BIOL 107 Part G - DNA and RNA (Questions)
1/38
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
39 Terms
The base unit of DNA is ___. Describe its structure (bonds, number of strands).
Nucleotides.
DNA is made of two strands, each made up of a chain of nucleotides. The chain of nucleotides within one strand are joined by covalent bonds. The two strands connect via hydrogen bonds between base pairs
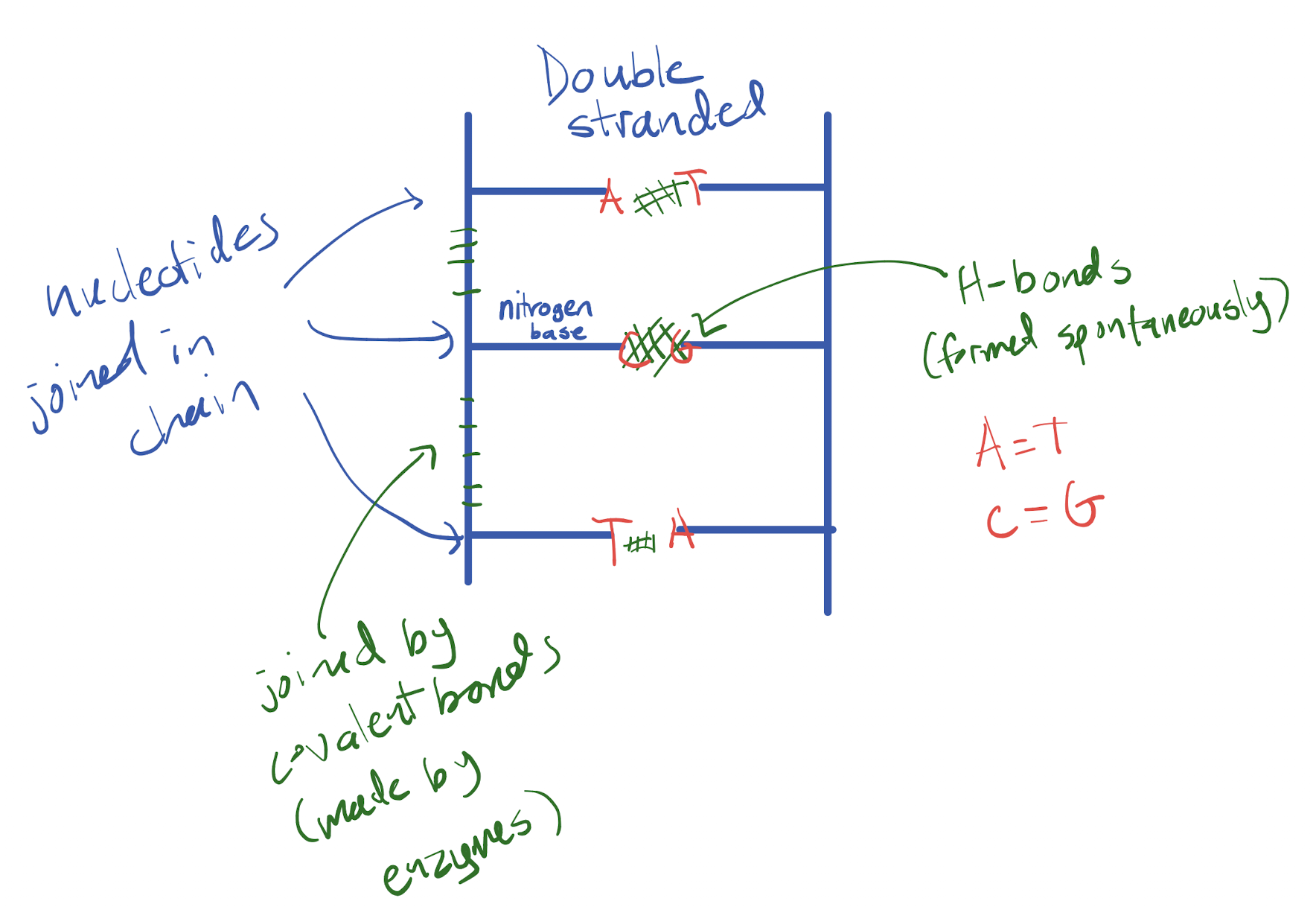
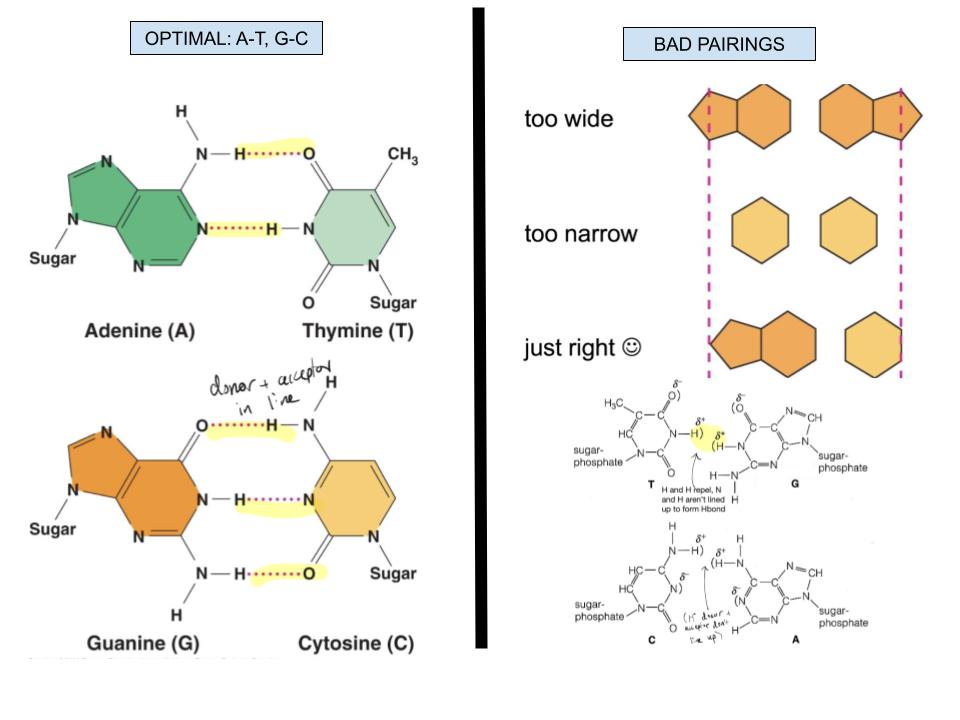
Which base pair combinations allow for hydrogen bonds between DNA molecules to be formed spontaneously? What makes for bad base pair combinations?
Complementary bases (A with T and G with C) have the correct shape and alignment for hydrogen bonds to form. Incompatible shapes may be too wide or too narrow to correctly bond, and the donor and receiver of electrons to form H bonds (H and electronegative atom) will not be aligned to form these bonds
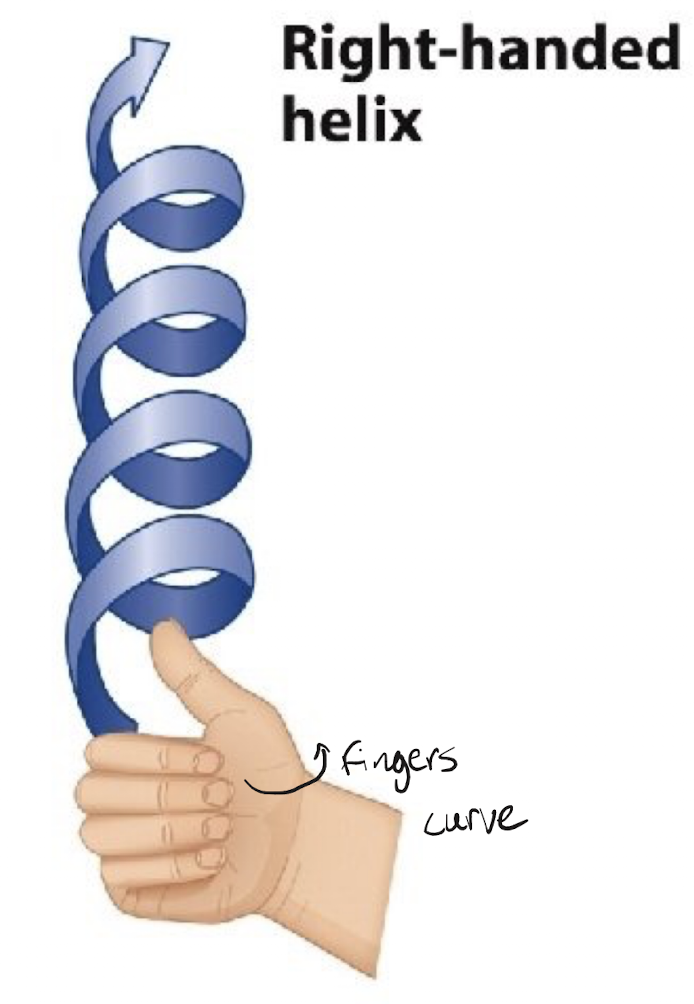
The orientation of DNA is a (right/left) handed helix. What is its size?
Right, 2 nm
Chromosomes and plasmids, nuclear chromosomes and organelle chromosomes.
Compare and contrast chromosome shapes, amounts and locations in prokaryotes and eukaryotes.
Prokaryotic chromosomes: circular, 1 to 2 copies per cell, middle of cytosol.
Eukaryotes have nuclear and organelle chromsomes:
Nuclear chromosomes are linear, have several (46 in humans) per cell, and found in nucleosomes fibres inside nucleus.
