lipids (forming glycolipids) or more commonly to proteins (glycoproteins)
Plasma membranes
separate what inside of cell (cytoplasm) from outside
selective permeability: certain molecules can passively diffuse across the membrane and larger molecules cannot (need active transport)
made up of phospholipids (amphipathic: hydrophobic and hydrophilic regions)
stable boundary between two aqueous compartments
contain domains that reside within and outside of PM
Fluid Mosaic Model
mosaic of protein molecules bobbing in a fluid bilayer of phospholipids
Mosaic model allows for proteins to shift/move in cell membrane due to the rearrangement of phospholipids
What determines/affects membrane fluidity
saturation of carbon molecules and cholesterol
fluid = unsaturated (double bond/kinked) (like olive oil)
viscous = saturated + packed together
cholesterol = reduced fluidity at moderate temps but hinders solidification at low temps
As temps cool, membranes shift from fluid to solid state
What organelles make the plasma membrane + its components
The ER and the Golgi apparatus (components include proteins, lipids, and carbohydrates)
more specifically: starts in the ER → golgi → vesicles → PM
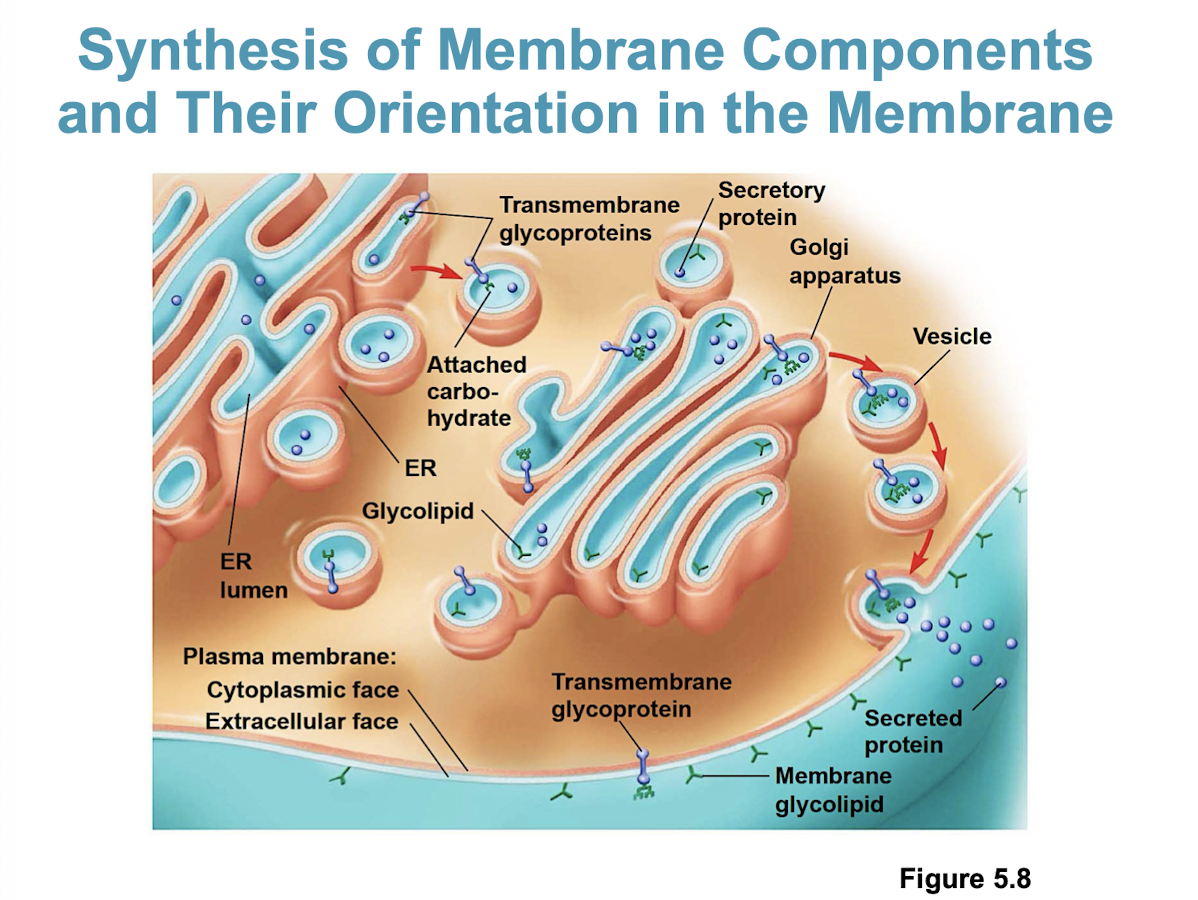
Movement in and of a membrane
most lipids and some proteins in a membrane can shift sideways
movement of phospholipids is rapid and proteins move much slower
