BIO417 Term test 2
1/151
Earn XP
Description and Tags
Lectures 7-11
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
152 Terms
Semiconservative replication
New DNA has one parental strand (conserved) and one new strand
Polymerase ε
Continuous replication on the leading strand; 5’ → 3’
Polymerase δ
Replicates the lagging strand one short fragment at a time
Polymerase ⍺
Synthesizes RNA-DNA hybrid primers (contains a primase)
Single stranded DNA binding protein (SSBP)
Binds to unwound DNA to stabilize it during replication (replication protein A)
Sliding clamp
A ring-shaped protein complex that anchors DNA polymerase to the DNA template, preventing it from detaching (Proliferating cell nuclear antigen, PCNA)
CMG helicase
Unwinds the double helix during DNA replication
Polymerase erros
Wrong nucleotide incorporated
Extra or missing nucleotides (repetitive sequences) from replication slippage
Polymerase error solutions
Proofreading from pol ε and δ
Mismatch repair
Simultaneous DNA replication and transcription erros
Encounters between replisome and transcriptional complexes
R-loops → ssDNA, mutations, nicks, fork stalling or collapse, genomic instability
R-loops
Three stranded nucleic acid structures that form when the RNA transcript anneals to DNA template (CG rich region)
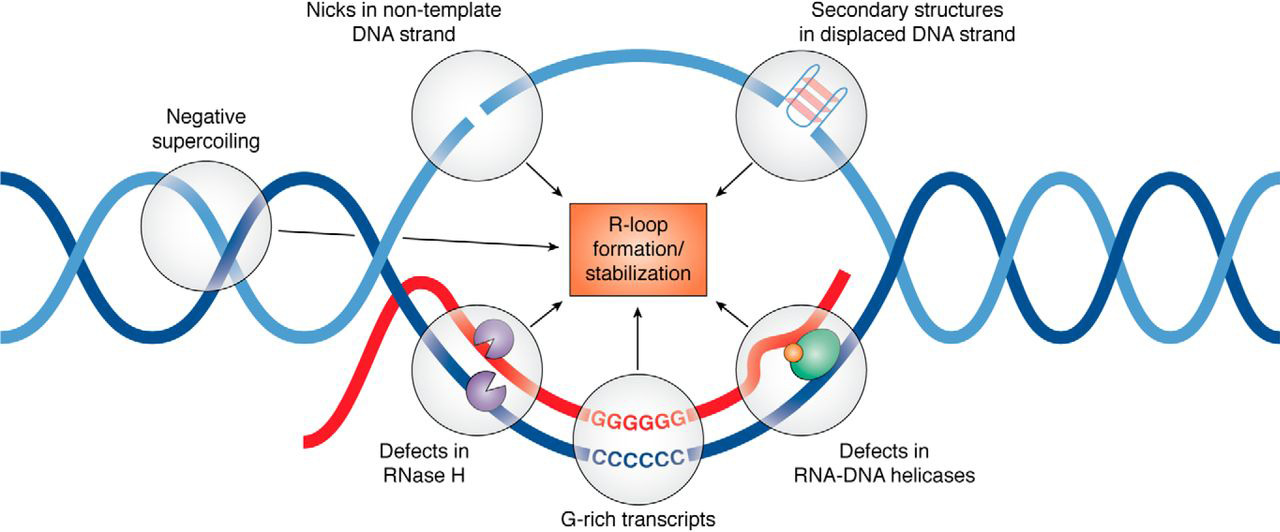
Simultaneous DNA replication and transcription error solutions
Temporal separation (early replication genes are transcribed later in the S phase
If transcribed throughout the S phase (e.g. rRNA genes), genes alternate between replication and transcription
Both don’t happen at the same time/place
Mismatch repair (MMR)
Repairs mispairing of nucleotides from tautomeric shifts, wobble, and strand slippage
Mispairing/wobble
Slight shifts in the position of nucleotides in space
e.g. causes wobble between a normal T & normal G
Non-tautomeric forms of bases → an additional proton on adenine causes a wobble in C:A
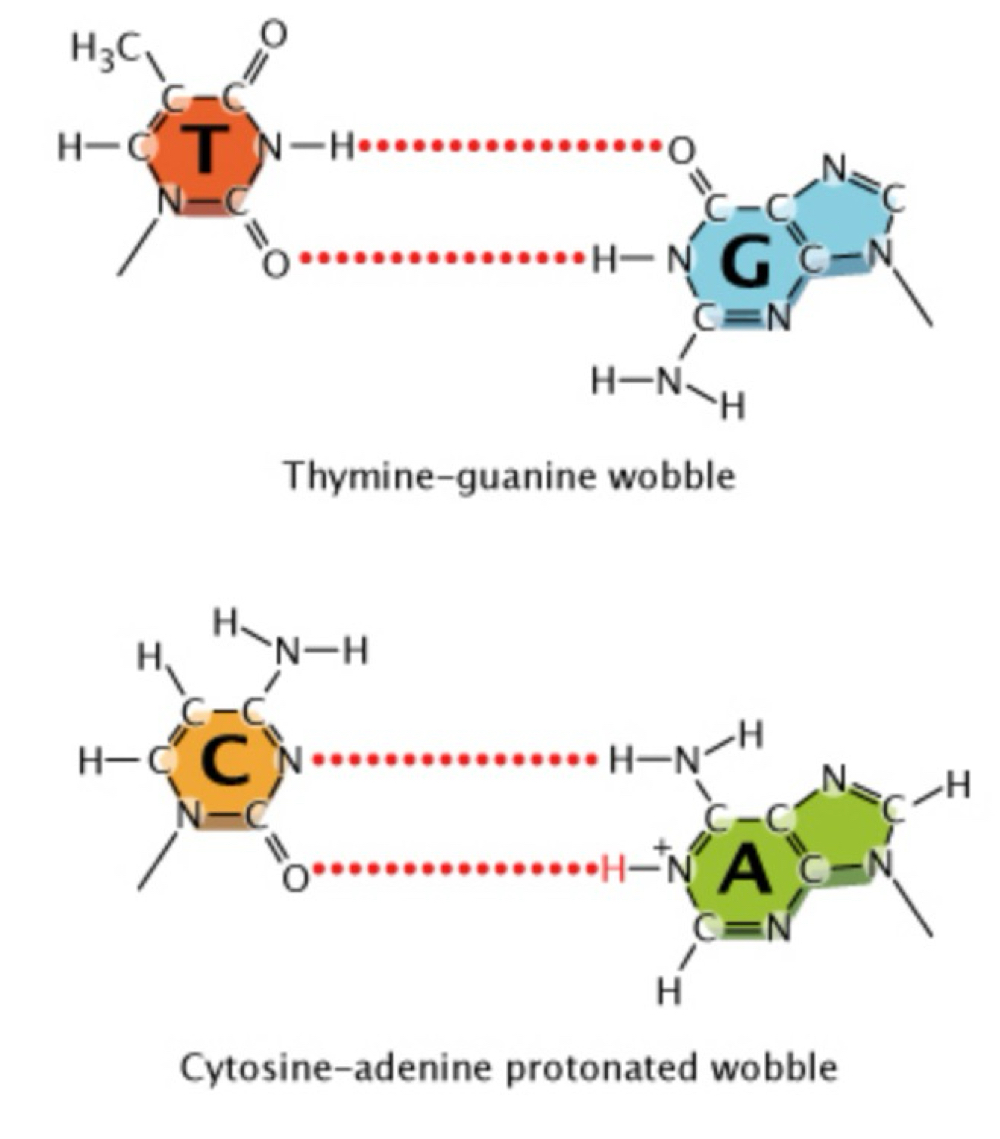
Strand slippage
A strand can loop out, causing an addition (newly synthesized strand loops out) or deletion (template strand loops out)
MMR genome maintenance mechanism
Recognition of replication errors
Cleaves newly synthesized strand
Exonuclease removes area around error
DNA polymerase fills gap and ligase seals break
Highly conserved
Key components of MMR
Mut proteins
MSH → MutS homolog complex, dimer
Associate with PCNA (sliding ring) and recruits:
MHL → MutL homolog complex (with endonuclease activity), dimer
Exonuclease (EXO1)