Organelle chromosomes are circular, have ~10 per organelle, found in stroma of chloroplasts or matrix of mitochondria
Describe the nucleic acid rules that apply to DNA and RNA
Made using triphosphate nucleotides: new DNA will contain dNTPs, RNA will contain NTPs
Nucleic acids will have a 3’ end and a 5’ end. New nucleic acids will always be made 5’ to 3’
Nucleic acids will be created using a single stranded DNA template
The new and template strand in DNA replication will pair antiparallel: the 5’ end of the template matches with the 3’ of the new and vice versa
Base pairs in nucleic acids will pair via complementary base pairing: the new strand will pair with the complementary base of the template strand
What supplies the energy needed for RNA synthesis? Why?
NTPs used to create the new strand, modification of triphosphates releases energy
All polymerases are able to ___. The ability specific to RNA polymerase is ___, while the ability specific to all DNA polymerases is ___.
Elongate. Initiate, proofread
Why does there need to be a separate leading and lagging strand created at each replication fork? How many DNA polymerases (III) are needed per DNA helix?
Leading strand can be made continuously as its template strand goes 3’ to 5’ from the fork, meaning it can be made from 5’ to 3’. However this means the other template strand (paired antiparallel) will be 5’ to 3’ from the fork, which means it is impossible to create a continuous strand via nucleic acid rules.
2 DNA pols needed: one at each strand
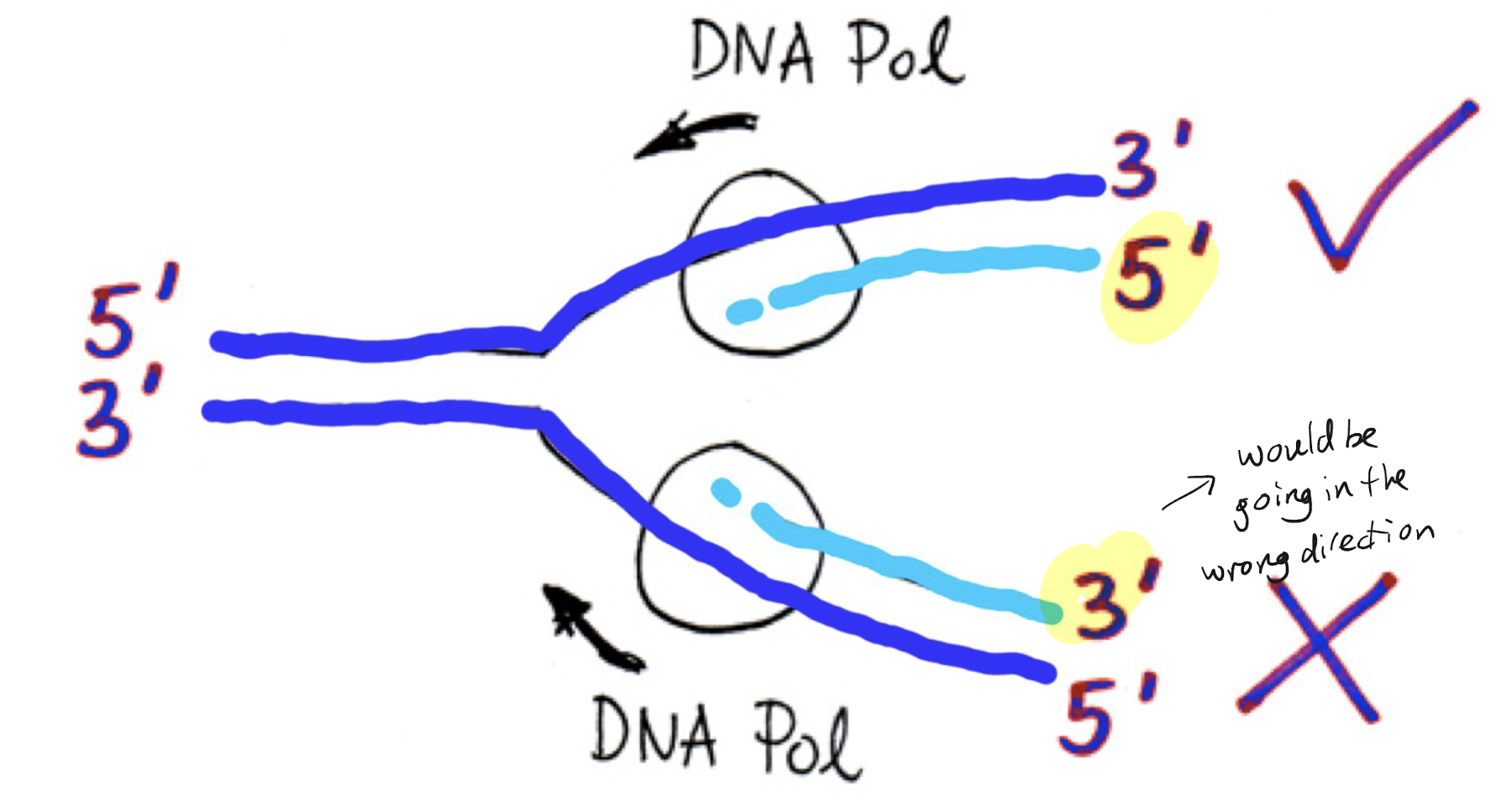
Which proteins/enzymes are needed to create the leading strand? Explain this process.
Primase and DNA pol III. Primase creates an initial primer at the ori, which allows DNA polymerase III to create a strand off of it. As the template strand is 3’ at the ori, based on antiparallel the leading strand is created at 5’ and can elongate continuously without issue
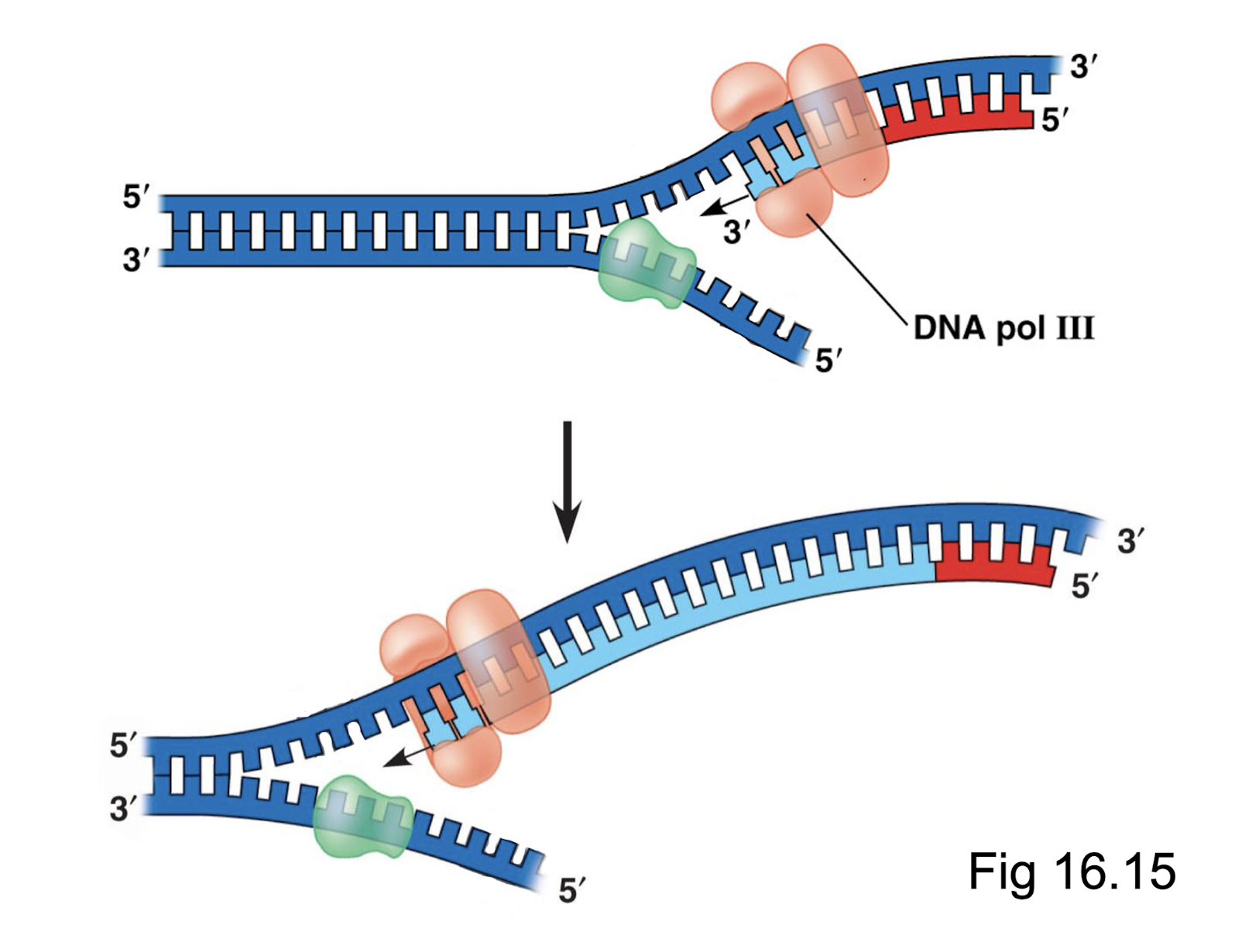
Which proteins/enzymes are needed to create the lagging strand? Explain this process.
Primase, DNA pol III, DNA pol I, DNA ligase.
As the template strand is 5’ at the ori, based on antiparallel the leading strand cannot start replication at the ori. As the DNA strands separate, primase creates an primer at the most newly exposed 3’ end of the template strand. DNA polymerase III creates an Okazaki fragment off of this primer towards the ori. As the DNA becomes progressively more unzipped, primase creates a new primer closer to the replication fork and DNA polymerase III builds off of this primer until it reaches the previous primer. More primers are created for the entirety of the template strand length.