some proteins move in directed manner, others anchored
some proteins simply drift in membrane
What can/can’t pass through membranes
Easily pass
small molecules (O2)
hydrophobic/nonpolar molecules (still have to be small)
Cant easily pass
polar molecules (sugars like glucose)
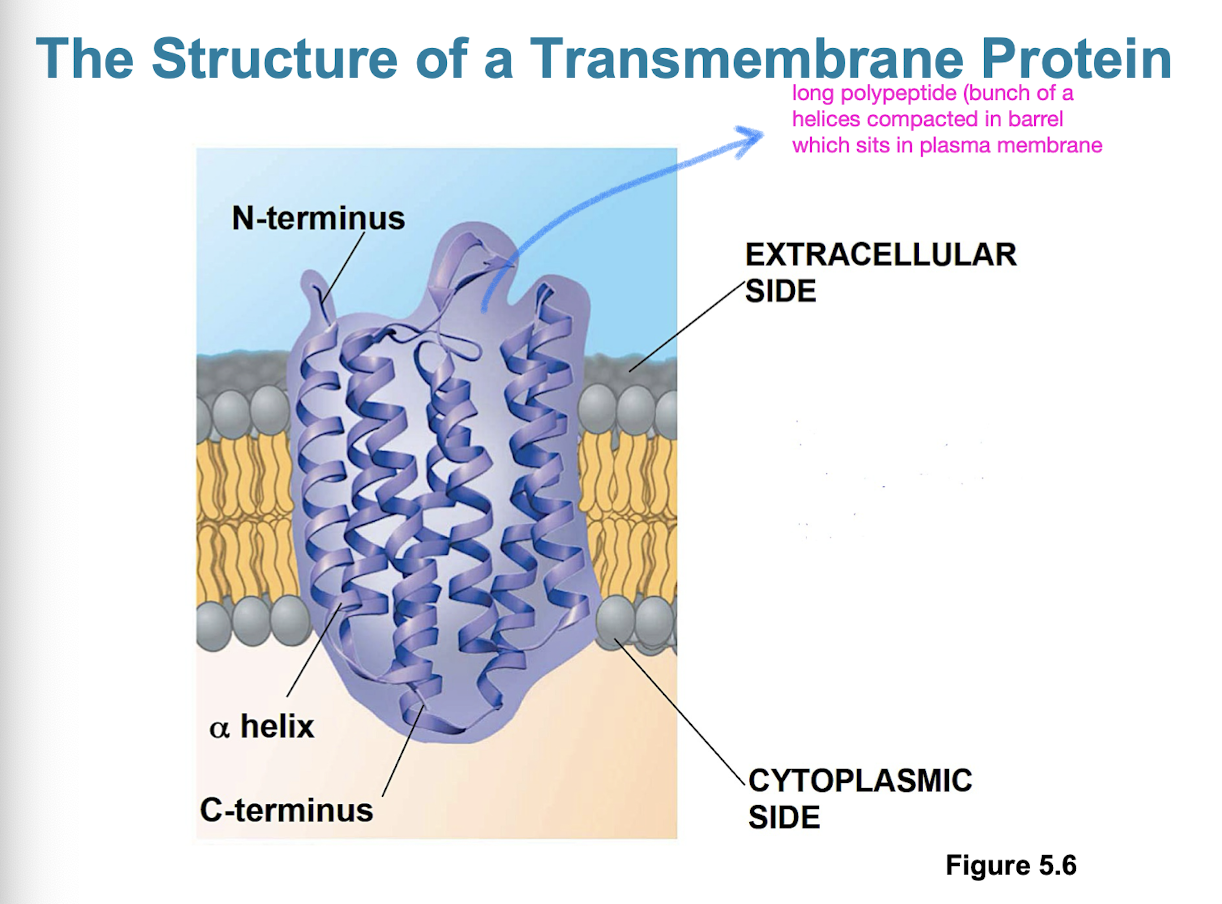
Characteristics of membrane proteins
most membrane proteins are amphipathic
determine most of the membranes specific functions
integral proteins: penetrate hydrophobic interior of lipid bilayer (also called transmembrane proteins)
peripheral proteins: loosely bound to surface of membrane (not actually embedded, could be hydrophilic)
Six major functions of membrane proteins
transport
enzymatic activity
signal transduction (ex. from outside to inside cell)
cell-cell recognition
intercellular joining
attachment to cytoskeleton + ECM
Transport proteins
allow passage of hydrophilic substances across membrane (H2O, amino acids)
specific for substance it moves
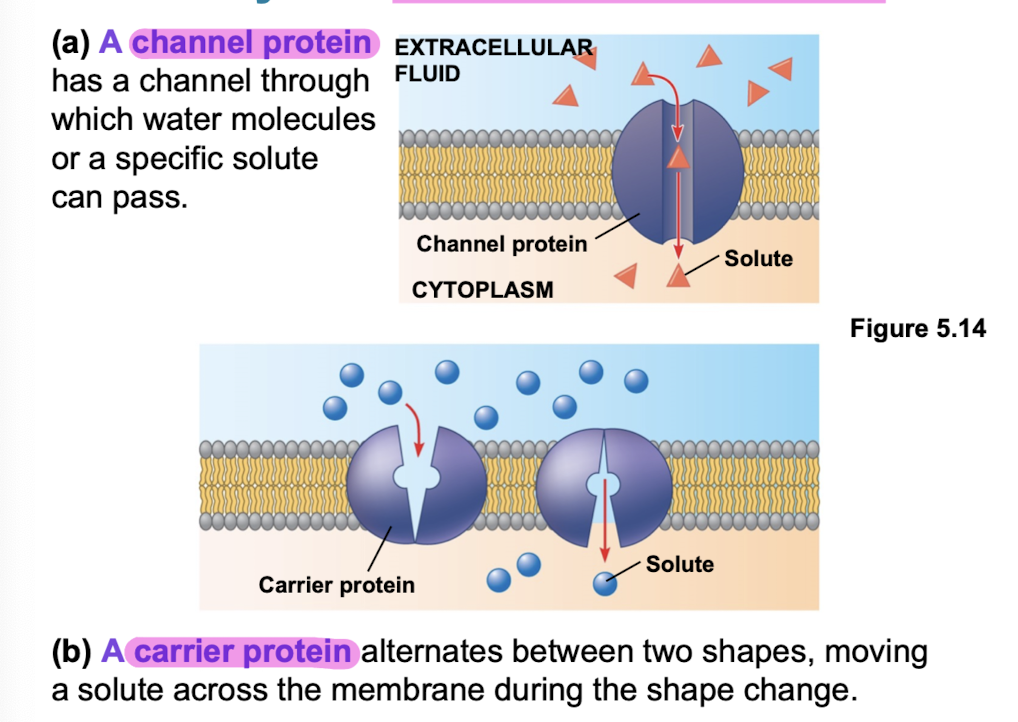
channel proteins have hydrophilic channel (aquaporins facilitate passage of water)
Carrier protiens
bind to molecules, change conformation (shape) and shuttle these molecules across the plasma membrane
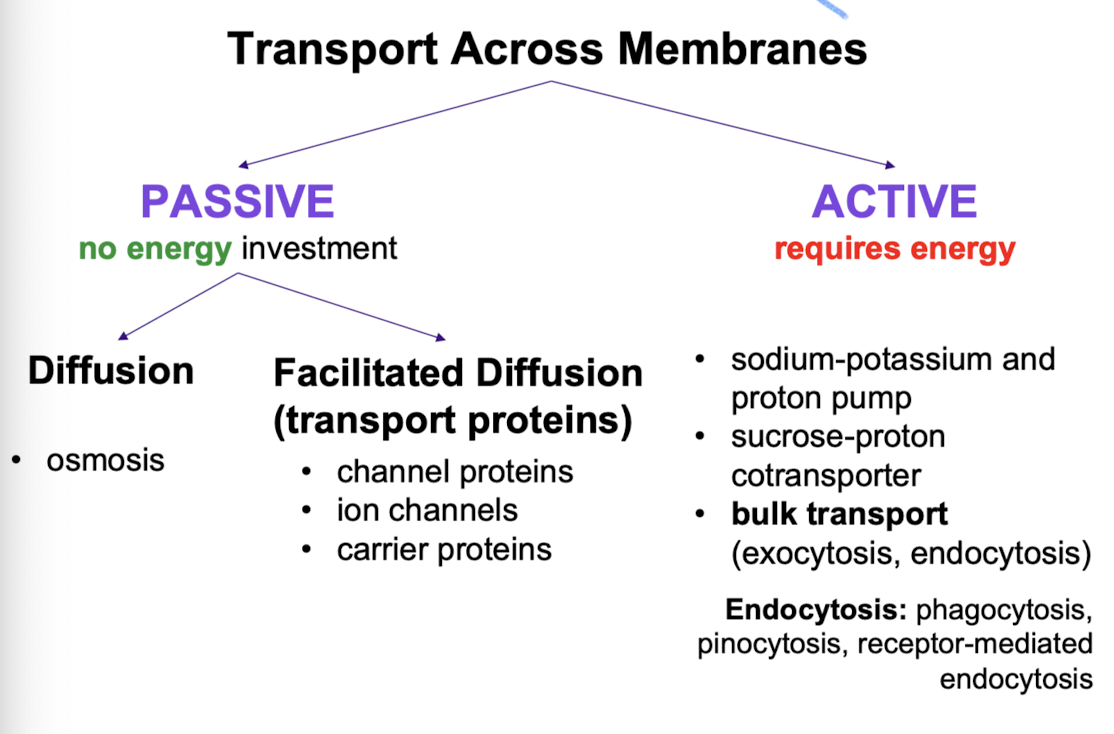
Passive transport
no energy investment
diffusion (osmosis; diffusion of water)
facilitated diffusion (transport proteins; channel proteins, ion channels, carrier proteins)