DNA polymerase
Ligase
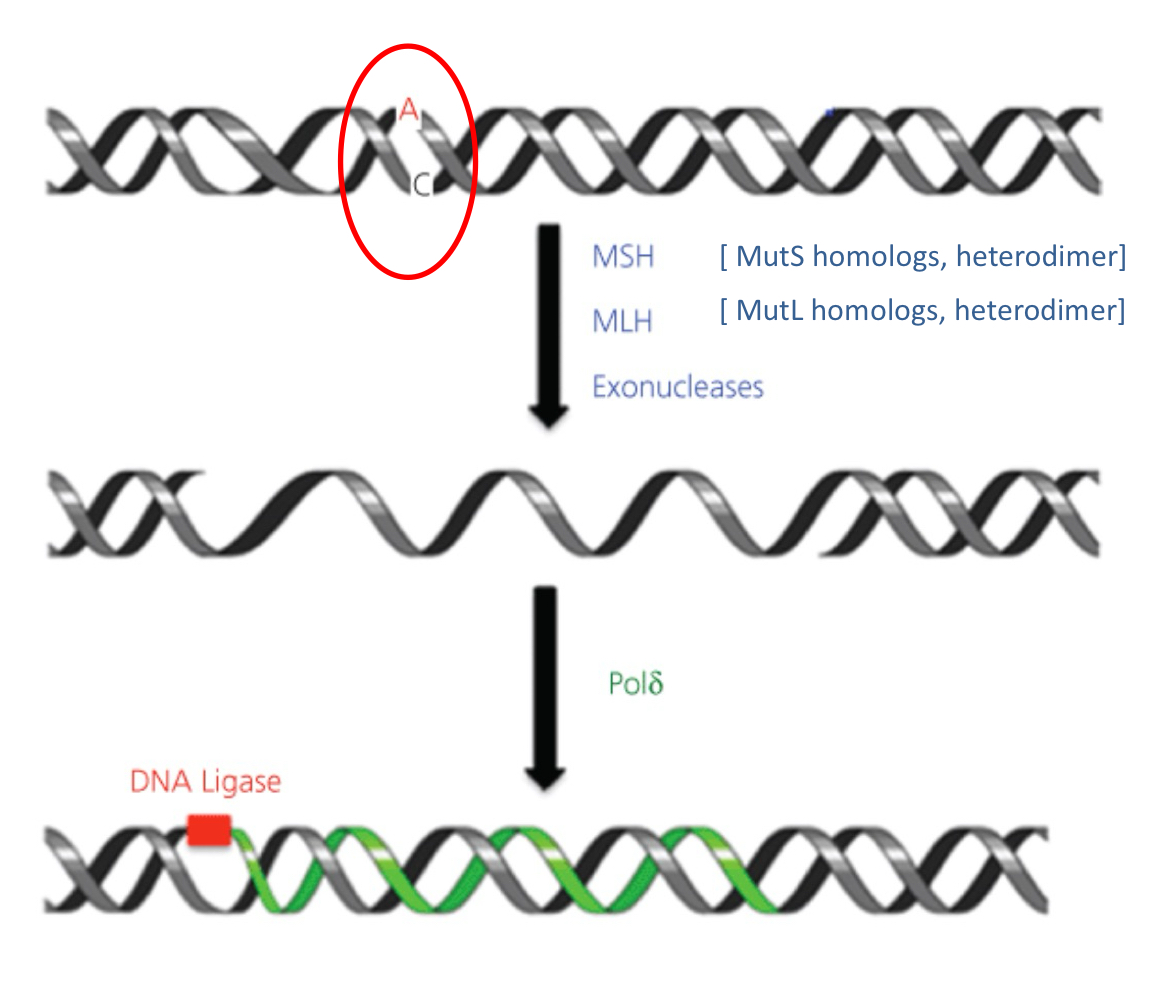
MMR process
MutS (mutator S) and MutL homologs (MSH, MLH) initiate MMR
MSH complex recognizes mismatch
Associates with PCNA (proliferating cell nuclear antigen)
MSH recruits MLH, w/ endonuclease activity
Newly synthesized strand is cleaved, region around mismatch is removed by EXO1
Gap filled by pol δ, nick sealed
Role of epigenetic marks in MMR
H3K23me3 (active chromatin) → recruits MSH complex (MutS 𝛂)
Recruitment of MSH to protect actively transcribed genes from mutation (prior to replication, followed by scan of newly synthesized strand)
Base excision repair (BER)
Removal of damaged or altered bases (non-bulky)
Altered by oxidation, methylation, deamination
Or excised by hydrolysis
Nucleotide excision repair (NER)
Repairs damage (chemicals, UV, radiation) that distorts DNA helix (pyrimidine dimers, DNA adducts); removal and re-synthesis of segments
Reactive oxygen species (ROS)
Byproducts of metabolism that can cause chemical modifications to bases
High reactivity → can cause damage as soon as it comes in contact w/ DNA
Can cause mispairing during replication if not repaired
BER mechanism
Constantly scan for errors
Glycosylases initiate excision of damage
Specific glycosylases for specific types of altered bases
Flips nucleotide out of helic, cleavage of abnormal bases
Creates apurinic or apyrimidinic sites (AP sites)
Recognized by AP endonucleases
Excise sugar phosphate groups
Addition of nuclotides:
Short patch (1 nucleotide) = Pol β
Long patch (2-10 nucleotides) = Pol ε/δ
Ligase seals nick
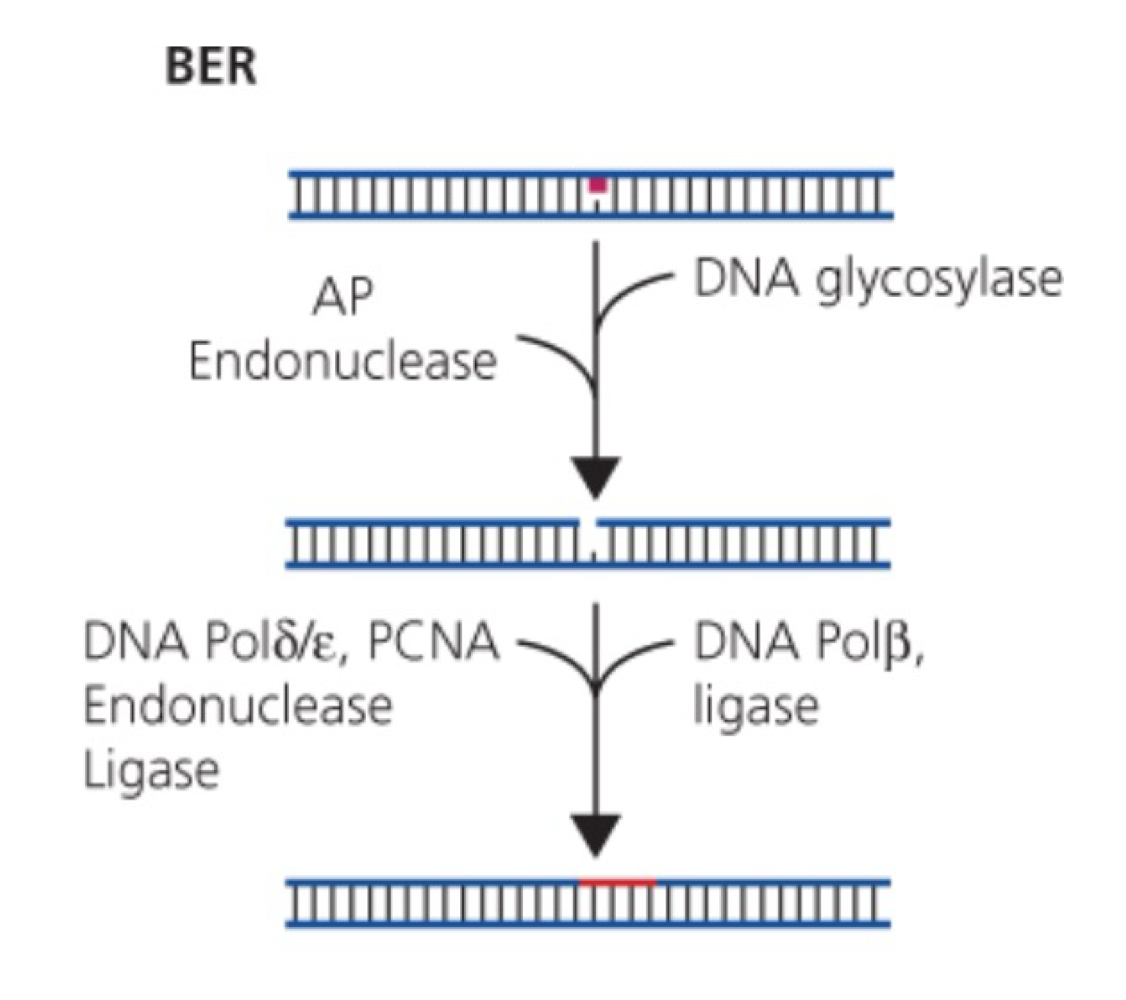
Glycosylases
Recognize and remove specific damaged or inappropriate bases
NER pathways
GG-NER (global genome NER)
TC-NER (transcription coupled NER)
Global genome NER (GG-NER)
Checks entire genome (actively transcribed and silent)
Ex) Xeroderma pigmentosum (XP)
Transcription coupled NER (TC-NER)
Removes blocks - RNA pol elongation
Ex) Cockayne syndrome (CS), XP
GG- and TC-NER…
Share components, but differ in the way damage is recognized
GG-NER damage recognition
XPC protein → recognizes damage anywhere in the genome
TC-NER damage recognition
Recognized by pol II stalling
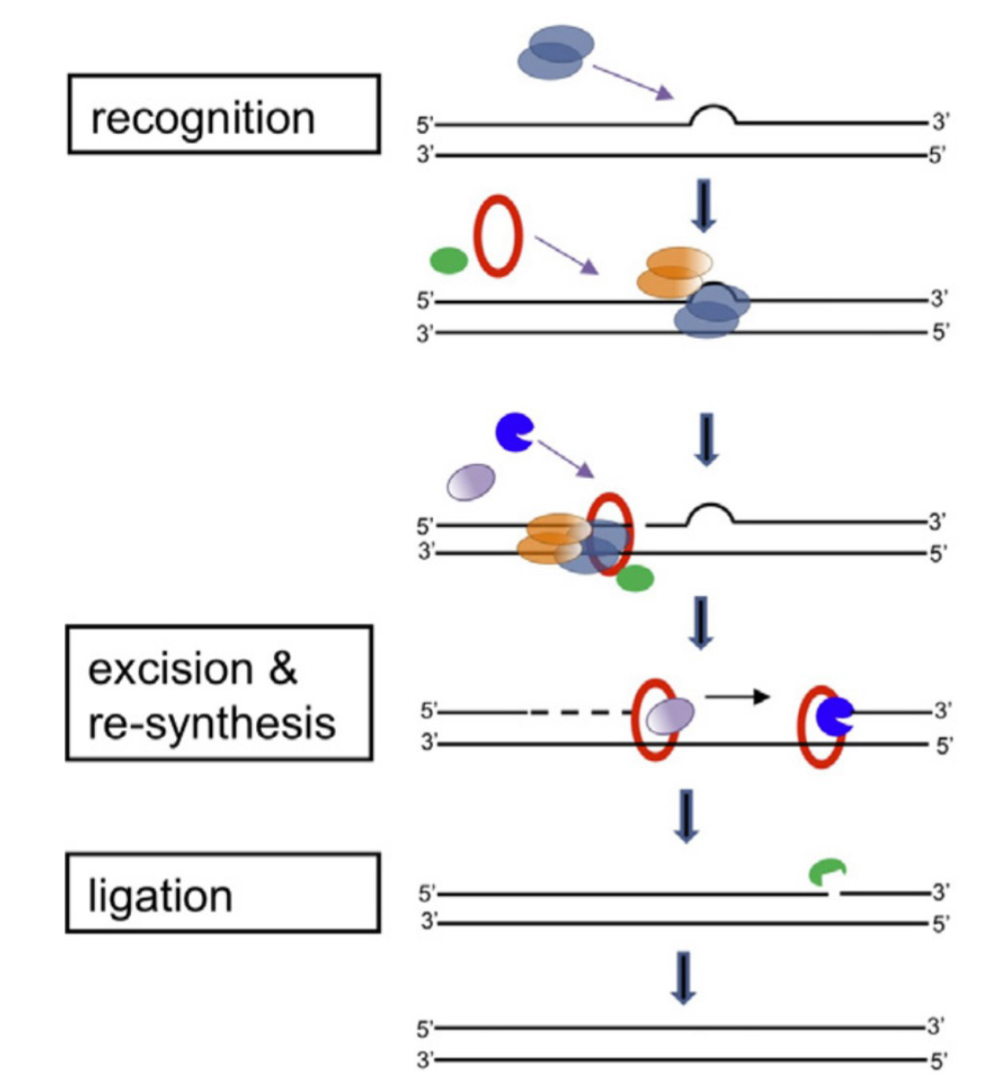
NER pathway
Recruitment of TFIIH complex (SU: XPB, XPD: helicases, associated XPA)
Unwinding
Cut by endonucleases (XPG, XPF):
25-30 nucleotides excised (excises region around damage)
DNA synthesis (pol δ/ε)
Ligation to seal nick
Role for DICER in chromatin decondenzation
Xeroderma pigmentosum (XP)