As the lagging strand is being created in Okazaki fragments, DNA polymerase I removes primers from between already created Okazaki fragments, replacing the RNA nucleotides with needed DNA nucleotides. DNA ligase attaches these separated fragments together
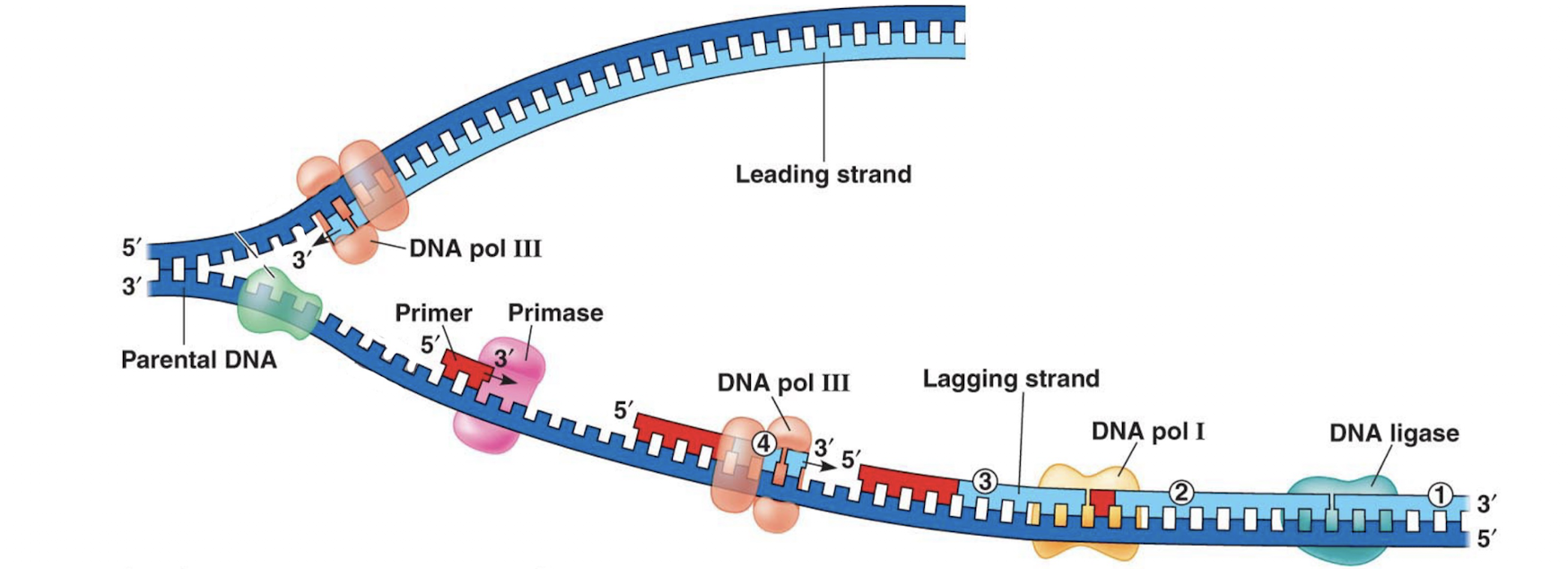
Is DNA ligase a type of polymerase? From your answer, does DNA ligase also need to work from 5’ to 3’ on the created strand?
No, it can glue nucleotides together from any direction because it does not follow polymerase rules
Explain how replication forks form to begin DNA replication at the ori. What is the role of each protein, and why?
Initiators attach to the ori and pull the double stranded DNA apart. Helicases move like a zipper attached to each single strand of DNA on the lagging side, separating the strands further from where initiators first pushed apart the strands.
Initiators start the transition from dsDNA to ssDNA, but cannot pull apart the strand in its entirety. Helicases are needed to further separate the strands but cannot bind to the DNA as dsDNA and thus needs the initiator.
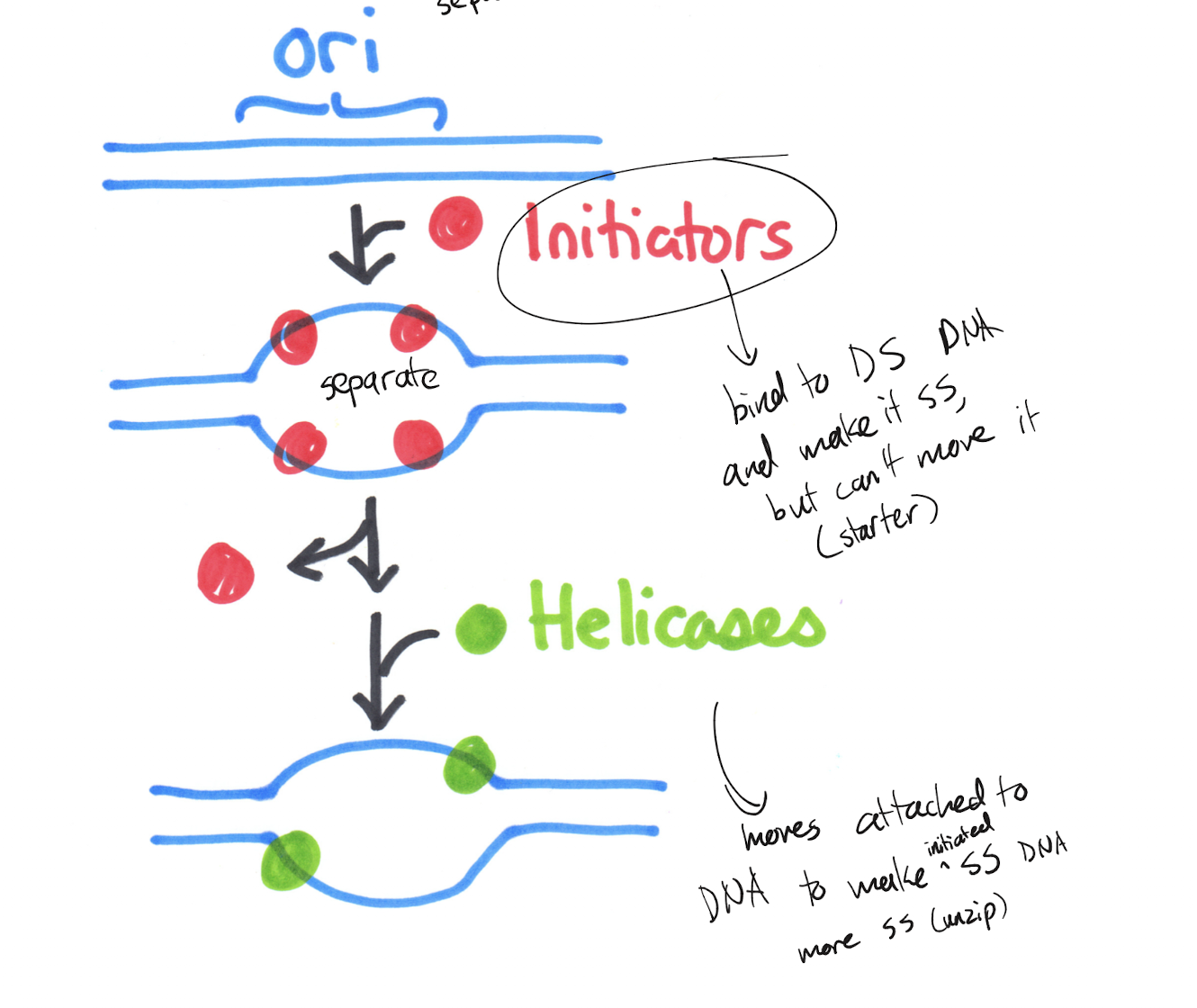
From one ori, how many replication forks are formed? What are the orientations of the leading/lagging strand at each replication fork?
2 formed, one in each direction. Leading and lagging strands are on opposite sides of each other, and their orientation is flipped for the replication fork in the opposite direction. If a strand goes from 5’ to 3’ left to right, the replication fork towards the left will start from 3’ and allow for the creation of the leading strand on this top strand while the replication fork towards the right starts from 5’ on the template and thus must create the lagging strand.