Active transport
Requires energy (usually ATP) + moves substances against concentration gradient. Allows cells to maintain concentration gradients that differ from surroundings.
sodium-potassium + proton pump (3 sodium “slots”, 2 potassium “slots”)
sucrose-proton cotransporter
bulk transport (exocytosis, endocytosis)
Ion Pumps
Sodium-potassium pump
exchanges Na+ for K+ across PM in animal cells
major electrogenic pump
generates voltage across a membrane (ions are charged)
sodium fills 3 slots → phosphorylation in cytoplasm of ATP (one phosphate on pump) → sodium ions released → potassium molecules from outside bind to transport → dephosphorylation of pump → release of potassium molecules
Proton pump
main electrogenic pump of plants, fungi, and bacteria
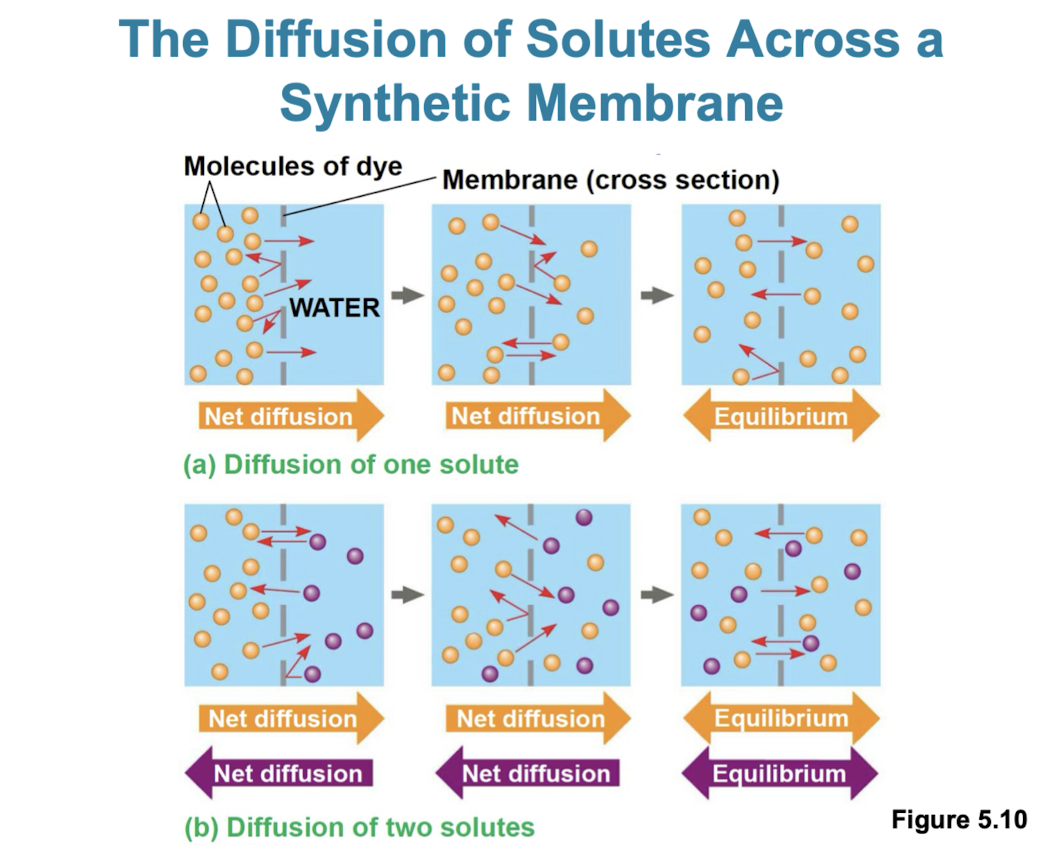
Diffusion
tendency for molecules to spread out evenly in available space
although molecules move randomly, it may be directional
at dynamic equilibrium there are an even amount of molecules crossing the membrane in one direction as the other
substances diffuse down their concentration gradient (more to less)
Facilitated diffusion
transport proteins speed up passive movement across PM
channel proteins provide corridors (aquaporins, ion channels)
carrier proteins undergo subtle shape change to move solute across membrane
no net energy input required!