Hypersensitivity to sunlight (UV, eyes, skin)
Autosomal recessive
Dry skin, freckles, skin coloring
Predisposition to cancer (skin and other types of cancer)
Development of neurological abnormalities
XPA-G, V
Cockayne syndrome (CS)
Neurodegenerative disease
No predisposition to cancer
Premature aging, growth deficiency
Photosensitivity
XPB (D, G), CSA-B
NER and chromatin
DNA damage, histone mods, and NER are interlinked
Ubiquiten ligase recruited to DNA damage → H2AUb
Attracts ZRF1 (Zutoin-related factor 1)
HMT recruited to damage in DICER-dependent manner → H4K20me2
Recruitment of XPA
Double strand breaks
Can arise from replication over SS DNA nicks
SS nicks are transient during replication or repair paths
Deleterious consequences: cytotoxic damage
Initiate damage response pathways
Can arrest cell cycle (G1/S, S, G2/M)
Involve master protein kinases: ATR, DNA-PK, ATM
SSB induced by
DNA replication errors → stalled replication forks
Ionizing radiation
ROS (reactive oxygen species)
BER
SSB can lead to
DSB (dividing cels)
RNA pol stalling and transcriptional defects (non-dividing cells)
Important to delay replication to allow time to repair before they become DSBs
Single strand break repair (SSBR)
SSBs detected by poly (ADP ribose) polymerase 1 (PARP-1) - first responder
Recruits a “scaffold” protein (X-ray cross complementing group 1 (XRCC1))
XRCC1 interacts w/ various proteions
Next steps: can use BER factors
Double strand break repair (DSBR) pathways
NHEJ and HR
Non-homologous end joining (NHEJ)
Major pathway for DSBs
Direct ligation of broken ends
Mostly G1, but throughout cell cycle
Faster and less accurate (error prone), straight forward re-ligation
Homologous recombination (HR)
Uses sister chromatid or homologous chromosome as template
Mostly S, G2 phase (sister chromatids available)
Transcribed genes
Usually error free
NHEJ steps
Break detection by Ku heterodimers
Processing of broken ends
Ku recruits DNA-PK (DNA-dependent protein kinase)
Complex protects and aligns broken ends
Platform for DNA repair enzymes
End processing enzymes (depending on damage), e.g.:
DNA cross-link repair 1C (5’-3’ exonuclease), ss overhang cleavage
DNA pol µ and λ (fill in gaps)
Polynucleotide kinase (PNK) (3’ phosphatase, 5’ kinase activity)
End ligation
Ku recruits XRCC4 (X-ray repair cross complementing 4) and XLF protein
Targets DNA ligase VI to broke ends
If DSB recognized by MRN…
HR pathway intiated
Key characteristics of HR
One strand is cut; MRN endonucleases function (and EXO1 and helicase)