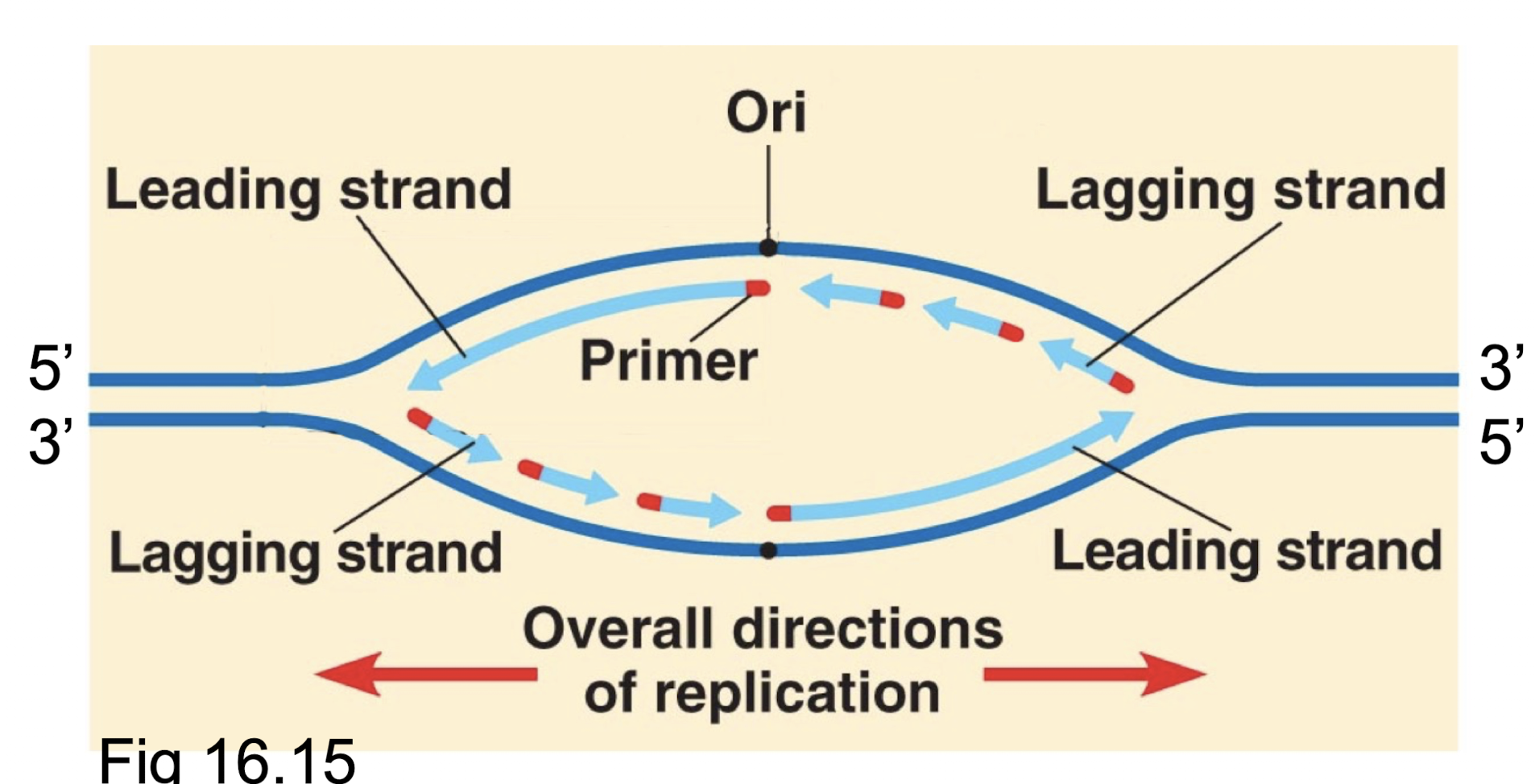
Explain how DNA polymerases proofread their own work. In what scenario does this occur?
Recognize a mistake, remove the incorrect nucleotide by changing direction briefly, replace the removed nucleotide with the correct one. Occurs during DNA replication
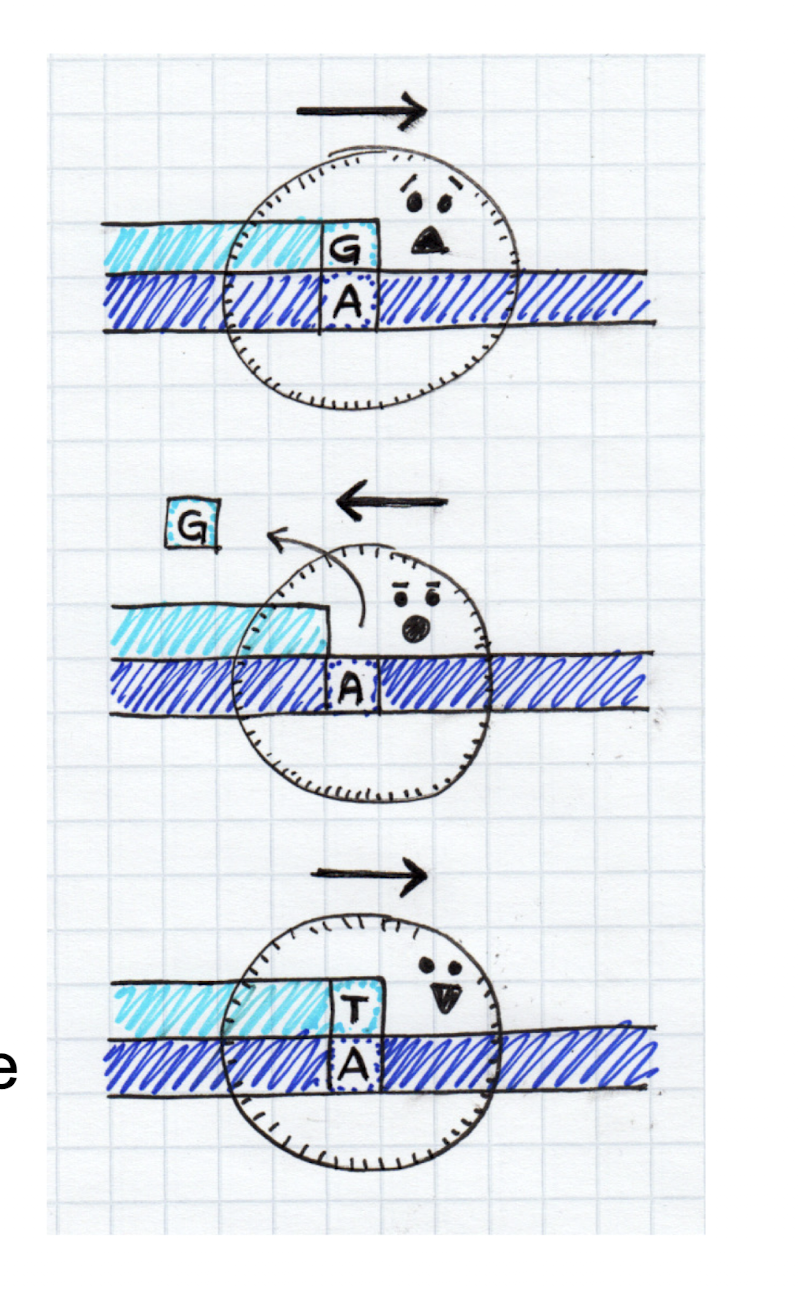
Explain how DNA repair enzymes fix damaged DNA. In what scenario does this occur?
Repair enzymes recognize and remove damaged sections. Then, DNA Pol I replaces the missing nucleotides, followed by DNA ligase joining the nucleotides. Repair will occur before or after DNA replication
Compare the proofreading ability of DNA pols with DNA repair enzymes.
DNA polymerases remove and replace (structurally sound) incorrect polymerases, while repair enzymes remove structurally damaged nucleotides that were correctly replicated
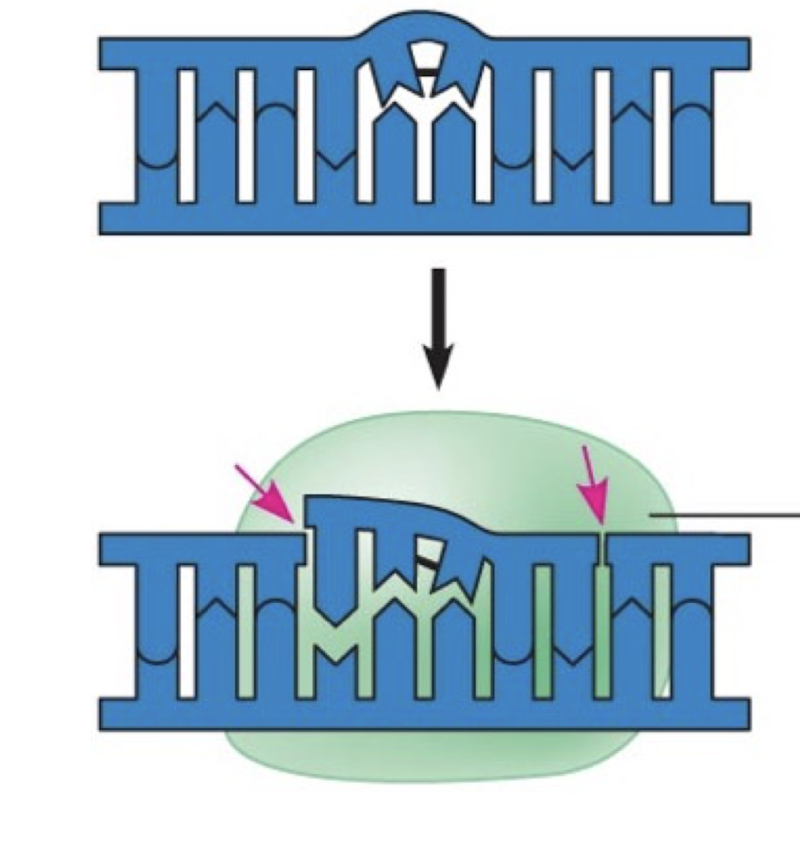
Explain the mechanism of DNA replication in circular DNA (number and location of ori, amount and behaviour of replication forks, termination). Which types of DNA use this, from which type of organisms?