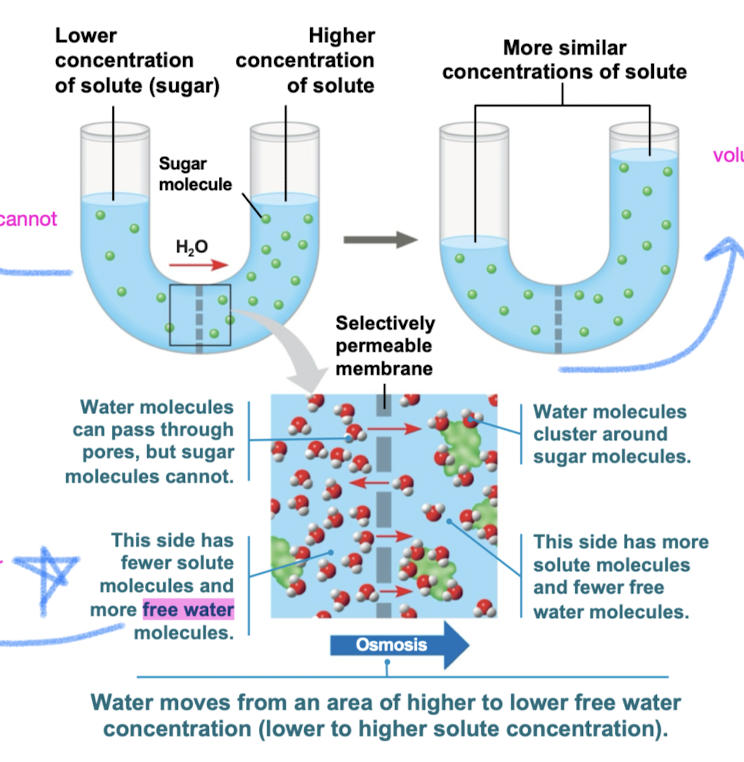
Osmosis
diffusion of free water (solvent) across selectively permeable membrane from diluted solution (fewer solutes) to concentrated solution (more solutes)
Flow of water is dependent on the solute concentration (not concentration of water)
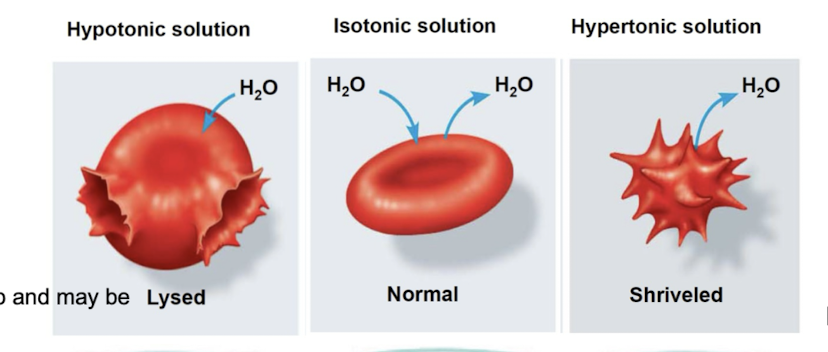
Tonicity
ability of surrounding solution to cause cell to gain/lose water
Isotonic: solute concentration same on either side: no net water movement
Hypertonic: solute concentration greater outside PM than inside of cell (water will move outside)
Hypotonic: solute concentration is greater inside cell (water will move in)
Osmoregulation
control of solute concentrations and water balance (necessary adaptation for life in such conditions)
ex. contractile vacuole in paramecium caudatum
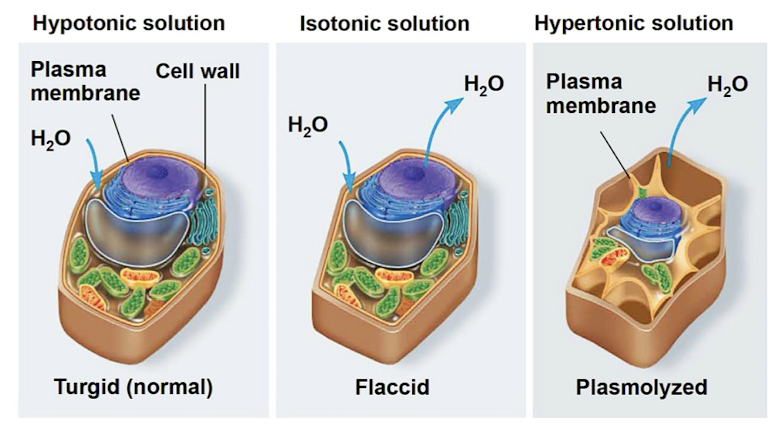
Water balance of plant cells
rigid cell walls (cellulose) helps maintain water balance
turgid (normal/very firm, hypotonic)
flaccid (limp, isotonic)
plasmolysis (lethal, hypertonic)
Cotransport
when a transport protein can couple the “downhill” diffusion of solute to “uphill” transport of second solute against its gradient
proton pumps couple hydrogen ion gradient to drive active transport of nutrients (pumped out → low concentration → contransporter brings hydrogen ions back in cell along w sucrose/other)
