Western Blot Protocol
1/21
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
22 Terms
Materials Provided
transfer buffer
methanol
filter paper
TBS: essential for washing, blocking, and antibody incubation
5% Tween-20: reduces non-specific antibody binding
25% Milk powder: blocking agent
PVDF membrane
primary antibody at 1:1000
colorimetric reagent
protein ladder
Transfer: Step 1
wet PVDF membrane, cut to gel size in 100% methanol
soak in transfer buffer for 30 mins
Transfer: Step 2
soak SDS-PAGE gel in transfer buffer for 5 mins
discard the buffer
repeat this step 2 more times (total 3 buffer changes)
Transfer: Step 3
soak 2 pieces of thick filter paper in transfer buffer
Transfer: Step 4
assemble the transfer stack on the platinum transfer apparatus anode, using a test tube or similar to roll out bubble between layers
the stack (bottom to top): thick filter paper, PVDF membrane, acrylamide gel, thick filter paper
Transfer: Step 5
replace the cathode and safety cover
run at 25V and 0.25A for 40 mins
Blocking and Probing of Membrane: Step 1
place the membrane into suitable container
ensuring the protein side is up, block non-specific sites by soaking in 20ml of 5% Milk powder in TBS-T for 60 mins
place the container on the shaking platform, ensure that the membrane is covered in solution
Blocking and Probing of Membrane: Step 2
remove the blocking solution (dispose in sink) and incubate with 15ml of primary antibody diluted to 1:1000 in 1% Milk powder/TBS-T for 30 min at room temp
Blocking and Probing of Membrane: Step 3
prepare 50ml of TBS-T (TBS w/ 0.1% Tween20)
Blocking and Probing of Membrane: Step 4
remove the antibody solution and wash 3 times with 15ml of TBS-T (1×1min, 2×5min)
each wash can be disposed of in the sink
Blocking and Probing of Membrane: Step 5
remove the membrane using forceps and place it protein side up on a plastic sheet
pipette 2ml of colorimetric reagent onto the membrane and incubate until color reaches desired intensity
Blocking and Probing of Membrane: Step 6
take a photo of the membrane of transilluminator
Fold purification
measures how much a protein has been enriched after a purification step compared to the initial mixture
calculated by determining the fraction of a protein of interest in the sample before the purification and comparing it to the fraction after purification
i.e the ratio of the purified protein in the sample after purification divided by the ratio of our protein in the sample before purification
requires:
protein concentrations from Bradford assay for the cleared lysate (before purification), and purified protein (after)
densitometry data on the Western blot image
the volume of the samples loaded on the gel
Fold purification: Example Calculation Data
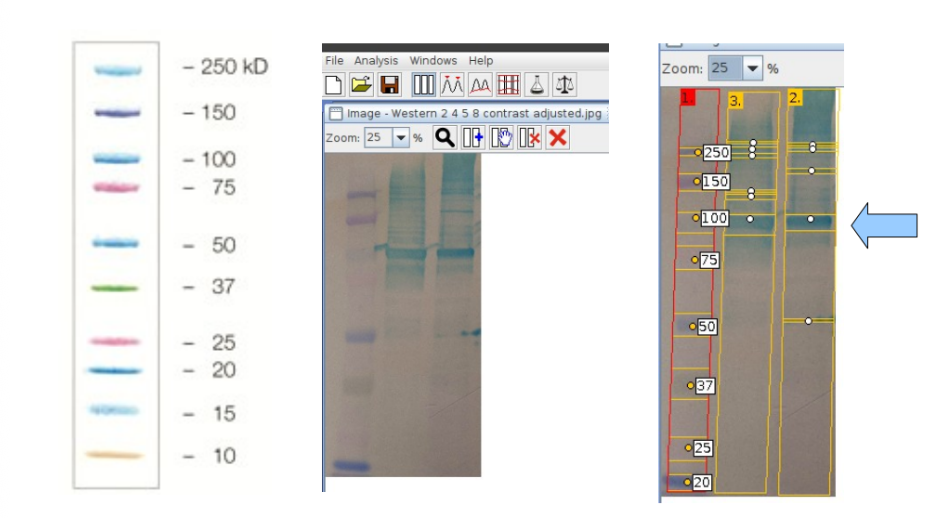
second lane = cleared lysate, third lane = purified protein
band of interest has a molecular weight of 104/102 kDA, which is close to the expected mass of 97kDA
therefore, pretty confident this band is correct
raw volume: lysate = 882, purified protein = 1084
therefore 1084/882 = 1.2 x as much recombinant protein in purified lane as in cleared lysate lane
the SDS-PAGE were loaded w/ 10µl of purified protein and ~3.8µl cleared lysate (3.8µl + 1.2 µl loading buffer)
Fold purification: Calculate Bradford Assay (this is now using our data lol)