Have a single ori on the circular chromosome. Will have two replication forks that spread from each side of the ori. Replication finishes when replication forks meet, effectively pinching off the two semi-circle forks to leave two circular chromosomes
Prokaryotic chromosomes and plasmids from prokaryotes. Organelle chromosomes from eukaryotes
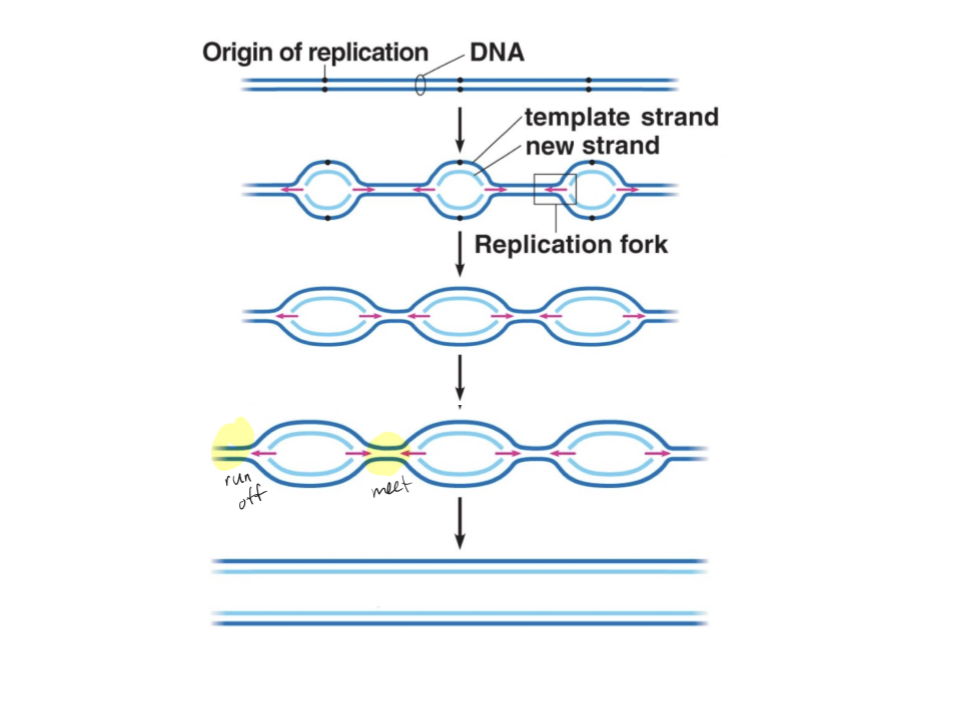
Explain the mechanism of DNA replication in linear DNA (number and location of ori, amount and behaviour of replication forks, termination). Which types of DNA use this, from which type of organisms?
One ori is located at every 40 000 bps, and replication simultaneously starts at these many oris. Two replication forks form for every ori, splitting farther out until a fork finds another fork or runs to the end of the molecule. The collision of two forks will cause the DNA to split completely. Replication finishes when all replication forks have collided or run off the ends.
Nuclear DNA from eukaryotes
Compare the speed of chromosome replication in human cells compared to prokaryotic chromosomes.
~8 hrs vs 40 mins because human chromosomes are magnitudes larger than prokaryotic chromosomes
Why do prokaryotic chromosomes, plasmid chromosomes, organelle chromosomes, and nuclear chromosomes replicate?
Prokaryotic and nuclear chromosomes replicate before cell reproduction. Plasmid chromosomes and organelle chromosomes replicate in order to maintain their number in the cell (~10 per organelle, ~100 per prokaryotic cell)
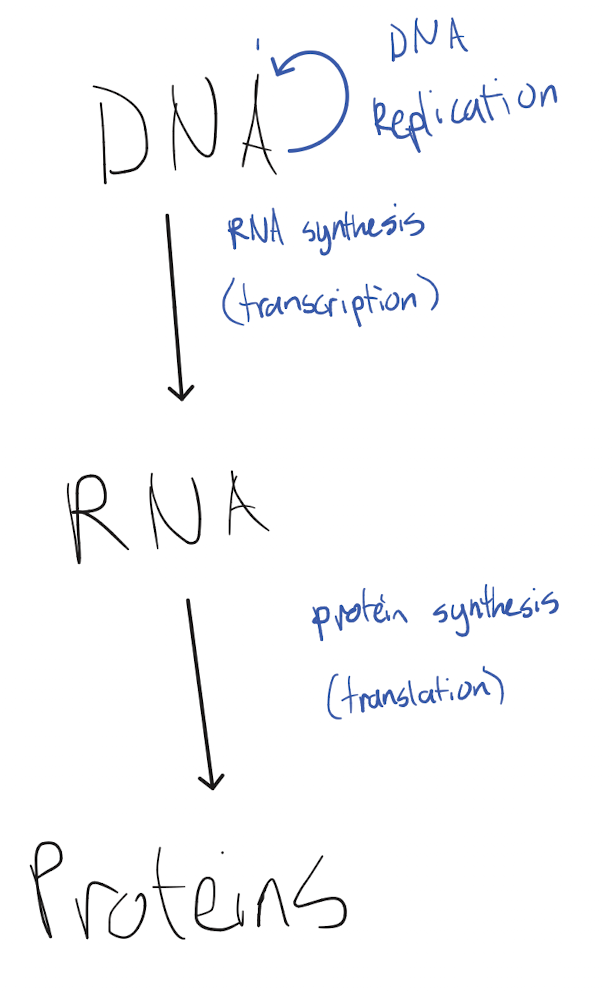
Explain the pathway of the central dogma of biology, including all relevant processes and locations.
Genetic information is stored in DNA in the nucleus, which is maintained via DNA replication. Transcription in the nucleus synthesizes an RNA molecule from the DNA, transferring the genetic information from the DNA molecule to RNA. The RNA molecule moves to the cytoplasm, where translation synthesizes a protein based on the RNA molecule, transferring the genetic information from RNA to proteins
What proportion of genes encode mRNAs? What about the remaining proportion?
50%, other 50% encode all other types of RNA
Which base pairs are used in DNA, and which are used in RNA?
A-T, C-G. A-U, C-G
Explain the process of translation initiation, including regions, proteins/enzymes.
RNA polymerase attaches to the promoter behind base pair +1, aided by a general TF protein to find the protein
RNA polymerase separates the strands of DNA at the promoter region
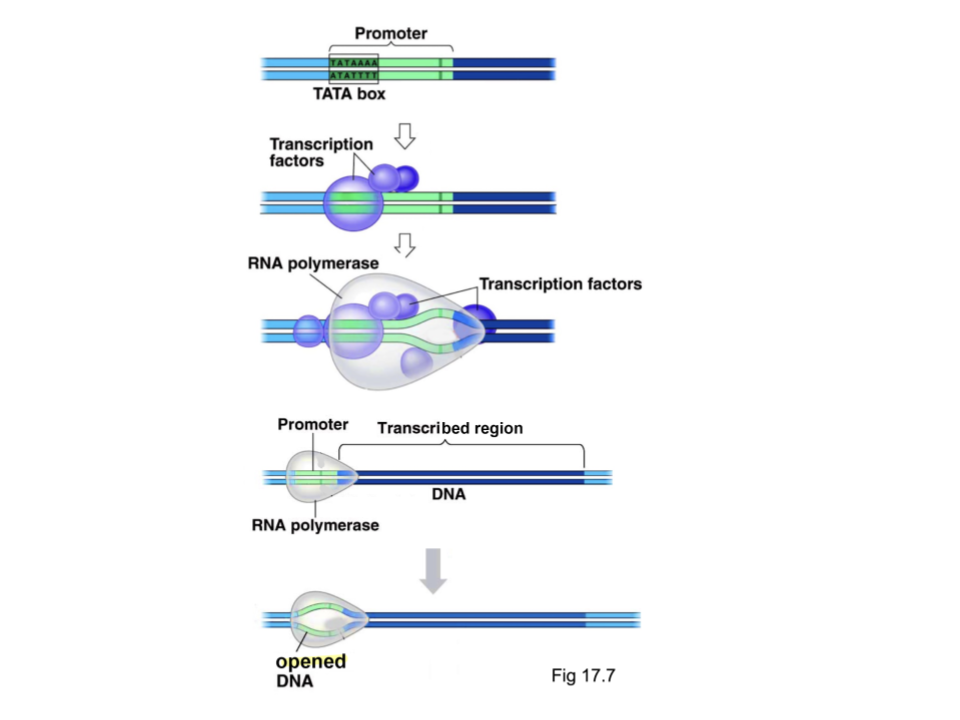
Explain the process of translation initiation, including regions, proteins/enzymes.
From the DNA opened by the RNA polymerase, RNA polymerase uses the 3’ to 5’ strand of the DNA as a template strand to create a 5’ to 3’ RNA
The RNA polymerase moves along the template DNA strand, attaching NTPs to create a strand of RNA
The DNA strands close behind the RNA polymerase once the NTPs are added and the template strand is no longer being used
Strong base pair bonds have how many H-bonds? How many does an A-U pairing have, and what does this imply?
Three. A-U has 2, meaning it is a weak pairing
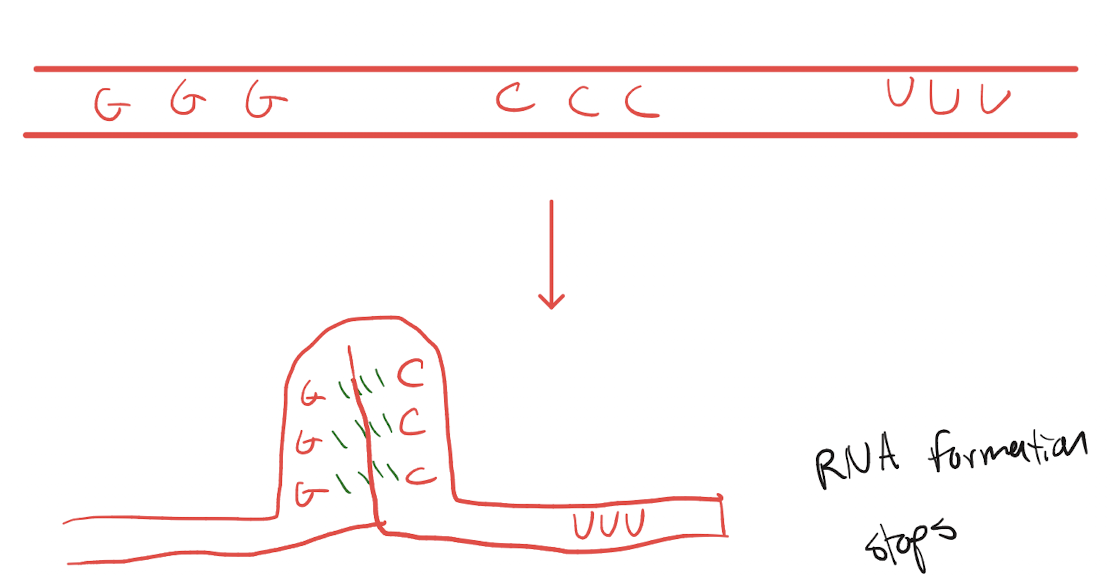
Explain the process of intrinsic termination, including regions, proteins/enzymes.