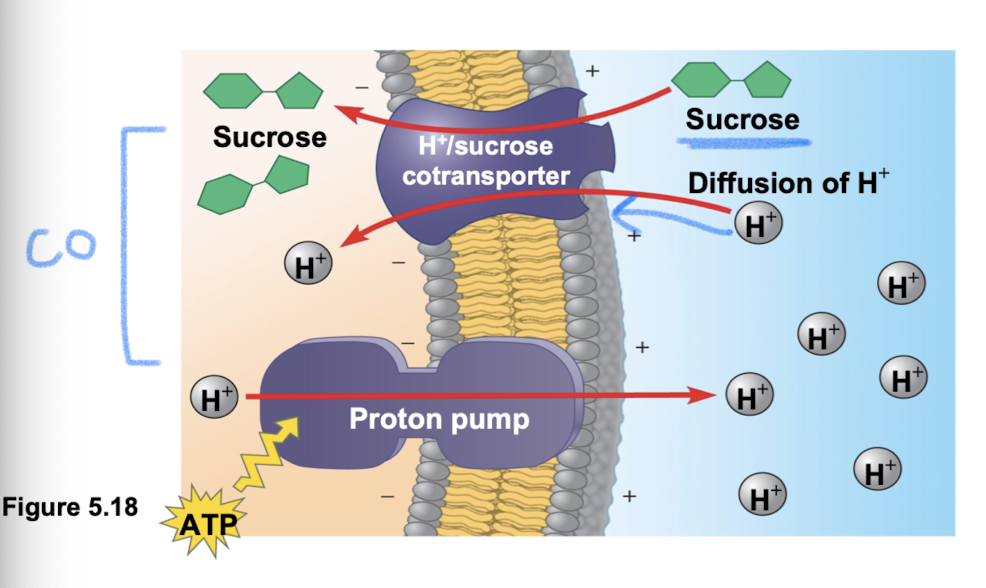
Bulk transport
large molecules (polysaccharides/proteins) cross membrane in bulk using vesicles (requires energy!)
Exocytosis
transport vesicles migrate to the membrane, fuse with it, and release their contents
many secretory cells use exocytosis to export products (ex. hormones, proteins, etc.)
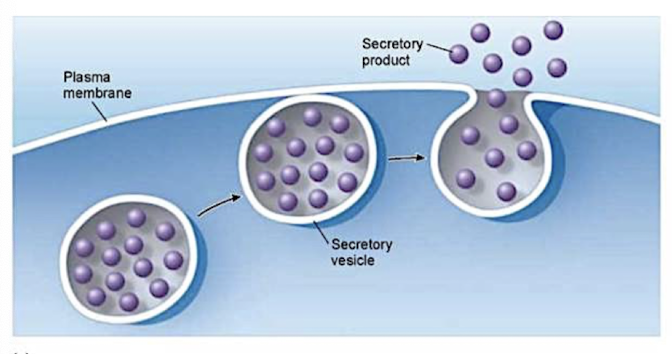
Endocytosis
cell takes in molecules/matter by forming new vesicles from PM (reversal of exocytosis + involves dif proteins)
Phagocytosis (cellular eating)
Pinocytosis (cellular drinking)
Receptor mediated endocytosis (receptor recognizes molecules + forms vesicles to bring content into cell)
ex. cholesterol travels in LDLs (lipoprotiens which bind to receptors) + familial hypercholesterolemia (too much in blood) occurs when receptor proteins are defective or missing → atherosclerosis
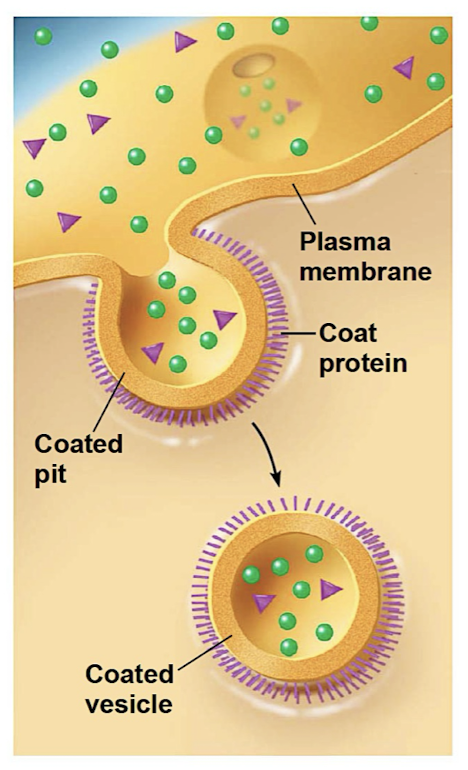
Metabolism
getting energy from food to fuel all chemical reactions (sum of all chemical reactions needed for life)
metabolism is an emergent property arising from orderly interactions between molecules (allows for fluctuations and adaptation to the environment)
Bioenergetics
study of how energy flows through living systems (environments and organisms)