Fold purification: Actual Western Blot Loading Volumes

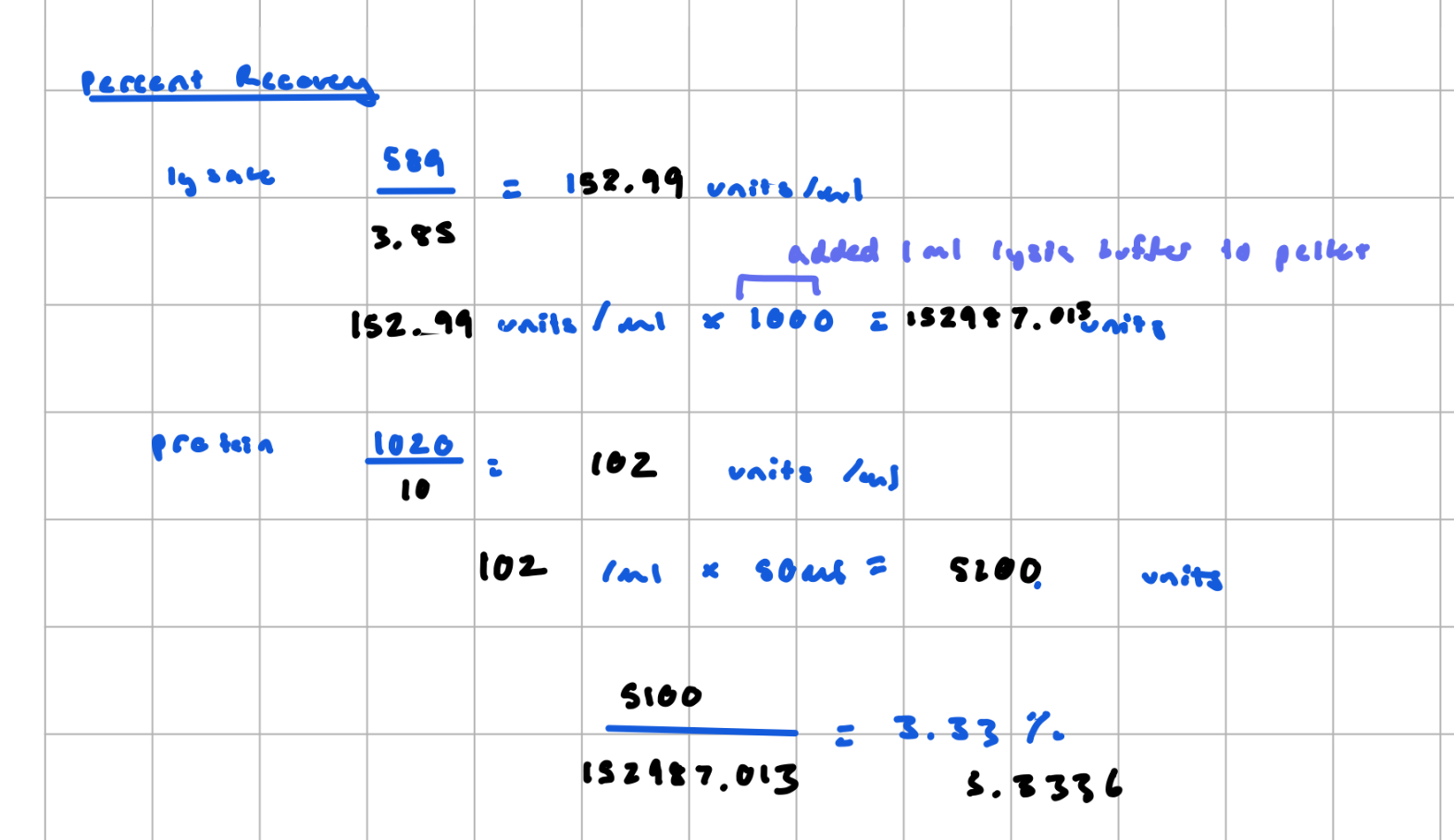
Fold purification

Percent Recovery
the yield, the percentage of total protein (or activity) you keep after purification step
requires:
total physical volumes of the purified protein and the cleared lysate solutions recovered (not just what was loaded on the gel)

Theory
detects/quantifies individual proteins within the mix
first step: protein components are separated based on molecular weight by electrophoresis in SDS-PAGE
proteins are then transferred to immobile substrate; the transfer process uses the same principle as SDS-PAGE, but the electric current is applied 90º to the gel and the proteins are migrated out of the gel onto the membrane
prior to incubation with the antibody solution, the membrane is blocked to reduce non specific (non-epitope) protein interactions between the membrane and antibody (we used non-fat dry milk)
small amount of detergent in the blocking and wash buffers also help to prevent non-specific binding
Primary Antibody
specific for protein of interest
incubated with the membrane in the blocking solution; at appropriate concentrations, the primary antibody should not bind any other proteins on the membrane (it frequently does tho)
Secondary Antibody
after washing the membrane in buffer/detergent solution to remove unbound primary antibody, a secondary antibody is incubated with the membrane
this antibody binds to one or more epitopes of the primary antibody and is typically linked to an enzyme that allows for visual identification by producing fluorescence or color change
the unbound secondary antibodies are washed away and the enzyme substrate is incubated with the membrane so that the positions of membrane-bound secondary antibodies can be detected
bands corresponding to detected protein of interest will appear and band densities in diff lanes can be compared
What is the difference between primary and secondary antibodies?
Primary:
directly binds to the target antigen (protein of interest), provides specificity, usually unlabeled
Secondary:
binds to the primary antibody (not the antigen)
conjugated to a detectable label (eg. enzyme, fluorophore)
provides signal amplification (multiple secondary antibodies can bind one primary)