Intrinsic terminator is reached in the DNA and translated into RNA
Terminator RNA sequence forms a hairpin loop, bonding with its own base pairs, that pulls away from the template DNA
RNA formation stops
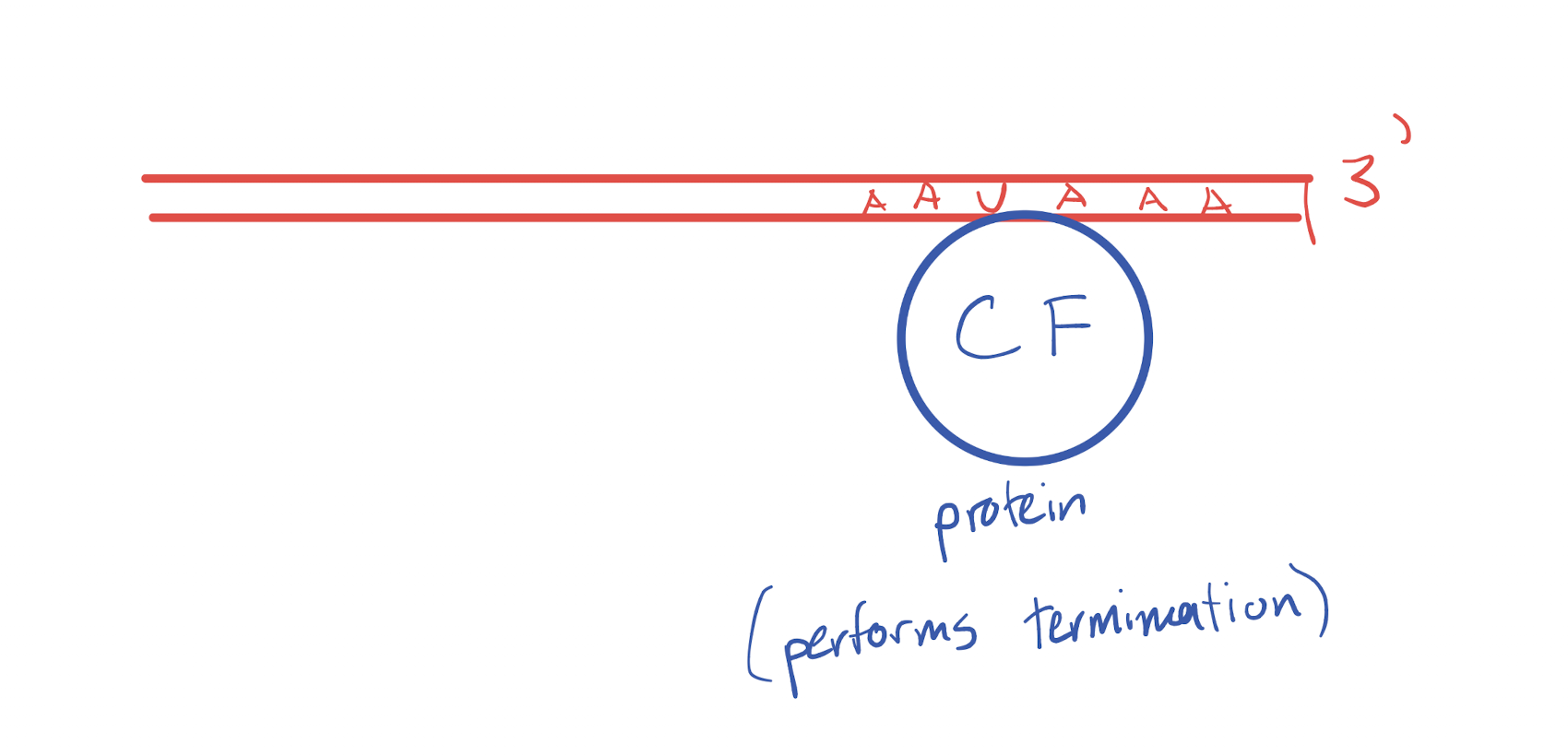
Explain the process of factor-dependent termination, including regions, proteins/enzymes.
Factor dependent terminator is reached in the DNA and translated into RNA
Terminator RNA sequence recruits a cleavage factor protein based on a sequence in the created strand and retains said sequence
RNA formation stops as RNA polymerase is stopped by the protein
Compare RNA synthesis and DNA replication in terms of:
Starting DNA region where polymerase is attached
Required amount and properties of template strands
Are primases and helicases required?
Ending DNA region where the polymerase stops
RNA starting region = promoter, DNA region = ori
RNA needs one template strand that goes 3’ to 5’, DNA uses both to create one leading and one lagging strand based on orientation
DNA synthesis needs primases for initiation and helicases to open DNA, RNA synthesis does not need helicase as the polymerase itself opens the DNA and does not need primase as TF proteins help to initiate the process
RNA ending region = terminator, DNA region = telomere
Prokaryotes perform which out of rRNA processing, mRNA processing, and tRNA processing? Eukaryotes?
Prokaryotes: rRNA, tRNA
Eukaryotes: all
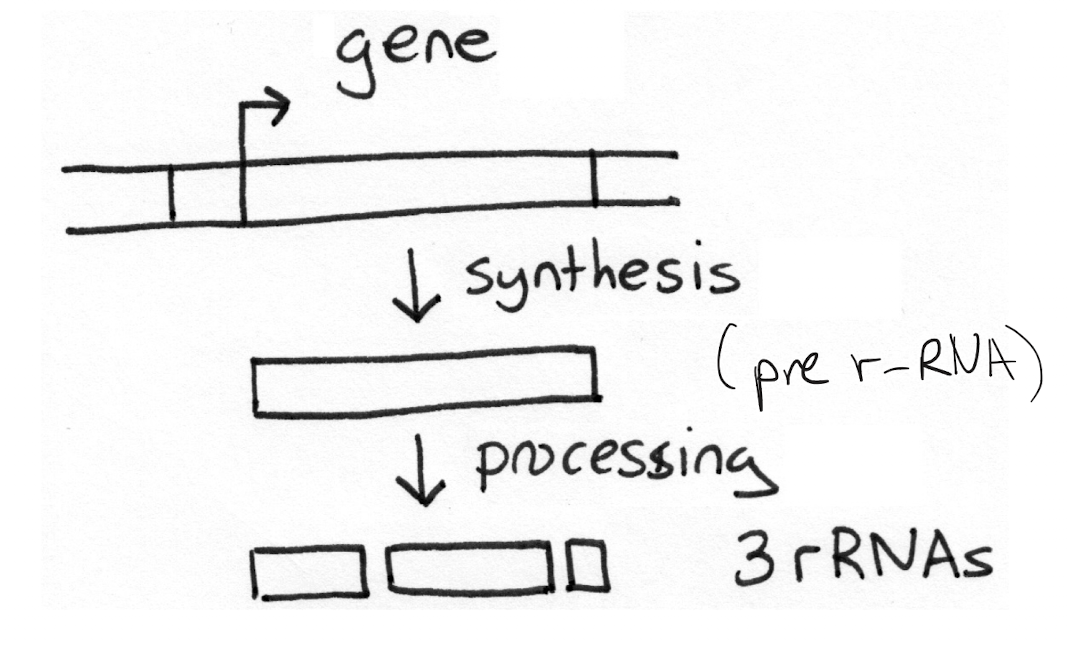
Explain how rRNA processing occurs. Why does this happen?
Pre-rRNA is cut into two or more different pieces. Done due to efficiency: all (multiple) rRNAs can be produced from a single gene instead of translation having to happen multiple times
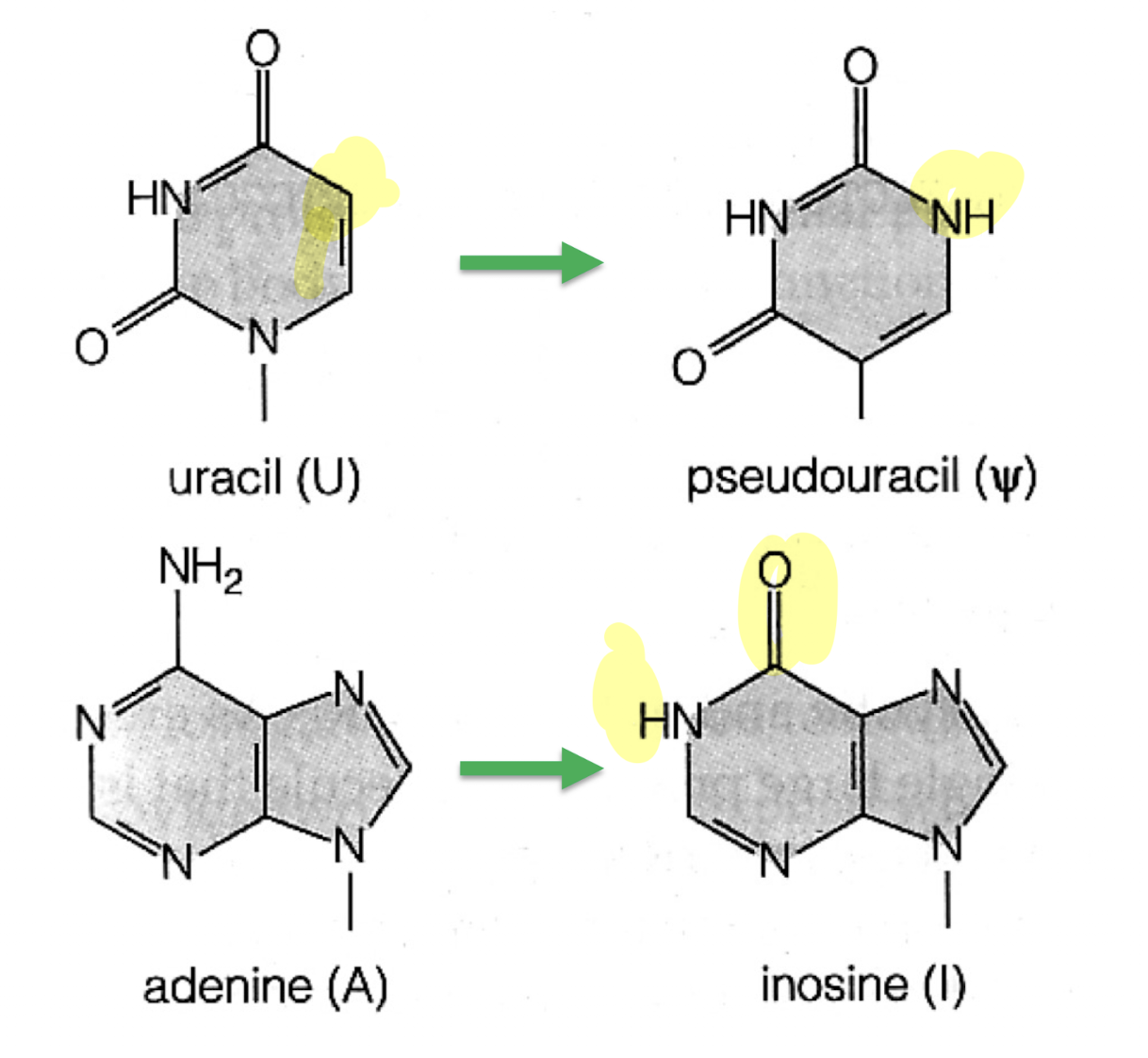
Explain how tRNA processing occurs. Why does this happen?