Metabolic pathways
starts with primary molecule → ends with product (ex. ATP)
enzymes essential for each step
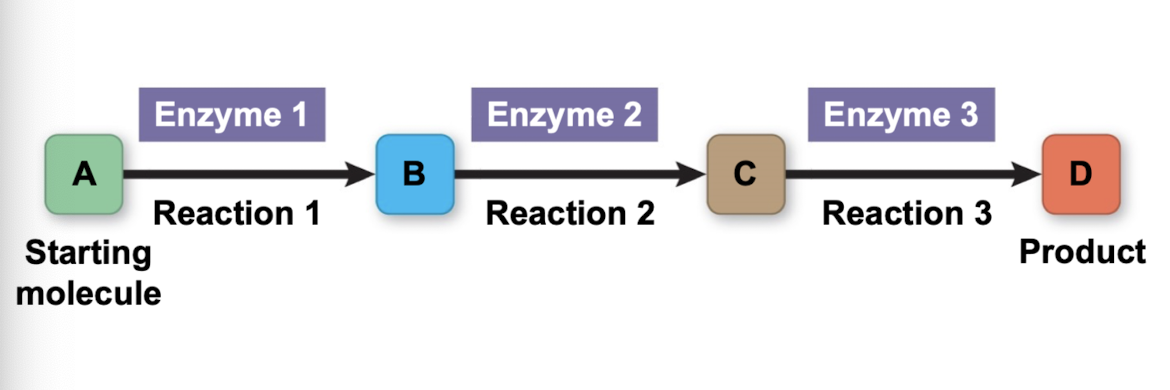
Catabolic pathways/catabolism
release energy by breaking down complex molecules into simpler compounds (DOWNHILL)
energy is then available to do cellular work (ex. cellular respiration; glucose + organic fuels broken down into CO2 and H2O)
Anabolic pathways/anabolism
consume energy to build complex molecules from simpler ones (called biosynthetic pathways) (ex. proteins synthesized from simpler amino acids) (UPHILL)
Energy
capacity to cause change
exists in various forms, some of which can perform work
Work
movement of matter against opposing forced (gravity/friction)
Forms of energy
Kinetic energy: energy associated with motion
Thermal energy: type of KE associated with random movement of atoms/molecules
Heat: transfer of thermal energy from one object to another (dif in temp)
Light: energy that can be harnessed to perform work
Potential energy: energy that matter possesses because of its location/structure
Chemical energy: potential energy available for release in a chemical reaction
energy can be converted from one form to another
Thermodynamics
organisms are open systems
First law: energy is neither created nor destroyed