Long 3’ ssDNA (longer) → invade homologous SNA mediated by recombination proteins
Formation of Holliday junction: need to be resolved or dissolved
Involves BRCA1 and BRCA2
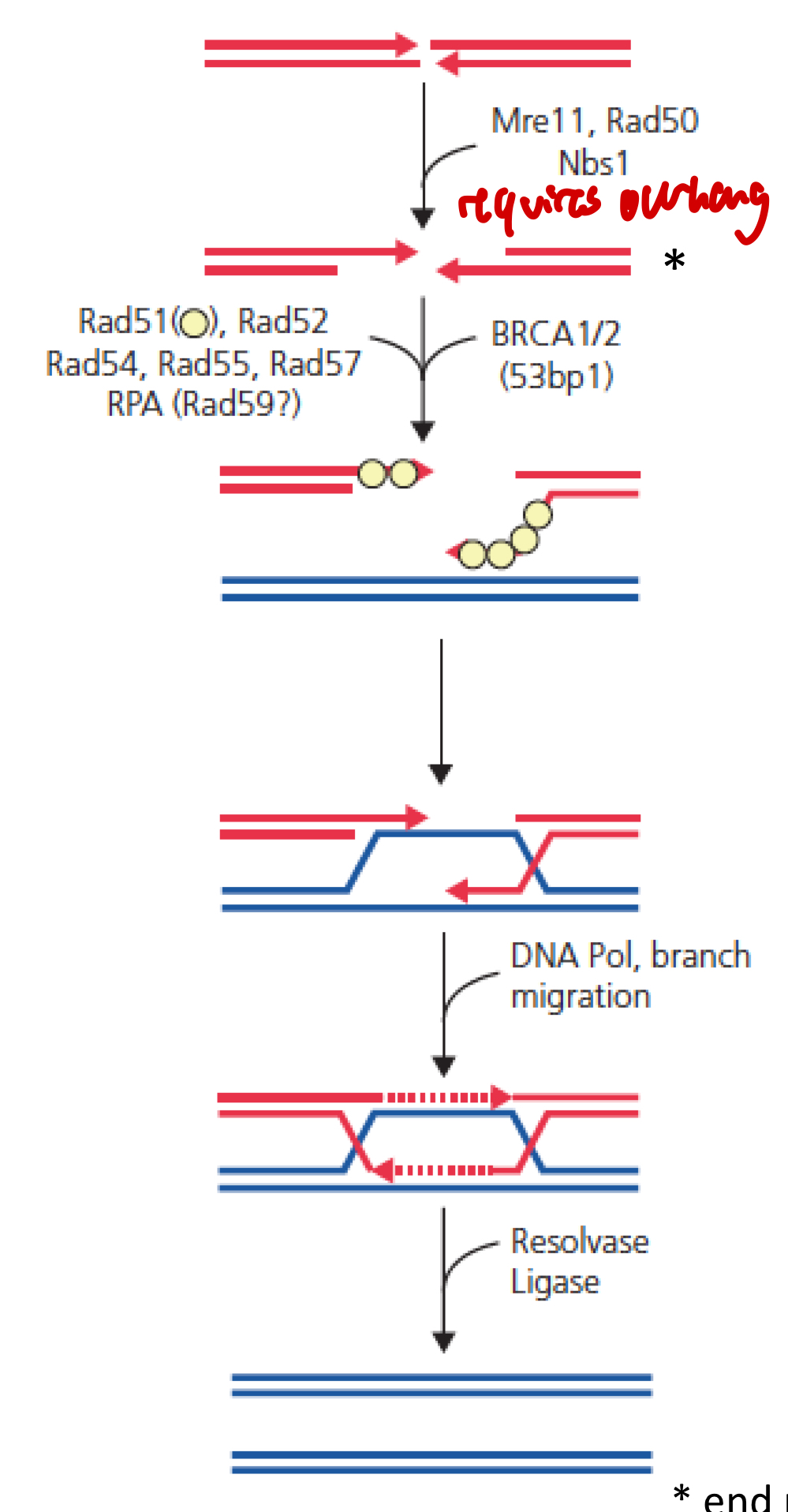
HR pathway
MRN complex detects DSB
MRN (+ EXO1 and helicase) → end procesing to generate ss tail: end resection
Processing of ends to create overhang
ssDNA tail covered by strand exchange protein: Rad51 (recA-type): filament for strand invasion
Stimulated by BRCA1 and BRCA2
D-loop → to invade homologous DNA strand
Formation of Holliday junction (4 connected helices, pol δ)
Or non-crossover synthesis-dependent strand annealing (SDSA)
Resolvase, dissolution
BRCA2 mutations
Germline mutations are heritable
Large “founder” effect in well-defined population
Cna theoretically be traced back to common ancestor
Iceland: single BRCA2 (999del5, frameshift, truncated form) accounts for almost all breast/ovarian cancer families
0.6% of gen population
Incomplete penetrance
5-methylcytosine
Transposable element silencing
Genome integrity and stability
Regulation of gene expression
Gene silencing
Protection of gene expression (gbM, moderately expressed, conserved genes)
Maintenance of DNA methylation
Plants = MET1
Mammals = DNMT1
Enzymes associate with PCNA/UHRF1 (ubiquitin-like with PHD and RING finger domains 1)
At replication fork
High affinity for hemimethylated DNA
Nucleosome removal at replication fork
H2A-H2B dimers removed before H3-H4 dimers
Replaced in opposite order
Synthesis of new histones required
Old and new histones associate with both strands
FACT (facilitates chromatin transcription) and ASF1 (anti-silencing function 1) help with disassembly
FACT re-establishes nucleosomes behind replication fork
+ other factors (CAF-1, Nap-1)
BER overview
Cause: ROS, X-rays, alkylating agents, spontaneous reactions
DNA damage: Oxidation, uracil, abasic site, SSB
Molecular components: DNA glycosylases, PARP 1, ATR kinase
MMR overview
Cause: Replication errors
DNA damage: A-G mismatch, T-C mismatch, insertion, deletion
Molecular components: MSH, MSL, pol δ
NER (GG- and TC-NER) overview
Cause: UV, light, polycyclic aromatic, hydrocarbons
DNA damage: Bulky adducts, intrastrand crosslink
Molecular components: XPC or pol II stalling; TFIIH
DSBR (NHEJ and NR) overview
Cause: X-rays, ionizing radiation, anti-tumor agents
DNA damage: DSB, interstrand crosslink
Molecular components: NHEJ = Ku dimer + DNA-PK; HR = MRN complex, ATM kinase, Rad51, BRCA 1/2, Holliday junction