Specific A, G, U, or C bases in pre-RNA are modified through changes to the base structure. Done to allow tRNAs to fold into complex shapes needed for its role
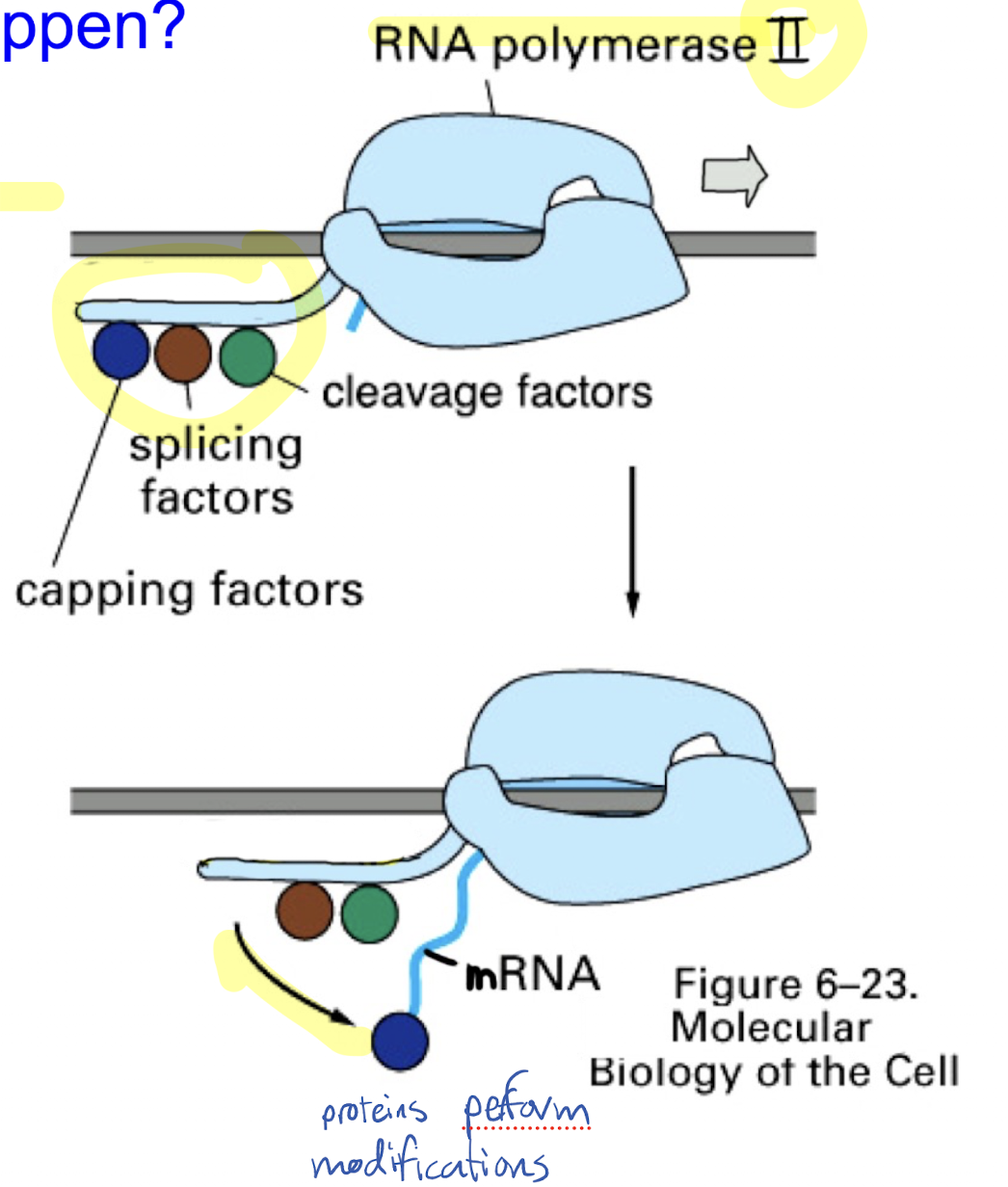
What are the three modifications made to pre-RNA during mRNA processing? Describe the general mechanism by which these modifications occur, including relevant proteins and enzymes.
Addition of 5’ cap at 5’ end
Addition of poly(A) tail at 3’ end
Removal of introns and splicing of exons
Factor proteins that perform modifications ride on RNA polymerase II, and as the pre-mRNA is created, the proteins will move onto the strand and modify it
Explain how a 5’ cap and poly(A) tail is added to pre-RNA during mRNA processing. Why are these modifications done?
Capping factor proteins attach a 5’ cap on the mRNA’s 5’ end. Cleavage factor section of RNA recruits poly(A) polymerase that attaches a poly(A) tail to the 3’ end of the mRNA.
Tail and cap helps export from nucleus, helps ribosomes bind, and extends lifespan of mRNA from minutes to hours by blocking RNA digestion enzymes (RNAses)
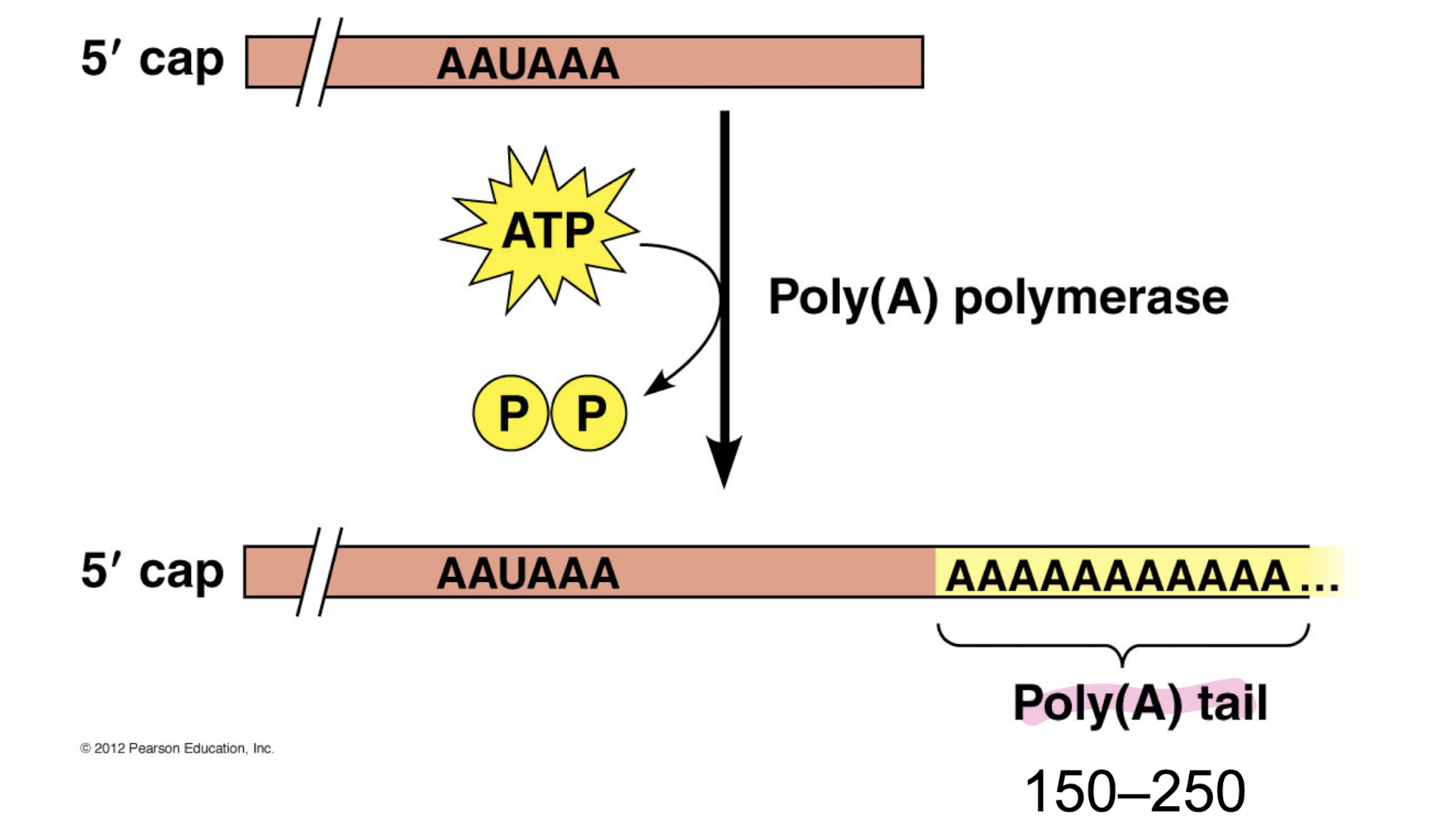
Are capping factors and poly(A) polymerases exceptions to the nucleic acid rules? Why or why not?