Second law: energy transfer/transformation increases entropy of the universe + some energy is lost to surroundings as heat

Entropy
measure of molecular disorder (dispersed energy in a system)
heat increases disorder of the surroundings
spontaneous processes require no energy input
non spontaneous require energy supplied + lead to decrease in entropy
Free energy (Gibbs)
the portion of a system’s energy that can perform work when temperature and pressure are uniform throughout the system (like a cell)
∆G = G final state - G initial state
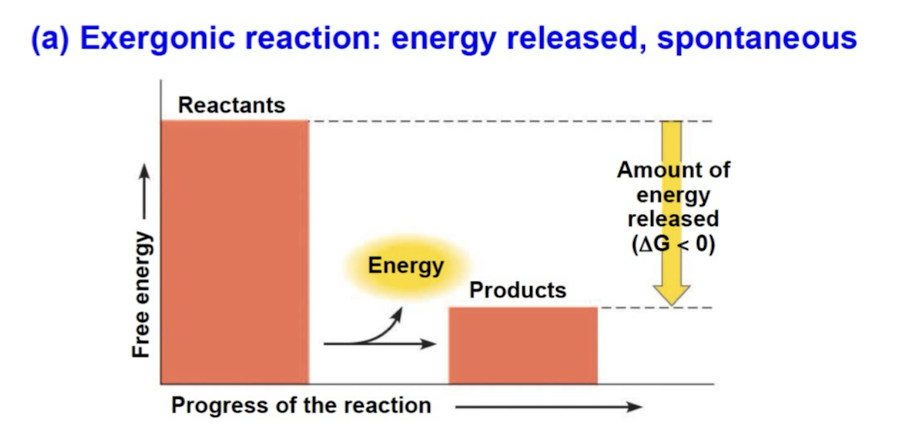
Exergonic reactions
spontaneous reaction with net release of free energy (∆G < 0) (energy exiting)
magnitude of ∆G represents max amount of work reaction can perform
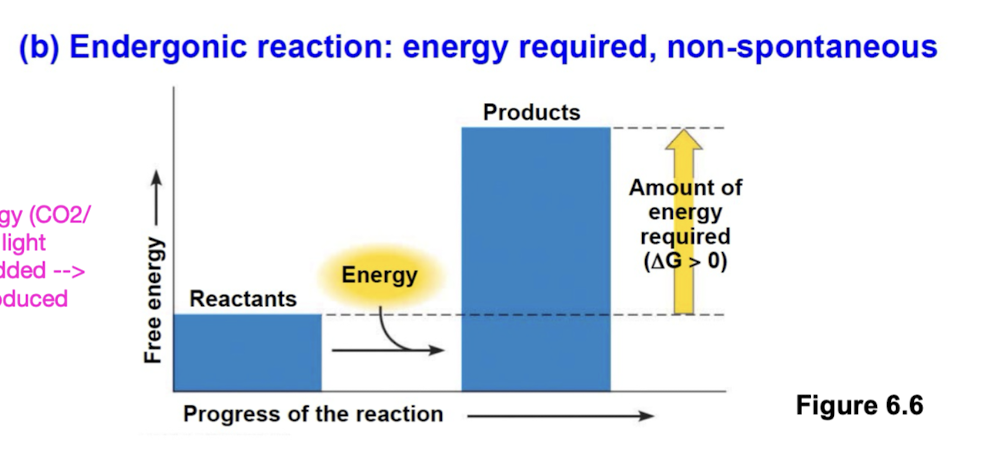
Endergonic reactions
non spontaneous and absorb free energy from surroundings (∆G > 0) (energy entering)
magnitude of ∆G represents quantity of energy required to drive reaction
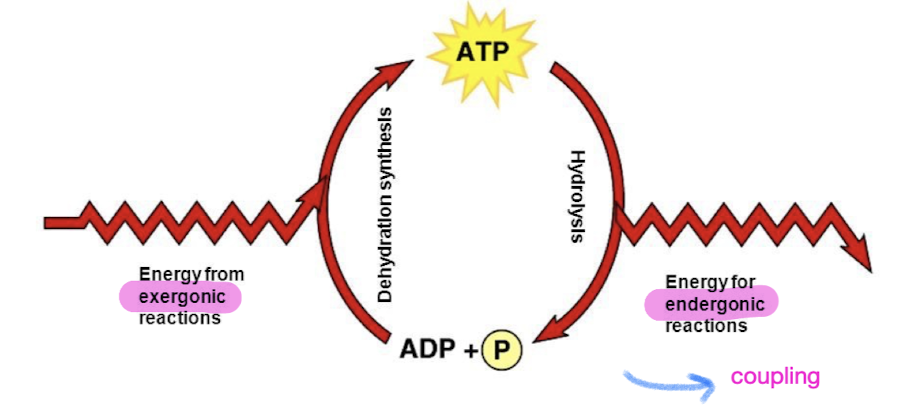
Energy coupling
use of exergonic process to drive an endergonic one to do work (most energy coupling in cells mediated by ATP)
3 main kinds of work done by a cell
Chemical (building biopolymer like protein/DNA)
Transport (ex. sodium potassium pump)