Repair pathways most active during G0/G1
NHEJ
Repair pathways most active during S phase
BER, MMR, TC-NER, HR
Repair pathways most active during G2
HR
Microirradiation experiments
Cell line
UV - micro laser beam to cause localized damage in G1
Pulse labeling with 3 H-thymidine (radioactive isotope)
Detect “unscheduled” DNA-synthesis
Subsequent mitosis
With autoradiography
Determined that the interphase nucleus is organized instead of intermized
Hybrid cell line and in situ hybridization
Provided further evidence of the organization of interphase nucleus
Selective detection of X chromosome only
Hybrid cell line (hamster x man hybrid; all human chrs usually lost)
Contains active human X chromosome as the only free human chromosome
Cells with no X chromosome are lost
Metaphase plate and two interphase nuclei
In situ hybridization with 3H-labeled human genomic data
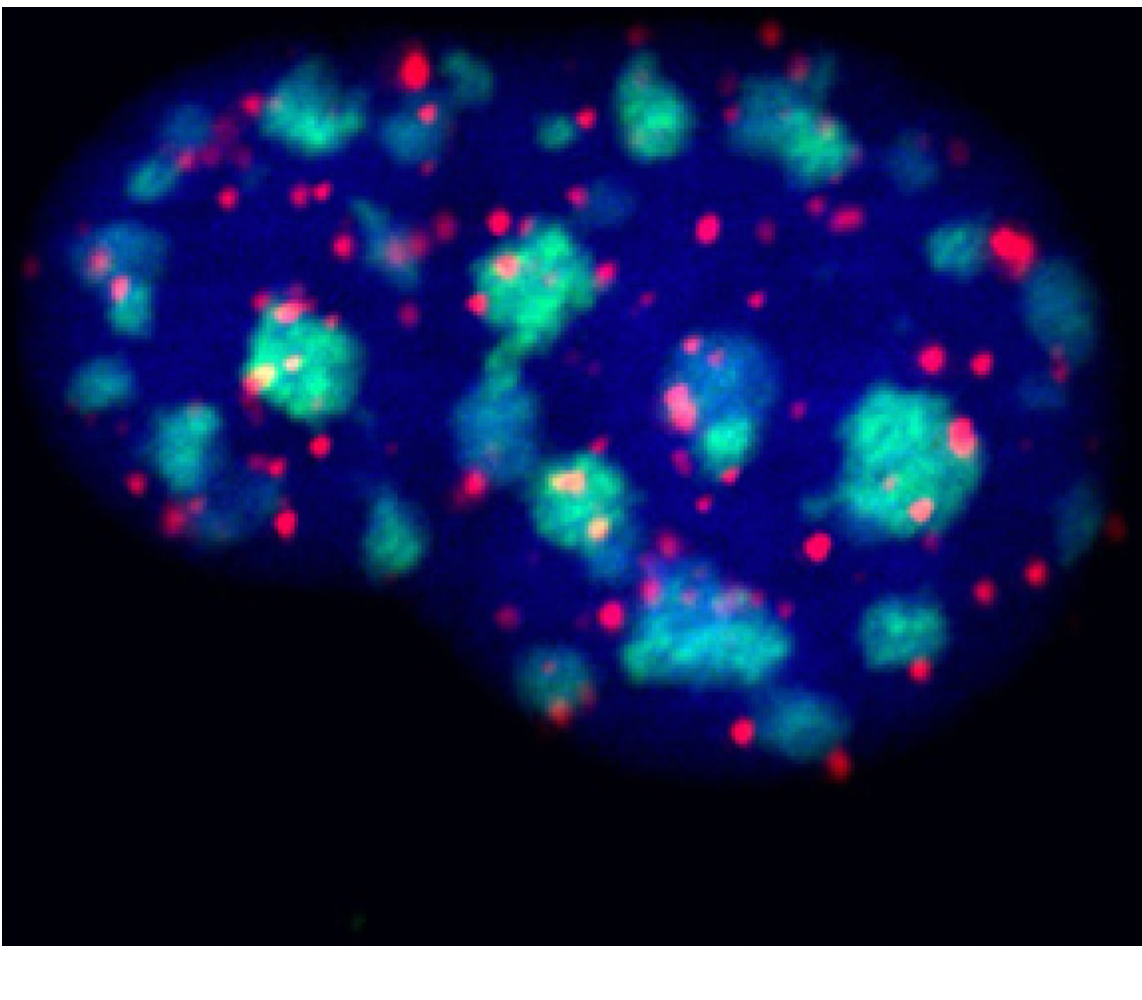
FISH as evidence for chromosome organization
Pancentromeric probe (green)
Telomeric probe (red)
DNA counterstained (blue)
HC11 (mouse mammary epithelium)
Chromosome territories (CTs)
Positioning: non-random (size, gene density)
Probabilistic, depending on tissue
Predominant configuration maintained during cell cycle
Relative position can strongly depend on cell types
Space between = splicing, transport, diffusible factors
CT subdivisions
Divided into active and inactive domains
Polycomb-repressed regions
Gradient
Heterochromatin towards outside, euchromatin towards middle
Topologically associated domains (TADs)
Structural building blocks of chromosomes (spatial structures)
Functional unites of gene regulation (co-expression)
kb-Mb in size
Separated by genetically defined boundary elements, recognized by architectural proteins
Housekeeping genes common at boarders
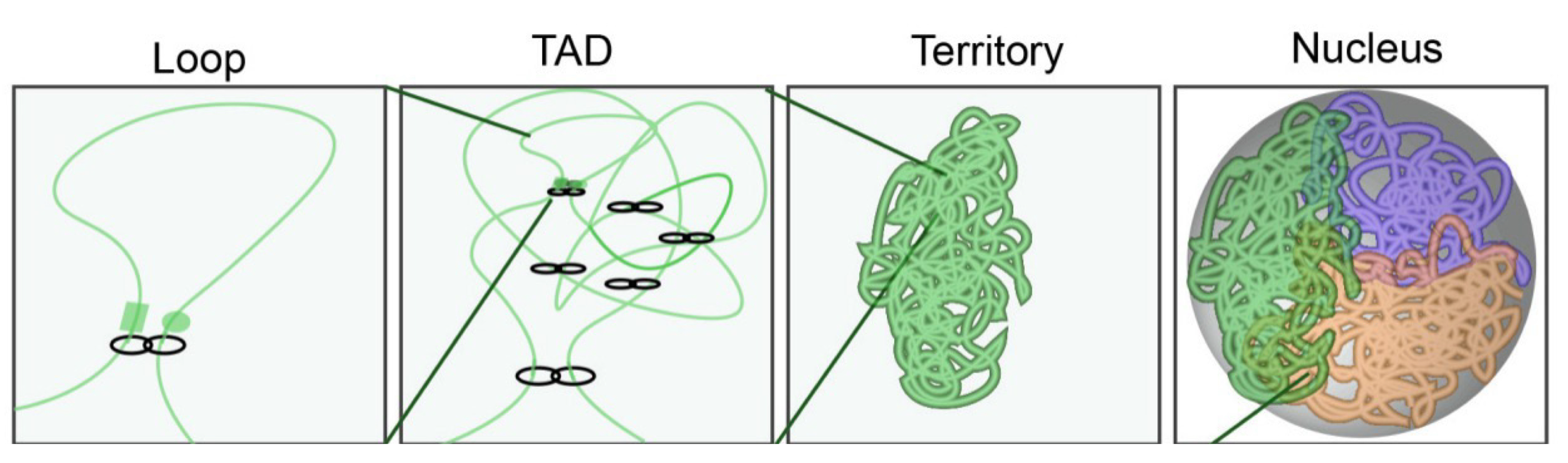
Loop formation in the genome
Human genome: 10,000 loops
Average loop size = 200,000 bp
Often similar chromatin marks within loop domain (e.g. H3K36me3, H3K27me3)
Domains w/ similar marks located next to each other (subcompartments)
Regulation of gene expression
Not fixed structure, can be dynamically adjusted
CTCF
Multidomain TF
11 zinc fingers (ZF)
Each ZF of ZF 3-7 contact 3 bases each of 15bp consensus
Shapes loops/subdomains
Fixes loop in place/stabilizes
Not symmetric
N-terminus → interacts w/ cohesion (stabilizing interaction)
C-terminus → No stable interaction w/ cohesion
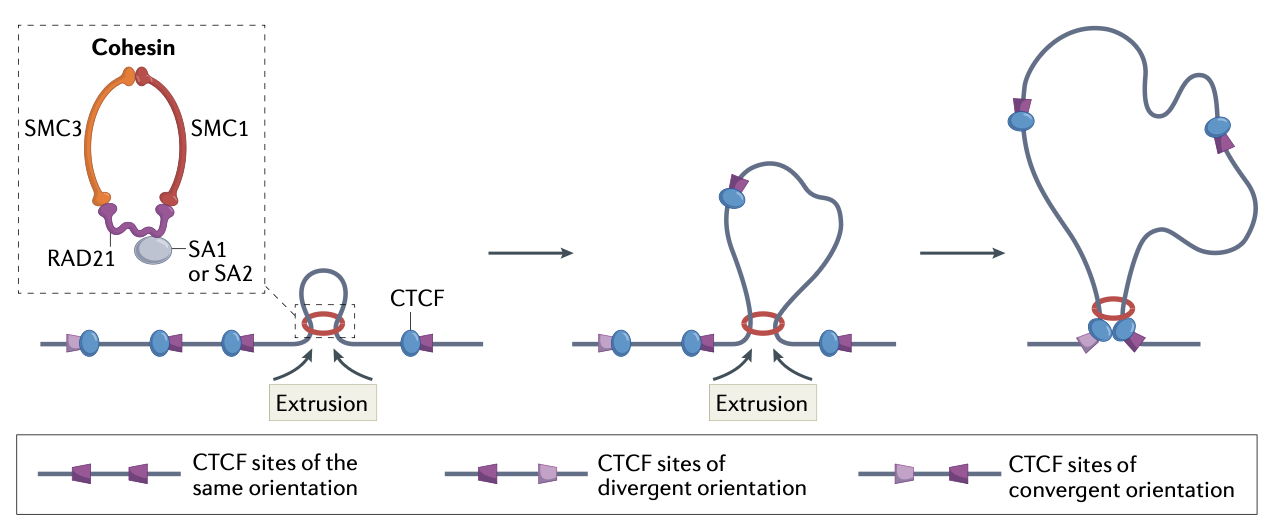
Cohesion
Forms a ring like structure that topologically encircles the DNA