Capping factors are not polymerases so the rules don’t apply. Can create nucleotides from 3’ of new strand
Poly(A) polymerase is not made using ssDNA template
Explain how introns are marked, then removed, and exons are spliced.
Splicing factors mark locations of introns during mRNA synthesis, identifying where introns start and end by jumping onto the boundaries to mark them
Once mRNA synthesis has finished, spliceosomes remove introns by cutting identified boundaries of introns and ligating remaining exons together
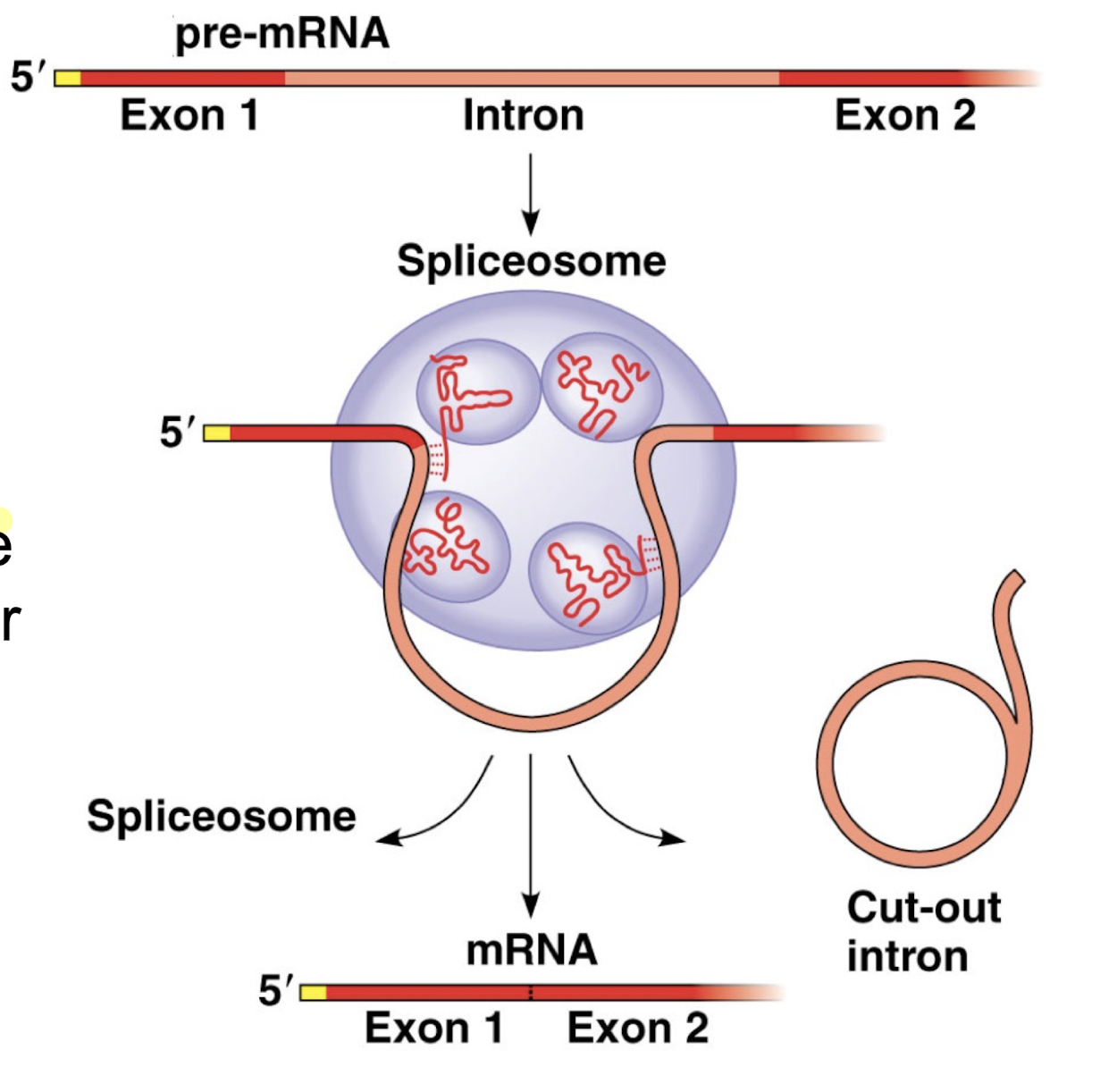
Why is having introns that end up being removed a disadvantage? Why does pre-mRNA still have introns despite this?
Introns make genes larger, meaning it takes more time to synthesize mRNAs. It also takes time to remove all the introns in processing.
Alternative splicing is the advantage: pre-RNA from one gene can be spliced multiple different ways, allowing for one gene to express multiple different proteins. This lets multicellular Eukaryotes make cell-specific versions of proteins (ex. Insulin receptor A found in the placenta vs insulin receptor B found in fat and muscle cells)
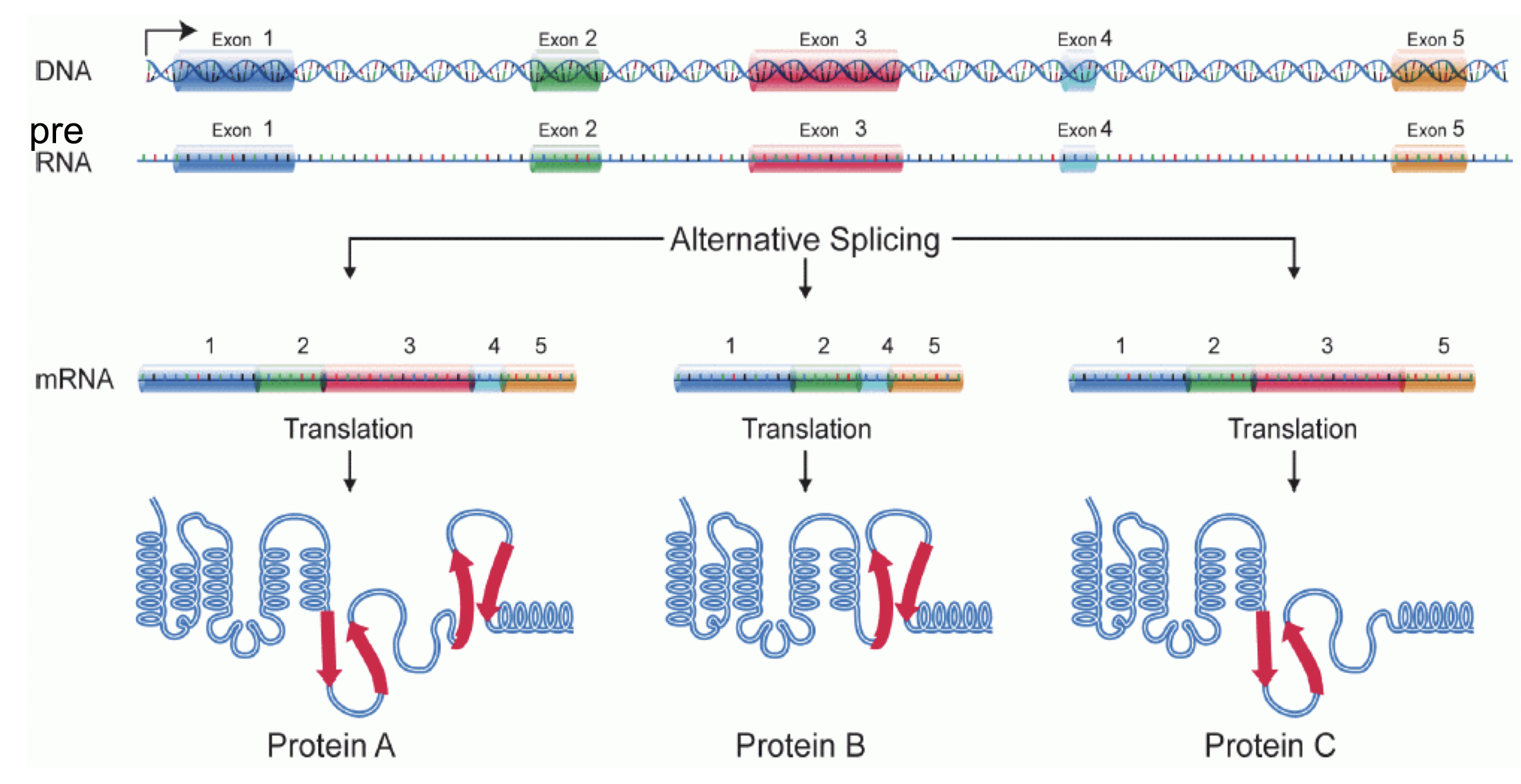
Explain the mechanism of alternative splicing.
RNA binding proteins will hide specific splice sites on pre-RNA marked by splicing factors, which will cause certain exons to be included in the intron boundaries that get removed from the mature mRNA. Different orientations of binding proteins will lead to different proteins