Mechanical (ex. protein motor walking on microtubule and carrying a vesicle)
Driven by ATP hydrolysis (energy released used for endergonic reactions)
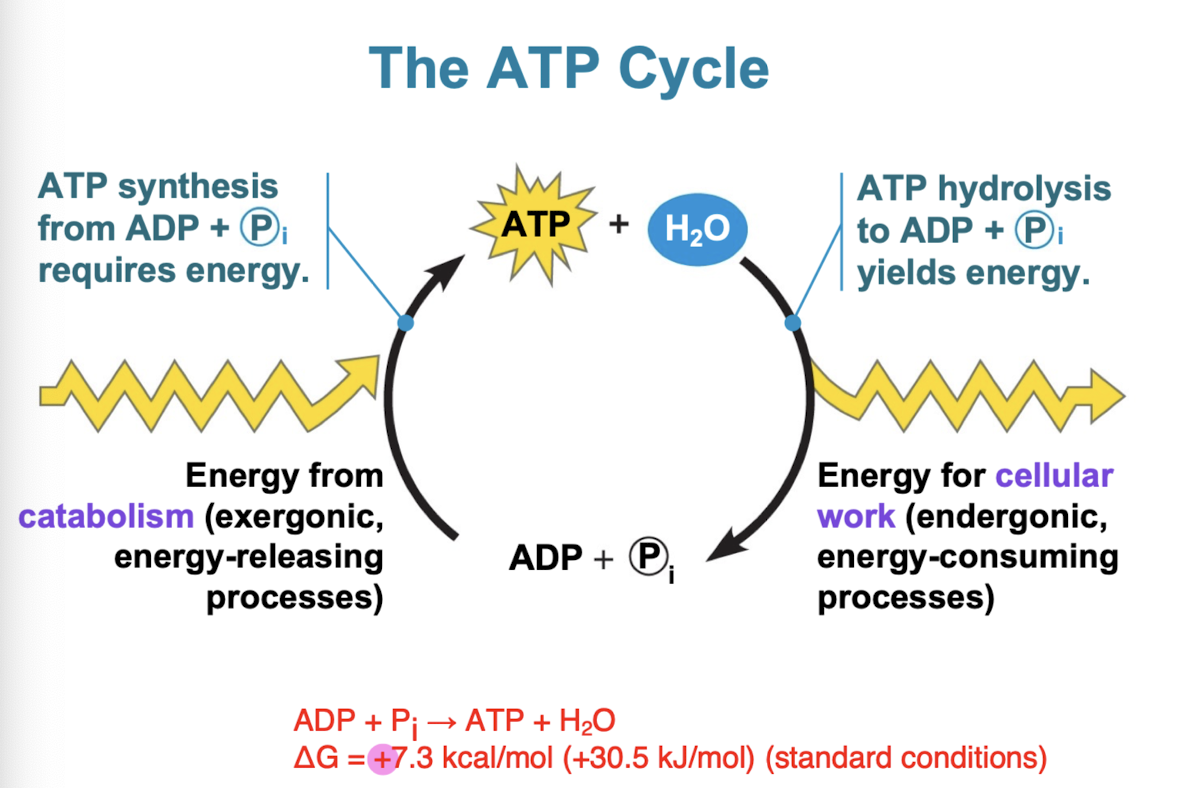
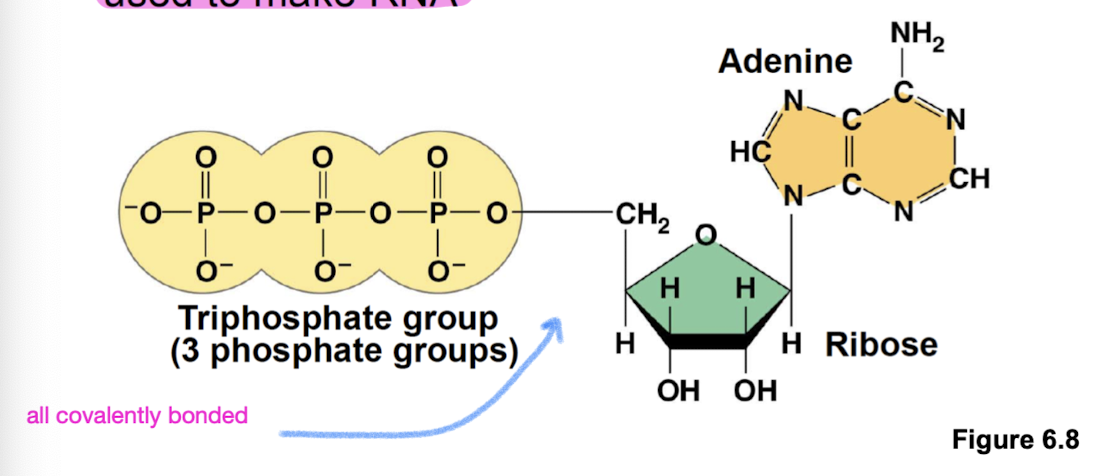
ATP
Adenosine triphosphate: composed of a ribose (sugar), adenine (nitrogenous base), and 3 phosphate groups
role in energy coupling (within the triphosphate group) and also used to make RNA (ribose backbone)
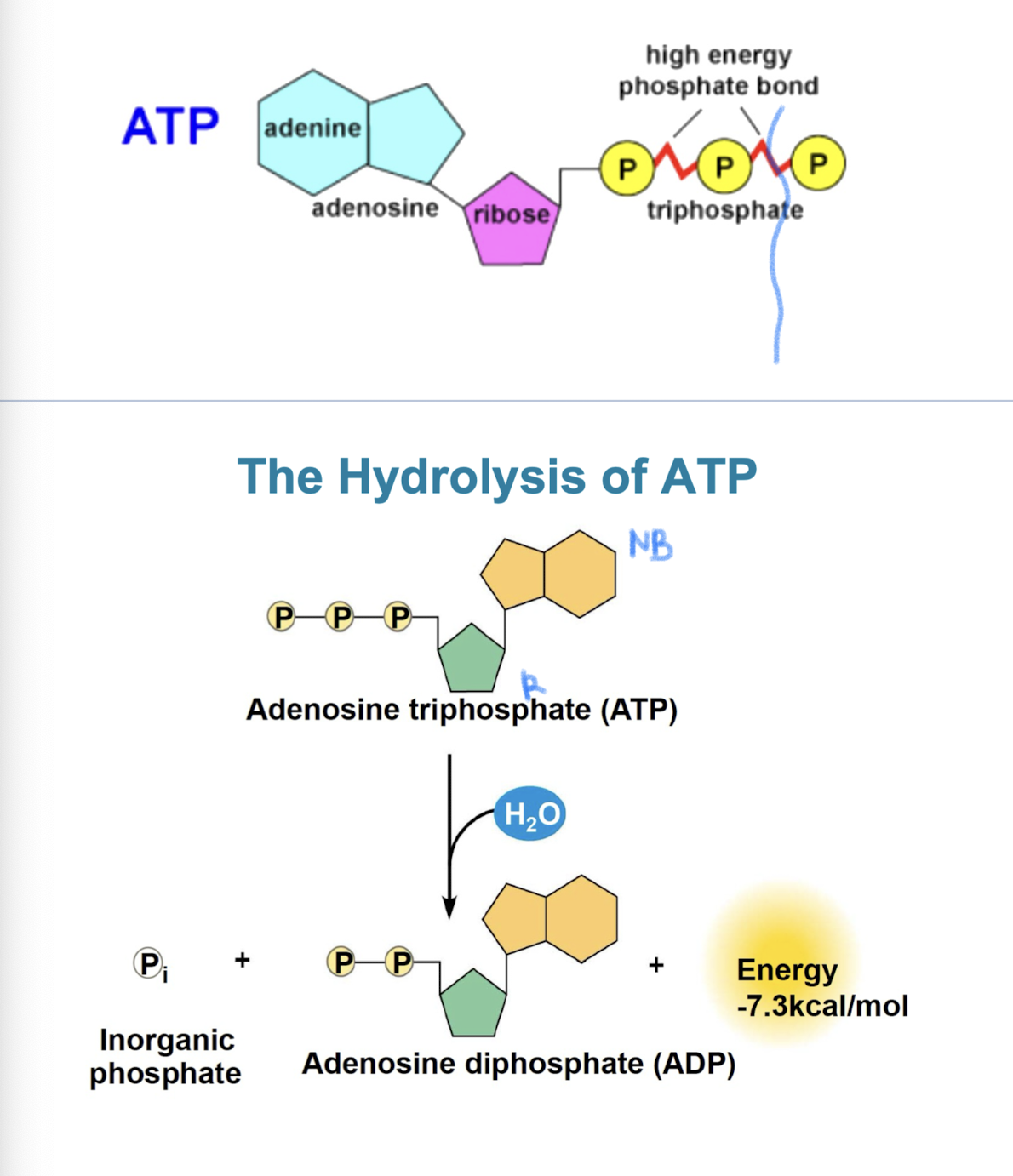
Hydrolysis of ATP
energy is released when terminal phosphate bond is broken (comes from chemical change to state of lower energy, not from bond itself) exergonic reaction
triphosphate tail of ATP like a compressed spring (hydrolysis releases a lot of energy due to repulsive forces)
ATP + H2O → ADP + Pi (ΔG = - 7.3 kcal/mol (- 30.5 kJ/mol))
How ATP drives transport + mechanical work
leads to a change in a transport proteins shape that allows transport of solutes (negatively charged phosphates)
ATP binds non covalently to motor proteins + then hydrolyzed —> shape change that walks the motor protein forward
Regeneration of ATP
ATP is a renewable resource that is regenerated by addition of phosphate group to ADP
energy to phosphorylate ADP comes from catabolic reactions in cell
overall: coupled reactions are exergonic (ΔG < 0)