Is an SMC complex (structural maintenance of chromosomes)
Uses ATP to translocate DNA and extrude loops (get larger and larger)
Reels DNA from both sides until barrier is met
CTCF orientation and loop extrusion process
If CTCF-bound to DNA encounters a cohesion ring, it pauses a bit (regardless of orientation)
If cohesion encounters N-terminus → forms a stable interaction and cohesion remains paused at the CTCF
If cohesion encounters C-terminus → no stable interaction is formed and the cohesion interface is partially opened/permissive and the DNA w/ CTCF passes through
Extrusion continues until each side of the cohesion complex encounters a DNA bound CTCF whose N-terminus faces the cohesion complex (i.e. a convergent pair)
Allows for stabilization on both sides
Ways to study the organization of the interphase DNA
FISH, 3C, DAPI stain
Examples for CTCF-mediated loops in the regulation of gene expression:
Human 𝛃-globulin locus: active chromatin hub
IGF2-H19 system: enhancer blocking
Hox-timer: Extrusion into sub TADs
Loops in human 𝛃-globulin locus
Active chromatin hub
Sequential activation of the β-globin gene cluster
5 gene, arranged in order of expression during development
One shared Locus Control Region
Complex enhancers, modulate chromatin structure
Recruitment of transcription factors
Recruitment of HAT activities: hyperacetylation of H3
Folding activates gene expression → ACH, loop
Loop formation and imprinting of IGF2-H19 system
Enhancer blocking and sub TADs
Gene is expressed based on whether the copy came from mom or dad
CTCF forms a loop to block enhancer and Igf2 interaction
Loop formation and Hox gene clusters
Subset of homeobox genes
Master regulators of development, control of body pain
Chromosomal organization: anterior-posterior expression
Co-linear gene activation: requires coordinated changes in higher order chromatin
Gradual looping out from chromosome territory coincides with gene expression
CT reorganization
Nocturnal animals use the physics of chromatin as a lens
Inverted nuclei: “microlenses” → channel photons to photosensitive ganglion cells
Dense in the center (heterochromatin), funnels light
Nuclear envelope
The interface between nucleus and cytoplasm
Protects the genome
Role in gene regulation
Structure of the nuclear membrane
Double membrane
Outer membrane: connected with ER
Inner membrane: associated with lamina
Nuclear core complex
Nuclear lamina
Mesh of protein fibers
Complex filamentous protein network associated with the inner nuclear membrane
Covers inner membrane
Consists of Lamins (4 types in mammals) + Lamina associated proteins
Plants: Nuclear Matrix Constituent Proteins (NMCPs)
Provides mechanical stability of the nucleus
Used as anchor proteins for chromatin
Attachment of the genome (e.g. via HP1)
Restricts CT movement; maintenance of chromosome positioning
Helps CTs stay in place
Defects can lead to age related disorders
Genes coding for proteins of the nuclear lamina
Most invertebrates: single lamin
Most vertebrates: 4 genes
Lamin B1, B2, A, LIII (lost in most mammals)
Mislocalization of centromeres
Mutation in human lamin A gene (LMNA)
Missense 433G>A (E145K)
Mutation in a domain of the protein (𝛂-helical central rod) that is required for polymerization
Abnormal localization of centromeres, mislocation of telomeres
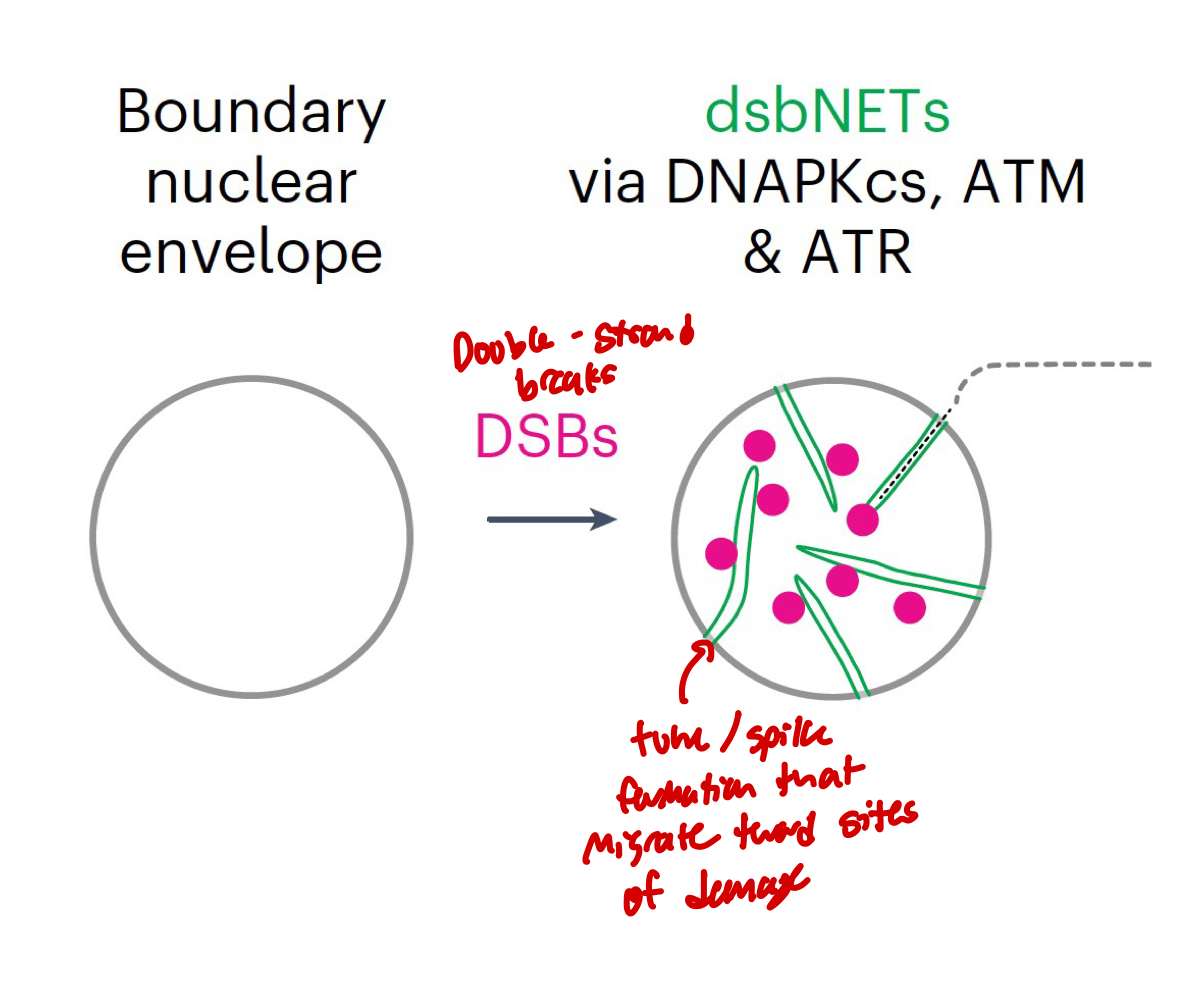
Link between nuclear morphology and DNA repair
Frequent DNA DSBs can induce nuclear envelope tubules: DSB-capturing nuclear envelope tubules (dsbNETs)
Contribute to genome stability
Nuclear structure-function relationships
Human cell lines
Nuclear core complex
Large multiprotein structure
Ca. 30 different proteins, in repeated subunits
transport requires nuclear localization sequence (import) and nuclear export seqeunce
Nuclear bodeis
Membraneless structures in the nucleus (i.e. Nucleolus, cajal bodies, PML bodies, speckles, transcription factories)
Nucleolus
Ribosome biogenesis
Cajal bodies
Telomerase biogenesis; RNA processing, small nuclear ribonucleoproteins snRNP
PML bodies
a.o. Promyelocytic Leukemia Protein genome maintenance
Speckles
Enriched in pre-mRNA splicing factors, CT borders
Biomolecular condensates
Arise through phase separation for membraneless compartment
Driven by multivalent macromolecular interactions
Separation into two phases:
One with higher concentration and once with lower
Need macromolecules and charges/polarity → changes the interactions
Can exist outside of nucleus
Liquid-liquid phase separation (LLPS) or condensation
Formation of membraneless compartments (membraneless bodies, biomolecular condensates, liquid assemblies)
Occurs in solutions of macromolecules automatically if conditions are right
Can separate into a dense phase (resembles liquid droplets) and a dilute phase
Functional benefits of three features common to condensates:
Compartmentalization → give substructure
Selective partitioning → bring similar molecules in close proximity
Concentration → act as reaction hubs/storage compartments
Transcription factories
Transcriptional hotspots
Pol II transcription organized into small structures
Fewer sites of active transcription than active genes and RNA pol II molecules (grouping of resources)
Brings co-expressed genes into direct contact
Hyperphosphorylated elongation form of RNA pol II
Ex) erythroid cells (𝛼- & β-globulin, Eraf [𝛼-globulin-stabilizing proteins]: closer together than other cell types)
Nucleolus
FC/DFC → center; rRNA gene transcription by pol I (except for 5s rRNA: pol III)
DFC → rRNA transcript processing
GC → outer region of nucleolus
Associated with re-imported ribosomal proteins
Final stages of ribosome assembly
Perinucleolar heterochromatin → silent rRNA genes loop out of nucleolus
Other processes:
Assembly of telomerase (to be transported to Cajal bodies)
miRNA storage
tRNA transcription by pol III
Forward genetics
Which gene(s) is/are necessary to support a phenomenon
No prior knowledge of the gene involved required:
Unbiased and powerful
Don’t need to know mechanisms of genes/pathways
Works well for single gene/phenotype
Things needed for forward genetics
Phenotype
Large data set or collection of random mutations
Screen for mutant phenotypes
Outcome/tools of forward genetics
Genetic linkage map
Physical map
Cytogenetic maps
How to get a mutant collection
Induce mutagenesis (use mutagen on organism)
Chemically, radiation, molecularly (T-DNA insertion, transposon mutagenesis)
Go out in field and collect different ecotypes (differ in different ways)
Spontaneous, varieties, true-to-type)
Sutton and Boveri
Found structure of metaphase chromosomes; Info comes in bundles (linked traits)
Back cross/test cross
Cross a heterozygous offspring with the homozygous recessive parent
Gamete ratio if characteristics are independent (on different chromosomes)
1:1:1:1
Gamete ratio if characteristics are coupled (on same chromosome)
2:0:0:2
Genetic/linkage map
Based on linkage and recombination frequency
1% recomb frequency = 1cM
Not a true physical distance
Physical maps
Genomic/chromosome fragment library, cloning, sequencing
Uses physical distance in bp, based on:
Restriction Fragment Length Polymorphisms (RFLPs)
Variable number of tandem repeats (VNTRs)
Single nucleotide polymorphisms (SNPs)
Requires overlap (contig construction, chromosome walking)
True physical representation of the genome
Modern application = optical maps (genome assembly, scaffolding)
Restriction Fragment Length Polymorphisms (RFLPs)
Cut DNA with restriction enzymes to get different fragment lengths to ID marks in the DNA; used to determine overall structure